Lessons from Genome-Wide Search for Disease-Related Genes with Special Reference to HLA-Disease Associations
Abstract
:1. Development of Genome-Wide Searches
2. Identified Susceptibility Genes to Multifactorial Diseases
2.1. Population Differences in Disease Susceptibility Genes
| Gen (SNP) | Population | Odds Ratio | p | Minor Allele Frequency |
|---|---|---|---|---|
| TCF7L2 (rs7903146) | European [8,9,10,11] | 1.37 | 1.0 × 10−48 | 0.31/0.25 |
| TCF7L2 (rs7903146) | Japanese [14] | 1.70 | 7.0 × 10−4 | 0.05/0.02 |
| KCNQ1 (rs2237892) | European [12] | 1.29 | 7.8 × 10−4 | 0.03/0.05 |
| KCNQ1 (rs2237892) | Japanese [12] | 1.43 | 3.0 × 10−29 | 0.31/0.40 |
2.2. Susceptibility Genes Common to Different Diseases
| Panel | Case: Minor Allele Frequency | Control: Minor Allele Frequency | p * | Odds Ratio |
|---|---|---|---|---|
| 1 | 0.237 | 0.167 | 5.8 × 10−3 | 2.33 (1.25–4.35) |
| 2 | 0.195 | 0.159 | 2.0 × 10−5 | 3.44 (1.89–6.25) |
| 3 | 0.224 | 0.174 | 8.7 × 10−6 | 2.39 (1.61–3.55) |
| Combined | 0.219 | 0.168 | 6.0 × 10−11 | 2.54 (1.91–3.40) |
2.3. Towards the Understanding of Pathogenic Mechanisms
3. Identified Response Genes to Drugs/Therapies
3.1. Development of New Gene Tests
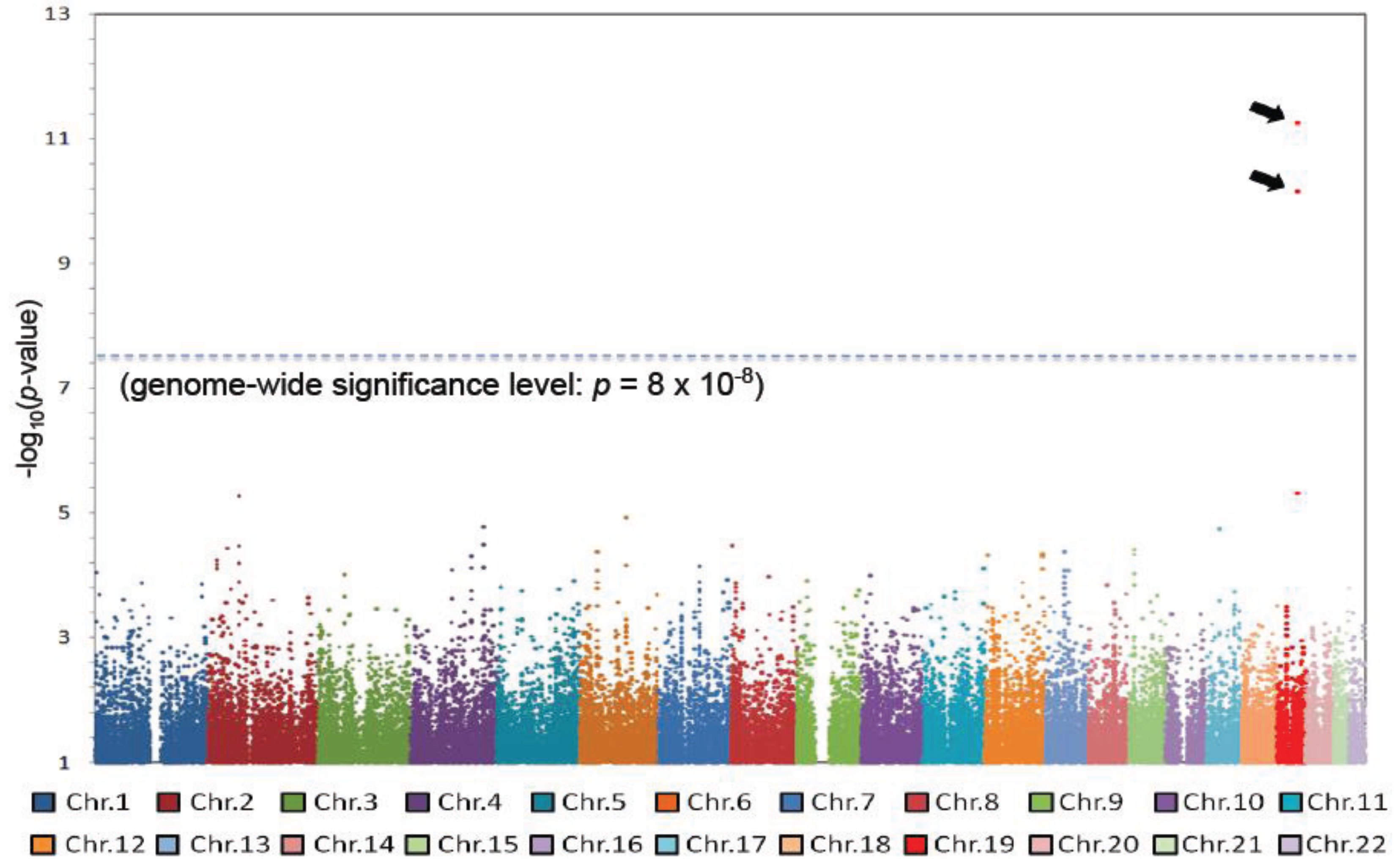
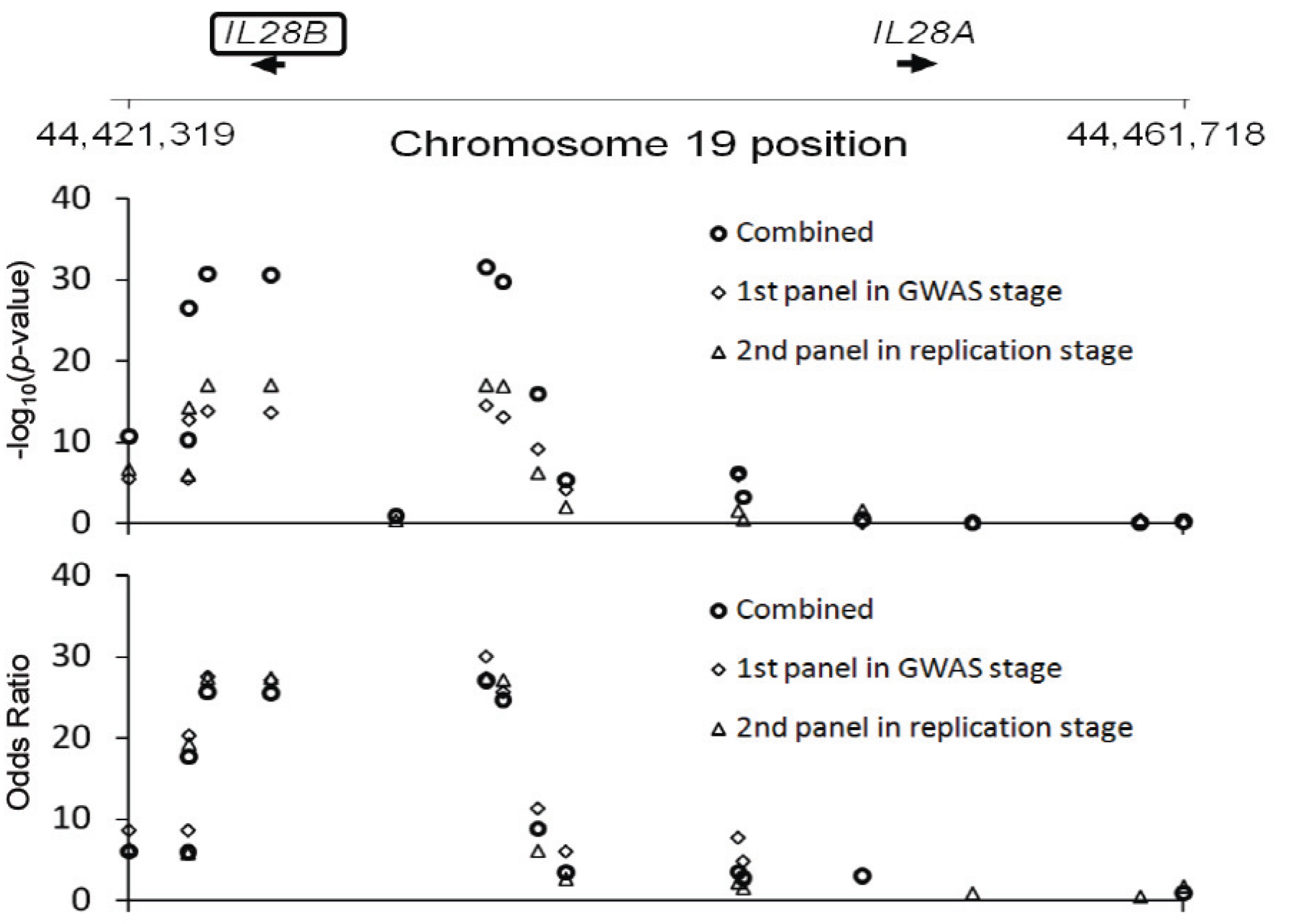
3.2. Identification of New Therapeutic Targets
4. Particular Importance of HLA
4.1. Immune-Mediated Diseases and HLA
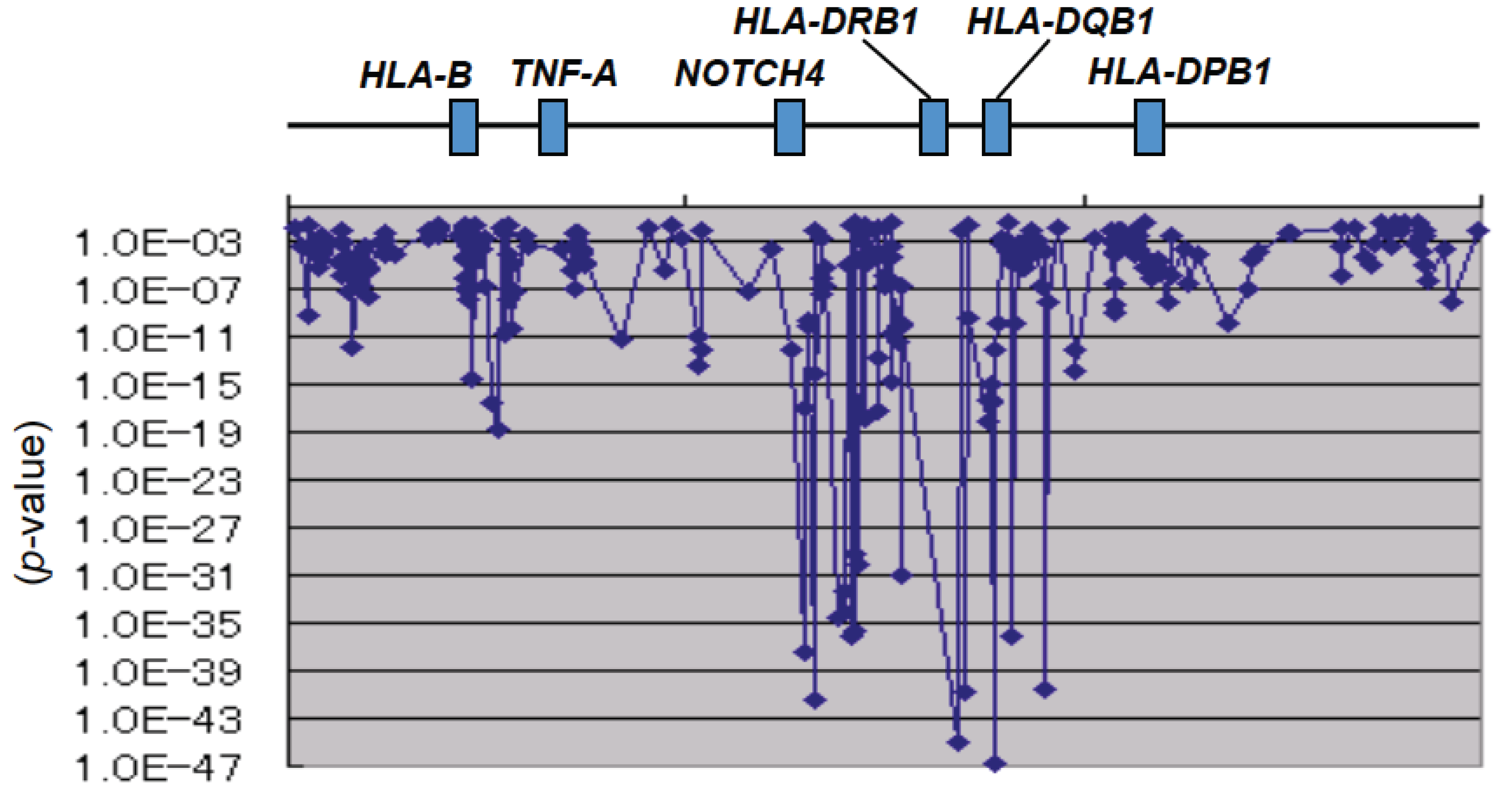
4.2. Drug Hypersensitivity and HLA
4.3. Characteristics of HLA and the Importance of HLA Typing
5. Conclusions and Issues for the Future
Acknowledgments
Conflicts of Interest
References
- Risch, N. Searching for genetic determinants in the millennium. Nature 2000, 405, 847–856. [Google Scholar] [CrossRef]
- Ozaki, K.; Ohnishi, Y.; Iida, A.; Sekine, A.; Yamada, R.; Tsunoda, T.; Sato, H.; Hori, M.; Nakamura, Y.; Tanaka, T. Functional SNPs in the lymphotoxin-alpha gene that are associated with susceptibility to myocardial infarction. Nat. Genet. 2002, 32, 650–654. [Google Scholar] [CrossRef]
