Discovering Host Genes Involved in the Infection by the Tomato Yellow Leaf Curl Virus Complex and in the Establishment of Resistance to the Virus Using Tobacco Rattle Virus-based Post Transcriptional Gene Silencing
Abstract
:1. Introduction
2. Analysis of gene expression in plants using a reverse genetics approach based on virus-induced gene silencing
3. Tomato yellow leaf curl viruses: a complex of begomoviruses infecting tomato plants worldwide
4. Identification of host genes involved in TYLCV infection
4.1. A Nicotiana benthamiana system to monitor TYLCSV infection in combination with host gene silencing
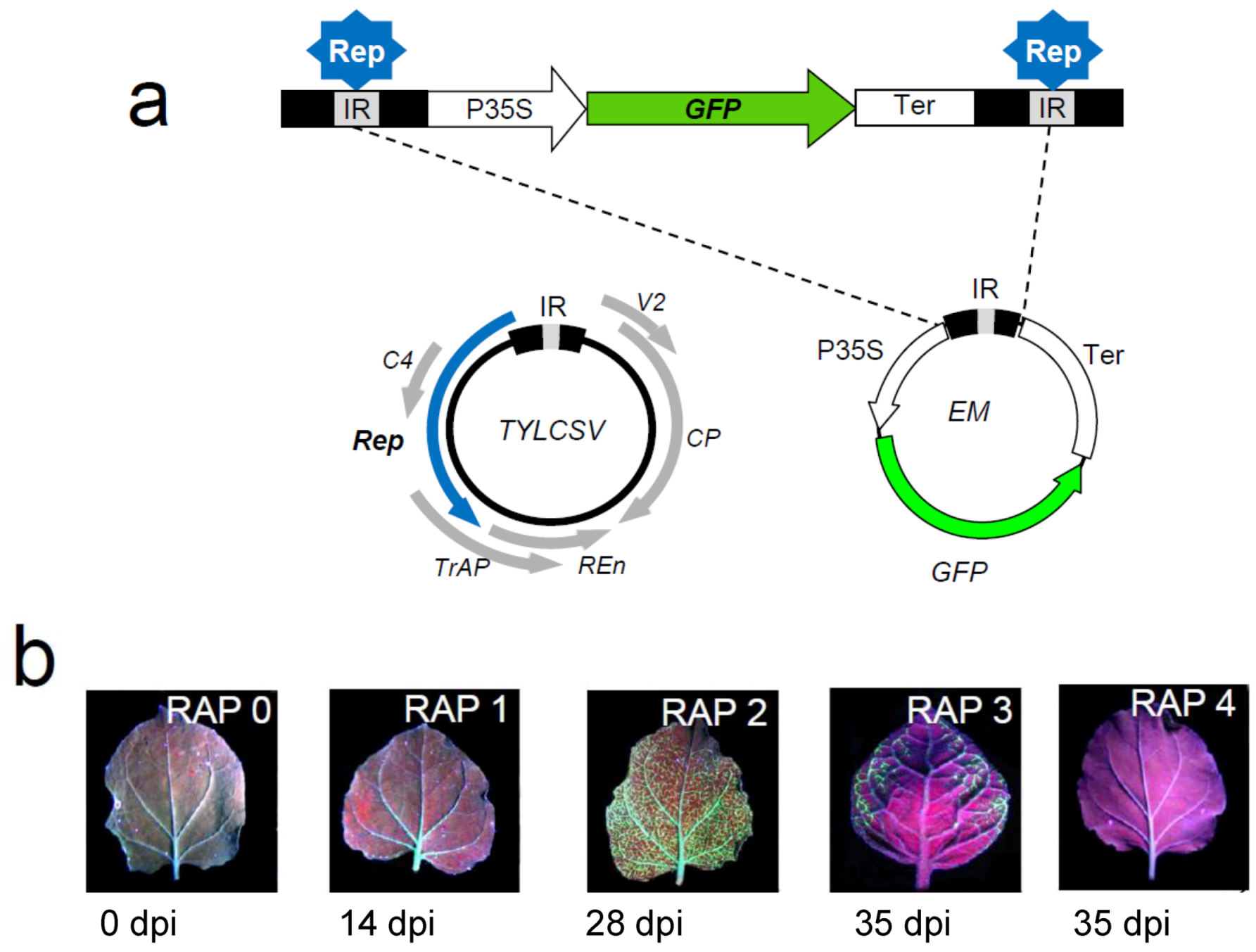
4.2. Selection and screening of candidate genes involved in TYLCSV infection
| Identity | ACC A. thaliana | GO Biological process | GO Cellular component | GO Molecular function | Selection criteria |
|---|---|---|---|---|---|
| Category A | |||||
| Bearskin 2 (BRN2) | AT4G10350 | Multicellular organismal development, positive regulation of gene expression, positive regulation of transcription, DNA-dependent, regulation of transcription, root cap development, secondary cell wall biogenesis | ND | Sequence-specific DNA binding transcription factor activity | Phloem over-expression |
| Importin alpha isoform 4 (IMPA-4) | AT1G09270 | Host response to induction by symbiont of tumor, nodule or growth in host, protein transport, symbiont intracellular protein transport in host | Cytosol, host cell, intracellular | Protein binding, protein transporter activity | Interaction with CP |
| Lactoylglutathione lyase (GLO1) | AT1G15380 | Carbohydrate metabolic process | ND | Lactoylglutathione lyase activity | Interaction with C3 |
| Replication protein A32 (RPA32/RPA2) | AT3G02920 | Unknown | ND | Nucleic acid binding | Interaction with Rep |
| Dehydration responsive 21 (RD21) | AT1G47128 | Metabolic process, response to water deprivation | Apoplast, chloroplast, plasmodesma, vacuole | Cysteine-type endopeptidase activity, protein binding | Interaction with V2 |
| RING-type E3 ubiquitin ligase (RHF2A) | AT5G22000 | Megagametogenesis, microgametogenesis, proteolysis involved in cellular protein catabolic process, regulation of cell cycle | Plasma membrane | Zinc ion binding | Transactived by TrAP/C2 |
| Ubiquitin activating enzyme (UBA1) | AT2G30110 | Metabolic process, protein ubiquitination, response to cadmium ion, response to other organism, ubiquitin-dependent protein catabolic process | Cytosol, plasma membrane, plasmodesma | Ubiquitin activating enzyme activity, ubiquitin-protein ligase activity | Interaction with TrAP/C2 |
