Journal Description
Genes
Genes
is a peer-reviewed, open access journal of genetics and genomics published monthly online by MDPI. The Spanish Society for Biochemistry and Molecular Biology (SEBBM) is affiliated with Genes and their members receive discounts on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, MEDLINE, PMC, Embase, PubAg, and other databases.
- Journal Rank: JCR - Q2 (Genetics & Heredity) / CiteScore - Q2 (Genetics)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 16.5 days after submission; acceptance to publication is undertaken in 2.3 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: Reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
Impact Factor:
3.5 (2022);
5-Year Impact Factor:
3.9 (2022)
Latest Articles
Altered Expression of PDE4 Genes in Schizophrenia: Insights from a Brain and Blood Sample Meta-Analysis and iPSC-Derived Neurons
Genes 2024, 15(5), 609; https://doi.org/10.3390/genes15050609 (registering DOI) - 10 May 2024
Abstract
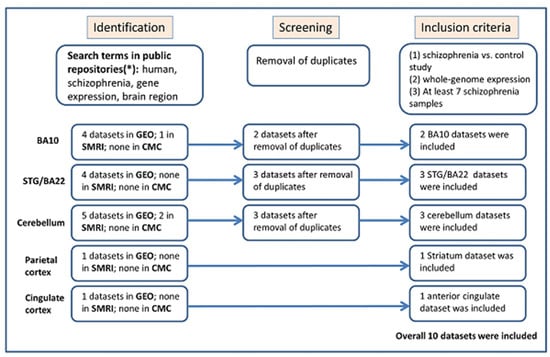
Schizophrenia symptomatology includes negative symptoms and cognitive impairment. Several studies have linked schizophrenia with the PDE4 family of enzymes due to their genetic association and function in cognitive processes such as long-term potentiation. We conducted a systematic gene expression meta-analysis of four PDE4
[...] Read more.
Schizophrenia symptomatology includes negative symptoms and cognitive impairment. Several studies have linked schizophrenia with the PDE4 family of enzymes due to their genetic association and function in cognitive processes such as long-term potentiation. We conducted a systematic gene expression meta-analysis of four PDE4 genes (PDE4A-D) in 10 brain sample datasets (437 samples) and three blood sample datasets (300 samples). Subsequently, we measured mRNA levels in iPSC-derived hippocampal dentate gyrus neurons generated from fibroblasts of three groups: healthy controls, healthy monozygotic twins (MZ), and their MZ siblings with schizophrenia. We found downregulation of PDE4B in brain tissues, further validated by independent data of the CommonMind consortium (515 samples). Interestingly, the downregulation signal was present in a subgroup of the patients, while the others showed no differential expression or even upregulation. Notably, PDE4A, PDE4B, and PDE4D exhibited upregulation in iPSC-derived neurons compared to healthy controls, whereas in blood samples, PDE4B was found to be upregulated while PDE4A was downregulated. While the precise mechanism and direction of altered PDE4 expression necessitate further investigation, the observed multilevel differential expression across the brain, blood, and iPSC-derived neurons compellingly suggests the involvement of PDE4 genes in the pathophysiology of schizophrenia.
Full article
(This article belongs to the Special Issue Genetic Basis Underlying Neuropsychiatric Disorders 2.0)
►
Show Figures
Open AccessArticle
Transcriptomic Analysis of Green Leaf Plants and White–Green Leaf Mutants in Haworthia cooperi var. pilifera
by
Peiling Li, Maofei Ren, Juanjuan Chen, Jianhua Yue, Songhu Liu, Qingsong Zhu and Zhiyong Wang
Genes 2024, 15(5), 608; https://doi.org/10.3390/genes15050608 (registering DOI) - 10 May 2024
Abstract
Haworthia cooperi var. pilifera is a succulent plant with ornamental value. The white–green leaf mutant (wl) showed a significant difference in leaf color from the wild-type plant (WT). In this study, we integrated the transcriptomes of wl and WT plants to
[...] Read more.
Haworthia cooperi var. pilifera is a succulent plant with ornamental value. The white–green leaf mutant (wl) showed a significant difference in leaf color from the wild-type plant (WT). In this study, we integrated the transcriptomes of wl and WT plants to screen differentially expressed genes related to leaf color variation. The results of transcriptome analysis showed that 84,163 unigenes were obtained after de novo assembly and the NR database annotated the largest number of unigenes, which accounted for 57.13%, followed by NT (43.02%), GO (39.84%), Swiss-Prot (39.25%), KEGG (36.06%), and COG (24.88%). Our finding showed that 2586 genes were differentially expressed in the two samples, including 1996 down-regulated genes and 590 up-regulated genes. GO analysis predicted that these differentially expressed genes (DEGs) participate in 12 cellular components, 20 biological processes, and 13 molecular function terms and KEGG analysis showed that metabolic pathways, plant–pathogen interaction, glycerophospholipid metabolism, endocytosis, plant hormone signal transduction, and ether lipid metabolism were enriched among all identified pathways. Through functional enrichment analysis of DEGs, we found that they were involved in chloroplast division and the biosynthesis of plant pigments, including chlorophyll, carotenoids, anthocyanin, and transcription factor families, which might be related to the formation mechanism of leaf color. Taken together, these results present insights into the difference in gene expression characteristics in leaves between WT and wl mutants and provide a new insight for breeding colorful leaf phenotypes in succulent plants.
Full article
(This article belongs to the Section Plant Genetics and Genomics)
►▼
Show Figures

