Journal Description
Microorganisms
Microorganisms
is a scientific, peer-reviewed, open access journal of microbiology, published monthly online by MDPI. The Hellenic Society Mikrobiokosmos (MBK), the Spanish Society for Nitrogen Fixation (SEFIN) and the Society for Microbial Ecology and Disease (SOMED) are affiliated with the Microorganisms, and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, PMC, PubAg, CAPlus / SciFinder, AGRIS, and other databases.
- Journal Rank: JCR - Q2 (Microbiology) / CiteScore - Q2 (Microbiology (medical))
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 15.1 days after submission; acceptance to publication is undertaken in 2.8 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Testimonials: See what our editors and authors say about the Microorganisms.
- Companion journal: Applied Microbiology.
Impact Factor:
4.5 (2022);
5-Year Impact Factor:
4.8 (2022)
Latest Articles
Bioaerosol Sampling Devices and Pretreatment for Bacterial Characterization: Theoretical Differences and a Field Experience in a Wastewater Treatment Plant
Microorganisms 2024, 12(5), 965; https://doi.org/10.3390/microorganisms12050965 (registering DOI) - 10 May 2024
Abstract
Studies on bioaerosol bacterial biodiversity have relevance in both ecological and health contexts, and molecular methods, such as 16S rRNA gene-based barcoded sequencing, provide efficient tools for the analysis of airborne bacterial communities. Standardized methods for sampling and analysis of bioaerosol DNA are
[...] Read more.
Studies on bioaerosol bacterial biodiversity have relevance in both ecological and health contexts, and molecular methods, such as 16S rRNA gene-based barcoded sequencing, provide efficient tools for the analysis of airborne bacterial communities. Standardized methods for sampling and analysis of bioaerosol DNA are lacking, thus hampering the comparison of results from studies implementing different devices and procedures. Three samplers that use gelatin filtration, swirling aerosol collection, and condensation growth tubes for collecting bioaerosol at an aeration tank of a wastewater treatment plant in Trieste (Italy) were used to determine the bacterial biodiversity. Wastewater samples were collected directly from the untreated sewage to obtain a true representation of the microbiological community present in the plant. Different samplers and collection media provide an indication of the different grades of biodiversity, with condensation growth tubes and DNA/RNA shieldTM capturing the richer bacterial genera. Overall, in terms of relative abundance, the air samples have a lower number of bacterial genera (64 OTUs) than the wastewater ones (75 OTUs). Using the metabarcoding approach to aerosol samples, we provide the first preliminary step toward the understanding of a significant diversity between different air sampling systems, enabling the scientific community to orient research towards the most informative sampling strategy.
Full article
(This article belongs to the Special Issue Advances in Bioaerosols)
Open AccessArticle
Clinical Evaluation of VITEK MS PRIME with PICKME Pen for Bacteria and Yeasts, and RUO Database for Filamentous Fungi
by
Hyeyoung Lee, Jehyun Koo, Junsang Oh, Sung-Il Cho, Hyunjoo Lee, Hyun Ji Lee, Gi-Ho Sung and Jayoung Kim
Microorganisms 2024, 12(5), 964; https://doi.org/10.3390/microorganisms12050964 (registering DOI) - 10 May 2024
Abstract
The VITEK MS PRIME (bioMérieux, Marcy-l’Étoile, France), a newly developed matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) system, alongside the VITEK PICKME pen (PICKME), offers easy sample preparation for bacteria and yeasts. The VITEK MS PRIME also offers two software platforms
[...] Read more.
The VITEK MS PRIME (bioMérieux, Marcy-l’Étoile, France), a newly developed matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) system, alongside the VITEK PICKME pen (PICKME), offers easy sample preparation for bacteria and yeasts. The VITEK MS PRIME also offers two software platforms for filamentous fungi: the IVD database and the RUO database. Our study evaluated its identification agreement on 320 clinical isolates of bacteria and yeasts, comparing PICKME and traditional wooden toothpick sampling techniques against MicroIDSys Elite (ASTA) results. Additionally, we assessed the IVD (v3.2) and SARAMIS (v4.16) RUO databases on 289 filamentous fungi against molecular sequencing. The concordance rates for species-level identification of bacteria and yeasts were about 89.4% (286/320) between the PICKME and wooden toothpick, and about 83.4–85.3% between the VITEK MS PRIME and ASTA MicroIDSys Elite. Retesting with PICKME improved concordance to 91.9%. For filamentous fungi, species-level identification reached 71.3% with the IVD database and 85.8% with RUO, which significantly enhanced basidiomycetes’ identification from 35.3% to 100%. Some strains in the IVD database, like Aspergillus versicolor, Exophiala xenobiotica, and Nannizzia gypsea, failed to be identified. The VITEK MS PRIME with PICKME offers reliable and efficient microorganism identification. For filamentous fungi, combined use of the RUO database can be beneficial, especially for basidiomycetes.
Full article
(This article belongs to the Section Medical Microbiology)
►▼
Show Figures