| Category B | |||||
| 4-coumarate:CoA ligase (AT4CL1) | AT1G51680 | Metabolic process, phenylpropanoid metabolic process, response to UV, response to fungus, response to wounding | Unknown | 4-coumarate-CoA ligase activity | Phloem over-expression |
| Allene oxide cyclase (AOC1) | AT3G25760 | Jasmonic acid biosynthetic process, metabolic process, response to desiccation | Chloroplast, chloroplast envelope, chloroplast thylakoid membrane | Allene-oxide cyclase activity | Phloem over-expression |
| Barely any meristem 1 (BAM1) | AT5G65700 | Anther development, floral organ development, gametophyte development, protein phosphorylation, regulation of meristem growth, regulation of meristem structural organization, trans-membrane receptor protein tyrosine kinase signaling pathway | Plasma membrane | Kinase activity, protein binding, protein self-association, protein serine/threonine kinase activity, receptor serine/threonine kinase binding | Interaction with C4 |
| Coatomer delta subunit (deltaCOP) | AT5G05010 | Intracellular protein transport, transport, vesicle-mediated transport | Cytosol, membrane, plasmodesma | ND | Interaction with C3 |
| COP9 signalosome subunit 3 (CSN3) | AT5G14250 | G2 phase of mitotic cell cycle, cullin deneddylation, photomorphogenesis | Cytosol, signalosome | Protein binding | Cellular process |
| Geminivirus Rep A-binding (GRAB2) | AT5G61430 | Multicellular organismal development, regulation of transcription, DNA-dependent | Unknown | sequence-specific DNA binding transcription factor | Interaction with Rep |
| Heat shock protein cognate 70 (HSC70) | AT5G02500 | Defense response to bacterium, defence response to fungus, negative regulation of seed germination, protein folding, response to cadmium ion, response to cold, response to heat, response to virus, stomatal closure | Apoplast, cell wall, chloroplast, cytoplasm, cytosol, membrane, nucleus, plasma membrane, plasmodesma | ATP binding, protease binding, protein binding | Phloem over-expression |
| Nuclear acetyltransferase (NSI) | AT1G32070 | Pathogenesis, spread of virus in host | Chloroplast, nucleus | N-acetyltransferase activity | Interaction with NSP |
| Patatin-like protein 2 (PLP2) | AT2G26560 | Cell death, cellular response to hypoxia, defence response to virus, lipid metabolic process, oxylipin biosynthetic process, plant-type hypersensitive response, response to cadmium ion | Cytoplasm, membrane | Lipase activity, nutrient reservoir activity | Phloem over-expression |
