Capturing Peptide–GPCR Interactions and Their Dynamics
Abstract
:1. Introduction
2. The Contact Interface
Crosslinking
3. A Ligand’s Perspective
3.1. Solution State NMR for Low-Affinity Ligands
3.2. Solid-State NMR of High-Affinity Ligands
4. A Receptor’s Perspective
4.1. Selection and Specific Labeling of Sites-of-Interest
4.2. NMR: Investigation of Conformational Equilibria
4.3. NMR: Contribution of Fast Side Chains and Segmental Dynamics?
4.4. Electron Paramagnetic Resonance (EPR)
4.5. Fluorescence
4.6. Fourier-Transform Infrared Spectroscopy (FTIR)
4.7. Hydrogen-Deuterium Exchange (HDX) and Hydroxyl Radical Footprinting (HRF)
5. Conclusions and Perspective
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| A | active |
| A2AR | adenosine A2A receptor |
| APJR | apelin receptor |
| AT1R | angiotensin receptor 1 |
| Azi | p-azido-phenylalanine |
| β2AR | β2 adrenergic receptor |
| B1/2R | bradykinin receptor 1/2 |
| BNPS-skatole | 3-bromo-3-methyl-2-(2-nitrophenylthio)-3H-indole |
| Bpa | p-benzoyl-phenylalanine |
| BRET | bioluminescence resonance energy transfer |
| BTFA | 3-bromo-1,1,1-trifluoroacetone |
| CCR5 | CC chemokine receptor 5 |
| CGRP | calcitonin gene-related peptide |
| CRF | corticopin-releasing factor |
| CRF1/2R | corticopin-releasing factor receptor 1/2 |
| CuAAC | copper-catalyzed azide-alkyne cycloaddition |
| CXCL12 | CXC chemokine ligand 12 |
| CXCR4 | CXC chemokine receptor 4 |
| CPMG | Carr-Purcell-Meiboom-Gill (pulse sequence) |
| cwEPR | continuous wave EPR |
| DDM | n-dodecyl-β-d-maltopyranoside |
| DEER | double electron-electron resonance |
| ECD | extracellular domain |
| ECL | extracellular loop |
| EM | electron microscopy |
| EPR | electron paramagnetic resonance |
| ETB | endothelin B receptor |
| FlAsH | fluorescein arsenical hairpin binder |
| FRET | fluorescence/Förster resonance energy transfer |
| FTIR | Fourier-transform infrared spectroscopy |
| GLP-1 | glucagon-like peptide-1 |
| GLP1R | glucagon-like peptide-1 receptor |
| GLR | glucagon receptor |
| GPCR | G protein-coupled receptor |
| HDX | hydrogen-deuterium exchange |
| HRF | hydroxyl radical footprinting |
| I | inactive |
| LRET | lanthanoid resonance energy transfer |
| MIP1α | macrophage inflammatory protein 1α (=CCL3) |
| MS | mass spectrometry |
| MTSL | (1-oxyl-2,2,5,5-tetramethyl-pyrroline-3-methyl) methanethiosulfonate |
| ncAA | non-canonical amino acid |
| NMR | nuclear magnetic resonance |
| NOE | nuclear Overhauser effect |
| NPY | neuropeptide Y |
| NTS1R | neurotensin receptor 1 |
| PACAP | pituitary adenylate cyclase-activating peptide |
| PAC1R | pituitary adenylate cyclase-activating peptide receptor 1 |
| PTH1R | parathyroid hormone receptor 1 |
| REDOR | rotational-echo double-resonance |
| SAR | structure activity relationship studies |
| SDS-PAGE | sodium dodecyl sulfate–polyacrylamide gel electrophoresis |
| SDSL | site-directed spin labeling |
| SPAAC | strain-promoted azide-alkyne cycloaddition |
| STD | saturation transfer difference spectroscopy |
| TCS | transferred cross saturation spectroscopy |
| TET | 2,2,2-trifluoro-ethanethiol |
| TM | transmembrane |
| trNOESY | transferred nuclear Overhauser effect spectroscopy |
| Uaa | unnatural amino acid |
| Ucn | Urocortin |
| Y1/2R | neuropeptide Y receptor 1/2 |
References
- Wu, F.; Song, G.; de Graaf, C.; Stevens, R.C. Structure and Function of Peptide-Binding G Protein-Coupled Receptors. J. Mol. Biol. 2017, 429, 2726–2745. [Google Scholar] [CrossRef] [PubMed]
- Fredriksson, R.; Lagerström, M.C.; Lundin, L.-G.; Schiöth, H.B. The G-protein-coupled receptors in the human genome form five main families. Phylogenetic analysis, paralogon groups, and fingerprints. Mol. Pharmacol. 2003, 63, 1256–1272. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hauser, A.S.; Attwood, M.M.; Rask-Andersen, M.; Schiöth, H.B.; Gloriam, D.E. Trends in GPCR drug discovery: New agents, targets and indications. Nat. Rev. Drug Discov. 2017, 16, 829–842. [Google Scholar] [CrossRef]
- Sriram, K.; Insel, P.A. G Protein-Coupled Receptors as Targets for Approved Drugs: How Many Targets and How Many Drugs? Mol. Pharmacol. 2018, 93, 251–258. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Henninot, A.; Collins, J.C.; Nuss, J.M. The Current State of Peptide Drug Discovery: Back to the Future? J. Med. Chem. 2018, 61, 1382–1414. [Google Scholar] [CrossRef] [PubMed]
- Davenport, A.P.; Scully, C.C.G.; de Graaf, C.; Brown, A.J.H.; Maguire, J.J. Advances in therapeutic peptides targeting G protein-coupled receptors. Nat. Rev. Drug Discov. 2020, 19, 389–413. [Google Scholar] [CrossRef]
- Egloff, P.; Hillenbrand, M.; Klenk, C.; Batyuk, A.; Heine, P.; Balada, S.; Schlinkmann, K.M.; Scott, D.J.; Schutz, M.; Pluckthun, A. Structure of signaling-competent neurotensin receptor 1 obtained by directed evolution in Escherichia coli. Proc. Natl. Acad. Sci. USA 2014, 111, E655–E662. [Google Scholar] [CrossRef] [Green Version]
- White, J.F.; Noinaj, N.; Shibata, Y.; Love, J.; Kloss, B.; Xu, F.; Gvozdenovic-Jeremic, J.; Shah, P.; Shiloach, J.; Tate, C.G.; et al. Structure of the agonist-bound neurotensin receptor. Nature 2012, 490, 508–513. [Google Scholar] [CrossRef] [Green Version]
- Kato, H.E.; Zhang, Y.; Hu, H.; Suomivuori, C.-M.; Kadji, F.M.N.; Aoki, J.; Krishna Kumar, K.; Fonseca, R.; Hilger, D.; Huang, W.; et al. Conformational transitions of a neurotensin receptor 1-Gi1 complex. Nature 2019, 572, 80–85. [Google Scholar] [CrossRef]
- Huang, W.; Masureel, M.; Qu, Q.; Janetzko, J.; Inoue, A.; Kato, H.E.; Robertson, M.J.; Nguyen, K.C.; Glenn, J.S.; Skiniotis, G.; et al. Structure of the neurotensin receptor 1 in complex with β-arrestin 1. Nature 2020, 579, 303–308. [Google Scholar] [CrossRef]
- Shihoya, W.; Nishizawa, T.; Okuta, A.; Tani, K.; Dohmae, N.; Fujiyoshi, Y.; Nureki, O.; Doi, T. Activation mechanism of endothelin ETB receptor by endothelin-1. Nature 2016, 537, 363–368. [Google Scholar] [CrossRef] [PubMed]
- Shihoya, W.; Izume, T.; Inoue, A.; Yamashita, K.; Kadji, F.M.N.; Hirata, K.; Aoki, J.; Nishizawa, T.; Nureki, O. Crystal structures of human ETB receptor provide mechanistic insight into receptor activation and partial activation. Nat. Commun. 2018, 9, 4711. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wingler, L.M.; McMahon, C.; Staus, D.P.; Lefkowitz, R.J.; Kruse, A.C. Distinctive Activation Mechanism for Angiotensin Receptor Revealed by a Synthetic Nanobody. Cell 2019, 176, 479–490.e12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wingler, L.M.; Skiba, M.A.; McMahon, C.; Staus, D.P.; Kleinhenz, A.L.W.; Suomivuori, C.-M.; Latorraca, N.R.; Dror, R.O.; Lefkowitz, R.J.; Kruse, A.C. Angiotensin and biased analogs induce structurally distinct active conformations within a GPCR. Science 2020, 367, 888–892. [Google Scholar] [CrossRef]
- Koehl, A.; Hu, H.; Maeda, S.; Zhang, Y.; Qu, Q.; Paggi, J.M.; Latorraca, N.R.; Hilger, D.; Dawson, R.; Matile, H.; et al. Structure of the µ-opioid receptor–Gi protein complex. Nature 2018, 558, 547–552. [Google Scholar] [CrossRef]
- Claff, T.; Yu, J.; Blais, V.; Patel, N.; Martin, C.; Wu, L.; Han, G.W.; Holleran, B.J.; Van der Poorten, O.; White, K.L.; et al. Elucidating the active δ-opioid receptor crystal structure with peptide and small-molecule agonists. Sci. Adv. 2019, 5, eaax9115. [Google Scholar] [CrossRef] [Green Version]
- Ma, Y.; Yue, Y.; Ma, Y.; Zhang, Q.; Zhou, Q.; Song, Y.; Shen, Y.; Li, X.; Ma, X.; Li, C.; et al. Structural Basis for Apelin Control of the Human Apelin Receptor. Structure 2017, 25, 858–866.e4. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.; Qiao, A.; Yang, L.; Eps, N.V.; Frederiksen, K.S.; Yang, D.; Dai, A.; Cai, X.; Zhang, H.; Yi, C.; et al. Structure of the glucagon receptor in complex with a glucagon analogue. Nature 2018, 553, 106–110. [Google Scholar] [CrossRef]
- Qiao, A.; Han, S.; Li, X.; Li, Z.; Zhao, P.; Dai, A.; Chang, R.; Tai, L.; Tan, Q.; Chu, X.; et al. Structural basis of Gs and Gi recognition by the human glucagon receptor. Science 2020, 367, 1346–1352. [Google Scholar] [CrossRef]
- Jazayeri, A.; Rappas, M.; Brown, A.J.H.; Kean, J.; Errey, J.C.; Robertson, N.J.; Fiez-Vandal, C.; Andrews, S.P.; Congreve, M.; Bortolato, A.; et al. Crystal structure of the GLP-1 receptor bound to a peptide agonist. Nature 2017, 546, 254–258. [Google Scholar] [CrossRef]
- Liang, Y.-L.; Khoshouei, M.; Glukhova, A.; Furness, S.G.B.; Zhao, P.; Clydesdale, L.; Koole, C.; Truong, T.T.; Thal, D.M.; Lei, S.; et al. Phase-plate cryo-EM structure of a biased agonist-bound human GLP-1 receptor-Gs complex. Nature 2018, 555, 121–125. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Sun, B.; Feng, D.; Hu, H.; Chu, M.; Qu, Q.; Tarrasch, J.T.; Li, S.; Sun Kobilka, T.; Kobilka, B.K.; et al. Cryo-EM structure of the activated GLP-1 receptor in complex with a G protein. Nature 2017, 546, 248–253. [Google Scholar] [CrossRef]