Figure 1
Open AccessArticle
Is CYP2C Haplotype Relevant for Efficacy and Bleeding Risk in Clopidogrel-Treated Patients?
by
Lana Ganoci, Jozefina Palić, Vladimir Trkulja, Katarina Starčević, Livija Šimičević, Nada Božina, Martina Lovrić-Benčić, Zdravka Poljaković and Tamara Božina
Genes 2024, 15(5), 607; https://doi.org/10.3390/genes15050607 - 10 May 2024
Abstract
A recently discovered haplotype—CYP2C:TG—determines the ultrarapid metabolism of several CYP2C19 substrates. The platelet inhibitor clopidogrel requires CYP2C19-mediated activation: the risk of ischemic events is increased in patients with a poor (PM) or intermediate (IM) CYP2C19 metabolizer phenotype (vs. normal, NM; rapid,
[...] Read more.
A recently discovered haplotype—CYP2C:TG—determines the ultrarapid metabolism of several CYP2C19 substrates. The platelet inhibitor clopidogrel requires CYP2C19-mediated activation: the risk of ischemic events is increased in patients with a poor (PM) or intermediate (IM) CYP2C19 metabolizer phenotype (vs. normal, NM; rapid, RM; or ultrarapid, UM). We investigated whether the CYP2C:TG haplotype affected efficacy/bleeding risk in clopidogrel-treated patients. Adults (n = 283) treated with clopidogrel over 3–6 months were classified by CYP2C19 phenotype based on the CYP2C19*2*17 genotype, and based on the CYP2C19/CYP2C cluster genotype, and regarding carriage of the CYP2:TG haplotype, and were balanced on a number of covariates across the levels of phenotypes/haplotype carriage. Overall, 45 (15.9%) patients experienced ischemic events, and 49 (17.3%) experienced bleedings. By either classification, the incidence of ischemic events was similarly numerically higher in PM/IM patients (21.6%, 21.8%, respectively) than in mutually similar NM, RM, and UM patients (13.2–14.8%), whereas the incidence of bleeding events was numerically lower (13.1% vs. 16.6–20.5%). The incidence of ischemic events was similar in CYP2C:TG carries and non-carries (14.1% vs. 16.1%), whereas the incidence of bleedings appeared mildly lower in the former (14.9% vs. 20.1%). We observed no signal to suggest a major effect of the CYP2C19/CYP2C cluster genotype or CYP2C:TG haplotype on the clinical efficacy/safety of clopidogrel.
Full article
(This article belongs to the Section Pharmacogenetics)
►▼
Show Figures

Figure 1
Open AccessArticle
miR-423-5p Regulates Skeletal Muscle Growth and Development by Negatively Inhibiting Target Gene SRF
by
Yanqin Pang, Jing Liang, Jianfang Huang, Ganqiu Lan, Fumei Chen, Hui Ji and Yunxiang Zhao
Genes 2024, 15(5), 606; https://doi.org/10.3390/genes15050606 - 10 May 2024
Abstract
The process of muscle growth directly affects the yield and quality of pork food products. Muscle fibers are created during the embryonic stage, grow following birth, and regenerate during adulthood; these are all considered to be phases of muscle development. A multilevel network
[...] Read more.
The process of muscle growth directly affects the yield and quality of pork food products. Muscle fibers are created during the embryonic stage, grow following birth, and regenerate during adulthood; these are all considered to be phases of muscle development. A multilevel network of transcriptional, post-transcriptional, and pathway levels controls this process. An integrated toolbox of genetics and genomics as well as the use of genomics techniques has been used in the past to attempt to understand the molecular processes behind skeletal muscle growth and development in pigs under divergent selection processes. A class of endogenous noncoding RNAs have a major regulatory function in myogenesis. But the precise function of miRNA-423-5p in muscle development and the related molecular pathways remain largely unknown. Using target prediction software, initially, the potential target genes of miR-423-5p in the Guangxi Bama miniature pig line were identified using various selection criteria for skeletal muscle growth and development. The serum response factor (SRF) was found to be one of the potential target genes, and the two are negatively correlated, suggesting that there may be targeted interactions. In addition to being strongly expressed in swine skeletal muscle, miR-423-5p was also up-regulated during C2C12 cell development. Furthermore, real-time PCR analysis showed that the overexpression of miR-423-5p significantly reduced the expression of myogenin and the myogenic differentiation antigen (p < 0.05). Moreover, the results of the enzyme-linked immunosorbent assay (ELISA) demonstrated that the overexpression of miR-423-5p led to a significant reduction in SRF expression (p < 0.05). Furthermore, miR-423-5p down-regulated the luciferase activities of report vectors carrying the 3′ UTR of porcine SRF, confirming that SRF is a target gene of miR-423-5p. Taken together, miR-423-5p’s involvement in skeletal muscle differentiation may be through the regulation of SRF.
Full article
(This article belongs to the Section Animal Genetics and Genomics)
►▼
Show Figures

Figure 1
Open AccessArticle
Molecular and Physiological Effects of 17α-methyltestosterone on Sex Differentiation of Black Rockfish, Sebastes schlegelii
by
Haijun Huang, Yuyan Liu, Qian Wang, Caichao Dong, Le Dong, Jingjing Zhang, Yu Yang, Xiancai Hao, Weijing Li, Ivana F. Rosa, Lucas B. Doretto, Xuebin Cao and Changwei Shao
Genes 2024, 15(5), 605; https://doi.org/10.3390/genes15050605 - 9 May 2024
Abstract
It is widely known that all-female fish production holds economic value for aquaculture. Sebastes schlegelii, a preeminent economic species, exhibits a sex dimorphism, with females surpassing males in growth. In this regard, achieving all-female black rockfish production could significantly enhance breeding profitability.
[...] Read more.
It is widely known that all-female fish production holds economic value for aquaculture. Sebastes schlegelii, a preeminent economic species, exhibits a sex dimorphism, with females surpassing males in growth. In this regard, achieving all-female black rockfish production could significantly enhance breeding profitability. In this study, we utilized the widely used male sex-regulating hormone, 17α-methyltestosterone (MT) at three different concentrations (20, 40, and 60 ppm), to produce pseudomales of S. schlegelii for subsequent all-female offspring breeding. Long-term MT administration severely inhibits the growth of S. schlegelii, while short term had no significant impact. Histological analysis confirmed sex reversal at all MT concentrations; however, both medium and higher MT concentrations impaired testis development. MT also influenced sex steroid hormone levels in pseudomales, suppressing E2 while increasing T and 11-KT levels. In addition, a transcriptome analysis revealed that MT down-regulated ovarian-related genes (cyp19a1a and foxl2) while up-regulating male-related genes (amh) in pseudomales. Furthermore, MT modulated the TGF-β signaling and steroid hormone biosynthesis pathways, indicating its crucial role in S. schlegelii sex differentiation. Therefore, the current study provides a method for achieving sexual reversal using MT in S. schlegelii and offers an initial insight into the underlying mechanism of sexual reversal in this species.
Full article
(This article belongs to the Special Issue Omic Study and Genes in Fish Sex Determination and Differentiation)
►▼
Show Figures

