Journal Description
Pathogens
Pathogens
is an international, peer-reviewed, open access journal on pathogens and pathogen-host interactions published monthly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, PMC, Embase, PubAg, CaPlus / SciFinder, AGRIS, and other databases.
- Journal Rank: JCR - Q2 (Microbiology) / CiteScore - Q2 (General Immunology and Microbiology)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 16.4 days after submission; acceptance to publication is undertaken in 2.6 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Companion journals for Pathogens include: Parasitologia, Bacteria and Zoonotic Diseases.
Impact Factor:
3.7 (2022);
5-Year Impact Factor:
3.7 (2022)
Latest Articles
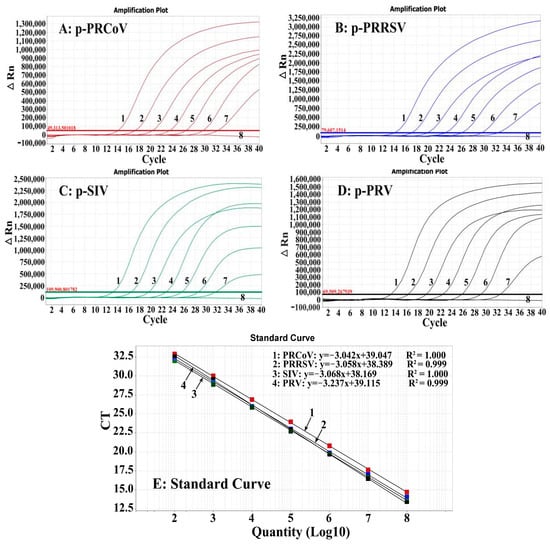
Simultaneous Detection of Porcine Respiratory Coronavirus, Porcine Reproductive and Respiratory Syndrome Virus, Swine Influenza Virus, and Pseudorabies Virus via Quadruplex One-Step RT-qPCR
Pathogens 2024, 13(4), 341; https://doi.org/10.3390/pathogens13040341 - 19 Apr 2024
Abstract
Porcine respiratory coronavirus (PRCoV), porcine reproductive and respiratory syndrome virus (PRRSV), swine influenza virus (SIV), and pseudorabies virus (PRV) are significant viruses causing respiratory diseases in pigs. Sick pigs exhibit similar clinical symptoms such as fever, cough, runny nose, and dyspnea, making it
[...] Read more.
Porcine respiratory coronavirus (PRCoV), porcine reproductive and respiratory syndrome virus (PRRSV), swine influenza virus (SIV), and pseudorabies virus (PRV) are significant viruses causing respiratory diseases in pigs. Sick pigs exhibit similar clinical symptoms such as fever, cough, runny nose, and dyspnea, making it very difficult to accurately differentially diagnose these diseases on site. In this study, a quadruplex one-step reverse-transcription real-time quantitative PCR (RT-qPCR) for the detection of PRCoV, PRRSV, SIV, and PRV was established. The assay showed strong specificity, high sensitivity, and good repeatability. It could detect only PRCoV, PRRSV, SIV, and PRV, without cross-reactions with TGEV, PEDV, PRoV, ASFV, FMDV, PCV2, PDCoV, and CSFV. The limits of detection (LODs) for PRCoV, PRRSV, SIV, and PRV were 129.594, 133.205, 139.791, and 136.600 copies/reaction, respectively. The intra-assay and inter-assay coefficients of variation (CVs) ranged from 0.29% to 1.89%. The established quadruplex RT-qPCR was used to test 4909 clinical specimens, which were collected in Guangxi Province, China, from July 2022 to September 2023. PRCoV, PRRSV, SIV, and PRV showed positivity rates of 1.36%, 10.17%, 4.87%, and 0.84%, respectively. In addition, the previously reported RT-qPCR was also used to test these specimens, and the agreement between these methods was higher than 99.43%. The established quadruplex RT-qPCR can accurately detect these four porcine respiratory viruses simultaneously, providing an accurate and reliable detection technique for clinical diagnosis.
Full article
(This article belongs to the Special Issue Veterinary Viral Infections and Host Immune Responses)
►
Show Figures
Open AccessArticle
Descriptive Epidemiology of Pathogens Associated with Acute Respiratory Infection in a Community-Based Study of K–12 School Children (2015–2023)
by
Cristalyne Bell, Maureen Goss, Derek Norton, Shari Barlow, Emily Temte, Cecilia He, Caroline Hamer, Sarah Walters, Alea Sabry, Kelly Johnson, Guanhua Chen, Amra Uzicanin and Jonathan Temte
Pathogens 2024, 13(4), 340; https://doi.org/10.3390/pathogens13040340 - 19 Apr 2024
Abstract
School-based outbreaks often precede increased incidence of acute respiratory infections in the greater community. We conducted acute respiratory infection surveillance among children to elucidate commonly detected pathogens in school settings and their unique characteristics and epidemiological patterns. The ORegon CHild Absenteeism due to
[...] Read more.
School-based outbreaks often precede increased incidence of acute respiratory infections in the greater community. We conducted acute respiratory infection surveillance among children to elucidate commonly detected pathogens in school settings and their unique characteristics and epidemiological patterns. The ORegon CHild Absenteeism due to Respiratory Disease Study (ORCHARDS) is a longitudinal, laboratory-supported, school-based, acute respiratory illness (ARI) surveillance study designed to evaluate the utility of cause-specific student absenteeism monitoring for early detection of increased activity of influenza and other respiratory viruses in schools from kindergarten through 12th grade. Eligible participants with ARIs provided demographic, epidemiologic, and symptom data, along with a nasal swab or oropharyngeal specimen. Multipathogen testing using reverse-transcription polymerase chain reaction (RT-PCR) was performed on all specimens for 18 respiratory viruses and 2 atypical bacterial pathogens (Chlamydia pneumoniae and Mycoplasma pneumoniae). Between 5 January 2015 and 9 June 2023, 3498 children participated. Pathogens were detected in 2455 of 3498 (70%) specimens. Rhinovirus/enteroviruses (36%) and influenza viruses A/B (35%) were most commonly identified in positive specimens. Rhinovirus/enteroviruses and parainfluenza viruses occurred early in the academic year, followed by seasonal coronaviruses, RSV, influenza viruses A/B, and human metapneumovirus. Since its emergence in 2020, SARS-CoV-2 was detected year-round and had a higher median age than the other pathogens. A better understanding of the etiologies, presentations, and patterns of pediatric acute respiratory infections can help inform medical and public health system responses.
Full article
(This article belongs to the Special Issue Recent Advances in Pediatric Infectious Diseases)
►▼
Show Figures
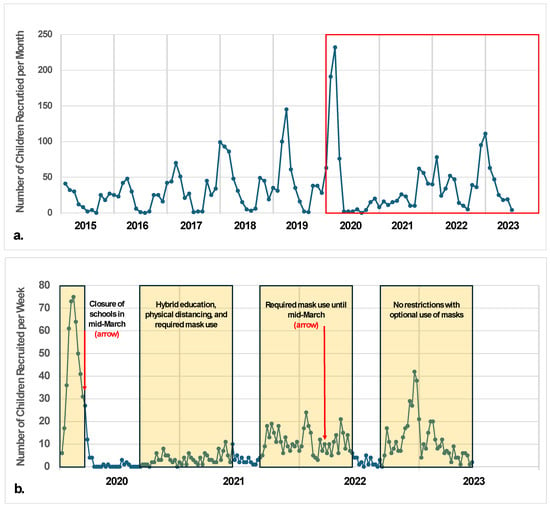
Figure 1
Open AccessReview
Chronic Hepatitis C Virus Infection, Extrahepatic Disease and the Impact of New Direct-Acting Antivirals
by
Nahum Méndez-Sánchez, Carlos E. Coronel-Castillo and Mariana Michelle Ramírez-Mejía
Pathogens 2024, 13(4), 339; https://doi.org/10.3390/pathogens13040339 - 19 Apr 2024
Abstract
Chronic hepatitis C virus infection is an important cause of liver cirrhosis, hepatocellular carcinoma and death. Furthermore, it is estimated that about 40–70% of patients develop non-hepatic alterations in the course of chronic infection. Such manifestations can be immune-related conditions, lymphoproliferative disorders and
[...] Read more.