| Shaggy-related kinase kappa (SK4-1/SKK) | AT1G09840 | Protein phosphorylation | Plasma membrane | ATP binding, protein serine/threonine kinase activity | Interaction with C4 |
| SKP1-like 2 (ASK2) | AT5G08590 | Phosphorylation, protein phosphorylation, response to osmotic stress, response to salt stress | Nucleus | Kinase activity, protein binding, protein kinase activity | Transactived by TrAP/C2 |
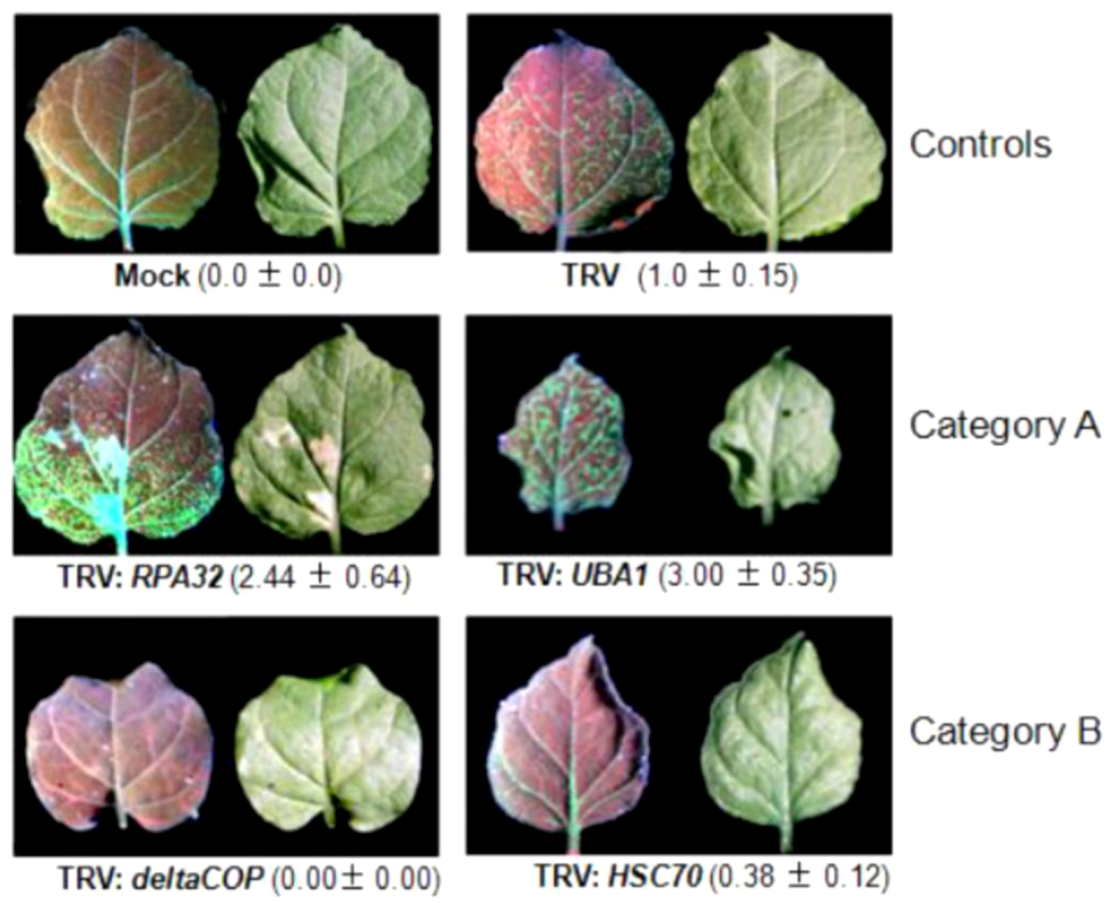
4.2.1. Genes with a known function in geminivirus infection
4.2.2. Genes involved in stress responses
4.2.3. Genes involved in post-translational modifications (PTMs)
5. Identification of genes involved in resistance to TYLCV
5.1. Genes preferentially expressed in TYLCV-resistant tomatoes and the effect of their silencing on resistance
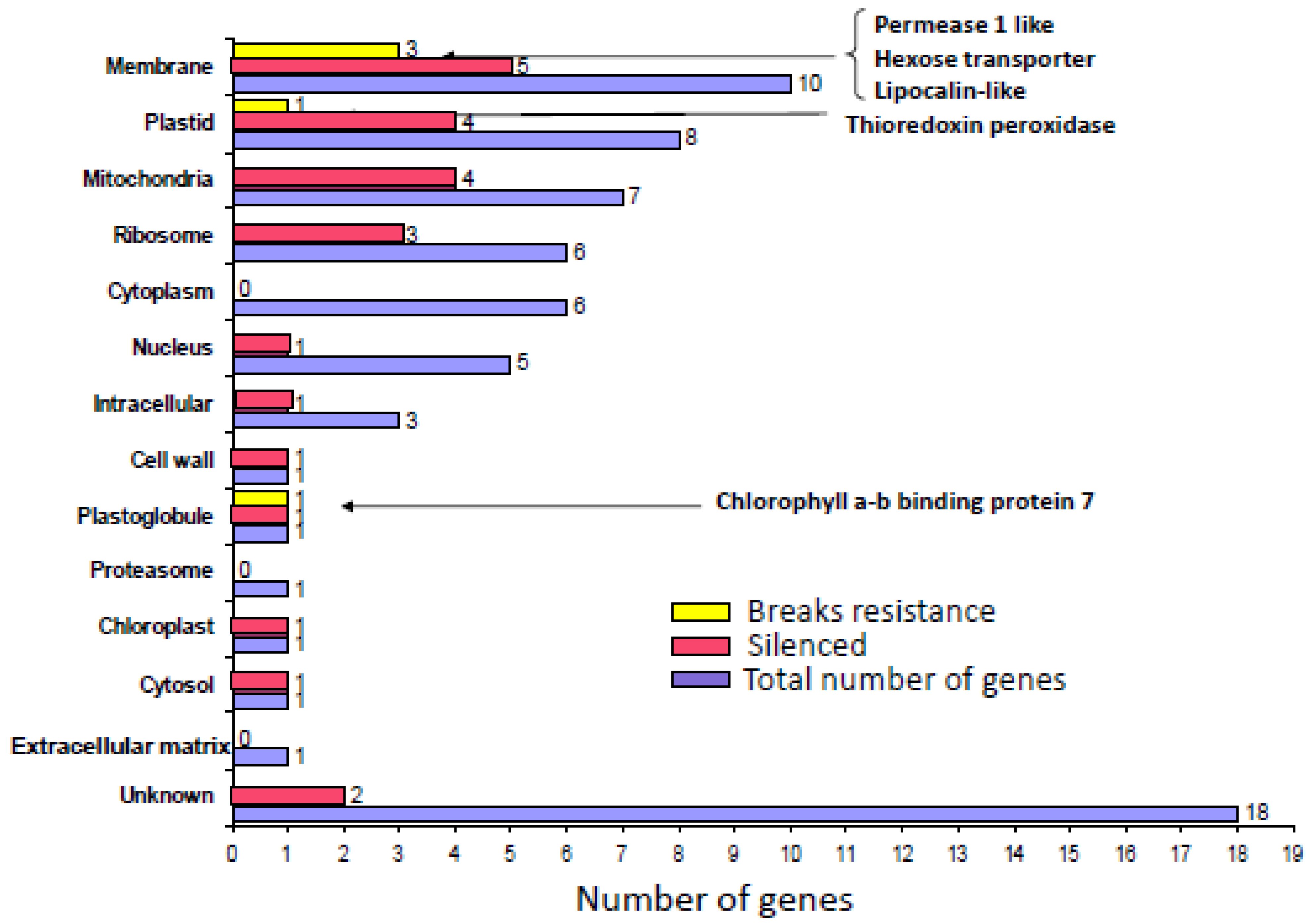
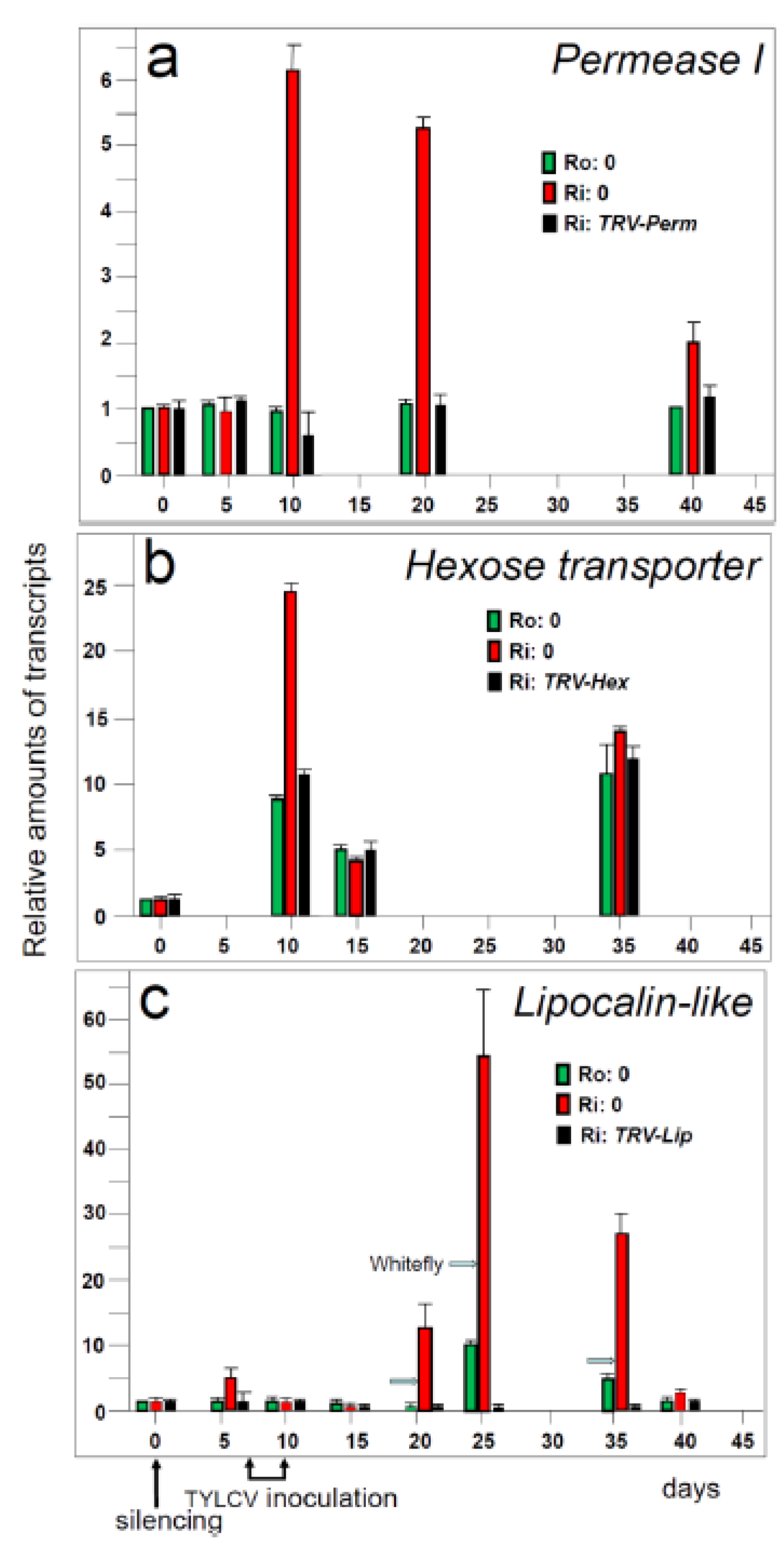
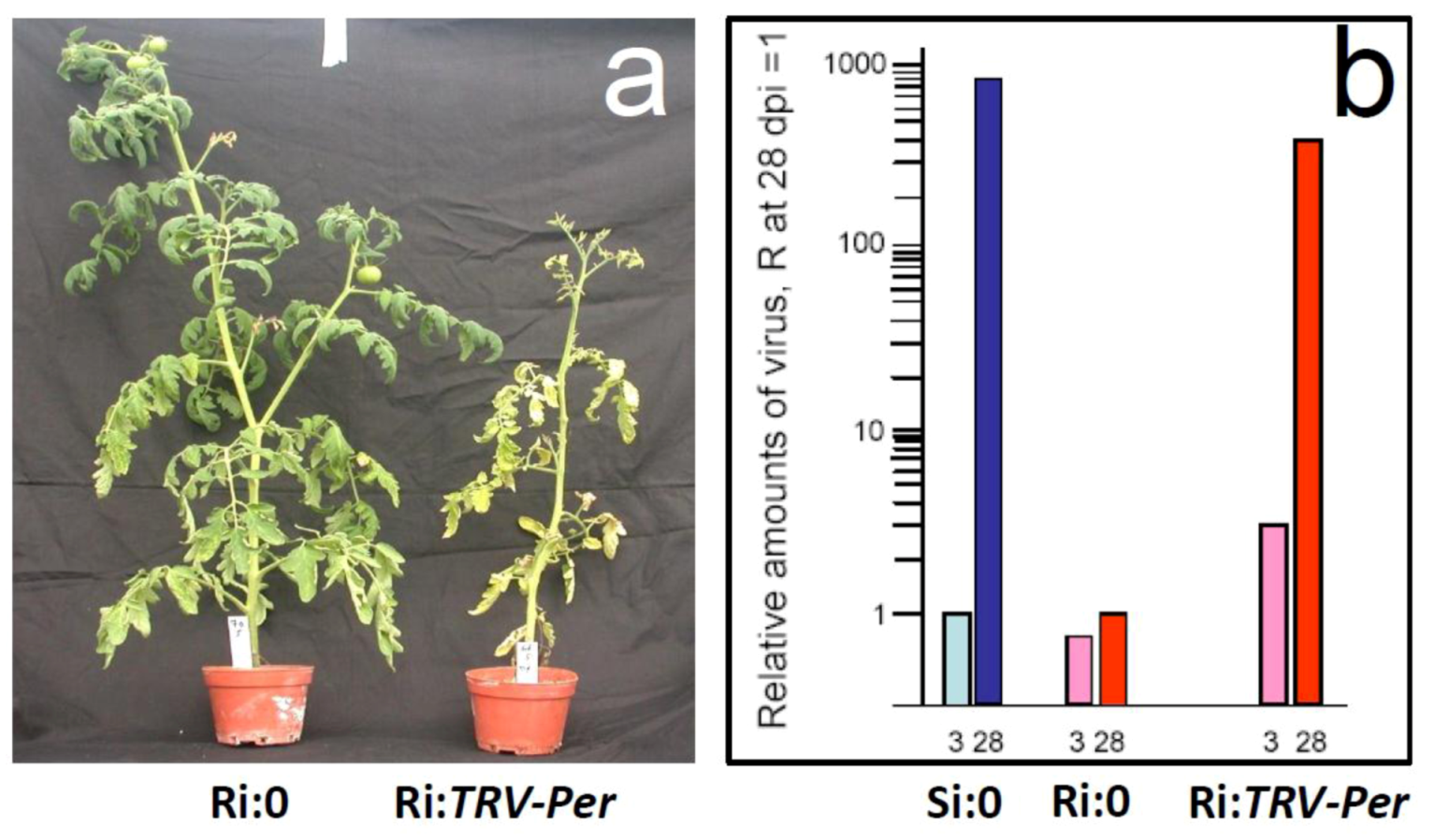
5.2. Hierarchy of genes involved in resistance to TYLCV
6. Discussion
Conflict of Interest
Acknowledgements
References
- Marathe, R.; Guan, Z.; Anandalakshmi, R.; Zhao, H.; Dinesh-Kumar, S.P. Study of Arabidopsis thaliana resistome in response to cucumber mosaic virus infection using whole genome microarray. Plant Mol. Biol. 2004, 55, 501–520. [Google Scholar] [CrossRef]
- Satoh, K.; Kondoh, H.; Sasaya, T.; Shimizu, T.; Choi, I.R.; Omura, T.; Kikuchi, S. Selective modification of rice (Oryza sativa) gene expression by rice stripe virus infection. J. Gen. Virol. 2010, 91, 294–305. [Google Scholar] [CrossRef]
- McGregor, C.E.; Miano, D.W.; Hoy, M.; Clark, C.A.; Rosa, G.J.M. Differential gene expression of resistant and susceptible sweetpotato plants after infection with the causal agents of sweet potato virus disease. J. Amer. Soc. Hort. Sci. 2009, 134, 658–666. [Google Scholar]
- Ascencio-Ibanez, J.T.; Sozzani, R.; Lee, T.J.; Chu, T.M.; Wolfinger, R.D.; Cella, R.; Hanley-Bowdoin, L. Global analysis of Arabidopsis gene expression uncovers a complex array of changes impacting pathogen response and cell cycle during geminivirus infection. Plant Physiol. 2008, 148, 436–454. [Google Scholar] [CrossRef]
- Eybishtz, A.; Peretz, Y.; Sade, D.; Akad, F.; Czosnek, H. Silencing of a single gene in tomato plants resistant to Tomato yellow leaf curl virus renders them susceptible to the virus. Plant Mol. Biol. 2009, 71, 157–171. [Google Scholar] [CrossRef]
- Lozano-Duran, R.; Rosas-Diaz, T.; Luna, A.P.; Bejarano, E.R. Identification of host genes involved in geminivirus infection using a reverse genetics approach. PLoS One 2011, 6, e22383. [Google Scholar]