- Ma, S.; Shen, Q.; Zhao, L.-H.; Mao, C.; Zhou, X.E.; Shen, D.-D.; de Waal, P.W.; Bi, P.; Li, C.; Jiang, Y.; et al. Molecular Basis for Hormone Recognition and Activation of Corticotropin-Releasing Factor Receptors. Mol. Cell 2020, 77, 669–680.e4. [Google Scholar] [CrossRef]
- Liang, Y.-L.; Belousoff, M.J.; Zhao, P.; Koole, C.; Fletcher, M.M.; Truong, T.T.; Julita, V.; Christopoulos, G.; Xu, H.E.; Zhang, Y.; et al. Toward a Structural Understanding of Class B GPCR Peptide Binding and Activation. Mol. Cell 2020, 77, 656–668.e5. [Google Scholar] [CrossRef]
- Zhao, L.-H.; Ma, S.; Sutkeviciute, I.; Shen, D.-D.; Zhou, X.E.; de Waal, P.W.; Li, C.-Y.; Kang, Y.; Clark, L.J.; Jean-Alphonse, F.G.; et al. Structure and dynamics of the active human parathyroid hormone receptor-1. Science 2019, 364, 148–153. [Google Scholar] [CrossRef]
- dal Maso, E.; Glukhova, A.; Zhu, Y.; Garcia-Nafria, J.; Tate, C.G.; Atanasio, S.; Reynolds, C.A.; Ramírez-Aportela, E.; Carazo, J.-M.; Hick, C.A.; et al. The Molecular Control of Calcitonin Receptor Signaling. ACS Pharmacol. Transl. Sci. 2019, 2, 31–51. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.-L.; Khoshouei, M.; Radjainia, M.; Zhang, Y.; Glukhova, A.; Tarrasch, J.; Thal, D.M.; Furness, S.G.B.; Christopoulos, G.; Coudrat, T.; et al. Phase-plate cryo-EM structure of a class B GPCR-G-protein complex. Nature 2017, 546, 118–123. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.-L.; Khoshouei, M.; Deganutti, G.; Glukhova, A.; Koole, C.; Peat, T.S.; Radjainia, M.; Plitzko, J.M.; Baumeister, W.; Miller, L.J.; et al. Cryo-EM structure of the active, Gs-protein complexed, human CGRP receptor. Nature 2018, 561, 492–497. [Google Scholar] [CrossRef]
- Venkatakrishnan, A.J.; Deupi, X.; Lebon, G.; Tate, C.G.; Schertler, G.F.; Babu, M.M. Molecular signatures of G-protein-coupled receptors. Nature 2013, 494, 185–194. [Google Scholar] [CrossRef]
- Venkatakrishnan, A.J.; Deupi, X.; Lebon, G.; Heydenreich, F.M.; Flock, T.; Miljus, T.; Balaji, S.; Bouvier, M.; Veprintsev, D.B.; Tate, C.G.; et al. Diverse activation pathways in class A GPCRs converge near the G-protein-coupling region. Nature 2016, 536, 484–487. [Google Scholar] [CrossRef]
- Zhou, Q.; Yang, D.; Wu, M.; Guo, Y.; Guo, W.; Zhong, L.; Cai, X.; Dai, A.; Jang, W.; Shakhnovich, E.I.; et al. Common activation mechanism of class A GPCRs. eLife 2019, 8. [Google Scholar] [CrossRef]
- Hilger, D.; Masureel, M.; Kobilka, B.K. Structure and dynamics of GPCR signaling complexes. Nat. Struct. Mol. Biol. 2018, 25, 4–12. [Google Scholar] [CrossRef] [PubMed]
- Hollenstein, K.; de Graaf, C.; Bortolato, A.; Wang, M.-W.; Marshall, F.H.; Stevens, R.C. Insights into the structure of class B GPCRs. Trends Pharmacol. Sci. 2014, 35, 12–22. [Google Scholar] [CrossRef] [Green Version]
- Karageorgos, V.; Venihaki, M.; Sakellaris, S.; Pardalos, M.; Kontakis, G.; Matsoukas, M.-T.; Gravanis, A.; Margioris, A.; Liapakis, G. Current understanding of the structure and function of family B GPCRs to design novel drugs. Hormones 2018, 17, 45–59. [Google Scholar] [CrossRef] [Green Version]
- Elgeti, M.; Rose, A.S.; Bartl, F.J.; Hildebrand, P.W.; Hofmann, K.-P.; Heck, M. Precision vs flexibility in GPCR signaling. J. Am. Chem. Soc. 2013, 135, 12305–12312. [Google Scholar] [CrossRef] [PubMed]
- Manglik, A.; Kobilka, B. The role of protein dynamics in GPCR function: Insights from the β2AR and rhodopsin. Curr. Opin. Cell Biol. 2014, 27, 136–143. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Unal, H.; Karnik, S.S. Domain coupling in GPCRs: The engine for induced conformational changes. Trends Pharmacol. Sci. 2012, 33, 79–88. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Flock, T.; Hauser, A.S.; Lund, N.; Gloriam, D.E.; Balaji, S.; Babu, M.M. Selectivity determinants of GPCR-G-protein binding. Nature 2017, 545, 317–322. [Google Scholar] [CrossRef]
- Kang, Y.; Kuybeda, O.; de Waal, P.W.; Mukherjee, S.; Van Eps, N.; Dutka, P.; Zhou, X.E.; Bartesaghi, A.; Erramilli, S.; Morizumi, T.; et al. Cryo-EM structure of human rhodopsin bound to an inhibitory G protein. Nature 2018, 558, 553–558. [Google Scholar] [CrossRef]
- Tsai, C.-J.; Marino, J.; Adaixo, R.; Pamula, F.; Muehle, J.; Maeda, S.; Flock, T.; Taylor, N.M.; Mohammed, I.; Matile, H.; et al. Cryo-EM structure of the rhodopsin-Gαi-βγ complex reveals binding of the rhodopsin C-terminal tail to the gβ subunit. eLife 2019, 8, e46041. [Google Scholar] [CrossRef]
- Du, Y.; Duc, N.M.; Rasmussen, S.G.F.; Hilger, D.; Kubiak, X.; Wang, L.; Bohon, J.; Kim, H.R.; Wegrecki, M.; Asuru, A.; et al. Assembly of a GPCR-G Protein Complex. Cell 2019, 177, 1232–1242.e11. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Xu, X.; Hilger, D.; Aschauer, P.; Tiemann, J.K.S.; Du, Y.; Liu, H.; Hirata, K.; Sun, X.; Guixà-González, R.; et al. Structural Insights into the Process of GPCR-G Protein Complex Formation. Cell 2019, 177, 1243–1251.e12. [Google Scholar] [CrossRef] [PubMed]
- Ward, A.B.; Sali, A.; Wilson, I.A. Biochemistry. Integrative structural biology. Science 2013, 339, 913–915. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xia, Y.; Fischer, A.W.; Teixeira, P.; Weiner, B.; Meiler, J. Integrated Structural Biology for α-Helical Membrane Protein Structure Determination. Structure 2018, 26, 657–666.e2. [Google Scholar] [CrossRef] [Green Version]
- Shimada, I.; Ueda, T.; Kofuku, Y.; Eddy, M.T.; Wüthrich, K. GPCR drug discovery: Integrating solution NMR data with crystal and cryo-EM structures. Nat. Rev. Drug Discov. 2019, 18, 59–82. [Google Scholar] [CrossRef] [PubMed]
- Kufareva, I.; Handel, T.M.; Abagyan, R. Experiment-Guided Molecular Modeling of Protein-Protein Complexes Involving GPCRs. G Protein-Coupled Recept. Drug Discov. 2015, 1335, 295–311. [Google Scholar] [CrossRef] [Green Version]
- Pándy-Szekeres, G.; Munk, C.; Tsonkov, T.M.; Mordalski, S.; Harpsøe, K.; Hauser, A.S.; Bojarski, A.J.; Gloriam, D.E. GPCRdb in 2018: Adding GPCR structure models and ligands. Nucleic Acids Res. 2018, 46, D440–D446. [Google Scholar] [CrossRef] [Green Version]
- Kornreich, W.D.; Galyean, R.; Hernandez, J.F.; Craig, A.G.; Donaldson, C.J.; Yamamoto, G.; Rivier, C.; Vale, W.; Rivier, J. Alanine series of ovine corticotropin releasing factor (oCRF): A structure-activity relationship study. J. Med. Chem. 1992, 35, 1870–1876. [Google Scholar] [CrossRef]
- Beck-Sickinger, A.G.; Wieland, H.A.; Wittneben, H.; Willim, K.D.; Rudolf, K.; Jung, G. Complete L-alanine scan of neuropeptide Y reveals ligands binding to Y1 and Y2 receptors with distinguished conformations. Eur. J. Biochem. 1994, 225, 947–958. [Google Scholar] [CrossRef]
- Beyermann, M.; Fechner, K.; Furkert, J.; Krause, E.; Bienert, M. A single-point slight alteration set as a tool for structure-activity relationship studies of ovine corticotropin releasing factor. J. Med. Chem. 1996, 39, 3324–3330. [Google Scholar] [CrossRef]
- Kirby, D.A.; Boublik, J.H.; Rivier, J.E. Neuropeptide Y: Y1 and Y2 affinities of the complete series of analogues with single D-residue substitutions. J. Med. Chem. 1993, 36, 3802–3808. [Google Scholar] [CrossRef] [PubMed]
- Grundemar, L.; Kahl, U.; Callréus, T.; Langel, Ü.; Bienert, M.; Beyermann, M. Ligand binding and functional effects of systematic double d-amino acid residue substituted neuropeptide Y analogs on Y1 and Y2 receptor types. Regul. Pept. 1996, 62, 131–136. [Google Scholar] [CrossRef]
- Gerling, U.I.M.; Brandenburg, E.; Berlepsch, H.V.; Pagel, K.; Koksch, B. Structure Analysis of an Amyloid-Forming Model Peptide by a Systematic Glycine and Proline Scan. Biomacromolecules 2011, 12, 2988–2996. [Google Scholar] [CrossRef] [PubMed]
- Rathmann, D.; Lindner, D.; DeLuca, S.H.; Kaufmann, K.W.; Meiler, J.; Beck-Sickinger, A.G. Ligand-mimicking Receptor Variant Discloses Binding and Activation Mode of Prolactin-releasing Peptide. J. Biol. Chem. 2012, 287, 32181–32194. [Google Scholar] [CrossRef] [Green Version]
- Horovitz, A. Double-mutant cycles: A powerful tool for analyzing protein structure and function. Fold. Des. 1996, 1, R121–R126. [Google Scholar] [CrossRef] [Green Version]
- Merten, N.; Lindner, D.; Rabe, N.; Römpler, H.; Mörl, K.; Schöneberg, T.; Beck-Sickinger, A.G. Receptor subtype-specific docking of Asp6.59 with C-terminal arginine residues in Y receptor ligands. J. Biol. Chem. 2007, 282, 7543–7551. [Google Scholar] [CrossRef] [Green Version]
- Kaiser, A.; Müller, P.; Zellmann, T.; Scheidt, H.A.; Thomas, L.; Bosse, M.; Meier, R.; Meiler, J.; Huster, D.; Beck-Sickinger, A.G.; et al. Unwinding of the C-Terminal Residues of Neuropeptide Y is critical for Y2 Receptor Binding and Activation. Angew. Chem. Int. Ed. 2015, 54, 7446–7449. [Google Scholar] [CrossRef] [Green Version]