Chronic hepatitis C virus infection is an important cause of liver cirrhosis, hepatocellular carcinoma and death. Furthermore, it is estimated that about 40–70% of patients develop non-hepatic alterations in the course of chronic infection. Such manifestations can be immune-related conditions, lymphoproliferative disorders and metabolic alterations with serious adverse events in the short and long term. The introduction of new Direct-Acting Antivirals has shown promising results, with current evidence indicating an improvement and remission of these conditions after a sustained virological response.
Full article
(This article belongs to the Special Issue Elimination Strategies for Viral Hepatitis in Latin America)
►▼
Show Figures
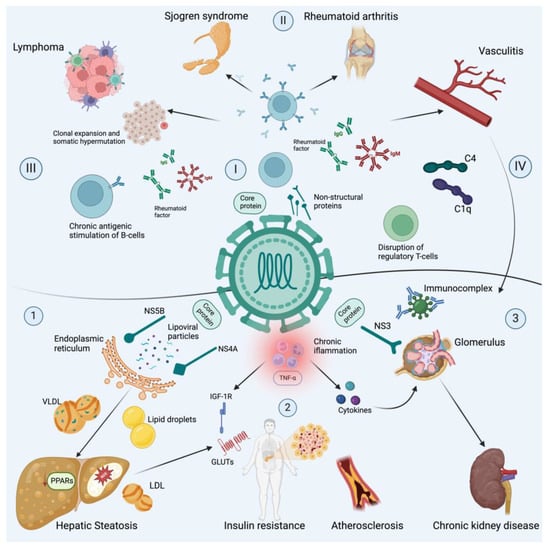
Figure 1
Open AccessArticle
Higher Prevalence of the Periodontal Pathogen Selenomonas noxia among Pediatric and Adult Patients May Be Associated with Overweight and Obesity
by
Austin Williams, Jace Porter, Karl Kingsley and Katherine M. Howard
Pathogens 2024, 13(4), 338; https://doi.org/10.3390/pathogens13040338 - 19 Apr 2024
Abstract
New evidence has suggested that oral and gut microflora may have significant impacts on the predisposition, development, and stability of obesity in adults over time—although less is known about this phenomenon in children. Compared with healthy-weight controls, overweight and obese adult patients are
[...] Read more.
New evidence has suggested that oral and gut microflora may have significant impacts on the predisposition, development, and stability of obesity in adults over time—although less is known about this phenomenon in children. Compared with healthy-weight controls, overweight and obese adult patients are now known to harbor specific pathogens, such as Selenomonas noxia (S. noxia), that are capable of digesting normally non-digestible cellulose and fibers that significantly increase caloric extraction from normal dietary intake. To evaluate this phenomenon, clinical saliva samples (N = 122) from subjects with a normal BMI (18–25) and a BMI over 25 (overweight, obese) from an existing biorepository were screened using qPCR. The prevalence of S. noxia in samples from normal-BMI participants were lower (21.4%) than in overweight-BMI (25–29; 46.1%) and obese-BMI (30 and above; 36.8%) samples—a strong, positive correlation that was not significantly affected by age or race and ethnicity. These data strongly suggest that S. noxia may be intricately associated with overweight and obesity among patients, and more research will be needed to determine the positive and negative feedback mechanisms that may be responsible for these observations as well as the interventions needed to remove or reduce the potential effects of this oral pathogen.
Full article
(This article belongs to the Special Issue Oral Microbiome and Human Systemic Health)
►▼
Show Figures
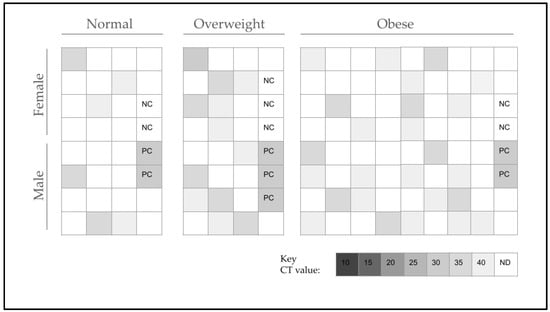
Figure 1
Open AccessArticle
Isospora and Lankesterella Parasites (Eimeriidae, Apicomplexa) of Passeriform Birds in Europe: Infection Rates, Phylogeny, and Pathogenicity
by
Carina Keckeisen, Alžbeta Šujanová, Tanja Himmel, Julia Matt, Nora Nedorost, Carolina R. F. Chagas, Herbert Weissenböck and Josef Harl
Pathogens 2024, 13(4), 337; https://doi.org/10.3390/pathogens13040337 - 18 Apr 2024
Abstract
Wild birds are common hosts to numerous intracellular parasites such as single-celled eukaryotes of the family Eimeriidae (order Eucoccidiorida, phylum Apicomplexa). We investigated the infection rates, phylogeny, and pathogenicity of Isospora and Lankesterella parasites in wild and captive passerine birds. Blood and tissue
[...] Read more.
Wild birds are common hosts to numerous intracellular parasites such as single-celled eukaryotes of the family Eimeriidae (order Eucoccidiorida, phylum Apicomplexa). We investigated the infection rates, phylogeny, and pathogenicity of Isospora and Lankesterella parasites in wild and captive passerine birds. Blood and tissue samples of 815 wild and 15 deceased captive birds from Europe were tested using polymerase chain reaction and partial sequencing of the mitochondrial cytochrome b and cytochrome c oxidase I and the nuclear 18S rRNA gene. The infection rate for Lankesterella in wild birds was 10.7% compared to 5.8% for Isospora. Chromogenic in situ hybridization with probes targeting the parasites’ 18S rRNA was employed to identify the parasites’ presence in multiple organs, and hematoxylin–eosin staining was performed to visualize the parasite stages and assess associated lesions. Isospora parasites were mainly identified in the intestine, spleen, and liver. Extraintestinal tissue stages of Isospora were accompanied by predominantly lymphohistiocytic inflammation of varying severity. Lankesterella was most frequently detected in the spleen, lung, and brain; however, infected birds presented only a low parasite burden without associated pathological changes. These findings contribute to our understanding of Isospora and Lankesterella parasites in wild birds.
Full article
(This article belongs to the Collection Pathology and Parasitic Diseases of Animals)
►▼
Show Figures
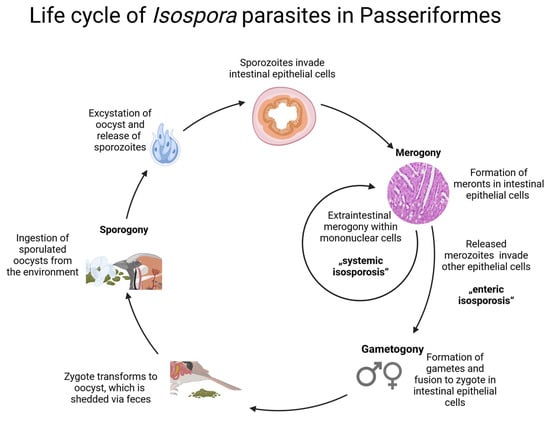
Figure 1
Open AccessCommunication
Pradofloxacin for Treatment of Bartonella henselae in Experimentally Inoculated Cats
by
Michael R. Lappin and Ronan Fitzgerald
Pathogens 2024, 13(4), 336; https://doi.org/10.3390/pathogens13040336 - 18 Apr 2024
Abstract
Bartonella henselae is associated with numerous clinical syndromes in people. Cats are the definitive hosts for B. henselae, develop high levels of bacteremia, and are associated with human infections, particularly in the presence of Ctenocephalides felis. Several antibiotic protocols used for
[...] Read more.