- Whitham, S.A.; Yang, C.; Goodin, M.M. Global impact: elucidating plant responses to viral infection. Mol. Plant-Microbe Inter. 2006, 19, 1207–1215. [Google Scholar] [CrossRef]
- Carrington, J.C.; Kasschau, K.D.; Johansen, L.K. Activation and suppression of RNA silencing by plant viruses. Virology 2001, 281, 1–5. [Google Scholar] [CrossRef]
- Hamilton, A.J.; Baulcombe, D.C. A species of small antisense RNA in posttranscriptional gene silencing in plants. Science 1999, 286, 950–952. [Google Scholar] [CrossRef]
- Pratt, A.J.; MacRae, I.J. The RNA-induced silencing complex: a versatile gene-silencing machine. J. Biol. Chem. 2009, 284, 17897–17901. [Google Scholar] [CrossRef]
- Burgyán, J.; Havelda, Z. Viral suppressors of RNA silencing. Trends Plant Sci. 2011, 16, 265–272. [Google Scholar] [CrossRef]
- Lu, R.; Martin-Hernández, A.M.; Peart, J.R.; Malcuit, I.; Baulcombe, D.C. Virus induced gene silencing in plants. Methods 2003, 30, 296–303. [Google Scholar] [CrossRef]
- Purkayastha, A.; Dasgupta, I. Virus-induced gene silencing: a versatile tool for discovery of gene functions in plants. Plant Physiol. Biochem. 2009, 47, 967–976. [Google Scholar] [CrossRef]
- Liu, Y.; Schiff, M.; Dinesh-Kumar, S.P. Virus-induced gene silencing in tomato. Plant J. 2002, 31, 777–786. [Google Scholar] [CrossRef]
- Brigneti, G.; Martin-Hernández, A.M.; Jin, H.; Vhen, J.; Baulcombe, D.C.; Baker, B.; Jones, J.D.G. Virus-induced gene silencing in Solanum species. Plant J. 2004, 39, 264–272. [Google Scholar] [CrossRef]
- Spitzer, B.; Moyal Ben Zvi, M.; Ovadis, M.; Marhevka, R.; Barkai, O.; Edelbaum, O.; Marton, I.; Masci, T.; Alon, M.; Morin, S.; Rogachev, I.; Aharoni, A.; Vainstein, A. Reverse genetics of floral scent: application of Tobacco rattle virus-based gene silencing in petunia. Plant Physiol. 2007, 145, 1241–1250. [Google Scholar] [CrossRef]
- Senthil-Kumar, M.; Rame Gowda, H.V.; Hema, R.; Mysore, K.S.; Udayakumar, M. Virus-induced gene silencing and its application in characterizing genes involved in water-deficit-stress tolerance. J. Plant Physiol. 2008, 165, 1404–1421. [Google Scholar] [CrossRef]
- Burch-Smith, T.; Schiff, M.; Liu, Y.; Dinesh-Kumar, S.P. Efficient virus-induced silencing in Arabidopsis. Plant Physiol. 2004, 142, 21–27. [Google Scholar]
- Schilmiller, A.L.; Charbonneau, A.L.; Last, R.L. Identification of a BAHD acetyltransferase that produces protective acyl sugars in tomato trichomes. Proc. Natl. Acad. Sci. U.S.A. 2012, 109, 16377–16382. [Google Scholar] [CrossRef]
- Duan, C.-G.; Wang, C.-H.; Guo, H.-S. Application of RNA silencing to plant disease resistance. Silence 2012, 3, 5. [Google Scholar] [CrossRef]