Figure 1
Open AccessArticle
Diversity and Composition of Soil Acidobacterial Communities in Different Temperate Forest Types of Northeast China
by
Feng Jiao, Lili Qian, Jinhua Wu, Dongdong Zhang, Junying Zhang, Mingyu Wang, Xin Sui and Xianbang Zhang
Microorganisms 2024, 12(5), 963; https://doi.org/10.3390/microorganisms12050963 (registering DOI) - 10 May 2024
Abstract
To gain an in-depth understanding of the diversity and composition of soil Acidobacteria in five different forest types in typical temperate forest ecosystems and to explore their relationship with soil nutrients. The diversity of soil Acidobacteria was determined by high-throughput sequencing technology. Soil
[...] Read more.
To gain an in-depth understanding of the diversity and composition of soil Acidobacteria in five different forest types in typical temperate forest ecosystems and to explore their relationship with soil nutrients. The diversity of soil Acidobacteria was determined by high-throughput sequencing technology. Soil Acidobacteria’s alpha-diversity index and soil nutrient content differed significantly among different forest types. β-diversity and the composition of soil Acidobacteria also varied across forest types. Acidobacterial genera, such as Acidobacteria_Gp1, Acidobacteria_Gp4, and Acidobacteria_Gp17, play key roles in different forests. The RDA analyses pointed out that the soil pH, available nitrogen (AN), carbon to nitrogen (C/N) ratio, available phosphorus (AP), total carbon (TC), and total phosphorus (TP) were significant factors affecting soil Acidobacteria in different forest types. In this study, the diversity and composition of soil Acidobacteria under different forest types in a temperate forest ecosystem were analyzed, revealing the complex relationship between them and soil physicochemical properties. These findings not only enhance our understanding of soil microbial ecology but also provide important guidance for ecological conservation and restoration strategies for temperate forest ecosystems.
Full article
(This article belongs to the Section Environmental Microbiology)
►▼
Show Figures

Figure 1
Open AccessArticle
Novel Tick-Borne Anaplasmataceae Genotypes in Tropical Birds from the Brazilian Pantanal Wetland
by
Amir Salvador Alabí Córdova, Alan Fecchio, Ana Cláudia Calchi, Clara Morato Dias, Anna Claudia Baumel Mongruel, Lorena Freitas das Neves, Daniel Antonio Braga Lee, Rosangela Zacarias Machado and Marcos Rogério André
Microorganisms 2024, 12(5), 962; https://doi.org/10.3390/microorganisms12050962 (registering DOI) - 10 May 2024
Abstract
Despite numerous reports of Anaplasmataceae agents in mammals worldwide, few studies have investigated their occurrence in birds. The present study aimed to investigate the occurrence and molecular identity of Anaplasmataceae agents in birds from the Pantanal wetland, Brazil. Blood samples were collected from
[...] Read more.
Despite numerous reports of Anaplasmataceae agents in mammals worldwide, few studies have investigated their occurrence in birds. The present study aimed to investigate the occurrence and molecular identity of Anaplasmataceae agents in birds from the Pantanal wetland, Brazil. Blood samples were collected from 93 different species. After DNA extraction, samples positive for the avian β-actin gene were subjected to both a multiplex quantitative real-time (q)PCR for Anaplasma and Ehrlichia targeting the groEL gene and to a conventional PCR for Anaplasmataceae agents targeting the 16S rRNA gene. As a result, 37 (7.4%) birds were positive for Anaplasma spp. and 4 (0.8%) for Ehrlichia spp. in the qPCR assay; additionally, 13 (2.6%) were positive for Anaplasmataceae agents in the PCR targeting the 16S rRNA gene. The Ehrlichia 16S rRNA sequences detected in Arundinicola leucocephala, Ramphocelus carbo, and Elaenia albiceps were positioned closely to Ehrlichia sp. Magellanica. Ehrlichia dsb sequences detected in Agelasticus cyanopus and Basileuterus flaveolus grouped with Ehrlichia minasensis. The 16S rRNA genotypes detected in Crax fasciolata, Pitangus sulphuratus and Furnarius leucopus grouped with Candidatus Allocryptoplasma. The 23S-5S genotypes detected in C. fasciolata, Basileuterus flaveolus, and Saltator coerulescens were related to Anaplasma phagocytophilum. In conclusion, novel genotypes of Anaplasma, Ehrlichia, and Candidatus Allocryptoplasma were detected in birds from the Pantanal wetland.
Full article
(This article belongs to the Section Parasitology)
►▼
Show Figures