Bartonella henselae is associated with numerous clinical syndromes in people. Cats are the definitive hosts for B. henselae, develop high levels of bacteremia, and are associated with human infections, particularly in the presence of Ctenocephalides felis. Several antibiotic protocols used for the treatment of B. henselae infection in cats have failed to clear bacteremia. The purpose of this study was to assess the safety and efficacy of a high-dose pradofloxacin protocol to eliminate B. henselae bacteremia. Bartonella henselae infection was initiated in 8 cats by intravenous inoculation of infected feline blood and then pradofloxacin was administered at 7.5 mg/kg, PO, twice daily for 28 days, starting 12 weeks after inoculation. Complete blood cell counts were performed prior to pradofloxacin administration and then every 2 weeks for 10 weeks. Bartonella PCR assay was performed prior to pradofloxacin administration and approximately every 2 weeks for 10 weeks and then weekly for 4 weeks. Methylprednisolone acetate (5 mg/kg) was administered by intramuscular injection to all cats on week 10. The cats remained normal and none developed a hematocrit, platelet count, lymphocyte count, or neutrophil count outside of the normal reference ranges. In the one month prior to pradofloxacin administration, all cats were PCR-positive for Bartonella DNA on at least two of four sample dates; after pradofloxacin administration, all cats were negative for B. henselae DNA in blood on all nine sample dates. The protocol appears to be safe and failure to amplify B. henselae DNA from the blood after the administration of pradofloxacin and one dose of methylprednisolone acetate suggests either an antibiotic effect or the organism was cleared spontaneously.
Full article
(This article belongs to the Special Issue The Expanding Clinical Spectrum of Bartonelloses)
Open AccessArticle
Development of a Targeted NGS Assay for the Detection of Respiratory Pathogens including SARS-CoV-2 in Felines
by
Jobin J. Kattoor, Mothomang Mlalazi-Oyinloye, Sarah M. Nemser and Rebecca P. Wilkes
Pathogens 2024, 13(4), 335; https://doi.org/10.3390/pathogens13040335 - 17 Apr 2024
Abstract
Acute respiratory diseases in felines can be attributed to a diverse range of pathogens. The recent emergence of novel viruses, particularly SARS-CoV-2 and its variants, has also been associated with respiratory ailments in cats and other pets, underscoring the need for a highly
[...] Read more.
Acute respiratory diseases in felines can be attributed to a diverse range of pathogens. The recent emergence of novel viruses, particularly SARS-CoV-2 and its variants, has also been associated with respiratory ailments in cats and other pets, underscoring the need for a highly sensitive diagnostic assay capable of concurrently detecting multiple respiratory pathogens. In this study, we developed a targeted next generation sequencing panel using Ion Torrent Ampliseq technology to detect multiple respiratory pathogens, including recent SARS-CoV-2 variants and Feline herpesvirus-1, Feline calicivirus, Bordetella bronchiseptica, Mycoplasmopsis (previously Mycoplasma) felis, and Chlamydia felis. A PCR amplification-based library preparation, employing primers designed for pathogen target regions, was synthesized and divided into two pools, followed by sequencing and assembly to a repertoire of target pathogen genomes. Analytical sensitivity was assessed based on Ct values from real-time PCR for the corresponding pathogens, indicating an equivalent detection limit. Most of the pathogens under study were positively identified to a limit of approximately Ct 36, whereas for Feline herpesvirus-1 and SARS-CoV-2, positive reads were observed in samples with a Ct of 37. Based on a limited number of samples, the diagnostic sensitivity values for the SARS-CoV-2, Feline herpesvirus-1, and M. felis samples were 100% with no false negative results. The diagnostic specificity of SARS-CoV-2, Feline herpesvirus-1, Feline calicivirus, and C. felis were 100%. Importantly, none of the target primers exhibited non-specific amplification, ensuring the absence of false positive results for other pathogens within the study. Additionally, the assay’s specificity was validated by cross-referencing the raw sequencing data with established databases like BLAST, affirming the high specificity of the targeted Next-Generation Sequencing (tNGS) assay. Variations in the sequencing reads of different pathogens were observed when subjected to diverse extraction methods. Rigorous assessment of the assay’s reliability involved reproducibility across testing personnel and repeated runs. The developed assay’s clinical applicability was tested using samples submitted to the diagnostic laboratory from cat shelters and suspected cases. The developed targeted next-generation sequencing methodology empowers the detection of multiple respiratory pathogens manifesting similar clinical symptoms while offering confirmation of results through genome sequencing.
Full article
(This article belongs to the Special Issue Diagnostics of Animal Viral Infectious Diseases)
Open AccessArticle
Phylodynamic and Evolution of the Hemagglutinin (HA) and Neuraminidase (NA) Genes of Influenza A(H1N1) pdm09 Viruses Circulating in the 2009 and 2023 Seasons in Italy
by
Fabio Scarpa, Leonardo Sernicola, Stefania Farcomeni, Alessandra Ciccozzi, Daria Sanna, Marco Casu, Marco Vitale, Alessia Cicenia, Marta Giovanetti, Chiara Romano, Francesco Branda, Massimo Ciccozzi and Alessandra Borsetti
Pathogens 2024, 13(4), 334; https://doi.org/10.3390/pathogens13040334 - 17 Apr 2024
Abstract
The influenza A(H1N1) pdm09 virus, which emerged in 2009, has been circulating seasonally since then. In this study, we conducted a comprehensive genome-based investigation to gain a detailed understanding of the genetic and evolutionary characteristics of the hemagglutinin (HA) and neuraminidase (NA) surface
[...] Read more.
The influenza A(H1N1) pdm09 virus, which emerged in 2009, has been circulating seasonally since then. In this study, we conducted a comprehensive genome-based investigation to gain a detailed understanding of the genetic and evolutionary characteristics of the hemagglutinin (HA) and neuraminidase (NA) surface proteins of A/H1N1pdm09 strains circulating in Italy over a fourteen-year period from 2009 to 2023 in relation to global strains. Phylogenetic analysis revealed rapid transmission and diversification of viral variants during the early pandemic that clustered in clade 6B.1. In contrast, limited genetic diversity was observed during the 2023 season, probably due to the genetic drift, which provides the virus with a constant adaptability to the host; furthermore, all isolates were split into two main groups representing two clades, i.e., 6B.1A.5a.2a and its descendant 6B.1A.5a.2a.1. The HA gene showed a faster rate of evolution compared to the NA gene. Using FUBAR, we identified positively selected sites 41 and 177 for HA and 248, 286, and 455 for NA in 2009, as well as sites 22, 123, and 513 for HA and 339 for NA in 2023, all of which may be important sites related to the host immune response. Changes in glycosylation acquisition/loss at prominent sites, i.e., 177 in HA and 248 in NA, should be considered as a predictive tool for early warning signs of emerging pandemics, and for vaccine and drug development.
Full article
(This article belongs to the Special Issue Advance in Influenza A and Influenza B Viruses)
►▼
Show Figures
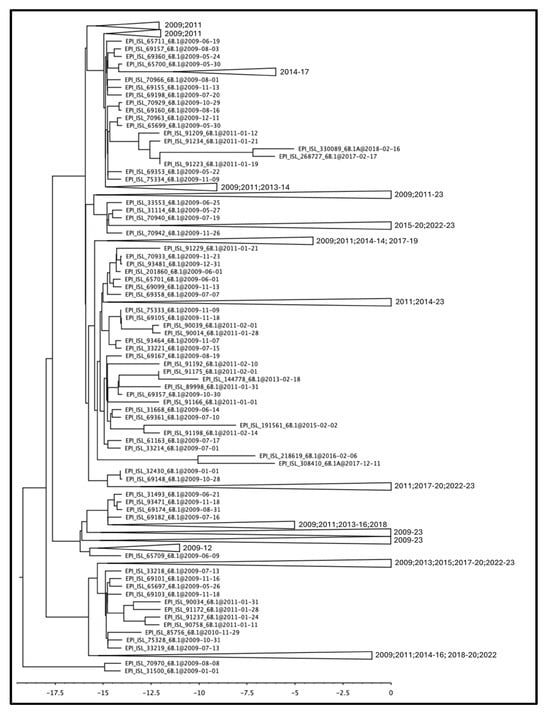
Figure 1
Open AccessArticle
Molecular Characterization of Non-H5 and Non-H7 Avian Influenza Viruses from Non-Mallard Migratory Waterbirds of the North American Flyways, 2006–2011
by
Shahan Azeem, John Baroch, Deepanker Tewari, Kristy L. Pabilonia, Mary Killian, Birgit Bradel-Tretheway, Dong Sun, Sara Ghorbani-Nezami and Kyoung-Jin Yoon
Pathogens 2024, 13(4), 333; https://doi.org/10.3390/pathogens13040333 - 17 Apr 2024
Abstract
The surveillance of migratory waterbirds (MWs) for avian influenza virus (AIV) is indispensable for the early detection of a potential AIV incursion into poultry. Surveying AIV infections and virus subtypes in understudied MW species could elucidate their role in AIV ecology. Oropharyngeal–cloacal (OPC)
[...] Read more.