- Unver, T.; Budak, H. Virus-induced gene silencing, a post transcriptional gene silencing method. Intl. J. Plant Genomics 2009, 2009, Article ID 198680. [Google Scholar] [CrossRef]
- Qin, G.; Gu, H.; Ma, L.; Peng, Y.; Deng, X.W.; Chen, Z.; Qu, L.J. Disruption of phytoene desaturase gene results in albino and dwarf phenotypes in Arabidopsis by impairing chlorophyll, carotenoid, and gibberellin biosynthesis. Cell Res. 2007, 17, 471–482. [Google Scholar] [CrossRef]
- Yuan, C.; Li, C.; Yan, L.; Jackson, A.O.; Liu, Z.; Huan, C.; Li, D. A high throughput Barley Stripe Mosaic Virus vector for virus induced gene silencing in monocots and dicots. PLoS ONE 2011, 6, e26468. [Google Scholar]
- Peele, C.; Jordan, C.V.; Muangsan, N.; Turnage, M.; Egelkrout, E.; Eagle, P.; Hanley-Bowdoin, L.; Robertson, D. Silencing of a meristematic gene using geminivirus-derived vectors. Plant J. 2001, 27, 357–366. [Google Scholar] [CrossRef]
- Pandey, P.; Choudhury, N.R.; Mukherjee, S.K. A geminiviral amplicon (VA) derived from Tomato leaf curl virus (ToLCV) can replicate in a wide variety of plant species and also acts as a VIGS vector. Virol. J. 2009, 6, 152. [Google Scholar] [CrossRef]
- Peretz, Y.; Mozes-Koch, R.; Akad, F.; Tanne, E.; Czosnek, H.; Sela, I. A universal expression/silencing vector in plants. Plant Physiol. 2007, 145, 1251–1263. [Google Scholar] [CrossRef]
- Muangsan, N.; Beclin, C.; Vaucheret, H.; Robertson, D. Geminivirus VIGS of endogenous genes requires SGS2/SDE1 and SGS3 and defines a new branch in the genetic pathway for silencing in plants. Plant J. 2004, 38, 1004–1014. [Google Scholar] [CrossRef]
- Huang, C.; Xie, Y.; Zhou, X. Efficient virus-induced gene silencing in plants using a modified geminivirus DNA1 component. Plant Biotech. J. 2009, 7, 254–265. [Google Scholar] [CrossRef]
- Fofana, I.B.; Sangare, A.; Collier, R.; Taylor, C.; Fauquet, C.M. A geminivirus-induced gene silencing system for gene function validation in cassava. Plant Mol. Biol. 2004, 56, 613–624. [Google Scholar] [CrossRef]
- Tuttle, J.R.; Idris, A.M.; Brown, J.K.; Haigler, C.H.; Robertson, D. Geminivirus-mediated gene silencing from Cotton leaf crumple virus is enhanced by low temperature in cotton. Plant Physiol. 2008, 148, 41–50. [Google Scholar] [CrossRef]
- Tomato yellow leaf curl virus disease: management, molecular biology, breeding for resistance; Czosnek, H. (Ed.) Springer: Dordrecht, The Netherlands, 2007.
- Bemisia, Bionomics and Management of a Global Pest; Stansly, P.A.; Naranjo, S.E. (Eds.) Springer: Dordrecht, The Netherlands, 2012.