Figure 1
Open AccessCommunication
Validation of a Loop-Mediated Isothermal Amplification-Based Kit for the Detection of Legionella pneumophila in Environmental Samples According to ISO/TS 12869:2012
by
Giorgia Caruso, Maria Anna Coniglio, Pasqualina Laganà, Teresa Fasciana, Giuseppe Arcoleo, Ignazio Arrigo, Paola Di Carlo, Mario Palermo and Anna Giammanco
Microorganisms 2024, 12(5), 961; https://doi.org/10.3390/microorganisms12050961 - 10 May 2024
Abstract
Legionella pneumophila is a freshwater opportunistic pathogen and the leading cause of severe pneumonia known as Legionnaires’ disease. It can be found in all water systems and survives in biofilms, free-living amoebae, and a wide variety of facilities, such as air conditioning and
[...] Read more.
Legionella pneumophila is a freshwater opportunistic pathogen and the leading cause of severe pneumonia known as Legionnaires’ disease. It can be found in all water systems and survives in biofilms, free-living amoebae, and a wide variety of facilities, such as air conditioning and showers in hospitals, hotels and spas. The reference cultural method allows for the isolation and identification in many days, and in addition, it does not detect viable but rather non-culturable bacteria, increasing the risk of infection. In this context, a new LAMP-based (loop-mediated isothermal amplification) kit was developed, allowing for the rapid, sensitive, and labor-saving detection of L. pneumophila. The kit, “Legionella pneumophila Glow”, was validated according to ISO/TS 12869:2012, testing sensitivity, inclusivity and exclusivity, and kit robustness. Sensitivity showed that the “Legionella pneumophila Glow” kit can detect up to 28 plasmid copies/µL. Robustness tests showed consistent results, with both contamination levels and the matrices used giving reproducible results. Furthermore, real samples were evaluated to compare the performance of the two methods. The LAMP kit “Legionella pneumophila Glow” proved a useful option for the rapid, efficient, and labor-saving screening of different typologies of water samples, offering significant advantages over the traditional method, as it is characterized by a high sensitivity, ease of use for laboratory testing, and a large reduction in analysis time, making it an asset to official controls.
Full article
(This article belongs to the Special Issue Clinical and Environmental Surveillance for the Prevention of Legionellosis 2.0)
Open AccessArticle
Genome-Wide Computational Prediction and Analysis of Noncoding RNAs in Oleidesulfovibrio alaskensis G20
by
Ram Nageena Singh and Rajesh K. Sani
Microorganisms 2024, 12(5), 960; https://doi.org/10.3390/microorganisms12050960 - 10 May 2024
Abstract
Noncoding RNAs (ncRNAs) play key roles in the regulation of important pathways, including cellular growth, stress management, signaling, and biofilm formation. Sulfate-reducing bacteria (SRB) contribute to huge economic losses causing microbial-induced corrosion through biofilms on metal surfaces. To effectively combat the challenges posed
[...] Read more.
Noncoding RNAs (ncRNAs) play key roles in the regulation of important pathways, including cellular growth, stress management, signaling, and biofilm formation. Sulfate-reducing bacteria (SRB) contribute to huge economic losses causing microbial-induced corrosion through biofilms on metal surfaces. To effectively combat the challenges posed by SRB, it is essential to understand their molecular mechanisms of biofilm formation. This study aimed to identify ncRNAs in the genome of a model SRB, Oleidesulfovibrio alaskensis G20 (OA G20). Three in silico approaches revealed genome-wide distribution of 37 ncRNAs excluding tRNAs in the OA G20. These ncRNAs belonged to 18 different Rfam families. This study identified riboswitches, sRNAs, RNP, and SRP. The analysis revealed that these ncRNAs could play key roles in the regulation of several pathways of biosynthesis and transport involved in biofilm formation by OA G20. Three sRNAs, Pseudomonas P10, Hammerhead type II, and sX4, which were found in OA G20, are rare and their roles have not been determined in SRB. These results suggest that applying various computational methods could enrich the results and lead to the discovery of additional novel ncRNAs, which could lead to understanding the “rules of life of OA G20” during biofilm formation.
Full article
(This article belongs to the Section Microbiomes)
►▼
Show Figures

Figure 1
Open AccessCommunication
The Heterogeneous Habitat of Taiga Forests Changes the Soil Microbial Functional Diversity
by
Tian Zhou, Song Wu, Mingliang Gao and Libin Yang
Microorganisms 2024, 12(5), 959; https://doi.org/10.3390/microorganisms12050959 - 10 May 2024
Abstract
The soil contains abundant and diverse microorganisms, which interrelate closely with the aboveground vegetation and impact the structure and function of the forest ecosystem. To explore the effect of vegetation diversity on soil microbial functional diversity in taiga forests, we selected significantly different
[...] Read more.
The soil contains abundant and diverse microorganisms, which interrelate closely with the aboveground vegetation and impact the structure and function of the forest ecosystem. To explore the effect of vegetation diversity on soil microbial functional diversity in taiga forests, we selected significantly different important values of Larix gmelinii as experimental grouping treatments based on plant investigation from fixed plots in Da Xing’anling Mountains. Following that, we collected soil samples and applied the Biolog-ECO microplate method to investigate differences in carbon source utilization, features of functional diversity in soil microorganisms, and factors influencing them in taiga forests. The AWCD decreased as the important value of Larix gmelinii grew, and soil microorganisms preferred carboxylic acids, amino acids, and carbohydrates over polymers, phenolic acids, and amines. The Shannon and McIntosh indexes decreased significantly with the increase of the important value of Larix gmelinii (p < 0.05) and were positively correlated with soil SOC, MBC, C/N, and pH, but negatively with TN, AP, and AN. Redundancy analysis revealed significant effects on soil microbial functional diversity from soil C/N, SOC, AP, MBC, TN, pH, AN, and WC. To sum up, heterogeneous habitats of taiga forests with different important values altered soil microbial functional diversity.
Full article
(This article belongs to the Section Environmental Microbiology)
►▼
Show Figures