The surveillance of migratory waterbirds (MWs) for avian influenza virus (AIV) is indispensable for the early detection of a potential AIV incursion into poultry. Surveying AIV infections and virus subtypes in understudied MW species could elucidate their role in AIV ecology. Oropharyngeal–cloacal (OPC) swabs were collected from non-mallard MWs between 2006 and 2011. OPC swabs (n = 1158) that molecularly tested positive for AIV (Cts ≤ 32) but tested negative for H5 and H7 subtypes were selected for virus isolation (VI). The selected samples evenly represented birds from all four North American flyways (Pacific, Central, Mississippi, and Atlantic). Eighty-seven low pathogenic AIV isolates, representing 31 sites in 17 states, were recovered from the samples. All isolates belonged to the North American lineage. The samples representing birds from the Central Flyway had the highest VI positive rate (57.5%) compared to those from the other flyways (10.3–17.2%), suggesting that future surveillance can focus on the Central Flyway. Of the isolates, 43.7%, 12.6%, and 10.3% were obtained from blue-winged teal, American wigeon, and American black duck species, respectively. Hatch-year MWs represented the majority of the isolates (70.1%). The most common H and N combinations were H3N8 (23.0%), H4N6 (18.4%), and H4N8 (18.4%). The HA gene between non-mallard and mallard MW isolates during the same time period shared 85.5–99.5% H3 identity and 89.3–99.7% H4 identity. Comparisons between MW (mallard and non-mallard) and poultry H3 and H4 isolates also revealed high similarity (79.0–99.0% and 88.7–98.4%), emphasizing the need for continued AIV surveillance in MWs.
Full article
(This article belongs to the Special Issue Pathogenesis, Epidemiology, and Control of Animal Influenza Viruses)
►▼
Show Figures
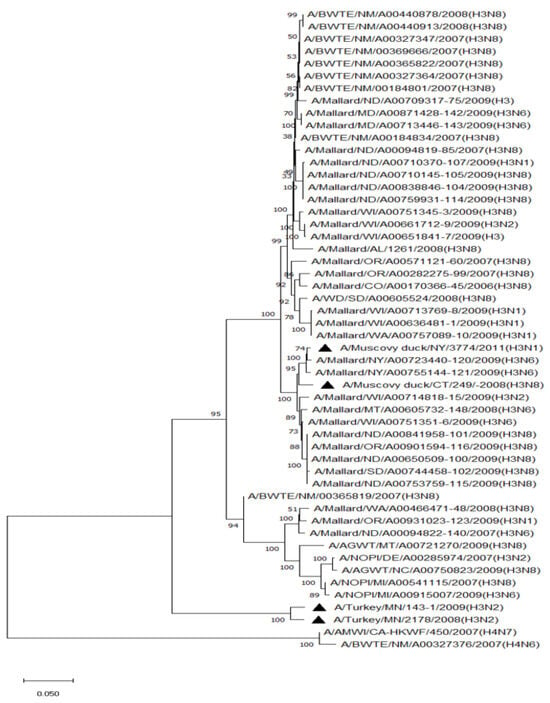
Figure 1
Open AccessReview
Dengue-Associated Hemophagocytic Lymphohistiocytosis: A Narrative Review of Its Identification and Treatment
by
Kay Choong See
Pathogens 2024, 13(4), 332; https://doi.org/10.3390/pathogens13040332 - 17 Apr 2024
Abstract
Dengue’s lack of specific treatments beyond supportive care prompts a focus on uncovering additional pathophysiological factors. Dengue-associated hemophagocytic lymphohistiocytosis (HLH), characterized by dysregulated macrophage activation and cytokine storm, remains underexplored despite its potential to worsen disease severity and mortality. While rare, dengue-associated HLH
[...] Read more.
Dengue’s lack of specific treatments beyond supportive care prompts a focus on uncovering additional pathophysiological factors. Dengue-associated hemophagocytic lymphohistiocytosis (HLH), characterized by dysregulated macrophage activation and cytokine storm, remains underexplored despite its potential to worsen disease severity and mortality. While rare, dengue-associated HLH disproportionately affects severe cases, significantly impacting mortality rates. To mitigate high mortality, early identification and familiarity with dengue-associated HLH are imperative for prompt treatment by clinicians. This narrative review therefore aims to examine the current clinical and therapeutic knowledge on dengue-associated HLH, and act as a resource for clinicians to improve their management of HLH associated with severe dengue. Dengue-associated HLH should be considered for all cases of severe dengue and may be suspected based on the presence of prolonged or recurrent fever for >7 days, or anemia without intravascular hemolysis or massive bleeding. Diagnosis relies on fulfilling at least five of the eight HLH-2004 criteria. Treatment predominantly involves short courses (3–4 days) of high-dose steroids (e.g., dexamethasone 10 mg/m2), with additional therapies considered in more severe presentations. Notably, outcomes can be favorable with steroid therapy alone.
Full article
(This article belongs to the Special Issue Impact of Bacterial and Viral Pathogens on Sepsis Prevalence and Outcome)
►▼
Show Figures
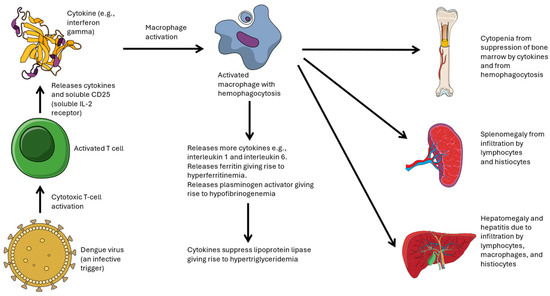
Figure 1
Open AccessReview
Biomarkers in Detection of Hepatitis C Virus Infection
by
Jungreem Woo and Youkyung Choi
Pathogens 2024, 13(4), 331; https://doi.org/10.3390/pathogens13040331 - 17 Apr 2024
Abstract
The hepatitis C virus (HCV) infection affects 58 million people worldwide. In the United States, the incidence rate of acute hepatitis C has doubled since 2014; during 2021, this increased to 5% from 2020. Acute hepatitis C is defined by any symptom of
[...] Read more.
The hepatitis C virus (HCV) infection affects 58 million people worldwide. In the United States, the incidence rate of acute hepatitis C has doubled since 2014; during 2021, this increased to 5% from 2020. Acute hepatitis C is defined by any symptom of acute viral hepatitis plus either jaundice or elevated serum alanine aminotransferase (ALT) activity with the detection of HCV RNA, the anti-HCV antibody, or hepatitis C virus antigen(s). However, most patients with acute infection are asymptomatic. In addition, ALT activity and HCV RNA levels can fluctuate, and a delayed detection of the anti-HCV antibody can occur among some immunocompromised persons with HCV infection. The detection of specific biomarkers can be of great value in the early detection of HCV infection at an asymptomatic stage. The high rate of HCV replication (which is approximately 1010 to 1012 virions per day) and the lack of proofreading by the viral RNA polymerase leads to enormous genetic diversity, creating a major challenge for the host immune response. This broad genetic diversity contributes to the likelihood of developing chronic infection, thus leading to the development of cirrhosis and liver cancer. Direct-acting antiviral (DAA) therapies for HCV infection are highly effective with a cure rate of up to 99%. At the same time, many patients with HCV infection are unaware of their infection status because of the mostly asymptomatic nature of hepatitis C, so they remain undiagnosed until the liver damage has advanced. Molecular mechanisms induced by HCV have been intensely investigated to find biomarkers for diagnosing the acute and chronic phases of the infection. However, there are no clinically verified biomarkers for patients with hepatitis C. In this review, we discuss the biomarkers that can differentiate acute from chronic hepatitis C, and we summarize the current state of the literature on the useful biomarkers that are detectable during acute and chronic HCV infection, liver fibrosis/cirrhosis, and hepatocellular carcinoma (HCC).