- Hanley-Bowdoin, L.; Settlage, S.B.; Orozco, B.M.; Nagar, S.; Robertson, D. Geminiviruses: models for plant DNA replication, transcription, and cell cycle regulation. Crit. Rev. Biochem. Mol. Biol. 2000, 35, 105–140. [Google Scholar]
- Díaz-Pendón, J.A.; Cañizares, M.C.; Moriones, E.; Bejarano, E.R.; Czosnek, H.; Navas-Castillo, J. Tomato yellow leaf curl viruses: ménage à trois between the virus complex, the plant, and the whitefly vector. Mol. Plant Pathol. 2010, 11, 441–450. [Google Scholar] [CrossRef]
- Shepherd, D.N.; Martin, D.P.; Thomson, J.A. Transgenic strategies for developing crops resistant to geminiviruses. Plant Sci. 2009, 176, 1–11. [Google Scholar] [CrossRef]
- Chen, H.; Fu, R.; Li, X. Tomato yellow leaf virus (TYLCV): The structure, ecotypes and the resistance germplasm resources in tomato. African J. Biotech. 2011, 10, 13361–13367. [Google Scholar]
- Zamir, D. Improving plant breeding with exotic genetic libraries. Nature Rev. Genetics 2001, 2, 983–989. [Google Scholar] [CrossRef]
- Katagiri, F. A global view of defense gene expression regulation—a highly interconnected signaling network. Curr. Opin. Plant Biol. 2004, 7, 506–511. [Google Scholar] [CrossRef]
- Lapidot, M.; Friedman, M. Breeding for resistance to whitefly-transmitted geminiviruses. Ann. Appl. Biol. 2002, 140, 109–127. [Google Scholar] [CrossRef]
- Anbinder, I.; Reuveni, M.; Azari, R.; Paran, I.; Nahon, S.; Shlomo, H.; Chen, L.; Lapidot, M.; Levin, I. Molecular dissection of Tomato yellow leaf curl virus (TYLCV) resistance in tomato line TY172 derived from Solanum peruvianum. Theor. Appl. Gen. 2009, 119, 519–530. [Google Scholar] [CrossRef]
- Morilla, G.; Castillo, A.G.; Preiss, W.; Jeske, H.; Bejarano, E.R. A versatile transreplication-based system to identify cellular proteins involved in geminivirus replication. J Virol. 2006, 80, 3624–3633. [Google Scholar] [CrossRef]
- Luna, A.P.; Morilla, G.; Voinnet, O.; Bejarano, E.R. Functional analysis of gene silencing suppressors from Tomato yellow leaf curl disease viruses. Mol. Plant Microbe Interact. 2012, 25, 1294–1306. [Google Scholar] [CrossRef]
- Trinks, D.; Rajeswaran, R.; Shivaprasad, P.V.; Akbergenov, R.; Oakeley, E.J.; Veluthambi, K.; Hohn, T.; Pooggin, M.M. Suppression of RNA silencing by a geminivirus nuclear protein, AC2, correlates with transactivation of host genes. J Virol. 2005, 79, 2517–2527. [Google Scholar] [CrossRef]
- McGarry, R.C.; Barron, Y.D.; Carvalho, M.F.; Hill, J.E.; Gold, D.; Cheung, E.; Kraus, W.L.; Lazarowitz, S.G. A novel Arabidopsis acetyltransferase interacts with the geminivirus movement protein NSP. Plant Cell 2003, 15, 1605–1618. [Google Scholar] [CrossRef]
- Xie, Q.; Sanz-Burgos, A.P.; Guo, H.; Garcia, J.A.; Gutierrez, C. GRAB proteins, novel members of the NAC domain family, isolated by their interaction with a geminivirus protein. Plant Mol. Biol. 1999, 39, 647–656. [Google Scholar] [CrossRef]
- Singh, D.K.; Islam, M.N.; Choudhury, N.R.; Karjee, S.; Mukherjee, S.K. The 32 kDa subunit of replication protein A (RPA) participates in the DNA replication of Mung bean yellow mosaic India virus (MYMIV) by interacting with the viral Rep protein. Nucleic Acids Res. 2006, 35, 755–770. [Google Scholar]
- Clément, M.; Leonhardt, N.; Droillard, M.J.; Reiter, I.; Montillet, J.L.; Genty, B.; Lauriere, C.; Nussaume, L.; Noel, L.D. The cytosolic/nuclear HSC70 and HSP90 molecular chaperones are important for stomatal closure and modulate abscisic acid-dependent physiological responses in Arabidopsis. Plant Physiol. 2011, 156, 1481–1492. [Google Scholar] [CrossRef]
- Aparicio, F.; Thomas, C.L.; Lederer, C.; Niu, Y.; Wang, D.; Maule, A.J. Virus induction of heat shock protein 70 reflects a general response to protein accumulation in the plant cytosol. Plant Physiol. 2005, 138, 529–536. [Google Scholar] [CrossRef]
- Noel, L.D.; Cagna, G.; Stuttmann, J.; Wirthmuller, L.; Betsuyaku, S.; Witte, C.P.; Bhat, R.; Pochon, N.; Colby, T.; Parker, J.E. Interaction between SGT1 and cytosolic/nuclear HSC70 chaperones regulates Arabidopsis immune responses. Plant Cell 2007, 19, 4061–4076. [Google Scholar] [CrossRef]
- Verchot, J. Cellular chaperones and folding enzymes are vital contributors to membrane bound replication and movement complexes during plant RNA virus infection. Frontiers Plant Sci. 2012, 3, 1–12. [Google Scholar]
- Shindo, T.; Misas-Villamil, J.C.; Horger, A.C.; Song, J.; van der Hoorn, R.A. A role in immunity for Arabidopsis cysteine protease RD21, the ortholog of the tomato immune protease C14. PLoS One 2012, 7, e29317. [Google Scholar]
- La Camera, S.; La Balague, C.; Gobel, C.; Geoffroy, P.; Legrand, M.; Feussner, I.; Roby, D.; Heitz, T. The Arabidopsis patatin-like protein 2 (PLP2) plays an essential role in cell death execution and differentially affects biosynthesis of oxylipins and resistance to pathogens. Mol. Plant Microbe Interact. 2009, 22, 469–481. [Google Scholar] [CrossRef]
- Yang, W.Y.; Zheng, Y.; Bahn, S.C.; Pan, X.Q.; Li, M.Y.; Vu, H.S.; Roth, M.R.; Scheu, B.; Welti, R.; Hong, Y.Y.; Wang, S.M. The patatin-containing phospholipase A pPLAIIalpha modulates oxylipin formation and water loss in Arabidopsis thaliana. Mol. Plant 2012, 5, 452–460. [Google Scholar] [CrossRef]