Figure 1
Open AccessCorrection
Correction: Nygren et al. Tick-Borne Encephalitis Risk Increases with Dog Ownership, Frequent Walks, and Gardening: A Case-Control Study in Germany 2018–2020. Microorganisms 2022, 10, 690
by
Teresa Marie Nygren, Antonia Pilic, Merle Margarete Böhmer, Christiane Wagner-Wiening, Ole Wichmann, Thomas Harder and Wiebke Hellenbrand
Microorganisms 2024, 12(5), 958; https://doi.org/10.3390/microorganisms12050958 (registering DOI) - 10 May 2024
Abstract
In the original publication [...]
Full article
(This article belongs to the Special Issue Epidemiology, Surveillance and Prevention of Tick-Borne Diseases)
Open AccessArticle
Exercise Affects Mucosa-Associated Microbiota and Colonic Tumor Formation Induced by Azoxymethane in High-Fat-Diet-Induced Obese Mice
by
Shogen Yo, Hiroshi Matsumoto, Tingting Gu, Momoyo Sasahira, Motoyasu Oosawa, Osamu Handa, Eiji Umegaki and Akiko Shiotani
Microorganisms 2024, 12(5), 957; https://doi.org/10.3390/microorganisms12050957 - 9 May 2024
Abstract
The only reliable factor that reduces the risk of colorectal carcinogenesis is physical activity. However, the underlying mechanisms remain unclear. In this study, we examined the effects of physical activity against gut microbiota, including mucosa-associated microbiota (MAM) on azoxymethane-induced colorectal tumors in obese
[...] Read more.
The only reliable factor that reduces the risk of colorectal carcinogenesis is physical activity. However, the underlying mechanisms remain unclear. In this study, we examined the effects of physical activity against gut microbiota, including mucosa-associated microbiota (MAM) on azoxymethane-induced colorectal tumors in obese mice. We divided the subjects into four groups: normal diet (ND), high-fat diet (HFD), ND + exercise (Ex), and HFD + Ex groups. The Ex group performed treadmill exercise for 20 weeks. Thereafter, fecal and colonic mucus samples were extracted for microbiota analysis. DNA was collected from feces and colonic mucosa, and V3–V4 amplicon sequencing analysis of the 16SrRNA gene was performed using MiSeq. The HFD group had significantly more colonic polyps than the ND group (ND 6.5 ± 1.3, HFD 11.4 ± 1.5, p < 0.001), and the addition of Ex suppressed the number of colonic polyps in ND and HFD groups (ND 6.5 ± 1.3, ND + Ex 2.8 ± 2.5, p < 0.05). The HFD group showed significantly lower concentrations of succinic, acetic, butyric, and propionic acids (mg/g) in feces, compared with the ND group (succinic acid HFD 0.59, ND 0.17; acetic acid HFD 0.63, ND 2.41; propionic acid HFD 0.10, ND 0.47; and N-butyric acid HFD 0.31, ND 0.93). In the case of ND, succinic acid and butyric acid tended to decrease with Ex (succinic acid ND 0.17, ND + Ex 0.12; N-butyric acid ND 0.93, ND + Ex 0.74 0.74). Succinic acid, acetic acid, butyric acid, and propionic acid levels in feces were significantly lower in the HFD group than in the ND group; in both feces and mucus samples, Butyricicoccus and Lactobacillus levels were significantly lower in the HFD group. Akkermansia was significantly increased in ND + Ex and HFD + Ex groups. Diet and exercise affected the number of colorectal tumors. Furthermore, diet and exercise alter intestinal MAM, which may be involved in colorectal tumor development.
Full article
(This article belongs to the Special Issue Gut Microbiota, Diet, and Gastrointestinal Cancer)
►▼
Show Figures