- Tamiya, G.; Shinya, M.; Imanishi, T.; Ikuta, T.; Makino, S.; Okamoto, K.; Furugaki, K.; Matsumoto, T.; Mano, S.; Ando, S.; et al. Whole genome association study of rheumatoid arthritis using 27039 microsatellites. Hum. Mol. Genet. 2005, 14, 2305–2321. [Google Scholar] [CrossRef]
- Database of Single Nucleotide Polymorphisms (dbSNP). Available online: http://www.ncbi.nlm.nih.gov/snp/ (accessed on 11 February 2014).
- Database of HapMap Project. Available online: http://hapmap.ncbi.nlm.nih.gov/ (accessed on 11 February 2014).
- Database of 1000 Genomes Project. Available online: http://www.1000genomes.org/ (accessed on 11 February 2014).
- A Catalog of Published Genome-Wide Association Studies. Available online: http://www.genome.gov/gwastudies/ (accessed on 11 February 2014).
- Sladek, R.; Rocheleau, G.; Rung, J.; Dina, C.; Shen, L.; Serre, D.; Boutin, P.; Vincent, D.; Belisle, A.; Hadjadj, S.; et al. Genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 2007, 445, 881–885. [Google Scholar] [CrossRef]
- Saxena, R.; Voight, B.F.; Lyssenko, V.; Burtt, N.P.; de Bakker, P.I.; Chen, H.; Roix, J.J.; Kathiresan, S.; Hirschhorn, J.N.; Daly, M.J.; et al. Genome-wide association analysis identifies loci for type 2 diabetes and triglyceride levels. Science 2007, 316, 1331–1336. [Google Scholar] [CrossRef]
- Scott, L.J.; Mohlke, K.L.; Bonnycastle, L.L.; Willer, C.J.; Li, Y.; Duren, W.L.; Erdos, M.R.; Stringham, H.M; Chines, P.S.; Jackson, A.U.; et al. A genome-wide association study of type 2 diabetes in Finns detects multiple susceptibility variants. Science 2007, 316, 1341–1345. [Google Scholar] [CrossRef]
- Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3000 shared controls. Nature 2007, 447, 661–678. [Google Scholar] [CrossRef]