- Yang, Z.; Han, S.; Keller, M.; Kaiser, A.; Bender, B.J.; Bosse, M.; Burkert, K.; Kögler, L.M.; Wifling, D.; Bernhardt, G.; et al. Structural basis of ligand binding modes at the neuropeptide Y Y1 receptor. Nature 2018, 556, 520–524. [Google Scholar] [CrossRef]
- Joedicke, L.; Mao, J.; Kuenze, G.; Reinhart, C.; Kalavacherla, T.; Jonker, H.R.A.; Richter, C.; Schwalbe, H.; Meiler, J.; Preu, J.; et al. The molecular basis of subtype selectivity of human kinin G-protein-coupled receptors. Nat. Chem. Biol. 2018, 14, 284–290. [Google Scholar] [CrossRef]
- Pal, K.; Melcher, K.; Xu, H.E. Structure and mechanism for recognition of peptide hormones by Class B G-protein-coupled receptors. Acta Pharmacol. Sin. 2012, 33, 300–311. [Google Scholar] [CrossRef]
- Sun, D.; Ostermaier, M.K.; Heydenreich, F.M.; Mayer, D.; Jaussi, R.; Standfuss, J.; Veprintsev, D.B. AAscan, PCRdesign and MutantChecker: A Suite of Programs for Primer Design and Sequence Analysis for High-Throughput Scanning Mutagenesis. PLoS ONE 2013, 8, e78878. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wootten, D.; Reynolds, C.A.; Smith, K.J.; Mobarec, J.C.; Koole, C.; Savage, E.E.; Pabreja, K.; Simms, J.; Sridhar, R.; Furness, S.G.B.; et al. The Extracellular Surface of the GLP-1 Receptor Is a Molecular Trigger for Biased Agonism. Cell 2016, 165, 1632–1643. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wootten, D.; Simms, J.; Miller, L.J.; Christopoulos, A.; Sexton, P.M. Polar transmembrane interactions drive formation of ligand-specific and signal pathway-biased family B G protein-coupled receptor conformations. Proc. Natl. Acad. Sci. USA 2013, 110, 5211–5216. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wootten, D.; Reynolds, C.A.; Koole, C.; Smith, K.J.; Mobarec, J.C.; Simms, J.; Quon, T.; Coudrat, T.; Furness, S.G.B.; Miller, L.J.; et al. A Hydrogen-Bonded Polar Network in the Core of the Glucagon-Like Peptide-1 Receptor Is a Fulcrum for Biased Agonism: Lessons from Class B Crystal Structures. Mol. Pharmacol. 2016, 89, 335–347. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wootten, D.; Reynolds, C.A.; Smith, K.J.; Mobarec, J.C.; Furness, S.G.B.; Miller, L.J.; Christopoulos, A.; Sexton, P.M. Key interactions by conserved polar amino acids located at the transmembrane helical boundaries in Class B GPCRs modulate activation, effector specificity and biased signalling in the glucagon-like peptide-1 receptor. Biochem. Pharmacol. 2016, 118, 68–87. [Google Scholar] [CrossRef] [Green Version]
- Wescott, M.P.; Kufareva, I.; Paes, C.; Goodman, J.R.; Thaker, Y.; Puffer, B.A.; Berdougo, E.; Rucker, J.B.; Handel, T.M.; Doranz, B.J. Signal transmission through the CXC chemokine receptor 4 (CXCR4) transmembrane helices. Proc. Natl. Acad. Sci. USA 2016, 113, 9928–9933. [Google Scholar] [CrossRef] [Green Version]
- Tanaka, Y.; Bond, M.R.; Kohler, J.J. Photocrosslinkers illuminate interactions in living cells. Mol. Biosyst. 2008, 4, 473. [Google Scholar] [CrossRef]
- Escher, E.H.; Dung, N.T.M.; Robert, H.; Regoli, D.C.; St. Pierre, S.A. Photoaffinity labeling of the angiotensin II receptor. 1. Synthesis and biological activities of the labeling peptides. J. Med. Chem. 1978, 21, 860–864. [Google Scholar] [CrossRef]
- Pham, V.; Sexton, P.M. Photoaffinity scanning in the mapping of the peptide receptor interface of class II G protein—Coupled receptors. J. Pept. Sci. Off. Publ. Eur. Pept. Soc. 2004, 10, 179–203. [Google Scholar] [CrossRef]
- Dong, M.; Lam, P.C.-H.; Pinon, D.I.; Hosohata, K.; Orry, A.; Sexton, P.M.; Abagyan, R.; Miller, L.J. Molecular basis of secretin docking to its intact receptor using multiple photolabile probes distributed throughout the pharmacophore. J. Biol. Chem. 2011, 286, 23888–23899. [Google Scholar] [CrossRef] [Green Version]
- Wittelsberger, A.; Corich, M.; Thomas, B.E.; Lee, B.-K.; Barazza, A.; Czodrowski, P.; Mierke, D.F.; Chorev, M.; Rosenblatt, M. The Mid-Region of Parathyroid Hormone (1−34) Serves as a Functional Docking Domain in Receptor Activation †. Biochemistry 2006, 45, 2027–2034. [Google Scholar] [CrossRef] [PubMed]
- Jacobsen, R.B.; Sale, K.L.; Ayson, M.J.; Novak, P.; Hong, J.; Lane, P.; Wood, N.L.; Kruppa, G.H.; Young, M.M.; Schoeniger, J.S. Structure and dynamics of dark-state bovine rhodopsin revealed by chemical cross-linking and high-resolution mass spectrometry. Protein Sci. 2006, 15, 1303–1317. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Muranaka, H.; Momose, T.; Handa, C.; Ozawa, T. Photoaffinity Labeling of the Human A 2A Adenosine Receptor and Cross-link Position Analysis by Mass Spectrometry. ACS Med. Chem. Lett. 2017, 8, 660–665. [Google Scholar] [CrossRef] [PubMed]
- Umanah, G.K.E.; Huang, L.; Ding, F.; Arshava, B.; Farley, A.R.; Link, A.J.; Naider, F.; Becker, J.M. Identification of Residue-to-residue Contact between a Peptide Ligand and Its G Protein-coupled Receptor Using Periodate-mediated Dihydroxyphenylalanine Cross-linking and Mass Spectrometry. J. Biol. Chem. 2010, 285, 39425–39436. [Google Scholar] [CrossRef] [Green Version]
- Umanah, G.K.E.; Son, C.; Ding, F.; Naider, F.; Becker, J.M. Cross-Linking of a DOPA-Containing Peptide Ligand into Its G Protein-Coupled Receptor †. Biochemistry 2009, 48, 2033–2044. [Google Scholar] [CrossRef] [Green Version]
- Liu, C.C.; Schultz, P.G. Adding New Chemistries to the Genetic Code. Annu. Rev. Biochem. 2010, 79, 413–444. [Google Scholar] [CrossRef] [Green Version]
- Lang, K.; Chin, J.W. Cellular Incorporation of Unnatural Amino Acids and Bioorthogonal Labeling of Proteins. Chem. Rev. 2014, 114, 4764–4806. [Google Scholar] [CrossRef]
- Coin, I. Application of non-canonical crosslinking amino acids to study protein-protein interactions in live cells. Curr. Opin. Chem. Biol. 2018, 46, 156–163. [Google Scholar] [CrossRef]
- Nguyen, T.-A.; Cigler, M.; Lang, K. Expanding the Genetic Code to Study Protein-Protein Interactions. Angew. Chem. Int. Ed. 2018, 57, 14350–14361. [Google Scholar] [CrossRef]
- Grunbeck, A.; Huber, T.; Sachdev, P.; Sakmar, T.P. Mapping the Ligand-Binding Site on a G Protein-Coupled Receptor (GPCR) Using Genetically Encoded Photocrosslinkers. Biochemistry 2011, 50, 3411–3413. [Google Scholar] [CrossRef] [Green Version]
- Coin, I.; Perrin, M.H.; Vale, W.W.; Wang, L. Photo-cross-linkers incorporated into G-protein-coupled receptors in mammalian cells: A ligand comparison. Angew. Chem. 2011, 50, 8077–8081. [Google Scholar] [CrossRef] [PubMed]
- Coin, I.; Katritch, V.; Sun, T.; Xiang, Z.; Siu, F.Y.; Beyermann, M.; Stevens, R.C.; Wang, L. Genetically encoded chemical probes in cells reveal the binding path of urocortin-I to CRF class B GPCR. Cell 2013, 155, 1258–1269. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grunbeck, A.; Huber, T.; Abrol, R.; Trzaskowski, B.; Goddard, W.A.; Sakmar, T.P. Genetically encoded photo-cross-linkers map the binding site of an allosteric drug on a G protein-coupled receptor. ACS Chem. Biol. 2012, 7, 967–972. [Google Scholar] [CrossRef]
- Valentin-Hansen, L.; Park, M.; Huber, T.; Grunbeck, A.; Naganathan, S.; Schwartz, T.W.; Sakmar, T.P. Mapping Substance P Binding Sites on the Neurokinin-1 Receptor Using Genetic Incorporation of a Photoreactive Amino Acid. J. Biol. Chem. 2014, 289, 18045–18054. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koole, C.; Reynolds, C.A.; Mobarec, J.C.; Hick, C.; Sexton, P.M.; Sakmar, T.P. Genetically encoded photocross-linkers determine the biological binding site of exendin-4 peptide in the N-terminal domain of the intact human glucagon-like peptide-1 receptor (GLP-1R). J. Biol. Chem. 2017, 292, 7131–7144. [Google Scholar] [CrossRef] [Green Version]
- Simms, J.; Uddin, R.; Sakmar, T.P.; Gingell, J.J.; Garelja, M.L.; Hay, D.L.; Brimble, M.A.; Harris, P.W.; Reynolds, C.A.; Poyner, D.R. Photoaffinity Cross-Linking and Unnatural Amino Acid Mutagenesis Reveal Insights into Calcitonin Gene-Related Peptide Binding to the Calcitonin Receptor-like Receptor/Receptor Activity-Modifying Protein 1 (CLR/RAMP1) Complex. Biochemistry 2018, 57, 4915–4922. [Google Scholar] [CrossRef] [Green Version]
- Seidel, L.; Zarzycka, B.; Zaidi, S.A.; Katritch, V.; Coin, I. Structural insight into the activation of a class B G-protein-coupled receptor by peptide hormones in live human cells. eLife 2017, 6. [Google Scholar] [CrossRef]
- Rannversson, H.; Andersen, J.; Sørensen, L.; Bang-Andersen, B.; Park, M.; Huber, T.; Sakmar, T.P.; Strømgaard, K. Genetically encoded photocrosslinkers locate the high-affinity binding site of antidepressant drugs in the human serotonin transporter. Nat. Commun. 2016, 7, 11261. [Google Scholar] [CrossRef] [Green Version]
- Rannversson, H.; Andersen, J.; Bang-Andersen, B.; Strømgaard, K. Mapping the Binding Site for Escitalopram and Paroxetine in the Human Serotonin Transporter Using Genetically Encoded Photo-Cross-Linkers. ACS Chem. Biol. 2017, 12, 2558–2562. [Google Scholar] [CrossRef]