Figure 1
Open AccessArticle
Molecular Regulation of Fetal Brain Development in Inbred and Congenic Mouse Strains Differing in Longevity
by
Maliha Islam and Susanta K. Behura
Genes 2024, 15(5), 604; https://doi.org/10.3390/genes15050604 - 9 May 2024
Abstract
The objective of this study was to investigate gene regulation of the developing fetal brain from congenic or inbred mice strains that differed in longevity. Gene expression and alternative splice variants were analyzed in a genome-wide manner in the fetal brain of C57BL/6J
[...] Read more.
The objective of this study was to investigate gene regulation of the developing fetal brain from congenic or inbred mice strains that differed in longevity. Gene expression and alternative splice variants were analyzed in a genome-wide manner in the fetal brain of C57BL/6J mice (long-lived) in comparison to B6.Cg-Cav1tm1Mls/J (congenic, short-lived) and AKR/J (inbred, short-lived) mice on day(d) 12, 15, and 17 of gestation. The analysis showed a contrasting gene expression pattern during fetal brain development in these mice. Genes related to brain development, aging, and the regulation of alternative splicing were significantly differentially regulated in the fetal brain of the short-lived compared to long-lived mice during development from d15 and d17. A significantly reduced number of splice variants was observed on d15 compared to d12 or d17 in a strain-dependent manner. An epigenetic clock analysis of d15 fetal brain identified DNA methylations that were significantly associated with single-nucleotide polymorphic sites between AKR/J and C57BL/6J strains. These methylations were associated with genes that show epigenetic changes in an age-correlated manner in mice. Together, the finding of this study suggest that fetal brain development and longevity are epigenetically linked, supporting the emerging concept of the early-life origin of longevity.
Full article
(This article belongs to the Section Molecular Genetics and Genomics)
►▼
Show Figures
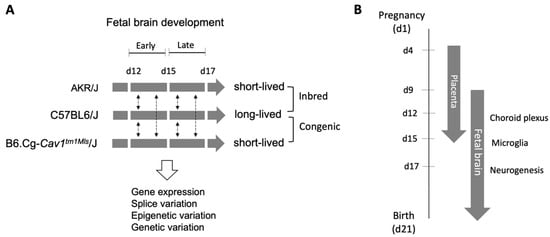
Figure 1
Open AccessArticle
PlantMine: A Machine-Learning Framework to Detect Core SNPs in Rice Genomics
by
Kai Tong, Xiaojing Chen, Shen Yan, Liangli Dai, Yuxue Liao, Zhaoling Li and Ting Wang
Genes 2024, 15(5), 603; https://doi.org/10.3390/genes15050603 - 9 May 2024
Abstract
As a fundamental global staple crop, rice plays a pivotal role in human nutrition and agricultural production systems. However, its complex genetic architecture and extensive trait variability pose challenges for breeders and researchers in optimizing yield and quality. Particularly to expedite breeding methods
[...] Read more.
As a fundamental global staple crop, rice plays a pivotal role in human nutrition and agricultural production systems. However, its complex genetic architecture and extensive trait variability pose challenges for breeders and researchers in optimizing yield and quality. Particularly to expedite breeding methods like genomic selection, isolating core SNPs related to target traits from genome-wide data reduces irrelevant mutation noise, enhancing computational precision and efficiency. Thus, exploring efficient computational approaches to mine core SNPs is of great importance. This study introduces PlantMine, an innovative computational framework that integrates feature selection and machine learning techniques to effectively identify core SNPs critical for the improvement of rice traits. Utilizing the dataset from the 3000 Rice Genomes Project, we applied different algorithms for analysis. The findings underscore the effectiveness of combining feature selection with machine learning in accurately identifying core SNPs, offering a promising avenue to expedite rice breeding efforts and improve crop productivity and resilience to stress.
Full article
(This article belongs to the Special Issue Genomic Studies of Plant Breeding)
►▼
Show Figures
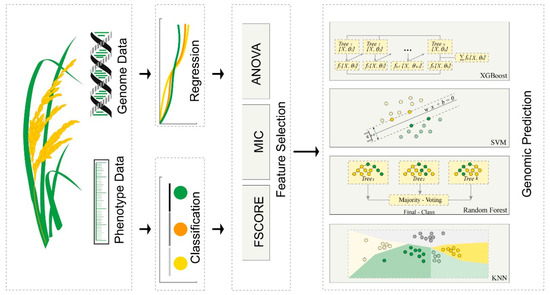
Figure 1
Open AccessArticle
Transcriptional Profiling of Early Defense Response to White Pine Blister Rust Infection in Pinus albicaulis (Whitebark Pine)
by
Laura Figueroa-Corona, Kailey Baesen, Akriti Bhattarai, Angelia Kegley, Richard A. Sniezko, Jill Wegrzyn and Amanda R. De La Torre
Genes 2024, 15(5), 602; https://doi.org/10.3390/genes15050602 - 9 May 2024
Abstract
Pathogen perception generates the activation of signal transduction cascades to host defense. White pine blister rust (WPBR) is caused by Cronartium ribicola J.C. Fisch and affects a number of species of Pinus. One of the most severely affected species is Pinus albicaulis
[...] Read more.
Pathogen perception generates the activation of signal transduction cascades to host defense. White pine blister rust (WPBR) is caused by Cronartium ribicola J.C. Fisch and affects a number of species of Pinus. One of the most severely affected species is Pinus albicaulis Engelm (whitebark pine). WPBR resistance in the species is a polygenic and complex trait that requires an optimized immune response. We identified early responses in 2-year-old seedlings after four days of fungal inoculation and compared the underlying transcriptomic response with that of healthy non-inoculated individuals. A de novo transcriptome assembly was constructed with 56,796 high quality-annotations derived from the needles of susceptible and resistant individuals in a resistant half-sib family. Differential expression analysis identified 599 differentially expressed transcripts, from which 375 were upregulated and 224 were downregulated in the inoculated seedlings. These included components of the initial phase of active responses to abiotic factors and stress regulators, such as those involved in the first steps of flavonoid biosynthesis. Four days after the inoculation, infected individuals showed an overexpression of chitinases, reactive oxygen species (ROS) regulation signaling, and flavonoid intermediates. Our research sheds light on the first stage of infection and emergence of disease symptoms among whitebark pine seedlings. RNA sequencing (RNA-seq) data encoding hypersensitive response, cell wall modification, oxidative regulation signaling, programmed cell death, and plant innate immunity were differentially expressed during the defense response against C. ribicola.
Full article
(This article belongs to the Section Genes & Environments)
►▼
Show Figures

