1. Introduction
Epithelioid Malignant Peritoneal Mesothelioma (EMPeM) is a rare type of cancer, with reported cases in industrialized countries ranging between 0.5 and 3 per one million for male patients and 0.2–2 per one million for female patients over a five-year period [
1]. The main treatment option typically involves Cytoreductive Surgery (CRS) followed by Hyperthermic Intraperitoneal Chemotherapy (HIPEC) [
2]. However, many EMPeM patients are not eligible for CRS [
2]. Hence, physicians must consider alternative treatment options, such as chemotherapy and immunotherapy [
2]. Physicians cannot confidently predict which patients will respond well to these treatments [
2]. The disease is diagnosed using biopsies [
2] and the digitization of these biopsy samples leads to the creation of Whole-Slide Images (WSIs). WSIs are gigapixel-sized images of histopathological tissue [
3].
Survival analysis is a broad field which traditionally relied on the Kaplan–Meier and the Cox proportional-hazards models to estimate patients’ expected lifespan [
4]. In particular, these statistical models sought to obtain an estimate of patients’ Overall Survival (OS). OS is formally defined as the amount of time from the initiation of treatment until death or the last follow-up for individuals that survive the disease [
5]. Machine Learning (ML) approaches, including Random Survival Forests (RSFs) [
6] and Survival Support Vector Machines (SVMs) [
7] were later presented.
However, the emergence of Deep Learning (DL) and specifically Convolutional Neural Networks (CNNs) enabled the development of architectures that can exploit WSI features to perform this task. A foundational model named DeepConvSurv is presented by Zhu et al. [
8]. It is a fully supervised approach and has the limitation of relying on patches extracted from manually annotated Regions of Interest. Authors in [
9] propose an improved method, where patches are sampled randomly and subsequently grouped into clusters. DeepConvSurv is then trained using each cluster as input to obtain a survival prediction [
9]. Authors in [
10] present a fully-supervised network utilizing a ResNet50 network as a feature extractor, followed by a series of fully connected layers.
Later works prioritized Weak Supervision as a more scalable solution, particularly when dealing with large datasets that would otherwise require extensive manual annotation. The majority of approaches use Multiple Instance Learning (MIL). A key reference in this direction is [
11], in which the authors present MesoNet, which was developed from WSIs of pleural mesothelioma patients.
Newer contributions to the field deploy attention mechanisms and transformer-based architectures. For instance, WSS-CNN is an architecture presented in [
4], which consists of a weakly supervised CNN with a visual attention mechanism, which includes a spatial and a channel module. Authors in [
12] combined CNNs and Transformers to develop a network capable of learning both local and global features for survival prediction.
All of the above-mentioned frameworks tackle survival estimation as a ranked regression problem as they essentially seek to place patients from the test set in the correct order. Their main goal is to achieve the highest possible concordance index [
13], which is the most widely used metric for this task.
Instead of ranking survival times, Wulczyn et al. in [
14] formulated the task as a classification problem. Survival times are assigned into discrete intervals, thus creating discrete classification bins [
14]. The model’s objective is thus to allocate each WSI to the correct bin. MIL is also used in this case as an average pooling layer aggregate feature from multiple patches belonging to a specific patient [
14]. Furthermore, the recently published Surformer network obtains both global and local WSI features and deploys self and cross-attention mechanisms along with a custom loss function to achieve a higher concordance index compared to other methods [
15].
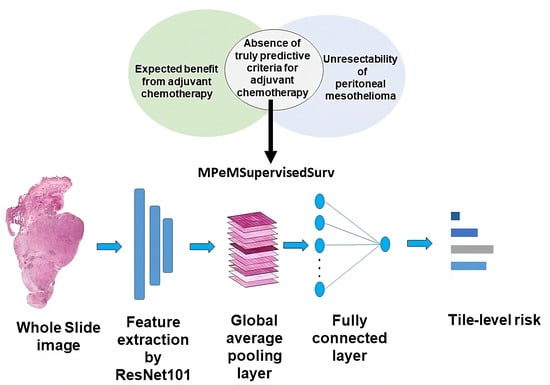
Our proposed network, MPeMSupervisedSurv, is a robust fully supervised network designed specifically to address the challenge of WSI-based survival estimation for EMPeM patients. Our approach is end-to-end and utilizes a neural network with a minimal number of layers, while avoiding the need for any manual annotations. Our contribution also includes a comprehensive analysis of the factors affecting the Overall Survival time of EMPeM patients. Our study is notable because it uniquely integrates clinical features with unstructured features extracted from Whole Slide Images (WSIs), providing clinicians with novel insights regarding this disease.