- Whittaker, J.; Whittaker, L.J.; Roberts, C.T.; Phillips, N.B.; Ismail-Beigi, F.; Lawrence, M.C.; Weiss, M.A. α-Helical element at the hormone-binding surface of the insulin receptor functions as a signaling element to activate its tyrosine kinase. Proc. Natl. Acad. Sci. USA 2012, 109, 11166–11171. [Google Scholar] [CrossRef] [Green Version]
- Pellequer, J.-L.; Chen, S.W. Multi-template approach to modeling engineered disulfide bonds. Proteins Struct. Funct. Bioinform. 2006, 65, 192–202. [Google Scholar] [CrossRef] [PubMed]
- Kufareva, I.; Gustavsson, M.; Holden, L.G.; Qin, L.; Zheng, Y.; Handel, T.M. Disulfide Trapping for Modeling and Structure Determination of Receptor: Chemokine Complexes. Methods Enzymol. 2016, 570, 389–420. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dong, M.; Lam, P.C.-H.; Orry, A.; Sexton, P.M.; Christopoulos, A.; Abagyan, R.; Miller, L.J. Use of Cysteine Trapping to Map Spatial Approximations between Residues Contributing to the Helix N-capping Motif of Secretin and Distinct Residues within Each of the Extracellular Loops of Its Receptor. J. Biol. Chem. 2016, 291, 5172–5184. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Monaghan, P.; Thomas, B.E.; Woznica, I.; Wittelsberger, A.; Mierke, D.F.; Rosenblatt, M. Mapping peptide hormone-receptor interactions using a disulfide-trapping approach. Biochemistry 2008, 47, 5889–5895. [Google Scholar] [CrossRef] [Green Version]
- Dong, M.; Xu, X.; Ball, A.M.; Makhoul, J.A.; Lam, P.C.-H.; Pinon, D.I.; Orry, A.; Sexton, P.M.; Abagyan, R.; Miller, L.J. Mapping spatial approximations between the amino terminus of secretin and each of the extracellular loops of its receptor using cysteine trapping. FASEB J. 2012, 26, 5092–5105. [Google Scholar] [CrossRef] [Green Version]
- Kufareva, I.; Stephens, B.S.; Holden, L.G.; Qin, L.; Zhao, C.; Kawamura, T.; Abagyan, R.; Handel, T.M. Stoichiometry and geometry of the CXC chemokine receptor 4 complex with CXC ligand 12: Molecular modeling and experimental validation. Proc. Natl. Acad. Sci. USA 2014, 111, E5363–E5372. [Google Scholar] [CrossRef] [Green Version]
- Ngo, T.; Stephens, B.S.; Gustavsson, M.; Holden, L.G.; Abagyan, R.; Handel, T.M.; Kufareva, I. Crosslinking-guided geometry of a complete CXC receptor-chemokine complex and the basis of chemokine subfamily selectivity. PLoS Biol. 2020, 18, e3000656. [Google Scholar] [CrossRef] [Green Version]
- Hamdan, F.F.; Ward, S.D.C.; Siddiqui, N.A.; Bloodworth, L.M.; Wess, J. Use of an in situ disulfide cross-linking strategy to map proximities between amino acid residues in transmembrane domains I and VII of the M3 muscarinic acetylcholine receptor. Biochemistry 2002, 41, 7647–7658. [Google Scholar] [CrossRef]
- Ward, S.D.C.; Hamdan, F.F.; Bloodworth, L.M.; Siddiqui, N.A.; Li, J.H.; Wess, J. Use of an in situ disulfide cross-linking strategy to study the dynamic properties of the cytoplasmic end of transmembrane domain VI of the M3 muscarinic acetylcholine receptor. Biochemistry 2006, 45, 676–685. [Google Scholar] [CrossRef]
- Ward, S.D.C.; Hamdan, F.F.; Bloodworth, L.M.; Wess, J. Conformational changes that occur during M3 muscarinic acetylcholine receptor activation probed by the use of an in situ disulfide cross-linking strategy. J. Biol. Chem. 2002, 277, 2247–2257. [Google Scholar] [CrossRef] [Green Version]
- Buck, E.; Wells, J.A. Disulfide trapping to localize small-molecule agonists and antagonists for a G protein-coupled receptor. Proc. Natl. Acad. Sci. USA 2005, 102, 2719–2724. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hagemann, I.S.; Miller, D.L.; Klco, J.M.; Nikiforovich, G.V.; Baranski, T.J. Structure of the complement factor 5a receptor-ligand complex studied by disulfide trapping and molecular modeling. J. Biol. Chem. 2008, 283, 7763–7775. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xue, L.; Rovira, X.; Scholler, P.; Zhao, H.; Liu, J.; Pin, J.-P.; Rondard, P. Major ligand-induced rearrangement of the heptahelical domain interface in a GPCR dimer. Nat. Chem. Biol. 2015, 11, 134–140. [Google Scholar] [CrossRef] [PubMed]
- Xue, L.; Sun, Q.; Zhao, H.; Rovira, X.; Gai, S.; He, Q.; Pin, J.-P.; Liu, J.; Rondard, P. Rearrangement of the transmembrane domain interfaces associated with the activation of a GPCR hetero-oligomer. Nat. Commun. 2019, 10, 2765. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, H.; Lee, B.-K.; Naider, F.; Becker, J.M. Identification of specific transmembrane residues and ligand-induced interface changes involved in homo-dimer formation of a yeast G protein-coupled receptor. Biochemistry 2009, 48, 10976–10987. [Google Scholar] [CrossRef] [Green Version]
- Knepp, A.M.; Periole, X.; Marrink, S.-J.; Sakmar, T.P.; Huber, T. Rhodopsin Forms a Dimer with Cytoplasmic Helix 8 Contacts in Native Membranes. Biochemistry 2012, 51, 1819–1821. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, X.E.; He, Y.; de Waal, P.W.; Gao, X.; Kang, Y.; Van Eps, N.; Yin, Y.; Pal, K.; Goswami, D.; White, T.A.; et al. Identification of Phosphorylation Codes for Arrestin Recruitment by G Protein-Coupled Receptors. Cell 2017, 170, 457–469.e13. [Google Scholar] [CrossRef] [Green Version]
- Xiang, Z.; Ren, H.; Hu, Y.S.; Coin, I.; Wei, J.; Cang, H.; Wang, L. Adding an unnatural covalent bond to proteins through proximity-enhanced bioreactivity. Nat. Methods 2013, 10, 885–888. [Google Scholar] [CrossRef] [Green Version]
- Xiang, Z.; Lacey, V.K.; Ren, H.; Xu, J.; Burban, D.J.; Jennings, P.A.; Wang, L. Proximity-enabled protein crosslinking through genetically encoding haloalkane unnatural amino acids. Angew. Chem. Int. Ed. 2014, 53, 2190–2193. [Google Scholar] [CrossRef]
- Cigler, M.; Müller, T.G.; Horn-Ghetko, D.; von Wrisberg, M.-K.; Fottner, M.; Goody, R.S.; Itzen, A.; Müller, M.P.; Lang, K. Proximity-Triggered Covalent Stabilization of Low-Affinity Protein Complexes In Vitro and In Vivo. Angew. Chem. Int. Ed. 2017, 56, 15737–15741. [Google Scholar] [CrossRef]
- Yang, B.; Wu, H.; Schnier, P.D.; Liu, Y.; Liu, J.; Wang, N.; DeGrado, W.F.; Wang, L. Proximity-enhanced SuFEx chemical cross-linker for specific and multitargeting cross-linking mass spectrometry. Proc. Natl. Acad. Sci. USA 2018, 115, 11162–11167. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, L. Genetically encoding new bioreactivity. New Biotechnol. 2017, 38, 16–25. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Q.; Chen, Q.; Klauser, P.C.; Li, M.; Zheng, F.; Wang, N.; Li, X.; Zhang, Q.; Fu, X.; Wang, Q.; et al. Developing Covalent Protein Drugs via Proximity-Enabled Reactive Therapeutics. Cell 2020, 182, 85–97.e16. [Google Scholar] [CrossRef] [PubMed]
- Seidel, L.; Zarzycka, B.; Katritch, V.; Coin, I. Exploring Pairwise Chemical Crosslinking To Study Peptide-Receptor Interactions. ChemBioChem 2019, 20, 683–692. [Google Scholar] [CrossRef] [PubMed]
- Hollenstein, K.; Kean, J.; Bortolato, A.; Cheng, R.K.Y.; Doré, A.S.; Jazayeri, A.; Cooke, R.M.; Weir, M.; Marshall, F.H. Structure of class B GPCR corticotropin-releasing factor receptor 1. Nature 2013, 499, 438–443. [Google Scholar] [CrossRef] [PubMed]
- Pal, K.; Melcher, K.; Xu, H.E. Structure modeling using genetically engineered crosslinking. Cell 2013, 155, 1207–1208. [Google Scholar] [CrossRef] [Green Version]
- Böttke, T.; Ernicke, S.; Serfling, R.; Ihling, C.; Burda, E.; Gurevich, V.V.; Sinz, A.; Coin, I. Exploring GPCR-arrestin interfaces with genetically encoded crosslinkers. EMBO Rep. 2020, e50437. [Google Scholar] [CrossRef]
- Dong, M.; Deganutti, G.; Piper, S.J.; Liang, Y.-L.; Khoshouei, M.; Belousoff, M.J.; Harikumar, K.G.; Reynolds, C.A.; Glukhova, A.; Furness, S.G.B.; et al. Structure and dynamics of the active Gs-coupled human secretin receptor. Nat. Commun. 2020, 11, 4137. [Google Scholar] [CrossRef]
- Monks, S.A.; Karagianis, G.; Howlett, G.J.; Norton, R.S. Solution structure of human neuropeptide Y. J. Biomol. NMR 1996, 8, 379–390. [Google Scholar] [CrossRef]
- Bader, R.; Bettio, A.; Beck-Sickinger, A.G.; Zerbe, O. Structure and Dynamics of Micelle-bound Neuropeptide Y: Comparison with Unligated NPY and Implications for Receptor Selection. J. Mol. Biol. 2001, 305, 307–329. [Google Scholar] [CrossRef]
- Miller, M.C.; Mayo, K.H. Chemokines from a Structural Perspective. Int. J. Mol. Sci. 2017, 18, 2088. [Google Scholar] [CrossRef] [Green Version]
- Nieto, J.L.; Rico, M.; Santoro, J.; Herranz, J.; Bermejo, F.J. Assignment and conformation of neurotensin in aqueous solution by 1H NMR. Int. J. Pept. Protein Res. 1986, 28, 315–323. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.-Y.; Deber, C.M. Conformations of neurotensin in solution and in membrane environments studied by 2-D NMR spectroscopy. Int. J. Pept. Protein Res. 1991, 37, 528–535. [Google Scholar] [CrossRef] [PubMed]