Figure 1
Open AccessArticle
Genetic Screening Revealed the Negative Regulation of miR-310~313 Cluster Members on Imd Pathway during Gram-Negative Bacterial Infection in Drosophila
by
Yao Li, Yixuan Sun, Ruimin Li, Hongjian Zhou, Shengjie Li and Ping Jin
Genes 2024, 15(5), 601; https://doi.org/10.3390/genes15050601 - 8 May 2024
Abstract
Innate immune response is the first line of host defense against pathogenic microorganisms, and its excessive or insufficient activation is detrimental to the organism. Many individual microRNAs (miRNAs) have emerged as crucial post-transcriptional regulators of immune homeostasis in Drosophila melanogaster. However, the
[...] Read more.
Innate immune response is the first line of host defense against pathogenic microorganisms, and its excessive or insufficient activation is detrimental to the organism. Many individual microRNAs (miRNAs) have emerged as crucial post-transcriptional regulators of immune homeostasis in Drosophila melanogaster. However, the synergistical regulation of miRNAs located within a cluster on the Imd-immune pathway remains obscured. In our study, a genetic screening with 52 transgenic UAS-miRNAs was performed to identify ten miRNAs or miRNA clusters, including the miR310~313 cluster, which may function on Imd-dependent immune responses. The miRNA RT-qPCR analysis showed that the expression of miR-310~313 cluster members exhibited an increase at 6–12 h post E. Coli infection. Furthermore, the overexpression of the miR-310~313 cluster impaired the Drosophila survival. And the overexpression of miR-310/311/312 reduced Dpt expression, an indication of Imd pathway induced by Gram-negative bacteria. Conversely, the knockdown of miR-310/311/312 led to increases in Dpt expression. The Luciferase reporter expression assays and RT-qPCR analysis confirmed that miR-310~313 cluster members directly co-targeted and inhibited Imd transcription. These findings reveal that the members of the miR-310~313 cluster synergistically inhibit Imd-dependent immune responses by co-targeting the Imd gene in Drosophila.
Full article
(This article belongs to the Topic MicroRNA: Mechanisms of Action, Physio-Pathological Implications, and Disease Biomarkers, 2nd Volume)
Open AccessReview
Genetic Causes of Qualitative Sperm Defects: A Narrative Review of Clinical Evidence
by
Andrea Graziani, Maria Santa Rocca, Cinzia Vinanzi, Giulia Masi, Giuseppe Grande, Luca De Toni and Alberto Ferlin
Genes 2024, 15(5), 600; https://doi.org/10.3390/genes15050600 - 8 May 2024
Abstract
Several genes are implicated in spermatogenesis and fertility regulation, and these genes are presently being analysed in clinical practice due to their involvement in male factor infertility (MFI). However, there are still few genetic analyses that are currently recommended for use in clinical
[...] Read more.
Several genes are implicated in spermatogenesis and fertility regulation, and these genes are presently being analysed in clinical practice due to their involvement in male factor infertility (MFI). However, there are still few genetic analyses that are currently recommended for use in clinical practice. In this manuscript, we reviewed the genetic causes of qualitative sperm defects. We distinguished between alterations causing reduced sperm motility (asthenozoospermia) and alterations causing changes in the typical morphology of sperm (teratozoospermia). In detail, the genetic causes of reduced sperm motility may be found in the alteration of genes associated with sperm mitochondrial DNA, mitochondrial proteins, ion transport and channels, and flagellar proteins. On the other hand, the genetic causes of changes in typical sperm morphology are related to conditions with a strong genetic basis, such as macrozoospermia, globozoospermia, and acephalic spermatozoa syndrome. We tried to distinguish alterations approved for routine clinical application from those still unsupported by adequate clinical studies. The most important aspect of the study was related to the correct identification of subjects to be tested and the correct application of genetic tests based on clear clinical data. The correct application of available genetic tests in a scenario where reduced sperm motility and changes in sperm morphology have been observed enables the delivery of a defined diagnosis and plays an important role in clinical decision-making. Finally, clarifying the genetic causes of MFI might, in future, contribute to reducing the proportion of so-called idiopathic MFI, which might indeed be defined as a subtype of MFI whose cause has not yet been revealed.
Full article
(This article belongs to the Special Issue Beyond the Basics: Genetic Insights into Male Infertility)
►▼
Show Figures
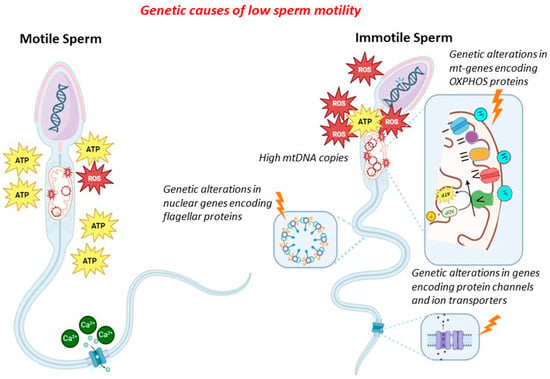
Figure 1
Open AccessArticle
Study on the Characteristics of Coarse Feeding Tolerance of Ding’an Pigs: Phenotypic and Candidate Genes Identification
by
Yanxia Song, Mingming Xue, Feng Wang, Qiguo Tang, Yabiao Luo, Meili Zheng, Yubei Wang, Pengxiang Xue, Ningqi Dong, Ruiping Sun and Meiying Fang
Genes 2024, 15(5), 599; https://doi.org/10.3390/genes15050599 - 8 May 2024
Abstract
Ding’an (DA) pig, a prominent local breed in Hainan Province, exhibits notable advantages in coarse feeding tolerance and high-quality meat. To explore the potential genetic mechanism of coarse feeding tolerance in DA pigs, 60-day-old full sibling pairs of DA and DLY (Duroc-Landrace-Yorkshire) pigs
[...] Read more.
Ding’an (DA) pig, a prominent local breed in Hainan Province, exhibits notable advantages in coarse feeding tolerance and high-quality meat. To explore the potential genetic mechanism of coarse feeding tolerance in DA pigs, 60-day-old full sibling pairs of DA and DLY (Duroc-Landrace-Yorkshire) pigs were subjected to fed normal (5%) and high (10%) crude fiber diets for 56 days, respectively. The findings showed that increasing the crude fiber level had no impact on the apparent digestibility of crude fiber, intramuscular fat, and marbling scores in DA pigs, whereas these factors were significantly reduced in DLY pigs (p < 0.05). Through differential expression analysis and Weighted Gene Co-expression Network Analysis (WGCNA) of the colonic mucosal transcriptome data, 65 and 482 candidate genes with coarse feeding tolerance in DA pigs were identified, respectively. Joint analysis screened four key candidate genes, including LDHB, MLC1, LSG1, and ESM1, potentially serving as key regulated genes for coarse feeding tolerance. Functional analysis revealed that the most significant pathway enriched in differential genes associated with coarse feeding tolerance in Ding’an pigs was the signaling receptor binding. The results hold substantial significance for advancing our understanding of the genetic mechanisms governing coarse feeding tolerance in Ding’an pigs.
Full article
(This article belongs to the Section Animal Genetics and Genomics)
►▼
Show Figures