This paper is organised as follows:
Section 2 describes the WSI EMPeM dataset used in our study and our proposed neural network architecture. The model’s achieved accuracy compared to similar methods is presented in
Section 3 along with its performance in specific patient stratification scenarios.
Section 4 discusses the results of our research and mentions limitations along with directions of future research. A summary of the findings and conclusions of our analysis is presented in
Section 5.
4. Discussion
In general, the concordance index typically falls within the range of
[
13]. A c-index of
is equivalent to a random guess [
13], while a value equal to 1 signifies total accuracy [
13]. Hence, any proposed model must demonstrate accuracy exceeding
and be as close to 1 as possible.
The results in
Table 2 demonstrate the significant performance improvement of deep learning over machine learning methods. SurvivalSVM and Random Survival Forests achieve a concordance index approximately equal to
. In contrast, Deep Learning approaches designed to handle images demonstrate more robust performance as they achieve a concordance index of at least
. The performance difference between DeepConvSurv and the other methods listed in
Table 2 is notable. It is attributed to the fact that DeepConvSurv does not utilize transfer learning, while EE-Surv and MPeMSupervisedSurv both use pre-trained ResNet networks for feature extraction. Transfer learning leverages knowledge gained from a source task so that higher accuracy can be achieved on the task of interest [
24]. Its efficacy is well documented in various fields, such as medicine and bioinformatics, among others [
24]. Subsequently, training the model with a deeper feature extractor and a single fully connected layer resulted in a slightly higher
c-index value for this dataset, compared to what could be achieved with the EE-Surv architecture.
Our findings regarding the factors affecting OS appear to agree with those of prior studies, whilst also providing novel insights on variables previously unexplored. The results in
Table 3 and
Table 4 suggest that providing the model with prior knowledge of demographic variables does not significantly enhance its predictive ability. This concurs with the analyses undertaken in [
25,
26], which do not indicate that demographic variables influence survival outcomes.
Authors in [
25] discuss the impact of neoadjuvant chemotherapy in EMPeM patients. Their multivariate analysis revealed that chemotherapy was not a good prognostic factor for OS. The observed results shown in
Table 5 do not significantly differ, which suggests that MPeMSupervisedSurv did not identify any additional features from either cohort that could enhance its predictive ability. A similar conclusion can be drawn from the results reported in
Table 7 regarding HIPEC treatment. In contrast, the reported concordance index results for cohorts stratified using adjuvant chemotherapy in
Table 6 differ significantly. This implies that the model identified more meaningful WSI features for OS estimation. It also suggests that MPeMSupervisedSurv could identify patients who respond to adjuvant chemotherapy.
With regards to patient stratification based on the PCI, our findings indicate that the model exhibited robust performance for both low and high PCI groups. The PCI is a score that describes the extent to which the disease has spread [
26]. Salo et al. in [
26] present statistical analysis which indicates that the PCI is a prognostic factor for OS. Our model’s robust performance across both low and high PCI groups indicates that meaningful features can be identified in WSIs of both cohorts. This therefore appears to confirm that PCI plays a role in survival prediction.
This work can play an important role regarding treatment planning in a clinical setting. There is currently limited evidence on the efficacy of chemotherapy combinations such as cisplatin and pemetrexed [
27]. As a result, clinicians often encounter uncertainty when deliberating between utilizing chemotherapy treatment with cisplatin and pemetrexed, considering alternative chemotherapy regimens, or opting for close monitoring. Our model, MPeMSupervisedSurv, serves as an objective tool to aid in this decision-making process. By incorporating chemotherapy data, our model offers insights that can guide clinicians in their decision making. This equips clinicians with valuable insights to optimize patient care and treatment outcomes.
Limitations of this work include the fact that MPeMSupervisedSurv was trained and tested for this particular EMPeM cohort. Thus, further validation with another EMPeM cohort would help improve the model and ensure that it can capture representative and meaningful features from a wide range of patient populations. Another limitation is that the results related to patient stratification based on chemotherapy should be interpreted cautiously because of the smaller training set size for each category. This implies that further validation is required.
Developing an architecture that uses only WSI-level labels for supervision and can extract both local and long-range features by leveraging spatial attention is the primary direction of future work. Our intent is to create a model to extract previously unknown features that can be more informative about OS and general disease outcomes. We aim to validate our future model on a larger and more diverse set of peritoneal mesothelioma cases to ensure minimal bias and applicability in clinical practice.