Full article
(This article belongs to the Special Issue Host Immune Responses to RNA Viruses, Volume II)
Open AccessArticle
MinION Sequencing of Fungi in Sub-Saharan African Air and a Novel LAMP Assay for Rapid Detection of the Tropical Phytopathogenic Genus Lasiodiplodia
by
Kevin M. King, Gail G. M. Canning and Jonathan S. West
Pathogens 2024, 13(4), 330; https://doi.org/10.3390/pathogens13040330 - 17 Apr 2024
Abstract
To date, there have been no DNA-based metabarcoding studies into airborne fungi in tropical Sub-Saharan Africa. In this initial study, 10 air samples were collected onto Vaseline-coated acrylic rods mounted on drones flown at heights of 15–50 meters above ground for 10–15 min
[...] Read more.
To date, there have been no DNA-based metabarcoding studies into airborne fungi in tropical Sub-Saharan Africa. In this initial study, 10 air samples were collected onto Vaseline-coated acrylic rods mounted on drones flown at heights of 15–50 meters above ground for 10–15 min at three sites in Ghana. Purified DNA was extracted from air samples, the internal transcribed spacer (ITS) region was amplified using fungal-specific primers, and MinION third-generation amplicon sequencing was undertaken with downstream bioinformatics analyses utilizing GAIA cloud-based software (at genus taxonomic level). Principal coordinate analyses based on Bray–Curtis beta diversity dissimilarity values found no clear evidence for the structuring of fungal air communities, nor were there significant differences in alpha diversity, based on geographic location (east vs. central Ghana), underlying vegetation type (cocoa vs. non-cocoa), or height above ground level (15–23 m vs. 25–50 m), and despite the short flight times (10–15 min), ~90 operational taxonomic units (OTUs) were identified in each sample. In Ghanaian air, fungal assemblages were skewed at the phylum taxonomic level towards the ascomycetes (53.7%) as opposed to basidiomycetes (24.6%); at the class level, the Dothideomycetes were predominant (29.8%) followed by the Agaricomycetes (21.8%). The most common fungal genus in Ghanaian air was cosmopolitan and globally ubiquitous Cladosporium (9.9% of reads). Interestingly, many fungal genera containing economically important phytopathogens of tropical crops were also identified in Ghanaian air, including Corynespora, Fusarium, and Lasiodiplodia. Consequently, a novel loop-mediated isothermal amplification (LAMP) assay, based on translation elongation factor-1α sequences, was developed and tested for rapid, sensitive, and specific detection of the fungal phytopathogenic genus Lasiodiplodia. Potential applications for improved tropical disease management are considered.
Full article
(This article belongs to the Special Issue Fungal Pathogens of Crops)
►▼
Show Figures
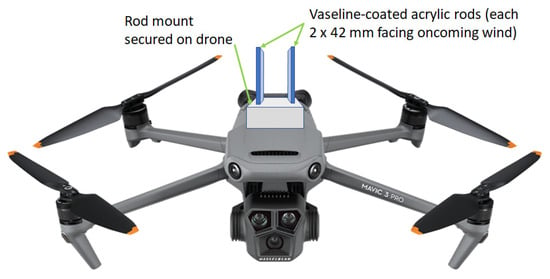
Figure 1
Open AccessArticle
A Set of Multiresistant Isolates of Mycoplasma bovis Subtype ST-1 with a Variable Susceptibility to Quinolones Are Also Circulating in Spain
by
Juan Carlos Corrales, Antonio Sánchez, Xóchitl Hernández, Joaquín Amores-Iniesta, Antón Esnal and Christian de la Fe
Pathogens 2024, 13(4), 329; https://doi.org/10.3390/pathogens13040329 - 16 Apr 2024
Abstract
Mycoplasma bovis (M. bovis) is one of the worldwide most important infectious agents involved in respiratory complex diseases (RCD). In Spain, the endemic presence of subtypes ST-2 and ST-3 with phenotypic differences linked to their susceptibility to fluoroquinolones opened the way
[...] Read more.
Mycoplasma bovis (M. bovis) is one of the worldwide most important infectious agents involved in respiratory complex diseases (RCD). In Spain, the endemic presence of subtypes ST-2 and ST-3 with phenotypic differences linked to their susceptibility to fluoroquinolones opened the way to develop control strategies focused on previous diagnosis of the subtype and the use of directed therapies when M. bovis were involved in RCD. Surprisingly, microbiological studies conducted during 2023 evidenced for the first time the presence of Spanish isolates of a new polC-subtype, previously classified as ST-1, recovered from calves with respiratory symptoms and pneumonia in different areas of the country (n = 16). Curiously, the minimum inhibitory concentration (MIC) to a panel of antimicrobials revealed phenotypic differences between these ST-1 isolates when using fluoroquinolones (FLQ). There is no geographical correlation between MIC profiles even for a set of 8 isolates recovered from different animals in the same flock. Sequencing of 4 genes (gyrA, gyrB, parC and parE) encoding quinolone resistance-determining regions (QRDR) evidenced the presence of accumulate mutations in 2 ST-1 isolates with high FLQ MICs, but not in all them (n = 3), thus suggesting that, as previously recorded for ST-2 isolates, other mechanisms should be involved in the acquisition of resistence to these antimicrobials. Additionally, as previously detected in the Spanish ST-2 and ST-3, subtype ST-1 isolates are also resistant to macrolides or lincosamides.
Full article
(This article belongs to the Special Issue Mycoplasmas in Respiratory Tract Infections of Cattle)
►▼
Show Figures
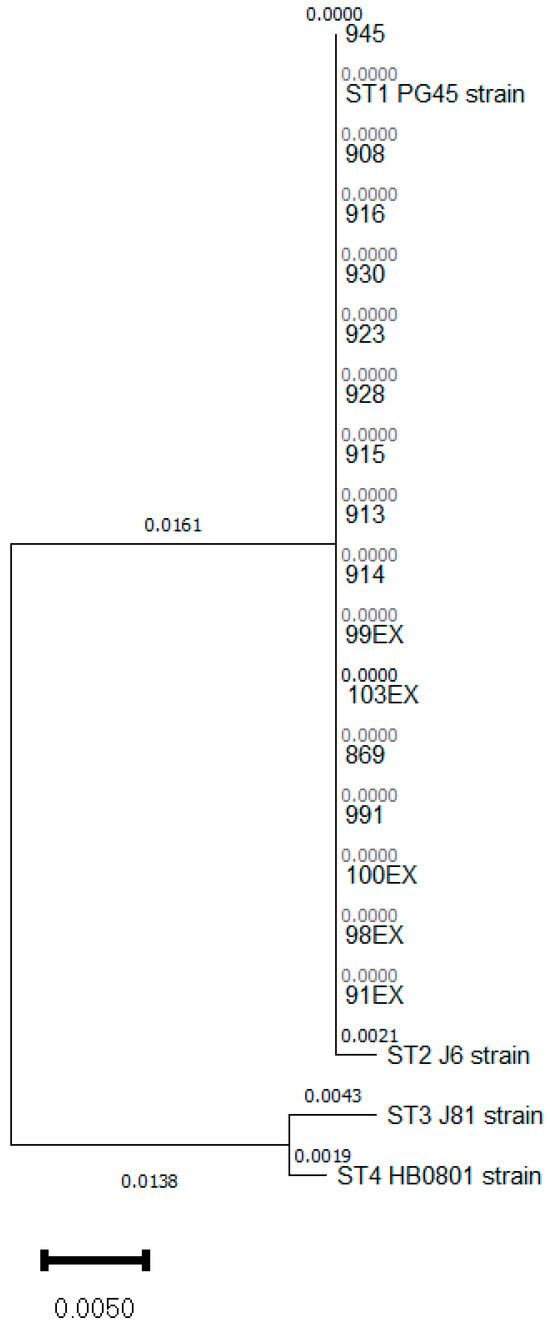
Figure 1
Open AccessCase Report
Mycobacteriosis in a Pet Ferret (Mustela putorius furo) Caused by Mycobacterium xenopi: A Case Report on Neglected Risk of Zoonotic Transmission
by
Željko Mihaljević, Irena Reil, Josipa Habuš, Zrinka Štritof, Šimun Naletilić, Gabrijela Jurkić Krsteska, Tajna Kovač, Maja Zdelar-Tuk, Sanja Duvnjak and Silvio Špičić
Pathogens 2024, 13(4), 328; https://doi.org/10.3390/pathogens13040328 - 16 Apr 2024
Abstract
Ferrets are highly susceptible to a wide range of mycobacteria, mainly M. bovis, M. avium, and M. triplex. Therefore, ferrets pose a risk of transmission of mycobacteriosis, especially zoonotically relevant tuberculosis. The aim of this study was to describe the
[...] Read more.