Figure 1
Open AccessArticle
Expression and Characterization of an Efficient Alginate Lyase from Psychromonas sp. SP041 through Metagenomics Analysis of Rotten Kelp
by
Ping Wang, Yi Cai, Hua Zhong, Ruiting Chen, Yuetao Yi, Yanrui Ye and Lili Li
Genes 2024, 15(5), 598; https://doi.org/10.3390/genes15050598 - 8 May 2024
Abstract
Alginate is derived from brown algae, which can be cultivated in large quantities. It can be broken down by alginate lyase into alginate oligosaccharides (AOSs), which exhibit a higher added value and better bioactivity than alginate. In this study, metagenomic technology was used
[...] Read more.
Alginate is derived from brown algae, which can be cultivated in large quantities. It can be broken down by alginate lyase into alginate oligosaccharides (AOSs), which exhibit a higher added value and better bioactivity than alginate. In this study, metagenomic technology was used to screen for genes that code for high-efficiency alginate lyases. The candidate alginate lyase gene alg169 was detected from Psychromonas sp. SP041, the most abundant species among alginate lyase bacteria on selected rotten kelps. The alginate lyase Alg169 was heterologously expressed in Escherichia coli BL21 (DE3), Ni-IDA-purified, and characterized. The optimum temperature and pH of Alg169 were 25 °C and 7.0, respectively. Metal ions including Mn2+, Co2+, Ca2+, Mg2+, Ni2+, and Ba2+ led to significantly increased enzyme activity. Alg169 exhibited a pronounced dependence on Na+, and upon treatment with Mn2+, its activity surged by 687.57%, resulting in the highest observed enzyme activity of 117,081 U/mg. Bioinformatic analysis predicted that Alg169 would be a double-domain lyase with a molecular weight of 65.58 kDa. It is a bifunctional enzyme with substrate specificity to polyguluronic acid (polyG) and polymannuronic acid (polyM). These results suggest that Alg169 is a promising candidate for the efficient manufacturing of AOSs from brown seaweed.
Full article
(This article belongs to the Special Issue Microbial Genome Engineering for Production of Natural Products and Biopolymers)
►▼
Show Figures

Figure 1
Open AccessCase Report
A Novel COL4A5 Pathogenic Variant Joins the Dots in a Family with a Synchronous Diagnosis of Alport Syndrome and Polycystic Kidney Disease
by
Ludovico Graziani, Chiara Minotti, Miriam Lucia Carriero, Mario Bengala, Silvia Lai, Alessandra Terracciano, Antonio Novelli and Giuseppe Novelli
Genes 2024, 15(5), 597; https://doi.org/10.3390/genes15050597 - 8 May 2024
Abstract
Alport Syndrome (AS) is the most common genetic glomerular disease, and it is caused by COL4A3, COL4A4, and COL4A5 pathogenic variants. The classic phenotypic spectrum associated with AS ranges from isolated hematuria to chronic kidney disease (CKD) with extrarenal abnormalities. Atypical
[...] Read more.
Alport Syndrome (AS) is the most common genetic glomerular disease, and it is caused by COL4A3, COL4A4, and COL4A5 pathogenic variants. The classic phenotypic spectrum associated with AS ranges from isolated hematuria to chronic kidney disease (CKD) with extrarenal abnormalities. Atypical presentation of the disorder is possible, and it can mislead the diagnosis. Polycystic kidney disease (PKD), which is most frequently associated with Autosomal Dominant PKD (ADPKD) due to PKD1 and PKD2 heterozygous variants, is emerging as a possible clinical manifestation in COL4A3-A5 patients. We describe a COL4A5 novel familial frameshift variant (NM_000495.5: c.1095dup p.(Leu366ValfsTer45)), which was associated with AS and PKD in the hemizygous proband, as well as with PKD, IgA glomerulonephritis and focal segmental glomerulosclerosis (FSGS) in the heterozygous mother. Establishing the diagnosis of AS can sometimes be difficult, especially in the context of misleading family history and atypical phenotypic features. This case study supports the emerging genotypic and phenotypic heterogeneity in COL4A3-A5-associated disorders, as well as the recently described association between PKD and collagen type IV (Col4) defects. We highlight the importance of the accurate phenotyping of all family members and the relevance of next-generation sequencing in the differential diagnosis of hereditary kidney disease.
Full article
(This article belongs to the Special Issue Genetics and Genomics of Rare Disorders Volume II)
►▼
Show Figures