- Silva Elipe, M.V.; Bednarek, M.A.; Gao, Y.D. 1H NMR structural analysis of human ghrelin and its six truncated analogs. Biopolymers 2001, 59, 489–501. [Google Scholar] [CrossRef]
- De Ricco, R.; Valensin, D.; Gaggelli, E.; Valensin, G. Conformation propensities of des-acyl-ghrelin as probed by CD and NMR. Peptides 2013, 43, 62–67. [Google Scholar] [CrossRef]
- Vortmeier, G.; DeLuca, S.H.; Els-Heindl, S.; Chollet, C.; Scheidt, H.A.; Beck-Sickinger, A.G.; Meiler, J.; Huster, D. Integrating solid-state NMR and computational modeling to investigate the structure and dynamics of membrane-associated ghrelin. PLoS ONE 2015, 10, e0122444. [Google Scholar] [CrossRef]
- Venkatakrishnan, A.; Flock, T.; Prado, D.E.; Oates, M.E.; Gough, J.; Madan Babu, M. Structured and disordered facets of the GPCR fold. Curr. Opin. Struct. Biol. 2014, 27, 129–137. [Google Scholar] [CrossRef] [PubMed]
- Babu, M.M.; Kriwacki, R.W.; Pappu, R.V. Versatility from Protein Disorder. Science 2012, 337, 1460–1461. [Google Scholar] [CrossRef] [Green Version]
- Fonin, A.V.; Darling, A.L.; Kuznetsova, I.M.; Turoverov, K.K.; Uversky, V.N. Multi-functionality of proteins involved in GPCR and G protein signaling: Making sense of structure-function continuum with intrinsic disorder-based proteoforms. Cell. Mol. Life Sci. CMLS 2019, 76, 4461–4492. [Google Scholar] [CrossRef] [PubMed]
- Luca, S.; White, J.F.; Sohal, A.K.; Filippov, D.V.; van Boom, J.H.; Grisshammer, R.; Baldus, M. The conformation of neurotensin bound to its G protein-coupled receptor. Proc. Natl. Acad. Sci. USA 2003, 100, 10706–10711. [Google Scholar] [CrossRef] [Green Version]
- Schwyzer, R. ACTH: A short introductory review. Ann. N. Y. Acad. Sci. 1977, 297, 3–26. [Google Scholar] [CrossRef] [PubMed]
- Schwyzer, R. Molecular mechanism of opioid receptor selection. Biochemistry 1986, 25, 6335–6342. [Google Scholar] [CrossRef] [PubMed]
- Sargent, D.F.; Schwyzer, R. Membrane lipid phase as catalyst for peptide-receptor interactions. Proc. Natl. Acad. Sci. USA 1986, 83, 5774–5778. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schwyzer, R. In search of the ‘bio-active conformation’—Is it induced by the target cell membrane? J. Mol. Recognit. 1995, 8, 3–8. [Google Scholar] [CrossRef]
- Schwyzer, R. Peptide–membrane interactions and a new principle in quantitative structure–activity relationships. Biopolymers 1991, 31, 785–792. [Google Scholar] [CrossRef]
- Bader, R.; Zerbe, O. Are hormones from the neuropeptide Y family recognized by their receptors from the membrane-bound state? ChemBioChem 2005, 6, 1520–1534. [Google Scholar] [CrossRef]
- Moroder, L.; Romano, R.; Guba, W.; Mierke, D.F.; Kessler, H.; Delporte, C.; Winand, J.; Christophe, J. New evidence for a membrane-bound pathway in hormone receptor binding. Biochemistry 1993, 32, 13551–13559. [Google Scholar] [CrossRef]
- Lopez, J.J.; Shukla, A.K.; Reinhart, C.; Schwalbe, H.; Michel, H.; Glaubitz, C. The structure of the neuropeptide bradykinin bound to the human G-protein coupled receptor bradykinin B2 as determined by solid-state NMR spectroscopy. Angew. Chem. Int. Ed. 2008, 47, 1668–1671. [Google Scholar] [CrossRef]
- Banères, J.-L.; Popot, J.-L.; Mouillac, B. New advances in production and functional folding of G-protein-coupled receptors. Trends Biotechnol. 2011, 29, 314–322. [Google Scholar] [CrossRef] [Green Version]
- Xiang, J.; Chun, E.; Liu, C.; Jing, L.; Al-Sahouri, Z.; Zhu, L.; Liu, W. Successful Strategies to Determine High-Resolution Structures of GPCRs. Trends Pharmacol. Sci. 2016, 37, 1055–1069. [Google Scholar] [CrossRef]
- Wiseman, D.N.; Otchere, A.; Patel, J.H.; Uddin, R.; Pollock, N.L.; Routledge, S.J.; Rothnie, A.J.; Slack, C.; Poyner, D.R.; Bill, R.M.; et al. Expression and purification of recombinant G protein-coupled receptors: A review. Protein Expr. Purif. 2020, 167, 105524. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.J.; Howell, S.C.; Van Horn, W.D.; Jeon, Y.H.; Sanders, C.R. Recent Advances in the Application of Solution NMR Spectroscopy to Multi-Span Integral Membrane Proteins. Prog. Nucl. Magn. Reson. Spectrosc. 2009, 55, 335–360. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Park, S.H.; Casagrande, F.; Das, B.B.; Albrecht, L.; Chu, M.; Opella, S.J. Local and global dynamics of the G protein-coupled receptor CXCR1. Biochemistry 2011, 50, 2371–2380. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Werner, K.; Richter, C.; Klein-Seetharaman, J.; Schwalbe, H. Isotope labeling of mammalian GPCRs in HEK293 cells and characterization of the C-terminus of bovine rhodopsin by high resolution liquid NMR spectroscopy. J. Biomol. NMR 2008, 40, 49–53. [Google Scholar] [CrossRef]
- Wiktor, M.; Morin, S.; Sass, H.-J.; Kebbel, F.; Grzesiek, S. Biophysical and structural investigation of bacterially expressed and engineered CCR5, a G protein-coupled receptor. J. Biomol. NMR 2013, 55, 79–95. [Google Scholar] [CrossRef] [Green Version]
- Mittermaier, A.K.; Kay, L.E. Observing biological dynamics at atomic resolution using NMR. Trends Biochem. Sci. 2009, 34, 601–611. [Google Scholar] [CrossRef]
- Laws, D.D.; Bitter, H.-M.L.; Jerschow, A. Solid-state NMR spectroscopic methods in chemistry. Angew. Chem. Int. Ed. 2002, 41, 3096–3129. [Google Scholar] [CrossRef]
- Park, S.H.; Lee, J.H. Dynamic G Protein-Coupled Receptor Signaling Probed by Solution NMR Spectroscopy. Biochemistry 2020, 59, 1065–1080. [Google Scholar] [CrossRef]
- Raingeval, C.; Krimm, I. NMR investigation of protein-ligand interactions for G-protein coupled receptors. Future Med. Chem. 2019, 11, 1811–1825. [Google Scholar] [CrossRef]
- Ueda, T.; Kofuku, Y.; Okude, J.; Imai, S.; Shiraishi, Y.; Shimada, I. Function-related conformational dynamics of G protein-coupled receptors revealed by NMR. Biophys. Rev. 2019, 11, 409–418. [Google Scholar] [CrossRef]
- Kumar, A.; Ernst, R.R.; Wüthrich, K. A two-dimensional nuclear Overhauser enhancement (2D NOE) experiment for the elucidation of complete proton-proton cross-relaxation networks in biological macromolecules. Biochem. Biophys. Res. Commun. 1980, 95, 1–6. [Google Scholar] [CrossRef]
- Mo, H.; Pochapsky, T.C. Intermolecular interactions characterized by nuclear Overhauser effects. Prog. Nucl. Magn. Reson. Spectrosc. 1997, 30, 1–38. [Google Scholar] [CrossRef]
- Balaram, P.; Bothner-By, A.A.; Dadok, J. Negative nuclear Overhauser effects as probes of macromolecular structure. J. Am. Chem. Soc. 1972, 94, 4015–4017. [Google Scholar] [CrossRef]
- Meyer, B.; Peters, T. NMR spectroscopy techniques for screening and identifying ligand binding to protein receptors. Angew. Chem. Int. Ed. 2003, 42, 864–890. [Google Scholar] [CrossRef] [PubMed]
- Inooka, H.; Ohtaki, T.; Kitahara, O.; Ikegami, T.; Endo, S.; Kitada, C.; Ogi, K.; Onda, H.; Fujino, M.; Shirakawa, M. Conformation of a peptide ligand bound to its G-protein coupled receptor. Nat. Struct. Biol. 2001, 8, 161–165. [Google Scholar] [CrossRef] [PubMed]
- O’Connor, C.; White, K.L.; Doncescu, N.; Didenko, T.; Roth, B.L.; Czaplicki, G.; Stevens, R.C.; Wüthrich, K.; Milon, A. NMR structure and dynamics of the agonist dynorphin peptide bound to the human kappa opioid receptor. Proc. Natl. Acad. Sci. USA 2015, 112, 11852–11857. [Google Scholar] [CrossRef] [Green Version]
- Chavkin, C.; Goldstein, A. Specific receptor for the opioid peptide dynorphin: Structure--activity relationships. Proc. Natl. Acad. Sci. USA 1981, 78, 6543–6547. [Google Scholar] [CrossRef] [Green Version]
- Ferré, G.; Louet, M.; Saurel, O.; Delort, B.; Czaplicki, G.; M’Kadmi, C.; Damian, M.; Renault, P.; Cantel, S.; Gavara, L.; et al. Structure and dynamics of G protein-coupled receptor-bound ghrelin reveal the critical role of the octanoyl chain. Proc. Natl. Acad. Sci. USA 2019, 116, 17525–17530. [Google Scholar] [CrossRef] [Green Version]
- Bender, B.J.; Vortmeier, G.; Ernicke, S.; Bosse, M.; Kaiser, A.; Els-Heindl, S.; Krug, U.; Beck-Sickinger, A.; Meiler, J.; Huster, D. Structural Model of Ghrelin Bound to its G Protein-Coupled Receptor. Structure 2019, 27, 537–544.e4. [Google Scholar] [CrossRef] [Green Version]
- Shimada, I. NMR techniques for identifying the interface of a larger protein-protein complex: Cross-saturation and transferred cross-saturation experiments. Methods Enzymol. 2005, 394, 483–506. [Google Scholar] [CrossRef]
- Kofuku, Y.; Yoshiura, C.; Ueda, T.; Terasawa, H.; Hirai, T.; Tominaga, S.; Hirose, M.; Maeda, Y.; Takahashi, H.; Terashima, Y.; et al. Structural basis of the interaction between chemokine stromal cell-derived factor-1/CXCL12 and its G-protein-coupled receptor CXCR4. J. Biol. Chem. 2009, 284, 35240–35250. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yoshiura, C.; Kofuku, Y.; Ueda, T.; Mase, Y.; Yokogawa, M.; Osawa, M.; Terashima, Y.; Matsushima, K.; Shimada, I. NMR analyses of the interaction between CCR5 and its ligand using functional reconstitution of CCR5 in lipid bilayers. J. Am. Chem. Soc. 2010, 132, 6768–6777. [Google Scholar] [CrossRef] [PubMed]