Author Contributions
Conceptualization, P.B., N.B. and F.N.K.K.; methodology, K.M.P.; software, K.M.P.; validation, K.M.P.; formal analysis, K.M.P. and P.B.; data curation, P.D., V.K., S.V.-D., C.I.-V., G.A., A.C., M.-H.L., L.V., O.G., S.I., J.H.-F. and N.B.; writing—original draft preparation, K.M.P.; writing—review and editing, K.M.P., P.B. and T.S.; supervision, T.S. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
This study was conducted in accordance with the 2013 Declaration of Helsinki. The regulations and ethical principles mentioned in the Declaration of Helsinki are adhered to by RENAPE (French Network For Rare Peritoneal Tumors). The Comité Consultatif sur le Traitement de l’Information en matière de Recherche dans le domaine de la Santé (CCTIRS en_10.257) approved the creation of the RENAPE database. This database is also registered with the Commission Nationale de l’Informatique et des Libertés (CNIL e no. DR-2010-297), which was completed on 4 October 2010. The registration was completed in accordance with the provisions contained in law no. 78-17 of the French Republic regarding data processing, files, and individuals’ right to privacy that was adopted on 6 January 1978 (ClinicalTrials.gov Identifier: NCT02834169 [
16]).
Informed Consent Statement
Informed consent was obtained from all subjects involved in the study.
Data Availability Statement
The data presented in this study may be provided on request from Dr. Nazim Benzerdjeb. This study uses a private dataset of Whole-Slide Images. Due to its large size, it cannot be made available on a cloud network.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| OS | Overall Survival |
| EMPeM | Epithelioid Malignant Peritoneal Mesothelioma |
| CNN | Convolutional Neural Network |
| WSI | Whole-Slide Image |
| PCI | Peritoneal Cancer Index |
| ResNet | Residual Network |
| HIPEC | Hyperthermic Intraperitoneal Chemotherapy |
| SVM | Support Vector Machine |
| RSF | Random Survival Forest |
References
- Boffetta, P. Epidemiology of peritoneal mesothelioma: A review. Ann. Oncol. 2007, 18, 985–990. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Bhagwandin, S.; Labow, D.M. Malignant peritoneal mesothelioma: A review. Ann. Transl. Med. 2017, 5, 236. [Google Scholar] [CrossRef] [PubMed]
- Van der Laak, J.; Litjens, G.; Ciompi, F. Deep learning in histopathology: The path to the clinic. Nat. Med. 2021, 27, 775–784. [Google Scholar] [CrossRef]
- Qaiser, T.; Lee, C.Y.; Vandenberghe, M.; Yeh, J.; Gavrielides, M.; Hipp, J.; Scott, M.; Reischl, J. Usability of deep learning and HE images predict disease outcome-emerging tool to optimize clinical trials. NPJ Precis. Oncol. 2022, 6, 37. [Google Scholar] [CrossRef]
- Hess, L.M.; Brnabic, A.; Mason, O.; Lee, P.; Barker, S. Relationship between Progression-free Survival and Overall Survival in Randomized Clinical Trials of Targeted and Biologic Agents in Oncology. J. Cancer 2019, 10, 3717–3727. [Google Scholar] [CrossRef] [PubMed]
- Ishwaran, H.; Kogalur, U.B.; Blackstone, E.H.; Lauer, M.S. Random survival forests. Ann. Appl. Stat. 2008, 2, 841–860. [Google Scholar] [CrossRef]
- Ding, Z. The application of support vector machine in survival analysis. In Proceedings of the 2011 2nd International Conference on Artificial Intelligence, Management Science and Electronic Commerce (AIMSEC), Zhengzhou, China, 8–10 August 2011; pp. 6816–6819. [Google Scholar]
- Zhu, X.; Yao, J.; Huang, J. Deep convolutional neural network for survival analysis with pathological images. In Proceedings of the 2016 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Shenzhen, China, 15–18 December 2016; pp. 544–547. [Google Scholar]
- Zhu, X.; Yao, J.; Zhu, F.; Huang, J. WSISA: Making Survival Prediction From Whole Slide Histopathological Images. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017; pp. 6855–6863. [Google Scholar]
- Laleh, N.G.; Echle, A.; Muti, H.S.; Hewitt, K.J.; Volkmar, S.; Kather, J.N. Deep Learning for interpretable end-to-end survival (E-ESurv) prediction in gastrointestinal cancer histopathology. In Proceedings of the MICCAI Workshop on Computational Pathology, Online, 27 September–1 October 2021; Volume 156, pp. 81–93. [Google Scholar]