Figure 1
Open AccessArticle
Interleukin-1β Polymorphisms Are Genetic Markers of Susceptibility to Periprosthetic Joint Infection in Total Hip and Knee Arthroplasty
by
Valentina Granata, Dario Strina, Valentina Possetti, Roberto Leone, Sonia Valentino, Katia Chiappetta, Mattia Loppini, Alberto Mantovani, Barbara Bottazzi, Rosanna Asselta, Cristina Sobacchi and Antonio Inforzato
Genes 2024, 15(5), 596; https://doi.org/10.3390/genes15050596 - 8 May 2024
Abstract
Periprosthetic joint infections (PJIs) are serious complications of prosthetic surgery. The criteria for the diagnosis of PJI integrate clinical and laboratory findings in a complex and sometimes inconclusive workflow. Host immune factors hold potential as diagnostic biomarkers in bone and joint infections. We
[...] Read more.
Periprosthetic joint infections (PJIs) are serious complications of prosthetic surgery. The criteria for the diagnosis of PJI integrate clinical and laboratory findings in a complex and sometimes inconclusive workflow. Host immune factors hold potential as diagnostic biomarkers in bone and joint infections. We reported that the humoral pattern-recognition molecule long pentraxin 3 (PTX3) predicts PJI in total hip and knee arthroplasty (THA and TKA, respectively). If and how genetic variation in PTX3 and inflammatory genes that affect its expression (IL-1β, IL-6, IL-10, and IL-17A) contributes to the risk of PJI is unknown. We conducted a case–control study on a Caucasian historic cohort of THA and TKA patients who had prosthesis explant due to PJI (cases) or aseptic complications (controls). Saliva was collected from 93 subjects and used to extract DNA and genotype PTX3, IL-1β, IL-6, IL-10, and IL-17A single-nucleotide polymorphisms (SNPs). Moreover, the concentration of IL-1β, IL-10, and IL-6 was measured in synovial fluid and plasma. No association was found between PTX3 polymorphisms and PJI; however, the AGG haplotype, encompassing rs2853550, rs1143634, and rs1143627 in IL-1β, was linked to the infection (p = 0.017). Also, synovial levels of all inflammatory markers were higher in cases than in controls, and a correlation emerged between synovial concentration of PTX3 and that of IL-1β in cases only (Spearman r = 0.67, p = 0.004). We identified a relationship between rs2853550 and the synovial concentration of IL-1β and PTX3. Our findings suggest that IL-1β SNPs could be used for the early identification of THA and TKA patients with a high risk of infection.
Full article
(This article belongs to the Special Issue Updates of DNA Variations in Evolution and Human Diseases)
►▼
Show Figures

Figure 1
Open AccessArticle
Neuropsychological Profile of 25 Brazilian Patients with 22q11.2 Deletion Syndrome: Effects of Clinical and Socioeconomic Variables
by
Larissa Salustiano Evangelista Pimenta, Claudia Berlim de Mello, Luciana Mello Di Benedetto, Diogo Cordeiro de Queiroz Soares, Leslie Domenici Kulikowski, Anelisa Gollo Dantas, Maria Isabel Melaragno and Chong Ae Kim
Genes 2024, 15(5), 595; https://doi.org/10.3390/genes15050595 - 8 May 2024
Abstract
The 22q11.2 deletion syndrome (22q11.2DS) is associated with a heterogeneous neurocognitive phenotype, which includes psychiatric disorders. However, few studies have investigated the influence of socioeconomic variables on intellectual variability. The aim of this study was to investigate the cognitive profile of 25 patients,
[...] Read more.
The 22q11.2 deletion syndrome (22q11.2DS) is associated with a heterogeneous neurocognitive phenotype, which includes psychiatric disorders. However, few studies have investigated the influence of socioeconomic variables on intellectual variability. The aim of this study was to investigate the cognitive profile of 25 patients, aged 7 to 32 years, with a typical ≈3 Mb 22q11.2 deletion, considering intellectual, adaptive, and neuropsychological functioning. Univariate linear regression analysis explored the influence of socioeconomic variables on intellectual quotient (IQ) and global adaptive behavior. Associations with relevant clinical conditions such as seizures, recurrent infections, and heart diseases were also considered. Results showed IQ scores ranging from 42 to 104. Communication, executive functions, attention, and visuoconstructive skills were the most impaired in the sample. The study found effects of access to quality education, family socioeconomic status (SES), and caregiver education level on IQ. Conversely, age at diagnosis and language delay were associated with outcomes in adaptive behavior. This characterization may be useful for better understanding the influence of social-environmental factors on the development of patients with 22q11.2 deletion syndrome, as well as for intervention processes aimed at improving their quality of life.
Full article
(This article belongs to the Special Issue Phenotype and Pathogenetic Mechanisms in 22q11.2 Deletion/DiGeorge Syndrome)
Open AccessArticle
Normalized Clinical Severity Scores Reveal a Correlation between X Chromosome Inactivation and Disease Severity in Rett Syndrome
by
Jonathan K. Merritt, Xiaolan Fang, Raymond C. Caylor, Steven A. Skinner, Michael J. Friez, Alan K. Percy and Jeffrey L. Neul
Genes 2024, 15(5), 594; https://doi.org/10.3390/genes15050594 - 8 May 2024
Abstract
Rett Syndrome (RTT) is a severe neurodevelopmental disorder predominately diagnosed in females and primarily caused by pathogenic variants in the X-linked gene Methyl-CpG Binding Protein 2 (MECP2). Most often, the disease causing the MECP2 allele resides on the paternal X chromosome
[...] Read more.
Rett Syndrome (RTT) is a severe neurodevelopmental disorder predominately diagnosed in females and primarily caused by pathogenic variants in the X-linked gene Methyl-CpG Binding Protein 2 (MECP2). Most often, the disease causing the MECP2 allele resides on the paternal X chromosome while a healthy copy is maintained on the maternal X chromosome with inactivation (XCI), resulting in mosaic expression of one allele in each cell. Preferential inactivation of the paternal X chromosome is theorized to result in reduced disease severity; however, establishing such a correlation is complicated by known MECP2 genotype effects and an age-dependent increase in severity. To mitigate these confounding factors, we developed an age- and genotype-normalized measure of RTT severity by modeling longitudinal data collected in the US Rett Syndrome Natural History Study. This model accurately reflected individual increase in severity with age and preserved group-level genotype specific differences in severity, allowing for the creation of a normalized clinical severity score. Applying this normalized score to a RTT XCI dataset revealed that XCI influence on disease severity depends on MECP2 genotype with a correlation between XCI and severity observed only in individuals with MECP2 variants associated with increased clinical severity. This normalized measure of RTT severity provides the opportunity for future discovery of additional factors contributing to disease severity that may be masked by age and genotype effects.
Full article
(This article belongs to the Section Human Genomics and Genetic Diseases)
►▼
Show Figures