- Lozano-Duran, R.; Bejarano, E.R. Geminivirus C2 protein might be the key player for geminiviral co- option of SCF-mediated ubiquitination. Plant Signal Behav. 2011, 6, 999–1001. [Google Scholar] [CrossRef]
- Mustafiz, A.; Sahoo, K.K.; Singla-Pareek, S.L.; Sopory, S.K. Metabolic engineering of glyoxalase pathway for enhancing stress tolerance in plants. Methods Mol. Biol. 2010, 639, 95–118. [Google Scholar] [CrossRef]
- Sun, W.; Xu, X.; Zhu, H.; Liu, A.; Liu, L.; Li, J.; Hua, X. Comparative transcriptomic profiling of a salt-tolerant wild tomato species and a salt-sensitive tomato cultivar. Plant Cell Physiol. 2010, 51, 997–1006. [Google Scholar] [CrossRef]
- Hakmaoui, A.; Pérez-Bueno, M.L.; García-Fontana, B.; Camejo, D.; Jiménez, A.; Sevilla, F.; Barón, M. Analysis of the antioxidant response of Nicotiana benthamiana to infection with two strains of Pepper mild mottle virus. J. Exp. Bot. 2012, 63, 5487–5496. [Google Scholar] [CrossRef]
- Chen, H.; Zhang, Z.; Teng, K.; Lai, J.; Zhang, Y.; Huang, Y.; Li, Y.; Liang, L.; Wang, Y.; Chu, C.; Guo, H.; Xie, Q. Up-regulation of LSB1/GDU3 affects geminivirus infection by activating the salicylic acid pathway. Plant J. 2010, 62, 12–23. [Google Scholar] [CrossRef]
- Dielen, A.S.; Badaoui, S.; Candresse, T.; German-Retana, S. The ubiquitin/26S proteasome system in plant-pathogen interactions: a never-ending hide-and-seek game. Mol. Plant Pathol. 2010, 11, 293–308. [Google Scholar] [CrossRef]
- Trujillo, M.; Shirasu, K. Ubiquitination in plant immunity. Curr. Opin. Plant Biol. 2010, 13, 402–408. [Google Scholar] [CrossRef]
- Alcaide-Loridan, C.; Jupin, I. Ubiquitin and plant viruses, let's play together ! Plant Physiol. 2012, 160, 72–82. [Google Scholar] [CrossRef]
- Eini, O.; Dogra, S.; Selth, L.A.; Dry, I.B.; Randles, J.W.; Rezaian, M.A. Interaction with a host ubiquitin-conjugating enzyme is required for the pathogenicity of a geminiviral DNA beta satellite. Mol. Plant Microbe Interact. 2009, 22, 737–746. [Google Scholar] [CrossRef]
- Goritschnig, S.; AZhang, Y.; Li, X. The ubiquitin pathway is required for innate immunity in Arabidopsis. Plant J. 2007, 49, 540–551. [Google Scholar] [CrossRef]
- Lozano-Duran, R.; Rosas-Diaz, T.; Gusmaroli, G.; Luna, A.P.; Taconnat, L.; Deng, X.W.; Bejarano, E.R. Geminiviruses subvert ubiquitination by altering CSN-mediated derubylation of SCF E3 ligase complexes and inhibit jasmonate signaling in Arabidopsis thaliana. Plant Cell 2011, 23, 1014–1032. [Google Scholar] [CrossRef]
- Hua, Z.; Zou, C.; Shiu, S.; Vierstra, R.D. Phylogenetic comparison of F-Box (FBX) gene superfamily within the plant kingdom reveals divergent evolutionary histories indicative of genomic drift. PLOS One 2011, 6, 1–20. [Google Scholar]
- Liu, F.; Ni, W.; Griffith, M.E.; Huang, Z.; Chang, C.; Peng, W.; Ma, H.; Xie, D. The ASK1 and ASK2 genes are essential for Arabidopsis early development. Plant Cell 2004, 16, 5–20. [Google Scholar] [CrossRef]
- Li, C.; Liu, Z.; Zhang, Q.; Wang, R.; Xiao, L.; Ma, H.; Chong, K.; Xu, Y. SKP1 is involved in abscisic acid signalling to regulate seed germination, stomatal opening and root growth in Arabidopsis thaliana. Plant Cell Environ. 2012, 35, 952–965. [Google Scholar] [CrossRef]
- Angot, A.; Peeters, N.; Lechner, E.; Vailleau, F.; Baud, C.; Gentzbittel, L.; Sartorel, E.; Genschik, P.; Boucher, C.; Genin, S. Ralstonia solanacearum requires F-box-like domain-containing type III effectors to promote disease on several host plants. Proc. Natl. Acad. Sci. USA 2006, 103, 14620–14625. [Google Scholar] [CrossRef]
- Schrammeijer, B.; Risseeuw, E.; Pansegrau, W.; Regensburg-Tuink, T.J.; Crosby, W.L.; Hooykaas, P.J. Interaction of the virulence protein VirF of Agrobacterium tumefaciens with plant homologs of the yeast Skp1 protein. Curr. Biol. 2001, 11, 258–262. [Google Scholar] [CrossRef]
- Glick, E.; Zrachya, A.; Levy, Y.; Mett, A.; Gidoni, D.; Belausov, E.; Citovsky, V.; Gafni, Y. Interaction with host SGS3 is required for suppression of RNA silencing by tomato yellow leaf curl virus V2 protein. Proc. Natl. Acad. Sci. USA 2008, 105, 157–161. [Google Scholar]
- Hotton, S.K.; Callis, J. Regulation of cullin RING ligases. Annu. Rev. Plant Biol. 2008, 59, 467–489. [Google Scholar] [CrossRef]
- Lozano-Duran, R.; Bejarano, E.R. Mutation in Arabidopsis CSN5A partially complements the lack of Beet curly top virus pathogenicity factor L2. J. Plant Pathol. Microbiol. 2011, 2, 3. [Google Scholar]
- Vidavski, F.; Czosnek, H. Tomato breeding lines immune and tolerant to Tomato yellow leaf curl virus (TYLCV) issued from Lycopersicon hirsutum. Phytopathology 1998, 88, 910–914. [Google Scholar] [CrossRef]
- Gear, M.; McPhillips, M.; Patrick, J.; McCurdy, D. Hexose transporters of tomato: molecular cloning, expression analysis and functional characterization. Plant Mol. Biol. 2000, 44, 687–697. [Google Scholar] [CrossRef]
- Bush, D.R. Proton-coupled sugar and amino acid transporters in plants. Ann. Rev. Plant Physiol. Plant Mol. Biol. 1993, 44, 513–542. [Google Scholar] [CrossRef]
- Büttner, M.; Sauer, N. Monosaccharide transporters in plants: structure, function and physiology. Biochim. Biophys. Acta 2000, 1465, 263–274. [Google Scholar] [CrossRef]