Ferrets are highly susceptible to a wide range of mycobacteria, mainly M. bovis, M. avium, and M. triplex. Therefore, ferrets pose a risk of transmission of mycobacteriosis, especially zoonotically relevant tuberculosis. The aim of this study was to describe the findings of M. xenopi mycobacteriosis in a pet ferret and emphasize its zoonotic potential. A pet ferret had a history of weight loss, apathy, hyporexia, and hair loss. Abdominal ultrasound revealed splenomegaly with two solid masses and cystic lesions of the liver. Fine-needle aspiration cytology revealed numerous acid-fast bacilli in epithelioid cells, thus leading to the suspicion of mycobacterial infection. Because of its poor general condition, the ferret was euthanized. Necropsy examination revealed generalized granulomatous lymphadenitis, pneumonia, myocarditis, splenitis, and hepatitis. Histologically, in all organs, there were multifocal to coalescing areas of inflammatory infiltration composed of epithelioid macrophages, a low number of lymphocytes, and plasma cells, without necrosis nor multinucleated giant cells. Ziehl–Neelsen staining detected the presence of numerous (multibacillary) acid-fast bacteria, which were PCR-typed as M. xenopi. This is the first study showing the antimicrobial susceptibility testing of M. xenopi in veterinary medicine, describing the resistance to doxycycline. Overall, our results could facilitate further diagnosis and provide guidelines for the treatment protocols for such infections.
Full article
(This article belongs to the Special Issue One Health and Neglected Zoonotic Diseases)
►▼
Show Figures
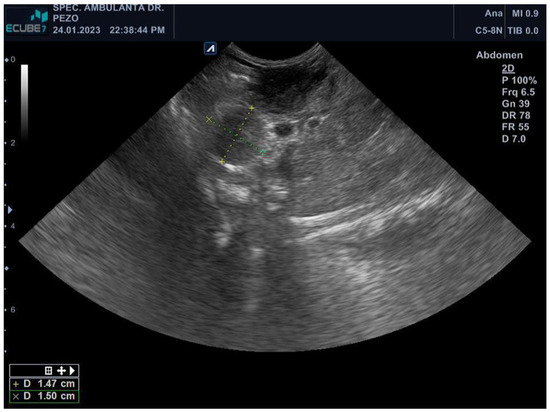
Figure 1
Open AccessArticle
Antibiotic Augmentation of Thermal Eradication of Staphylococcus epidermidis Biofilm Infections
by
Haydar A. S. Aljaafari, Nadia I. Abdulwahhab and Eric Nuxoll
Pathogens 2024, 13(4), 327; https://doi.org/10.3390/pathogens13040327 - 16 Apr 2024
Abstract
Staphylococcus epidermidis is a major contributor to bacterial infections on medical implants, currently treated by surgical removal of the device and the surrounding infected tissue at considerable morbidity and expense. In situ hyperthermia is being investigated as a non-invasive means of mitigating these
[...] Read more.
Staphylococcus epidermidis is a major contributor to bacterial infections on medical implants, currently treated by surgical removal of the device and the surrounding infected tissue at considerable morbidity and expense. In situ hyperthermia is being investigated as a non-invasive means of mitigating these bacterial biofilm infections, but minimizing damage to the surrounding tissue requires augmenting the thermal shock with other approaches such as antibiotics and discerning the minimum shock required to eliminate the biofilm. S. epidermidis biofilms were systematically shocked at a variety of temperatures (50–80 °C) and durations (1–10 min) to characterize their thermal susceptibility and compare it to other common nosocomial pathogens such as Staphylococcus aureus and Pseudomonas aeruginosa. Biofilms were also exposed to three classes of antibiotics (ciprofloxacin, tobramycin and erythromycin) separately at concentrations ranging from 0 to 128 μg mL−1 to evaluate their impact on the efficacy of thermal shock and the subsequent potential regrowth of the biofilm. S. epidermidis biofilms were shown to be more thermally susceptible to hyperthermia than other common bacterial pathogens. All three antibiotics substantially decreased the duration and/or temperature needed to eliminate the biofilms, though this augmentation did not meet the criteria of synergism immediately following thermal shock. Subsequent reincubation, however, revealed strong synergism on a longer timescale.
Full article
(This article belongs to the Section Bacterial Pathogens)
►▼
Show Figures
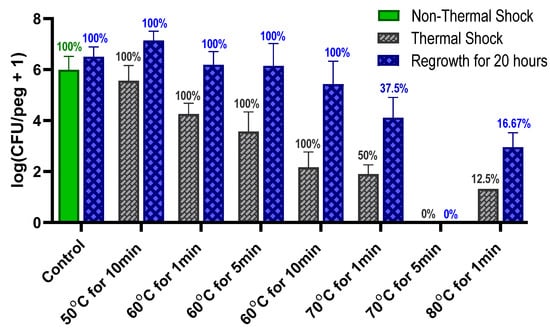
Figure 1
Open AccessArticle
Cold Plasma Deposition of Tobramycin as an Approach to Localized Antibiotic Delivery to Combat Biofilm Formation
by
Beatrice Olayiwola, Fiona O’Neill, Chloe Frewen, Darren F. Kavanagh, Rosemary O’Hara and Liam O’Neill
Pathogens 2024, 13(4), 326; https://doi.org/10.3390/pathogens13040326 - 16 Apr 2024
Abstract
Hospital-acquired infections (HAIs) remain a significant factor in hospitals, with implant surfaces often becoming contaminated by highly resistant strains of bacteria. Recent studies have shown that electrical plasma discharges can reduce bacterial load on surfaces, and this approach may help augment traditional antibiotic
[...] Read more.
Hospital-acquired infections (HAIs) remain a significant factor in hospitals, with implant surfaces often becoming contaminated by highly resistant strains of bacteria. Recent studies have shown that electrical plasma discharges can reduce bacterial load on surfaces, and this approach may help augment traditional antibiotic treatments. To investigate this, a cold atmospheric plasma was used to deposit tobramycin sulphate onto various surfaces, and the bacterial growth rate of K. pneumoniae in its planktonic and biofilm form was observed to probe the interactions between the plasma discharge and the antibiotic and to determine if there were any synergistic effects on the growth rate. The plasma-deposited tobramycin was still active after passing through the plasma field and being deposited onto titanium or polystyrene. This led to the significant inhibition of K. pneumoniae, with predictable antibiotic dose dependence. Separate studies have shown that the plasma treatment of the biofilm had a weak antimicrobial effect and reduced the amount of biofilm by around 50%. Combining a plasma pre-treatment on exposed biofilm followed by deposited tobramycin application proved to be somewhat effective in further reducing biofilm growth. The plasma discharge pre-treatment produced a further reduction in the biofilm load beyond that expected from just the antibiotic alone. However, the effect was not additive, and the results suggest that a complex interaction between plasma and antibiotic may be at play, with increasing plasma power producing a non-linear effect. This study may contribute to the treatment of infected surgical sites, with the coating of biomaterial surfaces with antibiotics reducing overall antibiotic use through the targeted delivery of therapeutics.