Figure 1
Open AccessArticle
Genotype–Phenotype Correlations in Alport Syndrome—A Single-Center Experience
by
Ștefan Nicolaie Lujinschi, Bogdan Marian Sorohan, Bogdan Obrișcă, Alexandra Vrabie, Gabriela Lupușoru, Camelia Achim, Andreea Gabriella Andronesi, Andreea Covic and Gener Ismail
Genes 2024, 15(5), 593; https://doi.org/10.3390/genes15050593 - 7 May 2024
Abstract
Background: Alport syndrome (AS) is a common and heterogeneous genetic kidney disease, that often leads to end-stage kidney disease (ESKD). Methods: This is a single-center, retrospective study that included 36 adults with type IV collagen (COL4) mutations. Our main scope was to describe
[...] Read more.
Background: Alport syndrome (AS) is a common and heterogeneous genetic kidney disease, that often leads to end-stage kidney disease (ESKD). Methods: This is a single-center, retrospective study that included 36 adults with type IV collagen (COL4) mutations. Our main scope was to describe how genetic features influence renal survival. Results: A total of 24 different mutations were identified, of which eight had not been previously described. Mutations affecting each of the type IV collagen α chains were equally prevalent (33.3%). Most of the patients had pathogenic variants (61.1%). Most patients had a family history of kidney disease (71%). The most prevalent clinical picture was nephritic syndrome (64%). One-third of the subjects had extrarenal manifestations, 41.6% of patients had ESKD at referral, and another 8.3% developed ESKD during follow-up. The median renal survival was 42 years (95% CI, 29.98–54.01). The COL4A4 group displayed better renal survival than the COL4A3 group (p = 0.027). Patients with missense variants had higher renal survival (p = 0.023). Hearing loss was associated with lower renal survival (p < 0.001). Conclusions: Patients with COL4A4 variants and those with missense mutations had significantly better renal survival, whereas those with COL4A3 variants and those with hearing loss had worse prognoses.
Full article
(This article belongs to the Special Issue Genetic and Phenotypic Correlation: Gene–Disease Validation Series II)
►▼
Show Figures

Figure 1
Open AccessArticle
The Association between Mutational Signatures and Clinical Outcomes among Patients with Early-Onset Breast Cancer
by
Robert B. Basmadjian, Dylan E. O’Sullivan, May Lynn Quan, Sasha Lupichuk, Yuan Xu, Winson Y. Cheung and Darren R. Brenner
Genes 2024, 15(5), 592; https://doi.org/10.3390/genes15050592 - 7 May 2024
Abstract
Early-onset breast cancer (EoBC), defined by a diagnosis <40 years of age, is associated with poor prognosis. This study investigated the mutational landscape of non-metastatic EoBC and the prognostic relevance of mutational signatures using 100 tumour samples from Alberta, Canada. The MutationalPatterns package
[...] Read more.
Early-onset breast cancer (EoBC), defined by a diagnosis <40 years of age, is associated with poor prognosis. This study investigated the mutational landscape of non-metastatic EoBC and the prognostic relevance of mutational signatures using 100 tumour samples from Alberta, Canada. The MutationalPatterns package in R/Bioconductor was used to extract de novo single-base substitution (SBS) and insertion–deletion (indel) mutational signatures and to fit COSMIC SBS and indel signatures. We assessed associations between these signatures and clinical characteristics of disease, in addition to recurrence-free (RFS) and overall survival (OS). Five SBS and two indel signatures were extracted. The SBS13-like signature had higher relative contributions in the HER2-enriched subtype. Patients with higher than median contribution tended to have better RFS after adjustment for other prognostic factors (HR = 0.29; 95% CI: 0.08–1.06). An unsupervised clustering algorithm based on absolute contribution revealed three clusters of fitted COSMIC SBS signatures, but cluster membership was not associated with clinical variables or survival outcomes. The results of this exploratory study reveal various SBS and indel signatures may be associated with clinical features of disease and prognosis. Future studies with larger samples are required to better understand the mechanistic underpinnings of disease progression and treatment response in EoBC.
Full article
(This article belongs to the Special Issue Bioinformatics and Computational Biology for Cancer Prediction and Prognosis)
►▼
Show Figures