Figure 1
Open AccessArticle
The Impact of Aboveground Epichloë Endophytic Fungi on the Rhizosphere Microbial Functions of the Host Melica transsilvanica
by
Chuanzhe Wang, Chong Shi, Wei Huang, Mengmeng Zhang and Jiakun He
Microorganisms 2024, 12(5), 956; https://doi.org/10.3390/microorganisms12050956 - 8 May 2024
Abstract
In nature, the symbiotic relationship between plants and microorganisms is crucial for ecosystem balance and plant growth. This study investigates the impact of Epichloë endophytic fungi, which are exclusively present aboveground, on the rhizosphere microbial functions of the host Melica transsilvanica. Using
[...] Read more.
In nature, the symbiotic relationship between plants and microorganisms is crucial for ecosystem balance and plant growth. This study investigates the impact of Epichloë endophytic fungi, which are exclusively present aboveground, on the rhizosphere microbial functions of the host Melica transsilvanica. Using metagenomic methods, we analyzed the differences in microbial functional groups and functional genes in the rhizosphere soil between symbiotic (EI) and non-symbiotic (EF) plants. The results reveal that the presence of Epichloë altered the community structure of carbon and nitrogen cycling-related microbial populations in the host’s rhizosphere, significantly increasing the abundance of the genes (porA, porG, IDH1) involved in the rTCA cycle of the carbon fixation pathway, as well as the abundance of nxrAB genes related to nitrification in the nitrogen-cycling pathway. Furthermore, the presence of Epichloë reduces the enrichment of virulence factors in the host rhizosphere microbiome, while significantly increasing the accumulation of resistance genes against heavy metals such as Zn, Sb, and Pb. This study provides new insights into the interactions among endophytic fungi, host plants, and rhizosphere microorganisms, and offers potential applications for utilizing endophytic fungi resources to improve plant growth and soil health.
Full article
(This article belongs to the Topic Microbe-Induced Abiotic Stress Alleviation in Plants)
Open AccessReview
Friend or Foe: Exploring the Relationship between the Gut Microbiota and the Pathogenesis and Treatment of Digestive Cancers
by
Monica Profir, Oana Alexandra Roşu, Sanda Maria Creţoiu and Bogdan Severus Gaspar
Microorganisms 2024, 12(5), 955; https://doi.org/10.3390/microorganisms12050955 - 8 May 2024
Abstract
Digestive cancers are among the leading causes of cancer death in the world. However, the mechanisms of cancer development and progression are not fully understood. Accumulating evidence in recent years pointing to the bidirectional interactions between gut dysbiosis and the development of a
[...] Read more.
Digestive cancers are among the leading causes of cancer death in the world. However, the mechanisms of cancer development and progression are not fully understood. Accumulating evidence in recent years pointing to the bidirectional interactions between gut dysbiosis and the development of a specific type of gastrointestinal cancer is shedding light on the importance of this “unseen organ”—the microbiota. This review focuses on the local role of the gut microbiota imbalance in different digestive tract organs and annexes related to the carcinogenic mechanisms. Microbiota modulation, either by probiotic administration or by dietary changes, plays an important role in the future therapies of various digestive cancers.
Full article
(This article belongs to the Special Issue Gut Microbiota, Diet, and Gastrointestinal Cancer)
►▼
Show Figures
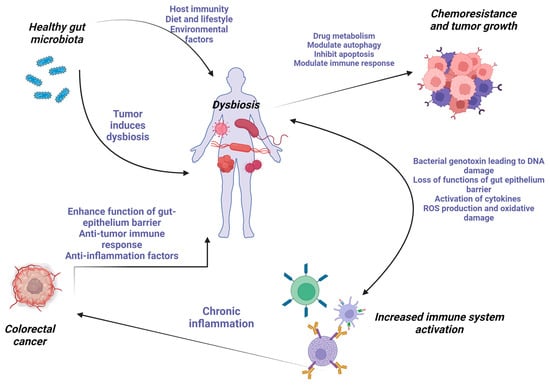
Figure 1
Open AccessArticle
Virulence and Antimicrobial Resistance of Listeria monocytogenes Isolated from Ready-to-Eat Food Products in Romania
by
Mihaela Niculina Duma, Laurenţiu Mihai Ciupescu, Sorin Daniel Dan, Oana Lucia Crisan-Reget and Alexandra Tabaran
Microorganisms 2024, 12(5), 954; https://doi.org/10.3390/microorganisms12050954 - 8 May 2024
Abstract
Listeria monocytogenes (L. monocytogenes) poses a significant threat to food safety due to its ability to cause severe human illness and its resistance to various antibiotics and environmental conditions. This study investigated the prevalence, serotype distribution, virulence gene profiles, and antimicrobial
[...] Read more.
Listeria monocytogenes (L. monocytogenes) poses a significant threat to food safety due to its ability to cause severe human illness and its resistance to various antibiotics and environmental conditions. This study investigated the prevalence, serotype distribution, virulence gene profiles, and antimicrobial resistance patterns of L. monocytogenes in ready-to-eat (RTE) food products from Romania. A total of 8151 samples were analyzed, including various processed dairy, bovine, poultry, pork, and fish products. Bacterial isolation was conducted using the classical standard method, followed by confirmation through biochemical and molecular testing. Among the isolated strains, serotypes 1/2a, 1/2b, and 1/2c were identified, with a prevalence of 75% for serotype 1/2a. Additionally, virulence genes specific to listeriolysin O (hlyA) and regulatory factor A (prfA) were detected in all isolates. Antimicrobial susceptibility testing revealed varying resistance patterns among the L. monocytogenes strains. Trimethoprim-sulfamethoxazole and oxacillin showed the highest prevalence of resistance at 26.92% and 23.07%, respectively. However, all strains remained susceptible to ciprofloxacin, levofloxacin, and moxifloxacin. Notably, 23.07% of the isolates exhibited multidrug resistance, with the most common pattern being resistance to oxacillin, penicillin, and tetracycline. Analysis of antimicrobial resistance genes identified tetracycline resistance genes, particularly tet(C), tet(M), and tet(K), in a significant proportion of isolates. The presence of ampC and dfrD genes was also notable, indicating potential mechanisms of resistance. These results emphasize the necessity for ongoing surveillance of L. monocytogenes in RTE foods and emphasize the importance of thorough monitoring of antimicrobial resistance to guide public health strategies within the European Union.
Full article
(This article belongs to the Special Issue Advances in Antibiotic and Antifungal Resistance and Related Alternative Therapies, Second Edition)
Open AccessArticle
Identification and Safety Assessment of Enterococcus casseliflavus KB1733 Isolated from Traditional Japanese Pickle Based on Whole-Genome Sequencing Analysis and Preclinical Toxicity Studies
by
Shohei Satomi, Shingo Takahashi, Takuro Inoue, Makoto Taniguchi, Mai Sugi, Masakatsu Natsume and Shigenori Suzuki
Microorganisms 2024, 12(5), 953; https://doi.org/10.3390/microorganisms12050953 - 8 May 2024
Abstract
The present study involves the precise identification and safety evaluation of Enterococcus casseliflavus KB1733, previously identified using 16S rRNA analysis, through whole-genome sequencing, phenotypic analysis, and preclinical toxicity studies. Analyses based on the genome sequencing data confirm the identity of KB1733 as E.
[...] Read more.
The present study involves the precise identification and safety evaluation of Enterococcus casseliflavus KB1733, previously identified using 16S rRNA analysis, through whole-genome sequencing, phenotypic analysis, and preclinical toxicity studies. Analyses based on the genome sequencing data confirm the identity of KB1733 as E. casseliflavus and show that the genes related to vancomycin resistance are only present on the chromosome, while no virulence factor genes are present on the chromosome or plasmid. Phenotypic analyses of antibiotic resistance and hemolytic activity also indicated no safety concerns. A bacterial reverse mutation test showed there was no increase in revertant colonies of heat-killed KB1733. An acute toxicity test employing heat-killed KB1733 at a dose of 2000 mg/kg body weight in rats resulted in no deaths and no weight gain or other abnormalities in the general condition of the animals, with renal depression foci and renal cysts only occurring at the same frequency as in the control. Taking the background data into consideration, the effects on the kidneys observed in the current study were not caused by KB1733. Our findings suggest that KB1733 is non-pathogenic to humans/animals, although further studies involving repeated oral toxicity tests and/or clinical tests are required.
Full article
(This article belongs to the Section Systems Microbiology)
►▼
Show Figures