Full article
(This article belongs to the Special Issue Innovative Strategies to Counteract Microbial Biofilm Growth)
►▼
Show Figures
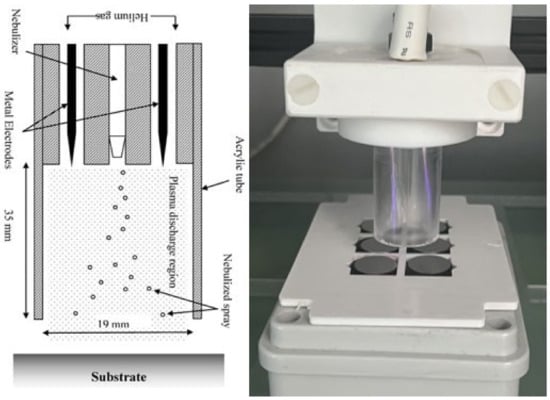
Figure 1
Open AccessReview
Are Kidneys Affected by SARS-CoV-2 Infection? An Updated Review on COVID-19-Associated AKI
by
Fabrizio Fabrizi, Luca Nardelli, Anna Regalia, Francesca Zanoni and Giuseppe Castellano
Pathogens 2024, 13(4), 325; https://doi.org/10.3390/pathogens13040325 - 16 Apr 2024
Abstract
Background: Human kidneys are an important target of SARS-CoV-2 infection, and many renal abnormalities have been found in patients with SARS-CoV-2 infection, including proteinuria, hematuria, and acute kidney injury. Acute kidney injury is now considered a common complication of COVID-19, and the epidemiology
[...] Read more.
Background: Human kidneys are an important target of SARS-CoV-2 infection, and many renal abnormalities have been found in patients with SARS-CoV-2 infection, including proteinuria, hematuria, and acute kidney injury. Acute kidney injury is now considered a common complication of COVID-19, and the epidemiology of AKI in SARS-CoV-2-infected patients continues to be controversial. Aim and Methods: We have carried out a narrative review to evaluate the frequency and risk factors for AKI among patients hospitalized due to COVID-19, and the latest surveys on this topic have been included. The mechanisms by which AKI occurs in COVID-19 patients have also been reviewed. Results: Multiple risk factors for the development of AKI in patients with SARS-CoV-2 infection have been identified; these have been classified in various groups (management and background factors, among others). SARS-CoV-2 targets the kidneys by indirect activity, but SARS-CoV-2 infects tubular epithelial cells and podocytes. We retrieved 24 reports (n = 502,593 unique patients with SARS-CoV-2 infection) and found an incidence of AKI of 31.8% (range, 0.5% to 56.9%). Only a minority (n = 2) of studies had a prospective design. We found that the AKI risk was greater in SARS-CoV-2 patients who underwent in-hospital deaths vs. those who survived; the summary estimate of the unadjusted RR of AKI was 2.63 (95% CI, 2.37; 2.93) (random-effects model). A stratified analysis showed that the incidence of AKI was greater in those reports where the frequency of COVID-19-positive patients having comorbidities (diabetes mellitus, arterial hypertension, and advanced age) was high. The unadjusted relative risk (aRR) of AKI was greater in SARS-CoV-2 patients who underwent ICU admission vs. those who did not; the pooled estimate of AKI risk was 2.64 (95% CI, 1.96; 3.56) according to the random-effects model. Conclusions: AKI is a common complication of hospitalized SARS-CoV-2-infected patients, and some comorbidities are important risk factors for it. The direct activity of the virus on the kidneys has been mentioned in the pathogenesis of AKI in SARS-CoV-2 patients. Further studies are ongoing in order to identify the mechanisms underlying the kidney injury in this population. The role of AKI on survival in SARS-CoV-2-infected patients is another area of active investigation.
Full article
(This article belongs to the Section Viral Pathogens)
►▼
Show Figures
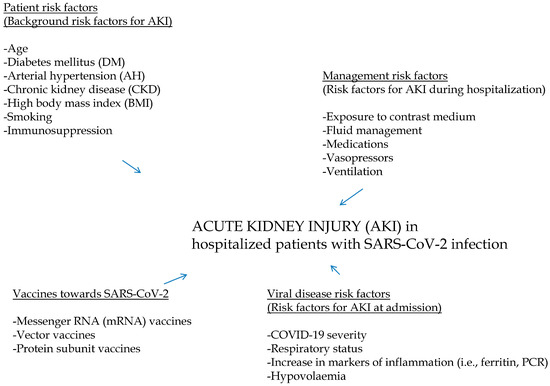
Figure 1
Open AccessArticle
Scientometrics Evaluation of Published Scientific Papers on the Use of Proteomics Technologies in Mastitis Research in Ruminants
by
Maria V. Bourganou, Dimitris C. Chatzopoulos, Daphne T. Lianou, George Th. Tsangaris, George C. Fthenakis and Angeliki I. Katsafadou
Pathogens 2024, 13(4), 324; https://doi.org/10.3390/pathogens13040324 - 15 Apr 2024
Abstract
The objective of this study was the presentation of quantitative characteristics regarding the scientific content and bibliometric details of the relevant publications. In total, 156 papers were considered. Most papers presented original studies (n = 135), and fewer were reviews (n
[...] Read more.
The objective of this study was the presentation of quantitative characteristics regarding the scientific content and bibliometric details of the relevant publications. In total, 156 papers were considered. Most papers presented original studies (n = 135), and fewer were reviews (n = 21). Most original articles (n = 101) referred to work involving cattle. Most original articles described work related to the diagnosis (n = 72) or pathogenesis (n = 62) of mastitis. Most original articles included field work (n = 75), whilst fewer included experimental (n = 31) or laboratory (n = 30) work. The tissue assessed most frequently in the studies was milk (n = 59). Milk was assessed more frequently in studies on the diagnosis (61.1% of relevant studies) or pathogenesis (30.6%) of the infection, but mammary tissue was assessed more frequently in studies on the treatment (31.0%). In total, 47 pathogens were included in the studies described; most were Gram-positive bacteria (n = 34). The three bacteria most frequently included in the studies were Staphylococcus aureus (n = 55 articles), Escherichia coli (n = 31) and Streptococcus uberis (n = 19). The proteomics technology employed more often in the respective studies was liquid chromatography-tandem mass spectrometry (LC-MS/MS), either on its own (n = 56) or in combination with other technologies (n = 40). The median year of publication of articles involving bioinformatics or LC-MS/MS and bioinformatics was the most recent: 2022. The 156 papers were published in 78 different journals, most frequently in the Journal of Proteomics (n = 16 papers) and the Journal of Dairy Science (n = 12). The median number of cited references in the papers was 48. In the papers, there were 1143 co-authors (mean: 7.3 ± 0.3 co-authors per paper, median: 7, min.–max.: 1–19) and 742 individual authors. Among them, 15 authors had published at least seven papers (max.: 10). Further, there were 218 individual authors who were the first or last authors in the papers. Most papers were submitted for open access (n = 79). The median number of citations received by the 156 papers was 12 (min.–max.: 0–339), and the median yearly number of citations was 2.0 (min.–max.: 0.0–29.5). The h-index of the papers was 33, and the m-index was 2. The increased number of cited references in papers and international collaboration in the respective study were the variables associated with most citations to published papers. This is the first ever scientometrics evaluation of proteomics studies, the results of which highlighted the characteristics of published papers on mastitis and proteomics. The use of proteomics in mastitis research has focused on the elucidation of pathogenesis and diagnosis of the infection; LC-MS/MS has been established as the most frequently used proteomics technology, although the use of bioinformatics has also emerged recently as a useful tool.