- Saitô, H. Conformation-dependent 13C chemical shifts: A new means of conformational characterization as obtained by high-resolution solid-state 13C NMR. Magn. Reson. Chem. 1986, 24, 835–852. [Google Scholar] [CrossRef]
- Spera, S.; Bax, A. Empirical correlation between protein backbone conformation and C.alpha. and C.beta. 13C nuclear magnetic resonance chemical shifts. J. Am. Chem. Soc. 1991, 113, 5490–5492. [Google Scholar] [CrossRef]
- Wishart, D.; Sykes, B. The 13C Chemical-Shift Index: A simple method for the identification of protein secondary structure using 13C chemical-shift data. J. Biomol. NMR 1994, 4. [Google Scholar] [CrossRef]
- Cornilescu, G.; Delaglio, F.; Bax, A. Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J. Biomol. NMR 1999, 13, 289–302. [Google Scholar] [CrossRef]
- Shen, Y.; Delaglio, F.; Cornilescu, G.; Bax, A. TALOS+: A hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J. Biomol. NMR 2009, 44, 213–223. [Google Scholar] [CrossRef]
- Rosay, M.; Lansing, J.C.; Haddad, K.C.; Bachovchin, W.W.; Herzfeld, J.; Temkin, R.J.; Griffin, R.G. High-frequency dynamic nuclear polarization in MAS spectra of membrane and soluble proteins. J. Am. Chem. Soc. 2003, 125, 13626–13627. [Google Scholar] [CrossRef]
- Mak-Jurkauskas, M.L.; Bajaj, V.S.; Hornstein, M.K.; Belenky, M.; Griffin, R.G.; Herzfeld, J. Energy transformations early in the bacteriorhodopsin photocycle revealed by DNP-enhanced solid-state NMR. Proc. Natl. Acad. Sci. USA 2008, 105, 883–888. [Google Scholar] [CrossRef] [Green Version]
- Lilly Thankamony, A.S.; Wittmann, J.J.; Kaushik, M.; Corzilius, B. Dynamic nuclear polarization for sensitivity enhancement in modern solid-state NMR. Prog. Nucl. Magn. Reson. Spectrosc. 2017, 102–103, 120–195. [Google Scholar] [CrossRef]
- Rajagopal, S.; Rajagopal, K.; Lefkowitz, R.J. Teaching old receptors new tricks: Biasing seven-transmembrane receptors. Nat. Rev. Drug Discov. 2010, 9, 373–386. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kenakin, T. Biased Receptor Signaling in Drug Discovery. Pharmacol. Rev. 2019, 71, 267–315. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wingler, L.M.; Elgeti, M.; Hilger, D.; Latorraca, N.R.; Lerch, M.T.; Staus, D.P.; Dror, R.O.; Kobilka, B.K.; Hubbell, W.L.; Lefkowitz, R.J. Angiotensin Analogs with Divergent Bias Stabilize Distinct Receptor Conformations. Cell 2019, 176, 468–478.e11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ye, L.; Van Eps, N.; Zimmer, M.; Ernst, O.P.; Prosser, R.S. Activation of the A2A adenosine G-protein-coupled receptor by conformational selection. Nature 2016, 533, 265–268. [Google Scholar] [CrossRef] [PubMed]
- Clark, L.D.; Dikiy, I.; Chapman, K.; Rödström, K.E.; Aramini, J.; LeVine, M.V.; Khelashvili, G.; Rasmussen, S.G.; Gardner, K.H.; Rosenbaum, D.M. Ligand modulation of sidechain dynamics in a wild-type human GPCR. eLife 2017, 6, e28505. [Google Scholar] [CrossRef] [Green Version]
- Schmidt, P.; Thomas, L.; Müller, P.; Scheidt, H.A.; Huster, D. The G-protein-coupled neuropeptide Y receptor type 2 is highly dynamic in lipid membranes as revealed by solid-state NMR spectroscopy. Chemistry 2014, 20, 4986–4992. [Google Scholar] [CrossRef]
- Schrottke, S.; Kaiser, A.; Vortmeier, G.; Els-Heindl, S.; Worm, D.; Bosse, M.; Schmidt, P.; Scheidt, H.A.; Beck-Sickinger, A.G.; Huster, D. Expression, Functional Characterization, and Solid-State NMR Investigation of the G Protein-Coupled GHS Receptor in Bilayer Membranes. Sci. Rep. 2017, 7, 46128. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thomas, L.; Kahr, J.; Schmidt, P.; Krug, U.; Scheidt, H.A.; Huster, D. The dynamics of the G protein-coupled neuropeptide Y2 receptor in monounsaturated membranes investigated by solid-state NMR spectroscopy. J. Biomol. NMR 2015, 61, 347–359. [Google Scholar] [CrossRef]
- Katritch, V.; Cherezov, V.; Stevens, R.C. Structure-function of the G protein-coupled receptor superfamily. Annu. Rev. Pharmacol. Toxicol. 2013, 53, 531–556. [Google Scholar] [CrossRef] [Green Version]
- Kobilka, B.; Schertler, G.F.X. New G-protein-coupled receptor crystal structures: Insights and limitations. Trends Pharmacol. Sci. 2008, 29, 79–83. [Google Scholar] [CrossRef]
- Deupi, X.; Standfuss, J. Structural insights into agonist-induced activation of G-protein-coupled receptors. Curr. Opin. Struct. Biol. 2011, 21, 541–551. [Google Scholar] [CrossRef] [PubMed]
- Yarnitzky, T.; Levit, A.; Niv, M.Y. Homology modeling of G-protein-coupled receptors with X-ray structures on the rise. Curr. Opin. Drug Discov. Dev. 2010, 13, 317–325. [Google Scholar]
- Costanzi, S.; Wang, K. The GPCR crystallography boom: Providing an invaluable source of structural information and expanding the scope of homology modeling. Adv. Exp. Med. Biol. 2014, 796, 3–13. [Google Scholar] [CrossRef] [PubMed]
- Bender, B.J.; Marlow, B.; Meiler, J. RosettaGPCR: Multiple Template Homology Modeling of GPCRs with Rosetta. bioRxiv 2019. [Google Scholar] [CrossRef] [Green Version]
- Verardi, R.; Traaseth, N.J.; Masterson, L.R.; Vostrikov, V.V.; Veglia, G. Isotope Labeling for Solution and Solid-State NMR Spectroscopy of Membrane Proteins. Adv. Exp. Med. Biol. 2012, 992, 35–62. [Google Scholar] [CrossRef] [Green Version]
- Lin, M.T.; Sperling, L.J.; Frericks Schmidt, H.L.; Tang, M.; Samoilova, R.I.; Kumasaka, T.; Iwasaki, T.; Dikanov, S.A.; Rienstra, C.M.; Gennis, R.B. A rapid and robust method for selective isotope labeling of proteins. Methods 2011, 55, 370–378. [Google Scholar] [CrossRef] [Green Version]
- Sobhanifar, S.; Reckel, S.; Junge, F.; Schwarz, D.; Kai, L.; Karbyshev, M.; Löhr, F.; Bernhard, F.; Dötsch, V. Cell-free expression and stable isotope labelling strategies for membrane proteins. J. Biomol. NMR 2010, 46, 33–43. [Google Scholar] [CrossRef]
- Krug, U.; Gloge, A.; Schmidt, P.; Becker-Baldus, J.; Bernhard, F.; Kaiser, A.; Montag, C.; Gauglitz, M.; Vishnivetskiy, S.A.; Gurevich, V.V.; et al. The Conformational Equilibrium of the Neuropeptide Y2 Receptor in Bilayer Membranes. Angew. Chem. Int. Ed. 2020. [Google Scholar] [CrossRef]
- Klare, J.P. Site-directed spin labeling EPR spectroscopy in protein research. Biol. Chem. 2013, 394, 1281–1300. [Google Scholar] [CrossRef] [PubMed]
- Sletten, E.M.; Bertozzi, C.R. Bioorthogonal Chemistry: Fishing for Selectivity in a Sea of Functionality. Angew. Chem. Int. Ed. 2009, 48, 6974–6998. [Google Scholar] [CrossRef] [Green Version]
- Resek, J.F.; Farahbakhsh, Z.T.; Hubbell, W.L.; Khorana, H.G. Formation of the meta II photointermediate is accompanied by conformational changes in the cytoplasmic surface of rhodopsin. Biochemistry 1993, 32, 12025–12032. [Google Scholar] [CrossRef] [PubMed]
- Gether, U.; Lin, S.; Ghanouni, P.; Ballesteros, J.A.; Weinstein, H.; Kobilka, B.K. Agonists induce conformational changes in transmembrane domains III and VI of the beta2 adrenoceptor. EMBO J. 1997, 16, 6737–6747. [Google Scholar] [CrossRef] [Green Version]
- Mary, S.; Damian, M.; Louet, M.; Floquet, N.; Fehrentz, J.-A.; Marie, J.; Martinez, J.; Baneres, J.-L. Ligands and signaling proteins govern the conformational landscape explored by a G protein-coupled receptor. Proc. Natl. Acad. Sci. USA 2012, 109, 8304–8309. [Google Scholar] [CrossRef] [Green Version]
- Witte, K.; Kaiser, A.; Schmidt, P.; Splith, V.; Thomas, L.; Berndt, S.; Huster, D.; Beck-Sickinger, A.G. Oxidative in vitro folding of a cysteine deficient variant of the G protein-coupled neuropeptide Y receptor type 2 improves stability at high concentration. Biol. Chem. 2013, 394, 1045–1056. [Google Scholar] [CrossRef] [PubMed]
- Lotze, J.; Reinhardt, U.; Seitz, O.; Beck-Sickinger, A.G. Peptide-tags for site-specific protein labelling in vitro and in vivo. Mol. Biosyst. 2016, 12, 1731–1745. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harmand, T.J.; Murar, C.E.; Bode, J.W. New chemistries for chemoselective peptide ligations and the total synthesis of proteins. Curr. Opin. Chem. Biol. 2014, 22C, 115–121. [Google Scholar] [CrossRef] [PubMed]
- Shiraishi, Y.; Natsume, M.; Kofuku, Y.; Imai, S.; Nakata, K.; Mizukoshi, T.; Ueda, T.; Iwaï, H.; Shimada, I. Phosphorylation-induced conformation of β2-adrenoceptor related to arrestin recruitment revealed by NMR. Nat. Commun. 2018, 9, 194. [Google Scholar] [CrossRef] [Green Version]
- Huber, T.; Sakmar, T.P. Chemical biology methods for investigating G protein-coupled receptor signaling. Chem. Biol. 2014, 21, 1224–1237. [Google Scholar] [CrossRef] [Green Version]
- Tian, H.; Fürstenberg, A.; Huber, T. Labeling and Single-Molecule Methods to Monitor G Protein-Coupled Receptor Dynamics. Chem. Rev. 2017, 117, 186–245. [Google Scholar] [CrossRef]