- Yasuda, K.; Miyake, K.; Horikawa, Y.; Hara, K.; Osawa, H.; Furuta, H.; Hirota, Y.; Mori, H.; Jonsson, A.; Sato, Y.; et al. Variants in KCNQ1 are associated with susceptibility to type 2 diabetes mellitus. Nat. Genet. 2008, 40, 1092–1097. [Google Scholar] [CrossRef]
- Unoki, H.; Takahashi, A.; Kawaguchi, T.; Hara, K.; Horikoshi, M.; Andersen, G.; Ng, D.P.; Holmkvist, J.; Borch-Johnsen, K.; Jørgensen, T.; et al. SNPs in KCNQ1 are associated with susceptibility to type 2 diabetes in East Asian and European populations. Nat. Genet. 2008, 40, 1098–1102. [Google Scholar] [CrossRef]
- Miyake, K.; Horikawa, Y.; Hara, K.; Yasuda, K.; Osawa, H.; Furuta, H.; Hirota, Y.; Yamagata, K.; Hinokio, Y.; Oka, Y.; et al. Association of TCF7L2 polymorphisms with susceptibility to type 2 diabetes in 4,087 Japanese subjects. J. Hum. Genet. 2008, 53, 174–180. [Google Scholar]
- Matsuki, K.; Juji, T.; Tokunaga, K.; Naohara, T.; Satake, M.; Honda, Y. Human histocompatibility leukocyte antigen (HLA) haplotype frequencies estimated from the data on HLA class I, II, and III antigens in 111 Japanese narcoleptics. J. Clin. Invest. 1985, 76, 2078–2083. [Google Scholar] [CrossRef]
- Miyagawa, T.; Kawashima, M.; Nishida, N.; Ohashi, J.; Kimura, R.; Fujimoto, A.; Shimada, M.; Morishita, S.; Shigeta, T.; Lin, L.; et al. Variant between CPT1B and CHKB associated with susceptibility to narcolepsy. Nat. Genet. 2008, 40, 1324–1328. [Google Scholar] [CrossRef]
- Mahasirimongkol, S.; Yanai, H.; Mushiroda, T.; Promphittayarat, W.; Wattanapokayakit, S.; Phromjai, J.; Yuliwulandari, R.; Wichukchinda, N.; Yowang, A.; Yamada, N.; et al. Genome-wide association studies of tuberculosis in Asians identify distinct at-risk locus foe young tuberculosis. J. Hum. Genet. 2012, 57, 363–367. [Google Scholar] [CrossRef]
- Okada, Y.; Terao, C.; Ikari, K.; Kochi, Y.; Ohmura, K.; Suzuki, A.; Kawaguchi, T.; Stahl, E.A.; Kurreeman, F.A.; Nishida, N.; et al. Meta-analysis of genome-wide association studies identifies multiple novel loci associated with rheumatoid arthritis in the Japanese population. Nat. Genet. 2012, 44, 511–516. [Google Scholar] [CrossRef]