Full article
(This article belongs to the Section Bacterial Pathogens)
►▼
Show Figures
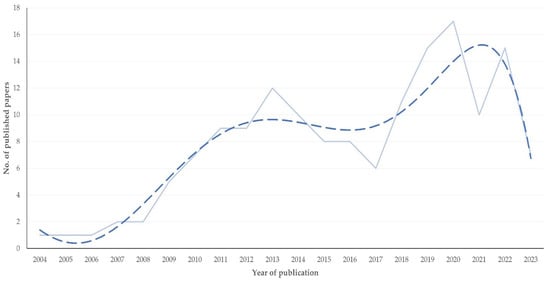
Figure 1
Open AccessReview
The Last Mile in Polio Eradication: Program Challenges and Perseverance
by
Rocio Lopez Cavestany, Martin Eisenhawer, Ousmane M. Diop, Harish Verma, Arshad Quddus and Ondrej Mach
Pathogens 2024, 13(4), 323; https://doi.org/10.3390/pathogens13040323 - 15 Apr 2024
Abstract
As the Global Polio Eradication Initiative (GPEI) strategizes towards the final steps of eradication, routine immunization schedules evolve, and high-quality vaccination campaigns and surveillance systems remain essential. New tools are consistently being developed, such as the novel oral poliovirus vaccine to combat outbreaks
[...] Read more.
As the Global Polio Eradication Initiative (GPEI) strategizes towards the final steps of eradication, routine immunization schedules evolve, and high-quality vaccination campaigns and surveillance systems remain essential. New tools are consistently being developed, such as the novel oral poliovirus vaccine to combat outbreaks more sustainably, as well as non-infectiously manufactured vaccines such as virus-like particle vaccines to eliminate the risk of resurgence of polio on the eve of a polio-free world. As the GPEI inches towards eradication, re-strategizing in the face of evolving challenges and preparing for unknown risks in the post-certification era are critical.
Full article
(This article belongs to the Special Issue Human Poliovirus)
►▼
Show Figures
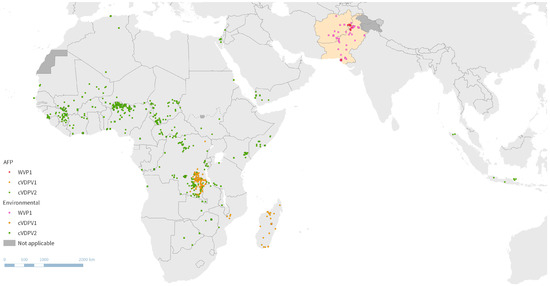
Figure 1
Open AccessArticle
Effect of Live and Fragmented Saccharomyces cerevisiae in the Feed of Pigs Challenged with Mycoplasma hyopneumoniae
by
Gabriela Vega-Munguía, Alejandro Vargas Sánchez, Juan E. Camacho-Medina, Luis Suárez-Vélez, Gabriela Bárcenas-Morales, David Quintar Guerrero, Abel Ciprian-Carrasco and Susana Mendoza Elvira
Pathogens 2024, 13(4), 322; https://doi.org/10.3390/pathogens13040322 - 14 Apr 2024
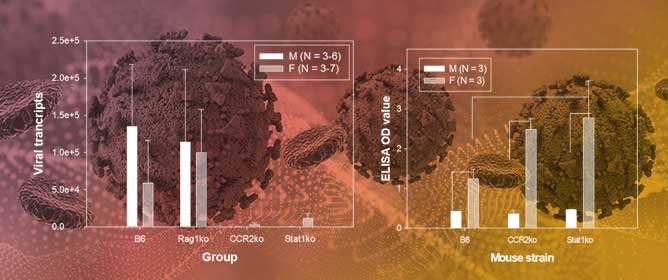
Abstract
►▼
Show Figures
Currently, the responsible use of antimicrobials in pigs has allowed the continuous development of alternatives to these antimicrobials. In this study, we describe the impact of treatments with two probiotics, one based on live Saccharomyces cerevisiae (S. cerevisiae) and another based
[...] Read more.
Currently, the responsible use of antimicrobials in pigs has allowed the continuous development of alternatives to these antimicrobials. In this study, we describe the impact of treatments with two probiotics, one based on live Saccharomyces cerevisiae (S. cerevisiae) and another based on fragmented S. cerevisiae (beta-glucans), that were administered to piglets at birth and at prechallenge with Mycoplasma hyopneumoniae. Thirty-two pigs were divided into four groups of eight animals each. The animals had free access to water and food. The groups were as follows: Group A, untreated negative control; Group B, inoculated by nebulization with M. hyopneumoniae positive control; Group C, first treated with disintegrated S. cerevisiae (disintegrated Sc) and inoculated by nebulization with M. hyopneumoniae; and Group D, treated with live S. cerevisiae yeast (live Sc) and inoculated by nebulization with M. hyopneumoniae. In a previous study, we found that on Days 1 and 21 of blood sampling, nine proinflammatory cytokines were secreted, and an increase in their secretion occurred for only five of them: TNF-α, INF-α, INF-γ, IL-10, and IL-12 p40. The results of the clinical evolution, the degree of pneumonic lesions, and the productive parameters of treated Groups C and D suggest that S. cerevisiae has an immunomodulatory effect in chronic proliferative M. hyopneumoniae pneumonia characterized by delayed hypersensitivity, which depends on the alteration or modulation of the respiratory immune response. The data presented in this study showed that S. cerevisiae contributed to the innate resistance of infected pigs.
Full article
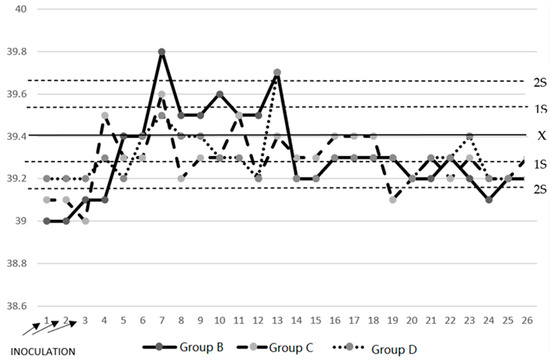
Figure 1

Journal Menu
► ▼ Journal Menu-
- Pathogens Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Biology, Cancers, Current Oncology, Diseases, JCM, Pathogens
Pathogenetic, Diagnostic and Therapeutic Perspectives in Head and Neck Cancer
Topic Editors: Shun-Fa Yang, Ming-Hsien ChienDeadline: 20 June 2024
Topic in
Diseases, IJMS, Microbiology Research, Pathogens, Vaccines
Advances in Human Pathogen Control—a 21st Century Challenge 2.0
Topic Editors: Jorge H. Leitão, Nitin Amdare, Joana R FelicianoDeadline: 30 June 2024
Topic in
Biomedicines, JCM, Pathogens, Vaccines, Viruses
Discovery and Development of Monkeypox Disease Treatments
Topic Editors: Mohd Imran, Ali A. RabaanDeadline: 31 August 2024
Topic in
Animals, Pathogens, Veterinary Sciences, Zoonotic Diseases
Zoonotic Vector-Borne Diseases of Companion Animals
Topic Editors: Anastasia Diakou, Donato TraversaDeadline: 20 December 2024

Conferences
Special Issues
Special Issue in
Pathogens
Novel Insights into Humoral Innate Resistance against Pathogens: Links with Extracellular Matrix and Haemostasis
Guest Editors: Andrea Doni, Raffaella Parente, Ying Jie MaDeadline: 20 April 2024
Special Issue in
Pathogens
Current Advances in Flavivirus and Other Arboviruses
Guest Editor: Jiahai LuDeadline: 30 April 2024
Special Issue in
Pathogens
Advances and Challenges in Understanding and Management of Diseases Caused by Oomycetes
Guest Editors: Anna Maria Vettraino, Benedetto T. LinaldedduDeadline: 15 May 2024
Topical Collections
Topical Collection in
Pathogens
Mastitis in Dairy Ruminants
Collection Editors: George Fthenakis, Maria Filippa Addis
Topical Collection in
Pathogens
Feature Papers on Parasitic Pathogens
Collection Editor: Sébastien Besteiro
Topical Collection in
Pathogens
Emerging and Re-emerging Pathogens
Collection Editors: Sheng-Fan Wang, Wen-Hung Wang, Arunee Thitithanyanont
Topical Collection in
Pathogens
New Insights into Bacterial Pathogenesis
Collection Editor: Carmelo Biondo