Figure 1
Open AccessReview
Genetic Variants in the ABCB1 and ABCG2 Gene Drug Transporters Involved in Gefitinib-Associated Adverse Reaction: A Systematic Review and Meta-Analysis
by
Mariana Vieira Morau, Cecília Souto Seguin, Marília Berlofa Visacri, Eder de Carvalho Pincinato and Patricia Moriel
Genes 2024, 15(5), 591; https://doi.org/10.3390/genes15050591 (registering DOI) - 7 May 2024
Abstract
This systematic review and meta-analysis aimed to verify the association between the genetic variants of adenosine triphosphate (ATP)-binding cassette subfamily B member 1 (ABCB1) and ATP-binding cassette subfamily G member 2 (ABCG2) genes and the presence and severity of
[...] Read more.
This systematic review and meta-analysis aimed to verify the association between the genetic variants of adenosine triphosphate (ATP)-binding cassette subfamily B member 1 (ABCB1) and ATP-binding cassette subfamily G member 2 (ABCG2) genes and the presence and severity of gefitinib-associated adverse reactions. We systematically searched PubMed, Virtual Health Library/Bireme, Scopus, Embase, and Web of Science databases for relevant studies published up to February 2024. In total, five studies were included in the review. Additionally, eight genetic variants related to ABCB1 (rs1045642, rs1128503, rs2032582, and rs1025836) and ABCG2 (rs2231142, rs2231137, rs2622604, and 15622C>T) genes were analyzed. Meta-analysis showed a significant association between the ABCB1 gene rs1045642 TT genotype and presence of diarrhea (OR = 5.41, 95% CI: 1.38–21.14, I2 = 0%), the ABCB1 gene rs1128503 TT genotype and CT + TT group and the presence of skin rash (OR = 4.37, 95% CI: 1.51–12.61, I2 = 0% and OR = 6.99, 95%CI: 1.61–30.30, I2= 0%, respectively), and the ABCG2 gene rs2231142 CC genotype and presence of diarrhea (OR = 3.87, 95% CI: 1.53–9.84, I2 = 39%). No ABCB1 or ABCG2 genes were positively associated with the severity of adverse reactions associated with gefitinib. In conclusion, this study showed that ABCB1 and ABCG2 variants are likely to exhibit clinical implications in predicting the presence of adverse reactions to gefitinib.
Full article
(This article belongs to the Section Human Genomics and Genetic Diseases)
►▼
Show Figures

Figure 1
Open AccessArticle
Genetic Analysis of the ts-Lethal Mutant Δpa0665/pTS-pa0665 Reveals Its Role in Cell Morphology and Oxidative Phosphorylation in Pseudomonas aeruginosa
by
Jiayin Zhu, Hulin Zhao and Zhili Yang
Genes 2024, 15(5), 590; https://doi.org/10.3390/genes15050590 - 7 May 2024
Abstract
Pa0665 in Pseudomonas aeruginosa shares homologous sequences with that of the essential A-type iron–sulfur (Fe-S) cluster insertion protein ErpA in Escherichia coli. However, its essentiality in P. aeruginosa and its complementation with E. coli erpA has not been experimentally examined. To fulfill this
[...] Read more.
Pa0665 in Pseudomonas aeruginosa shares homologous sequences with that of the essential A-type iron–sulfur (Fe-S) cluster insertion protein ErpA in Escherichia coli. However, its essentiality in P. aeruginosa and its complementation with E. coli erpA has not been experimentally examined. To fulfill this task, we constructed plasmid-based ts-mutant Δpa0665/pTS-pa0665 using a three-step protocol. The mutant displayed growth defects at 42 °C, which were complemented by expressing ec.erpA. Microscopic observations indicated a petite cell phenotype for Δpa0665/pTS-pa0665 at 42 °C, correlated with the downregulation of the oprG gene. RNA sequencing revealed significant transcriptional changes in genes associated with the oxidative phosphorylation (OXPHOS) system, aligning with reduced ATP levels in Δpa0665/pTS-pa0665 under 42 °C. Additionally, the ts-mutant showed heightened sensitivity to H2O2 at 42 °C. Overall, our study demonstrates the essential role of pa0665 for OXPHOS function and is complemented by ec.erpA. We propose that the plasmid-based ts-allele is useful for genetic analysis of essential genes of interest in P. aeruginosa.
Full article
(This article belongs to the Special Issue Genomics and Bioinformatics in Microbial Science)
►▼
Show Figures

Figure 1

Journal Menu
► ▼ Journal Menu-
- Genes Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Biology, BioMed, Cells, Genes
Applications of the Zebrafish Model
Topic Editors: De-Li Shi, Pengfei XuDeadline: 30 June 2024
Topic in
Biomedicines, Cells, CIMB, Genes, IJMS
Animal Models of Human Disease 2.0
Topic Editors: Sigrun Lange, Jameel M. InalDeadline: 31 August 2024
Topic in
Agriculture, Bioengineering, Genes, IJMS, Plants
Genetic Engineering in Agriculture
Topic Editors: Amy L. Klocko, Jianjun Chen, Haiwei LuDeadline: 30 September 2024
Topic in
Biology, BioMedInformatics, Cancers, Genes, IJMS
The 22nd International Conference on Bioinformatics (InCoB 2023): Translational Bioinformatics Transforming Life
Topic Editors: Jyotsna Batra, Srilakshmi Srinivasan, Shoba Ranganathan, Asif M. Khan, Harpreet SinghDeadline: 15 November 2024

Conferences
Special Issues
Special Issue in
Genes
Cotton Genes, Genetics, and Genomics
Guest Editor: Guanjing HuDeadline: 10 May 2024
Special Issue in
Genes
DNA Damage and Repair in Microorganisms, Plants and Mammalian Systems
Guest Editors: Ioly Kotta-Loizou, Nan Zhang, Xin WangDeadline: 20 May 2024
Special Issue in
Genes
Commemorating the Launch of the Section "Cytogenomics"
Guest Editor: Darren GriffinDeadline: 5 June 2024
Special Issue in
Genes
Genetic Markers and Liquid Biopsy for Kidney Diseases
Guest Editors: Guorong Li, Christophe MariatDeadline: 15 June 2024
Topical Collections
Topical Collection in
Genes
Study on Genotypes and Phenotypes of Pediatric Clinical Rare Diseases
Collection Editors: Livia Garavelli, Stefano Giuseppe Caraffi
Topical Collection in
Genes
Eukaryotic Non-coding RNAs: Diversity, Structure/Function, Implication in Cardiovascular Disease
Collection Editors: Morten Andre Høydal, Christiane Branlant
Topical Collection in
Genes
Feature Papers in ‘Animal Genetics and Genomics’
Collection Editors: Antonio Figueras, Raquel Vasconcelos