- Takamoto, M.; Kaburaki, T.; Mabuchi, A.; Araie, M.; Amano, S.; Aihara, M.; Tomidokoro, A.; Iwase, A.; Mabuchi, F.; Kashiwagi, K.; et al. Common variants on chromosome 9q21 are associated with normal tension glaucoma. PLoS One 2012, 7, e40107. [Google Scholar] [CrossRef]
- Nakamura, M.; Nishida, N.; Kawashima, M.; Aiba, Y.; Tanaka, A.; Yasunami, M.; Nakamura, H.; Komori, A.; Nakamuta, M.; Zeniya, M.; et al. Genome-wide association study identifies TNFSF15 and POU2AF1 as susceptibility loci for primary biliary cirrhosis in the Japanese population. Am. J. Hum. Genet. 2012, 91, 721–728. [Google Scholar] [CrossRef]
- Okamoto, K.; Tokunaga, K.; Doi, K.; Fujita, T.; Suzuki, H.; Katoh, T.; Watanabe, T.; Nishida, N.; Mabuchi, A.; Takahashi, A.; et al. Common variation in GPC5 is associated with acquired nephrotic syndrome. Nat. Genet. 2011, 43, 459–463. [Google Scholar] [CrossRef]
- Cotsapas, C.; Voight, B.F.; Rossin, E.; Lage, K.; Neale, B.M.; Wallace, C.; Abecasis, G.R.; Barrett, J.C.; Behrens, T.; Cho, J.; et al. Pervasive sharing of genetic effects in autoimmune disease. PLoS Genet. 2011, 7, e1002254. [Google Scholar] [CrossRef] [Green Version]
- Parkes, M.; Cortes, A.; van Heel, D.A.; Brown, M.A. Genetic insights into common pathways and complex relationships among immune-mediated diseases. Nat. Rev. Genet. 2013, 14, 661–673. [Google Scholar]
- Miyagawa, T.; Miyadera, H.; Tanaka, S.; Kawashima, M.; Shimada, M.; Honda, Y.; Tokunaga, K.; Honda, M. Abnormally low serum acylcarnitine level in narcolepsy. Sleep 2011, 34, 349–353. [Google Scholar]
- Miyagawa, T.; Kawamura, H.; Obuchi, M.; Ikesaki, A.; Ozaki, A.; Tokunaga, K.; Inoue, Y.; Honda, M. Effects of oral L-carnitine administration in narcolepsy patients: A randomized, double-blind, cross-over and placebo-controlled trial. PLoS One 2013, 8, e53707. [Google Scholar]