- Wahl, R.; Wippel, K.; Gioos, S.; Kämper, J.; Sauer, N. A novel high-affinity sucrose transporter is required for virulence of the plant pathogen Ustilago maydis. PLoS Biology 2010, 8, e10003030. [Google Scholar]
- Bolton, M.D. Primary metabolism and plant defense - fuel for the fire. Mol. Plant Microbe Interact. 2009, 22, 487–497. [Google Scholar] [CrossRef]
- Nørholm, M.H.; Nour-Eldin, H.H.; Brodersen, P.; Mundy, J.; Halkier, B.A. Expression of the Arabidopsis high-affinity hexose transporter STP13 correlates with programmed cell death. FEBS Lett. 2006, 580, 2381–2387. [Google Scholar] [CrossRef]
- Eybishtz, A.; Peretz, Y.; Sade, D.; Gorovits, R.; Czosnek, H. Tomato yellow leaf curl virus (TYLCV) infection of a resistant tomato line with a silenced sucrose transporter gene LeHT1 results in inhibition of growth, enhanced virus spread and necrosis. Planta 2010, 231, 537–548. [Google Scholar] [CrossRef]
- Greenberg, J.T.; Yao, N. The role and regulation of programmed cell death in plant-pathogen interactions. Cell. Microbiol. 2004, 6, 201–211. [Google Scholar] [CrossRef]
- Sade, D.; Eybishtz, A.; Gorovits, R.; Sobol, I.; Czosnek, H. A developmentally regulated lipocalin-like gene is overexpressed in Tomato yellow leaf curl virus-resistant tomato plants upon virus inoculation, and its silencing abolishes resistance. Plant Mol. Biol. 2012, 80, 273–287. [Google Scholar] [CrossRef]
- Charron, J.B.F.; Ouellet, F.; Pelletier, M.; Danyluk, J.; Chauve, C.; Sarhan, F. Identification; expression, and evolutionary analyses of plant lipocalins. Plant Physiol. 2005, 139, 2017–2028. [Google Scholar] [CrossRef]
- Levesque-Tremblay, G.; Havaux, M.; Ouellet, F. The chloroplastic lipocalin AtCHL prevents lipid peroxidation and protects Arabidopsis against oxidative stress. Plant J. 2009, 60, 691–702. [Google Scholar] [CrossRef]
- Thompson, G.A.; Goggin, F.L. Transcriptomics and functional genomics of plant defence induction by phloem-feeding insects. J. Exp. Bot. 2006, 57, 755–766. [Google Scholar] [CrossRef]
- Dorokhov, Y.L.; Frolova, O.Y.; Skurat, E.V.; Ivanov, P.A.; Gasanova, T.V.; Sheveleva, A.A.; Ravin, N.V.; Mäkinen, K.M.; Klimyuk, V.I.; Skryabin, K.G.; Gleba, Y.Y.; Atabekov, J.G. A novel function for a ubiquitous plant enzyme pectin methylesterase: the enhancer of RNA silencing. FEBS Lett. 2006, 580, 3872–3878. [Google Scholar] [CrossRef]
- Chen, M.; Citovsky, V. Systemic movement of a tobamovirus requires host cell pectin methylesterase. Plant J. 2003, 35, 386–392. [Google Scholar] [CrossRef]
- McCurdy, D.W.; Dibley, S.; Cahyanegara, R.; Martin, A.; Patrick, J.W. Functional characterization and RNAi-mediated suppression reveals roles for hexose transporters in sugar accumulation by tomato fruit. Mol. Plant 2010, 3, 1049–1063. [Google Scholar] [CrossRef]
- Rolland, F.; Moore, B.; Sheen, J. Sugar sensing and signaling in plants. Plant Cell 2002, S185–S205. [Google Scholar]
- Atkinson, N.J.; Urwin, P.E. The interaction of plant biotic and abiotic stresses: from genes to the field. J. Exp. Bot. 2012, 63, 3523–3543. [Google Scholar] [CrossRef]
- Sade, D.; Brotman, Y.; Eybishtz, A.; Cuadros-Inostroza, A.; Fernie, A.R.; Willmitzer, L.; Czosnek, H. Involvement of the hexose transporter gene LeHT1 and of sugars in resistance of tomato to Tomato yellow leaf curl virus. Mol. Plant 2013. accepted.. [Google Scholar]
- He, P.; Shan, L.; Sheen, J. Elicitation and suppression of microbe-associated molecular pattern-triggered immunity in plant-microbe interactions. Cell Microbiol. 2007, 9, 1385–1396. [Google Scholar] [CrossRef]
- Bombarely, A.; Rosli, H.G.; Vrebalov, J.; Moffett, P.; Mueller, L.; Martin, G. A draft genome sequence of Nicotiana benthamiana to enhance molecular plant-microbe biology research. Mol. Plant Microbe Interact. 2012, 25, 1523–1530. [Google Scholar] [CrossRef]
- Tomato-Genome-Consortium. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 2012, 485, 635–641.[Green Version]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Czosnek, H.; Eybishtz, A.; Sade, D.; Gorovits, R.; Sobol, I.; Bejarano, E.; Rosas-Díaz, T.; Lozano-Durán, R. Discovering Host Genes Involved in the Infection by the Tomato Yellow Leaf Curl Virus Complex and in the Establishment of Resistance to the Virus Using Tobacco Rattle Virus-based Post Transcriptional Gene Silencing. Viruses 2013, 5, 998-1022. https://doi.org/10.3390/v5030998
Czosnek H, Eybishtz A, Sade D, Gorovits R, Sobol I, Bejarano E, Rosas-Díaz T, Lozano-Durán R. Discovering Host Genes Involved in the Infection by the Tomato Yellow Leaf Curl Virus Complex and in the Establishment of Resistance to the Virus Using Tobacco Rattle Virus-based Post Transcriptional Gene Silencing. Viruses. 2013; 5(3):998-1022. https://doi.org/10.3390/v5030998
Chicago/Turabian StyleCzosnek, Henryk, Assaf Eybishtz, Dagan Sade, Rena Gorovits, Iris Sobol, Eduardo Bejarano, Tábata Rosas-Díaz, and Rosa Lozano-Durán. 2013. "Discovering Host Genes Involved in the Infection by the Tomato Yellow Leaf Curl Virus Complex and in the Establishment of Resistance to the Virus Using Tobacco Rattle Virus-based Post Transcriptional Gene Silencing" Viruses 5, no. 3: 998-1022. https://doi.org/10.3390/v5030998