Figure 1
Open AccessArticle
Genetic Diversity of Human Respiratory Syncytial Virus during COVID-19 Pandemic in Yaoundé, Cameroon, 2020–2021
by
Moïse Henri Moumbeket Yifomnjou, Gwladys Chavely Monamele, Abdou Fatawou Modiyinji, Mohamadou Njankouo-Ripa, Boyomo Onana and Richard Njouom
Microorganisms 2024, 12(5), 952; https://doi.org/10.3390/microorganisms12050952 - 8 May 2024
Abstract
Worldwide, human respiratory syncytial virus (HRSV) is a major cause of severe infections of the lower respiratory system, affecting individuals of all ages. This study investigated the genetic variability of HRSV during the COVID-19 outbreak in Yaoundé; nasopharyngeal samples positive for HRSV were
[...] Read more.
Worldwide, human respiratory syncytial virus (HRSV) is a major cause of severe infections of the lower respiratory system, affecting individuals of all ages. This study investigated the genetic variability of HRSV during the COVID-19 outbreak in Yaoundé; nasopharyngeal samples positive for HRSV were collected from different age groups between July 2020 and October 2021. A semi-nested RT-PCR was performed on the second hypervariable region of the G gene of detected HRSV, followed by sequencing and phylogenetic assessment. Throughout the study, 40 (37.7%) of the 106 HRSV-positive samples successfully underwent G-gene amplification. HRSV A and HRSV B co-circulated at rates of 47.5% and 52.5%, respectively. HRSV A clustered in the GA2.3.5 genetic lineage (ON1) and HRSV B clustered in the GB5.0.5a genetic lineage (BA9). Differences in circulating genotypes were observed between pre- and post-pandemic years for HRSV A. Predictions revealed potential N-glycosylation sites at positions 237-318 of HRSV A and positions 228-232-294 of HRSV B. This study reports the molecular epidemiology of HRSV in Cameroon during the COVID-19 pandemic. It describes the exclusive co-circulation of two genetic lineages. These findings highlight the importance of implementing comprehensive molecular surveillance to prevent the unexpected emergence of other diseases.
Full article
(This article belongs to the Special Issue Emerging and Re-emerging Respiratory Viruses)
►▼
Show Figures

Figure 1
Open AccessArticle
Copper Resistance Mechanism and Copper Response Genes in Corynebacterium crenatum
by
Mingzhu Huang, Wenxin Liu, Chunyan Qin, Yang Xu, Xu Zhou, Qunwei Wen, Wenbin Ma, Yanzi Huang and Xuelan Chen
Microorganisms 2024, 12(5), 951; https://doi.org/10.3390/microorganisms12050951 - 8 May 2024
Abstract
Heavy metal resistance mechanisms and heavy metal response genes are crucial for microbial utilization in heavy metal remediation. Here, Corynebacterium crenatum was proven to possess good tolerance in resistance to copper. Then, the transcriptomic responses to copper stress were investigated, and the vital
[...] Read more.
Heavy metal resistance mechanisms and heavy metal response genes are crucial for microbial utilization in heavy metal remediation. Here, Corynebacterium crenatum was proven to possess good tolerance in resistance to copper. Then, the transcriptomic responses to copper stress were investigated, and the vital pathways and genes involved in copper resistance of C. crenatum were determined. Based on transcriptome analysis results, a total of nine significantly upregulated DEGs related to metal ion transport were selected for further study. Among them, GY20_RS0100790 and GY20_RS0110535 belong to transcription factors, and GY20_RS0110270, GY20_RS0100790, and GY20_RS0110545 belong to copper-binding peptides. The two transcription factors were studied for the function of regulatory gene expression. The three copper-binding peptides were displayed on the C. crenatum surface for a copper adsorption test. Furthermore, the nine related metal ion transport genes were deleted to investigate the effect on growth in copper stress. This investigation provided the basis for utilizing C. crenatum in copper bioremediation.
Full article
(This article belongs to the Special Issue Application of Microbes in Environmental Remediation)
►▼
Show Figures

Figure 1
Open AccessArticle
Two Novel Alkaliphilic Species Isolated from Saline-Alkali Soil in China: Halalkalibacter flavus sp. nov., and Halalkalibacter lacteus sp. nov
by
Pin-Jiao Jin, Lei Sun, Yong-Hong Liu, Kang-Kang Wang, Manik Prabhu Narsing Rao, Osama Abdalla Abdelshafy Mohamad, Bao-Zhu Fang, Li Li, Lei Gao, Wen-Jun Li and Shuang Wang
Microorganisms 2024, 12(5), 950; https://doi.org/10.3390/microorganisms12050950 - 8 May 2024
Abstract
The degradation of farmland in China underscores the need for developing and utilizing saline-alkali soil. Soil health relies on microbial activity, which aids in the restoration of the land’s ecosystem, and hence it is important to understand microbial diversity. In the present study,
[...] Read more.
The degradation of farmland in China underscores the need for developing and utilizing saline-alkali soil. Soil health relies on microbial activity, which aids in the restoration of the land’s ecosystem, and hence it is important to understand microbial diversity. In the present study, two Gram-stain-positive strains HR 1-10T and J-A-003T were isolated from saline-alkali soil. Preliminary analysis suggested that these strains could be a novel species. Therefore, the taxonomic positions of these strains were evaluated using polyphasic analysis. Phylogenetic and 16S rRNA gene sequence analysis indicated that these strains should be assigned to the genus Halalkalibacter. Cell wall contained meso-2,6-diaminopimelic acid. The polar lipids present in both strains were diphosphatidyl-glycerol, phosphatidylglycerol, and an unidentified phospholipid. The major fatty acids (>10%) were anteiso-C15:0, C16:0 and iso-C15:0. Average nucleotide identity and digital DNA#x2013;DNA hybridization values were below the threshold values (95% and 70%, respectively) for species delineation. Based on the above results, the strains represent two novel species of the genus Halalkalibacter, for which the names Halalkalibacter flavus sp. nov., and Halalkalibacter lacteus sp. nov., are proposed. The type strains are HR 1-10T (=GDMCC 1.2946T = MCCC 1K08312T = JCM 36285T), and J-A-003T (=GDMCC 1.2949T = MCCC 1K08417T = JCM 36286T).
Full article
(This article belongs to the Section Systems Microbiology)
►▼
Show Figures