- Hallmayer, J.; Faraco, J.; Lin, L.; Hesselson, S.; Winkelmann, J.; Kawashima, M.; Mayer, G.; Plazzi, G.; Nevsimalova, S.; Bourgin, P.; et al. Narcolepsy is strongly associated with the TCR alpha locus. Nat. Genet. 2009, 41, 708–711. [Google Scholar] [CrossRef]
- Kornum, B.R.; Kawashima, M.; Faraco, J.; Lin, L.; Rico, T.J.; Hesselson, S.; Axtell, R.C.; Kuipers, H.; Weiner, K.; Hamacher, A.; et al. Common variants in P2RY11 are associated with narcolepsy. Nat. Genet. 2011, 43, 66–71. [Google Scholar] [CrossRef]
- Tanaka, Y.; Nishida, N.; Sugiyama, M.; Kurosaki, M.; Matsuura, K.; Sakamoto, N.; Nakagawa, M.; Korenaga, M.; Hino, K.; Hige, S.; et al. Genome-wide association of IL28B with response to pegylated interferon-alpha and ribavirin therapy for chronic hepatitis C. Nat. Genet. 2009, 41, 1105–1109. [Google Scholar] [CrossRef]
- Nishida, N.; Sawai, H.; Matsuura, K.; Sugiyama, M.; Ahn, S.H.; Park, J.Y.; Hige, S.; Kang, J.H.; Suzuki, K.; Kurosaki, M.; et al. Genome-wide association study confirming association of HLA-DP with protection against chronic hepatitis B and viral clearance in Japanese and Korean. PLoS One 2012, 7, e39175. [Google Scholar] [CrossRef]
- Juji, T.; Satake, M.; Honda, Y.; Doi, Y. HLA antigens in Japanese patients with narcolepsy. All patients were DR2 positive. Tissue Antigens 1984, 24, 316–319. [Google Scholar]
- Hirschfield, G.M.; Liu, X.; Xu, C.; Lu, Y.; Xie, G.; Lu, Y.; Gu, X.; Walker, E.J.; Jing, K.; Juran, B.D.; et al. Primary biliary cirrhosis associated with HLA, IL12A, and IL12RB2 variants. N. Engl. J. Med. 2009, 360, 2544–2555. [Google Scholar] [CrossRef]
- Nakamura, M.; Yasunami, M.; Kondo, H.; Horie, H.; Aiba, Y.; Komori, A.; Migita, K.; Yatsuhashi, H.; Ito, M.; Shimoda, S.; et al. Analysis of HLA-DRB1 polymorphisms in Japanese patients with primary biliary cirrhosis (PBC): The HLA-DRB1 polymorphism determines the relative risk of antinuclear antibodies for disease progression in PBC. Hepatol. Res. 2010, 40, 494–504. [Google Scholar]