- Courtiol, P.; Maussion, C.; Moarii, M.; Pronier, E.; Pilcer, S.; Sefta, M.; Manceron, P.; Toldo, S.; Zaslavskiy, M.; Le Stang, N.; et al. Deep learning-based classification of mesothelioma improves prediction of patient outcome. Nat. Med. 2019, 25, 1519–1525. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Liu, L.; Tang, Z.; Chen, Z.; Ma, G.; Dong, J.; Zhang, X.; Yang, L.; Zheng, Q. Explainable Survival Analysis with Convolution-Involved Vision Transformer. In Proceedings of the AAAI Conference on Artificial Intelligence, Singapore, China, 17–20 June 2022; Volume 36, pp. 2207–2215. [Google Scholar]
- Steck, H.; Krishnapuram, B.; Dehing-oberije, C.; Lambin, P.; Raykar, V.C. On Ranking in Survival Analysis: Bounds on the Concordance Index. In Proceedings of the Advances in Neural Information Processing Systems, Vancouver, BC, Canada, 3–6 December 2007; Volume 20, pp. 1209–1216. [Google Scholar]
- Wulczyn, E.; Steiner, D.F.; Xu, Z.; Sadhwani, A.; Wang, H.; Flament-Auvigne, I.; Mermel, C.H.; Chen, P.H.C.; Liu, Y.; Stumpe, M.C. Deep learning-based survival prediction for multiple cancer types using histopathology images. PLOS ONE 2020, 15, e0233678. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Gao, Q.; Yi, X.; Zhang, X.; Zhang, Y.; Zhang, D.; Liò, P.; Bain, C.; Bassed, R.; Li, S.; et al. Surformer: An interpretable pattern-perceptive survival transformer for cancer survival prediction from histopathology whole slide images. Comput. Methods Programs Biomed. 2023, 241, 107733. [Google Scholar] [CrossRef] [PubMed]
- French National Registry of Rare Peritoneal Surface Malignancies (RENAPE). Available online: https://clinicaltrials.gov/study/NCT02834169 (accessed on 10 September 2023).
- Echle, A.; Rindtorff, N.T.; Brinker, T.J.; Luedde, T.; Pearson, A.T.; Kather, J.N. Deep learning in cancer pathology: A new generation of clinical biomarkers. Br. J. Cancer 2021, 124, 686–696. [Google Scholar] [CrossRef] [PubMed]
- Wsi-Tile-Cleanup. Available online: https://github.com/lucasrla/wsi-tile-cleanup (accessed on 23 September 2023).
- Jiang, J.; Trundle, P.; Ren, J. Medical image analysis with artificial neural networks. Comput. Med. Imaging Graph. 2010, 34, 617–631. [Google Scholar] [CrossRef] [PubMed]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Toğaçar, M.; Ergen, B.; Tümen, V. Use of dominant activations obtained by processing OCT images with the CNNs and slime mold method in retinal disease detection. Biocybern. Biomed. Eng. 2022, 42, 646–666. [Google Scholar] [CrossRef]
- Mormont, R.; Geurts, P.; Marée, R. Multi-Task Pre-Training of Deep Neural Networks for Digital Pathology. IEEE J. Biomed. Health Inform. 2021, 25, 412–421. [Google Scholar] [CrossRef] [PubMed]
- Scikit-Survival. Available online: https://scikit-survival.readthedocs.io/en/stable/ (accessed on 13 December 2023).
- Hosna, A.; Merry, E.; Gyalmo, J.; Alom, Z.; Aung, Z.; Azim, M.A. Transfer learning: A friendly introduction. J. Big Data 2022, 9, 102. [Google Scholar] [CrossRef] [PubMed]
- Naffouje, S.A.; Tulla, K.A.; Salti, G.I. The impact of chemotherapy and its timing on survival in malignant peritoneal mesothelioma treated with complete debulking. Med. Oncol. 2018, 35, 69. [Google Scholar] [CrossRef] [PubMed]
- Salo, S.; Lantto, E.; Robinson, E.; Myllarniemi, M.; Laaksonen, S.; Salo, J.; Rantanen, T.; Ilonen, I. Prognostic role of radiological peritoneal cancer index in malignant peritoneal mesothelioma: National cohort study. Sci. Rep. 2020, 10, 13257. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, S.; Asao, T.; Tsutsumi, S.; Fujii, T.; Yamauchi, H.; Kigure, W.; Morita, H.; Yajima, R.; Suto, T.; Kuwano, H. Effectiveness of Intraperitoneal Hyperthermo-Chemotherapy for Malignant Peritoneal Mesothelioma and Estimation of its Effect by Repeated FDG-PET: A Case Report. Hepato-Gastroentorology 2011, 58, 861–864. [Google Scholar]
| Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).