- Serfling, R.; Seidel, L.; Bock, A.; Lohse, M.J.; Annibale, P.; Coin, I. Quantitative Single-Residue Bioorthogonal Labeling of G Protein-Coupled Receptors in Live Cells. ACS Chem. Biol. 2019, 14, 1141–1149. [Google Scholar] [CrossRef]
- Schmidt, M.J.; Fedoseev, A.; Bücker, D.; Borbas, J.; Peter, C.; Drescher, M.; Summerer, D. EPR Distance Measurements in Native Proteins with Genetically Encoded Spin Labels. ACS Chem. Biol. 2015, 10, 2764–2771. [Google Scholar] [CrossRef]
- Fleissner, M.R.; Brustad, E.M.; Kálai, T.; Altenbach, C.; Cascio, D.; Peters, F.B.; Hideg, K.; Peuker, S.; Schultz, P.G.; Hubbell, W.L. Site-directed spin labeling of a genetically encoded unnatural amino acid. Proc. Natl. Acad. Sci. USA 2009, 106, 21637–21642. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schmidt, M.J.; Borbas, J.; Drescher, M.; Summerer, D. A genetically encoded spin label for electron paramagnetic resonance distance measurements. J. Am. Chem. Soc. 2014, 136, 1238–1241. [Google Scholar] [CrossRef] [PubMed]
- Ye, S.; Huber, T.; Vogel, R.; Sakmar, T.P. FTIR analysis of GPCR activation using azido probes. Nat. Chem. Biol. 2009, 5, 397–399. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kofuku, Y.; Ueda, T.; Okude, J.; Shiraishi, Y.; Kondo, K.; Maeda, M.; Tsujishita, H.; Shimada, I. Efficacy of the β2-adrenergic receptor is determined by conformational equilibrium in the transmembrane region. Nat. Commun. 2012, 3, 1045. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nygaard, R.; Zou, Y.; Dror, R.O.; Mildorf, T.J.; Arlow, D.H.; Manglik, A.; Pan, A.C.; Liu, C.W.; Fung, J.J.; Bokoch, M.P.; et al. The dynamic process of β(2)-adrenergic receptor activation. Cell 2013, 152, 532–542. [Google Scholar] [CrossRef] [Green Version]
- Kofuku, Y.; Ueda, T.; Okude, J.; Shiraishi, Y.; Kondo, K.; Mizumura, T.; Suzuki, S.; Shimada, I. Functional Dynamics of Deuterated β2 -Adrenergic Receptor in Lipid Bilayers Revealed by NMR Spectroscopy. Angew. Chem. Int. Ed. 2014, 53, 13376–13379. [Google Scholar] [CrossRef]
- Okude, J.; Ueda, T.; Kofuku, Y.; Sato, M.; Nobuyama, N.; Kondo, K.; Shiraishi, Y.; Mizumura, T.; Onishi, K.; Natsume, M.; et al. Identification of a Conformational Equilibrium That Determines the Efficacy and Functional Selectivity of the μ-Opioid Receptor. Angew. Chem. Int. Ed. 2015, 54, 15771–15776. [Google Scholar] [CrossRef] [Green Version]
- Sounier, R.; Mas, C.; Steyaert, J.; Laeremans, T.; Manglik, A.; Huang, W.; Kobilka, B.K.; Déméné, H.; Granier, S. Propagation of conformational changes during μ-opioid receptor activation. Nature 2015, 524, 375–378. [Google Scholar] [CrossRef] [Green Version]
- Solt, A.S.; Bostock, M.J.; Shrestha, B.; Kumar, P.; Warne, T.; Tate, C.G.; Nietlispach, D. Insight into partial agonism by observing multiple equilibria for ligand-bound and Gs-mimetic nanobody-bound β1-adrenergic receptor. Nat. Commun. 2017, 8, 1795. [Google Scholar] [CrossRef]
- Ruschak, A.M.; Kay, L.E. Methyl groups as probes of supra-molecular structure, dynamics and function. J. Biomol. NMR 2010, 46, 75–87. [Google Scholar] [CrossRef] [PubMed]
- Kurauskas, V.; Schanda, P.; Sounier, R. Methyl-Specific Isotope Labelling Strategies for NMR studies of Membrane Proteins. Membr. Protein Struct. Funct. Charact. 2017, 1635, 109–123. [Google Scholar] [CrossRef] [Green Version]
- Ballesteros, J.A.; Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors. In Methods in Neurosciences; Sealfon, S.C., Ed.; Receptor Molecular Biology; Academic Press: Cambridge, MA, USA, 1995; Volume 25, pp. 366–428. ISBN 1043-9471. [Google Scholar]
- Manglik, A.; Kim, T.H.; Masureel, M.; Altenbach, C.; Yang, Z.; Hilger, D.; Lerch, M.T.; Kobilka, T.S.; Thian, F.S.; Hubbell, W.L.; et al. Structural Insights into the Dynamic Process of β2-Adrenergic Receptor Signaling. Cell 2015, 161, 1101–1111. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- DeVree, B.T.; Mahoney, J.P.; Vélez-Ruiz, G.A.; Rasmussen, S.G.F.; Kuszak, A.J.; Edwald, E.; Fung, J.-J.; Manglik, A.; Masureel, M.; Du, Y.; et al. Allosteric coupling from G protein to the agonist-binding pocket in GPCRs. Nature 2016, 535, 182–186. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Staus, D.P.; Strachan, R.T.; Manglik, A.; Pani, B.; Kahsai, A.W.; Kim, T.H.; Wingler, L.M.; Ahn, S.; Chatterjee, A.; Masoudi, A.; et al. Allosteric nanobodies reveal the dynamic range and diverse mechanisms of G-protein-coupled receptor activation. Nature 2016, 535, 448–452. [Google Scholar] [CrossRef] [PubMed]
- Eddy, M.T.; Lee, M.-Y.; Gao, Z.-G.; White, K.L.; Didenko, T.; Horst, R.; Audet, M.; Stanczak, P.; McClary, K.M.; Han, G.W.; et al. Allosteric Coupling of Drug Binding and Intracellular Signaling in the A2A Adenosine Receptor. Cell 2018, 172, 68–80.e12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Frei, J.N.; Broadhurst, R.W.; Bostock, M.J.; Solt, A.; Jones, A.J.Y.; Gabriel, F.; Tandale, A.; Shrestha, B.; Nietlispach, D. Conformational plasticity of ligand-bound and ternary GPCR complexes studied by 19F NMR of the β1-adrenergic receptor. Nat. Commun. 2020, 11, 669. [Google Scholar] [CrossRef] [PubMed]
- Casiraghi, M.; Damian, M.; Lescop, E.; Point, E.; Moncoq, K.; Morellet, N.; Levy, D.; Marie, J.; Guittet, E.; Banères, J.-L.; et al. Functional Modulation of a G Protein-Coupled Receptor Conformational Landscape in a Lipid Bilayer. J. Am. Chem. Soc. 2016, 138, 11170–11175. [Google Scholar] [CrossRef]
- Gregorio, G.G.; Masureel, M.; Hilger, D.; Terry, D.S.; Juette, M.; Zhao, H.; Zhou, Z.; Perez-Aguilar, J.M.; Hauge, M.; Mathiasen, S.; et al. Single-molecule analysis of ligand efficacy in β2AR-G-protein activation. Nature 2017, 547, 68–73. [Google Scholar] [CrossRef]
- Larda, S.T.; Bokoch, M.P.; Evanics, F.; Prosser, R.S. Lysine methylation strategies for characterizing protein conformations by NMR. J. Biomol. NMR 2012, 54, 199–209. [Google Scholar] [CrossRef]
- Tugarinov, V.; Hwang, P.M.; Ollerenshaw, J.E.; Kay, L.E. Cross-correlated relaxation enhanced 1H[bond]13C NMR spectroscopy of methyl groups in very high molecular weight proteins and protein complexes. J. Am. Chem. Soc. 2003, 125, 10420–10428. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.J.; Horst, R.; Katritch, V.; Stevens, R.C.; Wuthrich, K. Biased Signaling Pathways in 2-Adrenergic Receptor Characterized by 19F-NMR. Science 2012, 335, 1106–1110. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chung, K.Y.; Kim, T.H.; Manglik, A.; Alvares, R.; Kobilka, B.K.; Prosser, R.S. Role of detergents in conformational exchange of a G protein-coupled receptor. J. Biol. Chem. 2012, 287, 36305–36311. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Horst, R.; Liu, J.J.; Stevens, R.C.; Wüthrich, K. β2-adrenergic receptor activation by agonists studied with 19F NMR spectroscopy. Angew. Chem. Int. Ed. 2013, 52, 10762–10765. [Google Scholar] [CrossRef] [Green Version]
- Loewen, M.C.; Klein-Seetharaman, J.; Getmanova, E.V.; Reeves, P.J.; Schwalbe, H.; Khorana, H.G. Solution 19F nuclear Overhauser effects in structural studies of the cytoplasmic domain of mammalian rhodopsin. Proc. Natl. Acad. Sci. USA 2001, 98, 4888–4892. [Google Scholar] [CrossRef] [Green Version]
- Klein-Seetharaman, J.; Getmanova, E.V.; Loewen, M.C.; Reeves, P.J.; Khorana, H.G. NMR spectroscopy in studies of light-induced structural changes in mammalian rhodopsin: Applicability of solution (19)F NMR. Proc. Natl. Acad. Sci. USA 1999, 96, 13744–13749. [Google Scholar] [CrossRef] [Green Version]
- Sušac, L.; Eddy, M.T.; Didenko, T.; Stevens, R.C.; Wüthrich, K. A2A adenosine receptor functional states characterized by 19F-NMR. Proc. Natl. Acad. Sci. USA 2018, 115, 12733–12738. [Google Scholar] [CrossRef] [Green Version]
- Kim, T.H.; Chung, K.Y.; Manglik, A.; Hansen, A.L.; Dror, R.O.; Mildorf, T.J.; Shaw, D.E.; Kobilka, B.K.; Prosser, R.S. The Role of Ligands on the Equilibria between Functional States of a G Protein-Coupled Receptor. J. Am. Chem. Soc. 2013, 135, 9465–9474. [Google Scholar] [CrossRef] [Green Version]
- Eddy, M.T.; Didenko, T.; Stevens, R.C.; Wüthrich, K. β2-Adrenergic Receptor Conformational Response to Fusion Protein in the Third Intracellular Loop. Structure 2016, 24, 2190–2197. [Google Scholar] [CrossRef] [Green Version]
- Didenko, T.; Liu, J.J.; Horst, R.; Stevens, R.C.; Wüthrich, K. Fluorine-19 NMR of integral membrane proteins illustrated with studies of GPCRs. Curr. Opin. Struct. Biol. 2013, 23, 740–747. [Google Scholar] [CrossRef] [Green Version]
- Danielson, M.A.; Falke, J.J. Use of 19F to probe protein structure and conformational changes. Annu. Rev. Biophys. Biomol. Struct. 1996, 25, 163–195. [Google Scholar] [CrossRef] [PubMed]
- Crocker, E.; Eilers, M.; Ahuja, S.; Hornak, V.; Hirshfeld, A.; Sheves, M.; Smith, S.O. Location of Trp265 in metarhodopsin II: Implications for the activation mechanism of the visual receptor rhodopsin. J. Mol. Biol. 2006, 357, 163–172. [Google Scholar] [CrossRef] [PubMed]