- Begovich, A.B.; Klitz, W.; Moonsamy, P.V.; van de Water, J.; Peltz, G.; Gershwin, M.E. Genes within the HLA class II region confer both predisposition and resistance to primary biliary cirrhosis. Tissue Antigens 1994, 43, 71–77. [Google Scholar] [CrossRef]
- Mallal, S.; Nolan, D.; Witt, C.; Masel, G.; Martin, A.M.; Moore, C.; Sayer, D.; Castley, A.; Mamotte, C.; Maxwell, D.; et al. Association between presence of HLA-B*5701, HLA-DR7 and HLA-DQ3 and hypersensitivity to HIV-1 reverse-transcriptase inhibitor abacavir. Lancet 2002, 359, 727–732. [Google Scholar] [CrossRef]
- Chung, W.H.; Hung, S.I.; Hong, H.S.; Hsih, M.S.; Yang, L.C.; Ho, H.C.; Wu, J.Y.; Chen, Y.T. Medical genetics: A marker for Stevens-Johnson syndrome. Nature 2004, 428, 486. [Google Scholar]
- Ozeki, T.; Mushiroda, T.; Yowang, A.; Takahashi, A.; Kubo, M.; Shirakata, Y.; Ikezawa, Z.; Iijima, M.; Shiohara, T.; Hashimoto, K.; et al. Genome-wide association study identifies HLA-A*3101 allele as a genetic risk factor for carbamazepine-induced cutaneous adverse drug reactions in Japanese population. Hum. Mol. Genet. 2011, 20, 1034–1041. [Google Scholar] [CrossRef]
- Ueta, M.; Tokunaga, K.; Sotozono, C.; Inatomi, T.; Yabe, T.; Matsushita, M.; Mitsuishi, Y.; Kinoshita, S. HLA class I and II gene polymorphisms in Stevens-Johnson syndrome with ocular complications in Japanese. Mol. Vis. 2008, 14, 550–555. [Google Scholar]
- Dilthey, A.; Leslie, S.; Moutsianas, L.; Shen, J.; Cox, C.; Nelson, M.R.; McVean, G. Multi-population classical HLA type imputation. PLoS Comput. Biol. 2013, 9, e1002877. [Google Scholar] [CrossRef]
- Zheng, X.; Shen, J.; Cox, C.; Wakefield, J.C.; Ehm, M.G.; Nelson, M.R.; Weir, B.S. HIBAG-HLA genotype imputation with attribute bagging. Pharmacogenomics J. 2013. [Google Scholar] [CrossRef]
- Jia, X.; Han, B.; Onengut-Gumuscu, S.; Chen, W.M.; Concannon, P.J.; Rich, S.S.; Raychaudhuri, S.; de Bakker, P.I. Imputing amino acid polymorphisms in human leucocyte antigens. PLoS One 2013, 8, e64683. [Google Scholar]