Figure 1
Open AccessArticle
Green Manuring Enhances Soil Multifunctionality in Tobacco Field in Southwest China
by
Yu Feng, Hua Chen, Libo Fu, Mei Yin, Zhiyuan Wang, Yongmei Li and Weidong Cao
Microorganisms 2024, 12(5), 949; https://doi.org/10.3390/microorganisms12050949 - 7 May 2024
Abstract
The use of green manure can substantially increase the microbial diversity and multifunctionality of soil. Green manuring practices are becoming popular for tobacco production in China. However, the influence of different green manures in tobacco fields has not yet been clarified. Here, smooth
[...] Read more.
The use of green manure can substantially increase the microbial diversity and multifunctionality of soil. Green manuring practices are becoming popular for tobacco production in China. However, the influence of different green manures in tobacco fields has not yet been clarified. Here, smooth vetch (SV), hairy vetch (HV), broad bean (BB), common vetch (CV), rapeseed (RS), and radish (RD) were selected as green manures to investigate their impact on soil multifunctionality and evaluate their effects on enhancing soil quality for tobacco cultivation in southwest China. The biomass of tobacco was highest in the SV treatment. Soil pH declined, and soil organic matter (SOM), total nitrogen (TN), and dissolved organic carbon (DOC) content in CV and BB and activity of extracellular enzymes in SV and CV treatments were higher than those in other treatments. Fungal diversity declined in SV and CV but did not affect soil multifunctionality, indicating that bacterial communities contributed more to soil multifunctionality than fungal communities. The abundance of Firmicutes, Rhizobiales, and Micrococcales in SV and CV treatments increased and was negatively correlated with soil pH but positively correlated with soil multifunctionality, suggesting that the decrease in soil pH contributed to increases in the abundance of functional bacteria. In the bacteria–fungi co-occurrence network, the relative abundance of key ecological modules negatively correlated with soil multifunctionality and was low in SV, CV, BB, and RS treatments, and this was associated with reductions in soil pH and increases in the content of SOM and nitrate nitrogen (NO3−-N). Overall, we found that SV and CV are more beneficial for soil multifunctionality, and this was driven by the decrease in soil pH and the increase in SOM, TN, NO3−-N, and C- and N-cycling functional bacteria.
Full article
(This article belongs to the Special Issue Interactions of Mycorrhizal Fungi and Other Soil Microorganisms with Plants)
►▼
Show Figures

Figure 1
Open AccessArticle
Fast Bacterial Succession Associated with the Decomposition of Larix gmelinii Litter in Wudalianchi Volcano
by
Lihong Xie, Jiahui Cheng, Hongjie Cao, Fan Yang, Mingyue Jiang, Maihe Li and Qingyang Huang
Microorganisms 2024, 12(5), 948; https://doi.org/10.3390/microorganisms12050948 - 7 May 2024
Abstract
In order to understand the role of microorganisms in litter decomposition and the nutrient cycle in volcanic forest ecosystems, the dominant forest species Larix gmelinii in the volcanic lava plateau of the Wudalianchi volcano was considered as the research object. We analyzed the
[...] Read more.
In order to understand the role of microorganisms in litter decomposition and the nutrient cycle in volcanic forest ecosystems, the dominant forest species Larix gmelinii in the volcanic lava plateau of the Wudalianchi volcano was considered as the research object. We analyzed the response of bacterial community structure and diversity to litter decomposition for 1 year, with an in situ decomposition experimental design using litter bags and Illumina MiSeq high-throughput sequencing. The results showed that after 365 days, the litter quality residual rate of Larix gmelinii was 77.57%, and the litter N, P, C:N, C:P, and N:P showed significant differences during the decomposition period (p < 0.05). The phyla Cyanobacteria and the genus unclassified_o_Chloroplast were the most dominant groups in early decomposition (January and April). The phyla Proteobacteria, Actinobacteriota, and Acidobacteriota and the genera Massilia, Pseudomonas, and Sphingomona were higher in July and October. The microbial communities showed extremely significant differences during the decomposition period (p < 0.05), with PCoa, RDA, and litter QRR, C:P, and N as the main factors driving litter bacteria succession. Microbial functional prediction analysis showed that Chloroplasts were the major functional group in January and April. Achemoheterotrophy and aerobic chemoheterotrophy showed a significant decrease as litter decomposition progressed.
Full article
(This article belongs to the Topic Litter Decompositions: From Individuals to Ecosystems)
►▼
Show Figures