- Maher, B. Personal genomes: The case of the missing heritability. Nature 2008, 456, 18–21. [Google Scholar] [CrossRef]
- Dinu, I.; Mahasirimongkol, S.; Liu, Q.; Yanai, H.; El-Din, N.S.; Kreiter, E.; Wu, X.; Jabbari, S.; Tokunaga, K.; Yasui, Y. SNP-SNP interactions discovered by logic regression explain Crohn’s disease genetics. PLoS One 2012, 7, e43035. [Google Scholar] [CrossRef]
- Database of Genotypes and Phenotypes (dbGaP). Available online: http://www.ncbi.nlm.nih.gov/gap/ (accessed on 11 February 2014).
- European Genome-Phenome Archive (EGA). Available online: https://www.ebi.ac.uk/ega/ (accessed on 11 February 2014).
- GWAS Central. Available online: https://www.gwascentral.org/ (accessed on 11 February 2014).
- Koike, A.; Nishida, N.; Inoue, I.; Tsuji, S.; Tokunaga, K. Genome-wide association database developed in the Japanese Integrated Database Project. J. Hum. Genet. 2009, 54, 543–546. [Google Scholar] [CrossRef]
- Koike, A.; Nishida, N.; Yamashita, D.; Tokunaga, K. Comparative analysis of copy number variation detection methods and database construction. BMC Genet. 2011, 12, e29. [Google Scholar]
- Human Genome Variation Database. Available online: https://gwas.biosciencedbc.jp/index.html/ (accessed on 11 February 2014).
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Tokunaga, K. Lessons from Genome-Wide Search for Disease-Related Genes with Special Reference to HLA-Disease Associations. Genes 2014, 5, 84-96. https://doi.org/10.3390/genes5010084
Tokunaga K. Lessons from Genome-Wide Search for Disease-Related Genes with Special Reference to HLA-Disease Associations. Genes. 2014; 5(1):84-96. https://doi.org/10.3390/genes5010084
Chicago/Turabian StyleTokunaga, Katsushi. 2014. "Lessons from Genome-Wide Search for Disease-Related Genes with Special Reference to HLA-Disease Associations" Genes 5, no. 1: 84-96. https://doi.org/10.3390/genes5010084