Figure 1
Open AccessArticle
Do Weather Conditions Still Have an Impact on the COVID-19 Pandemic? An Observation of the Mid-2022 COVID-19 Peak in Taiwan
by
Wan-Yi Lin, Hao-Hsuan Lin, Shih-An Chang, Tai-Chi Chen Wang, Juei-Chao Chen and Yu-Sheng Chen
Microorganisms 2024, 12(5), 947; https://doi.org/10.3390/microorganisms12050947 - 7 May 2024
Abstract
Since the onset of the COVID-19 pandemic in 2019, the role of weather conditions in influencing transmission has been unclear, with results varying across different studies. Given the changes in border policies and the higher vaccination rates compared to earlier conditions, this study
[...] Read more.
Since the onset of the COVID-19 pandemic in 2019, the role of weather conditions in influencing transmission has been unclear, with results varying across different studies. Given the changes in border policies and the higher vaccination rates compared to earlier conditions, this study aimed to reassess the impact of weather on COVID-19, focusing on local climate effects. We analyzed daily COVID-19 case data and weather factors such as temperature, humidity, wind speed, and a diurnal temperature range from 1 March to 15 August 2022 across six regions in Taiwan. This study found a positive correlation between maximum daily temperature and relative humidity with new COVID-19 cases, whereas wind speed and diurnal temperature range were negatively correlated. Additionally, a significant positive correlation was identified between the unease environmental condition factor (UECF, calculated as RH*Tmax/WS), the kind of Climate Factor Complex (CFC), and confirmed cases. The findings highlight the influence of local weather conditions on COVID-19 transmission, suggesting that such factors can alter environmental comfort and human behavior, thereby affecting disease spread. We also introduced the Fire-Qi Period concept to explain the cyclic climatic variations influencing infectious disease outbreaks globally. This study emphasizes the necessity of considering both local and global climatic effects on infectious diseases.
Full article
(This article belongs to the Special Issue Advances in Epidemiology and Modeling)
►▼
Show Figures

Figure 1
Open AccessArticle
Characterizing Glycosylation of Adeno-Associated Virus Serotype 9 Capsid Proteins Generated from HEK293 Cells through Glycopeptide Mapping and Released Glycan Analysis
by
Yu Zhou, Sonal Priya and Joseph Y. Ong
Microorganisms 2024, 12(5), 946; https://doi.org/10.3390/microorganisms12050946 - 7 May 2024
Abstract
Recombinant adeno-associated viral (AAV) vectors have emerged as prominent gene delivery vehicles for gene therapy. AAV capsid proteins determine tissue specificity and immunogenicity and play important roles in receptor binding, the escape of the virus from the endosome, and the transport of the
[...] Read more.
Recombinant adeno-associated viral (AAV) vectors have emerged as prominent gene delivery vehicles for gene therapy. AAV capsid proteins determine tissue specificity and immunogenicity and play important roles in receptor binding, the escape of the virus from the endosome, and the transport of the viral DNA to the nuclei of target cells. Therefore, the comprehensive characterization of AAV capsid proteins is necessary for a better understanding of the vector assembly, stability, and transduction efficiency of AAV gene therapies. Glycosylation is one of the most common post-translational modifications (PTMs) and may affect the tissue tropism of AAV gene therapy. However, there are few studies on the characterization of the N- and O-glycosylation of AAV capsid proteins. In this study, we identified the N- and O-glycosylation sites and forms of AAV9 capsid proteins generated from HEK293 cells using liquid chromatography–tandem mass spectrometry (LC-MS)-based glycopeptide mapping and identified free N-glycans released from AAV9 capsid proteins by PNGase F using hydrophilic interaction (HILIC) LC-MS and HILIC LC-fluorescence detection (FLD) methods. This study demonstrates that AAV9 capsids are sprinkled with sugars, including N- and O-glycans, albeit at low levels. It may provide valuable information for a better understanding of AAV capsids in supporting AAV-based gene therapy development.
Full article
(This article belongs to the Special Issue Adeno-Associated Virus Biology and AAV Vector-Mediated Gene Therapy)
►▼
Show Figures

Graphical abstract

Journal Menu
► ▼ Journal Menu-
- Microorganisms Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Aquaculture Journal, Fishes, Microorganisms, Water
Women in Aquaculture Research
Topic Editors: Camino Ordás, Patrícia Díaz-RosalesDeadline: 30 June 2024
Topic in
Agronomy, Environments, Microorganisms, Pollutants, Sustainability, Water
Soil and Water Pollution Process and Remediation Technologies, 2nd Volume
Topic Editors: Hongbiao Cui, Ru Wang, Yu Shi, Haiying Lu, Lin ChenDeadline: 15 July 2024
Topic in
Antibiotics, Antioxidants, JoF, Microbiology Research, Microorganisms
Redox in Microorganisms, 2nd Edition
Topic Editors: Michal Letek, Volker BehrendsDeadline: 31 July 2024
Topic in
Molecules, Pharmaceutics, Antibiotics, Microorganisms, Biomolecules, Marine Drugs, Polymers, IJMS
Antimicrobial Agents and Nanomaterials
Topic Editors: Sandra Pinto, Vasco D. B. BonifácioDeadline: 30 September 2024

Conferences
Special Issues
Special Issue in
Microorganisms
Human Skin Microbiota 2.0
Guest Editors: Holger Brüggemann, Rolf LoodDeadline: 15 May 2024
Special Issue in
Microorganisms
The Urban Microbiome
Guest Editors: Haruo Suzuki, Soojin JangDeadline: 31 May 2024
Special Issue in
Microorganisms
Emerging Pathogens Causing Acute Hepatitis
Guest Editors: Ibrahim M. Sayed, Sayed F. Abdelwahab, Ahmed El-ShamyDeadline: 15 June 2024
Special Issue in
Microorganisms
Monkeypox—Current Knowledge and Future Perspectives
Guest Editors: Krisztián Bányai, Jakab FerencDeadline: 30 June 2024
Topical Collections
Topical Collection in
Microorganisms
Trends in Yeast Biochemistry and Biotechnology
Collection Editors: Seiji Shibasaki, Mitsuyoshi Ueda
Topical Collection in
Microorganisms
Epidemiology and Pathogenicity of Animal-Adapted Streptococci
Collection Editors: Peter Valentin-Weigand, Marcus Fulde
Topical Collection in
Microorganisms
Biodegradation and Environmental Microbiomes
Collection Editors: Shuangjiang Liu, Hongzhi Tang, Jiandong Jiang, Xiaolei Wu