- Stehle, J.; Silvers, R.; Werner, K.; Chatterjee, D.; Gande, S.; Scholz, F.; Dutta, A.; Wachtveitl, J.; Klein-Seetharaman, J.; Schwalbe, H. Characterization of the simultaneous decay kinetics of metarhodopsin states II and III in rhodopsin by solution-state NMR spectroscopy. Angew. Chem. Int. Ed. 2014, 53, 2078–2084. [Google Scholar] [CrossRef]
- Eddy, M.T.; Gao, Z.-G.; Mannes, P.; Patel, N.; Jacobson, K.A.; Katritch, V.; Stevens, R.C.; Wüthrich, K. Extrinsic Tryptophans as NMR Probes of Allosteric Coupling in Membrane Proteins: Application to the A2A Adenosine Receptor. J. Am. Chem. Soc. 2018. [Google Scholar] [CrossRef] [PubMed]
- Isogai, S.; Deupi, X.; Opitz, C.; Heydenreich, F.M.; Tsai, C.-J.; Brueckner, F.; Schertler, G.F.X.; Veprintsev, D.B.; Grzesiek, S. Backbone NMR reveals allosteric signal transduction networks in the β1-adrenergic receptor. Nature 2016, 530, 237–241. [Google Scholar] [CrossRef]
- Frederick, K.K.; Marlow, M.S.; Valentine, K.G.; Wand, A.J. Conformational entropy in molecular recognition by proteins. Nature 2007, 448, 325–329. [Google Scholar] [CrossRef]
- Marlow, M.S.; Dogan, J.; Frederick, K.K.; Valentine, K.G.; Wand, A.J. The role of conformational entropy in molecular recognition by calmodulin. Nat. Chem. Biol. 2010, 6, 352–358. [Google Scholar] [CrossRef] [Green Version]
- Klare, J.P.; Steinhoff, H.-J. Spin labeling EPR. Photosynth. Res. 2009, 102, 377–390. [Google Scholar] [CrossRef]
- Claxton, D.P.; Kazmier, K.; Mishra, S.; Mchaourab, H.S. Navigating membrane protein structure, dynamics, and energy landscapes using spin labeling and EPR spectroscopy. Methods Enzymol. 2015, 564, 349–387. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Berliner, L.J.; Grunwald, J.; Hankovszky, H.O.; Hideg, K. A novel reversible thiol-specific spin label: Papain active site labeling and inhibition. Anal. Biochem. 1982, 119, 450–455. [Google Scholar] [CrossRef]
- Hubbell, W.L.; Cafiso, D.S.; Altenbach, C. Identifying conformational changes with site-directed spin labeling. Nat. Struct. Biol. 2000, 7, 735–739. [Google Scholar] [CrossRef] [PubMed]
- Farahbakhsh, Z.T.; Ridge, K.D.; Khorana, H.G.; Hubbell, W.L. Mapping light-dependent structural changes in the cytoplasmic loop connecting helices C and D in rhodopsin: A site-directed spin labeling study. Biochemistry 1995, 34, 8812–8819. [Google Scholar] [CrossRef] [PubMed]
- Altenbach, C.; Yang, K.; Farrens, D.L.; Farahbakhsh, Z.T.; Khorana, H.G.; Hubbell, W.L. Structural features and light-dependent changes in the cytoplasmic interhelical E-F loop region of rhodopsin: A site-directed spin-labeling study. Biochemistry 1996, 35, 12470–12478. [Google Scholar] [CrossRef] [PubMed]
- Dijkman, P.M.; Muñoz-García, J.C.; Lavington, S.R.; Kumagai, P.S.; Dos Reis, R.I.; Yin, D.; Stansfeld, P.J.; Costa-Filho, A.J.; Watts, A. Conformational dynamics of a G protein–coupled receptor helix 8 in lipid membranes. Sci. Adv. 2020, 6, eaav8207. [Google Scholar] [CrossRef] [PubMed]
- Rabenstein, M.D.; Shin, Y.K. Determination of the distance between two spin labels attached to a macromolecule. Proc. Natl. Acad. Sci. USA 1995, 92, 8239–8243. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McHaourab, H.S.; Steed, P.R.; Kazmier, K. Toward the fourth dimension of membrane protein structure: Insight into dynamics from spin-labeling EPR spectroscopy. Structure 2011, 19, 1549–1561. [Google Scholar] [CrossRef] [Green Version]
- Van Eps, N.; Caro, L.N.; Morizumi, T.; Kusnetzow, A.K.; Szczepek, M.; Hofmann, K.P.; Bayburt, T.H.; Sligar, S.G.; Ernst, O.P.; Hubbell, W.L. Conformational equilibria of light-activated rhodopsin in nanodiscs. Proc. Natl. Acad. Sci. USA 2017, 114, E3268–E3275. [Google Scholar] [CrossRef] [Green Version]
- Altenbach, C.; Kusnetzow, A.K.; Ernst, O.P.; Hofmann, K.P.; Hubbell, W.L. High-resolution distance mapping in rhodopsin reveals the pattern of helix movement due to activation. Proc. Natl. Acad. Sci. USA 2008, 105, 7439–7444. [Google Scholar] [CrossRef] [Green Version]
- Van Eps, N.; Caro, L.N.; Morizumi, T.; Ernst, O.P. Characterizing rhodopsin signaling by EPR spectroscopy: From structure to dynamics. Photochem. Photobiol. Sci. 2015, 14, 1586–1597. [Google Scholar] [CrossRef] [PubMed]
- Knierim, B.; Hofmann, K.P.; Ernst, O.P.; Hubbell, W.L. Sequence of late molecular events in the activation of rhodopsin. Proc. Natl. Acad. Sci. USA 2007, 104, 20290–20295. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Suomivuori, C.-M.; Latorraca, N.R.; Wingler, L.M.; Eismann, S.; King, M.C.; Kleinhenz, A.L.W.; Skiba, M.A.; Staus, D.P.; Kruse, A.C.; Lefkowitz, R.J.; et al. Molecular mechanism of biased signaling in a prototypical G protein-coupled receptor. Science 2020, 367, 881–887. [Google Scholar] [CrossRef] [PubMed]
- Van Eps, N.; Oldham, W.M.; Hamm, H.E.; Hubbell, W.L. Structural and dynamical changes in an alpha-subunit of a heterotrimeric G protein along the activation pathway. Proc. Natl. Acad. Sci. USA 2006, 103, 16194–16199. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van Eps, N.; Anderson, L.L.; Kisselev, O.G.; Baranski, T.J.; Hubbell, W.L.; Marshall, G.R. Electron paramagnetic resonance studies of functionally active, nitroxide spin-labeled peptide analogues of the C-terminus of a G-protein alpha subunit. Biochemistry 2010, 49, 6877–6886. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van Eps, N.; Preininger, A.M.; Alexander, N.; Kaya, A.I.; Meier, S.; Meiler, J.; Hamm, H.E.; Hubbell, W.L. Interaction of a G protein with an activated receptor opens the interdomain interface in the alpha subunit. Proc. Natl. Acad. Sci. USA 2011, 108, 9420–9424. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van Eps, N.; Altenbach, C.; Caro, L.N.; Latorraca, N.R.; Hollingsworth, S.A.; Dror, R.O.; Ernst, O.P.; Hubbell, W.L. Gi- and Gs-coupled GPCRs show different modes of G-protein binding. Proc. Natl. Acad. Sci. USA 2018, 115, 2383–2388. [Google Scholar] [CrossRef] [Green Version]
- Hanson, S.M.; Van Eps, N.; Francis, D.J.; Altenbach, C.; Vishnivetskiy, S.A.; Arshavsky, V.Y.; Klug, C.S.; Hubbell, W.L.; Gurevich, V.V. Structure and function of the visual arrestin oligomer. EMBO J. 2007, 26, 1726–1736. [Google Scholar] [CrossRef] [Green Version]
- Zhuo, Y.; Vishnivetskiy, S.A.; Zhan, X.; Gurevich, V.V.; Klug, C.S. Identification of Receptor Binding-induced Conformational Changes in Non-visual Arrestins. J. Biol. Chem. 2014, 289, 20991–21002. [Google Scholar] [CrossRef] [Green Version]
- Kang, Y.; Zhou, X.E.; Gao, X.; He, Y.; Liu, W.; Ishchenko, A.; Barty, A.; White, T.A.; Yefanov, O.; Han, G.W.; et al. Crystal structure of rhodopsin bound to arrestin by femtosecond X-ray laser. Nature 2015, 523, 561–567. [Google Scholar] [CrossRef] [Green Version]
- Ghanouni, P.; Steenhuis, J.J.; Farrens, D.L.; Kobilka, B.K. Agonist-induced conformational changes in the G-protein-coupling domain of the beta 2 adrenergic receptor. Proc. Natl. Acad. Sci. USA 2001, 98, 5997–6002. [Google Scholar] [CrossRef] [Green Version]
- Swaminath, G.; Xiang, Y.; Lee, T.W.; Steenhuis, J.; Parnot, C.; Kobilka, B.K. Sequential Binding of Agonists to the 2 Adrenoceptor: Kinetic Evidence for Intermediate Conformational States. J. Biol. Chem. 2004, 279, 686–691. [Google Scholar] [CrossRef] [Green Version]
- Swaminath, G.; Deupi, X.; Lee, T.W.; Zhu, W.; Thian, F.S.; Kobilka, T.S.; Kobilka, B. Probing the 2 Adrenoceptor Binding Site with Catechol Reveals Differences in Binding and Activation by Agonists and Partial Agonists. J. Biol. Chem. 2005, 280, 22165–22171. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kahsai, A.W.; Wisler, J.W.; Lee, J.; Ahn, S.; Cahill Iii, T.J.; Dennison, S.M.; Staus, D.P.; Thomsen, A.R.B.; Anasti, K.M.; Pani, B.; et al. Conformationally selective RNA aptamers allosterically modulate the β2-adrenoceptor. Nat. Chem. Biol. 2016, 12, 709–716. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yao, X.; Parnot, C.; Deupi, X.; Ratnala, V.R.P.; Swaminath, G.; Farrens, D.; Kobilka, B. Coupling ligand structure to specific conformational switches in the beta2-adrenoceptor. Nat. Chem. Biol. 2006, 2, 417–422. [Google Scholar] [CrossRef]
- Ghanouni, P.; Gryczynski, Z.; Steenhuis, J.J.; Lee, T.W.; Farrens, D.L.; Lakowicz, J.R.; Kobilka, B.K. Functionally different agonists induce distinct conformations in the G protein coupling domain of the beta 2 adrenergic receptor. J. Biol. Chem. 2001, 276, 24433–24436. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mielke, T.; Alexiev, U.; Gläsel, M.; Otto, H.; Heyn, M.P. Light-induced changes in the structure and accessibility of the cytoplasmic loops of rhodopsin in the activated MII state. Biochemistry 2002, 41, 7875–7884. [Google Scholar] [CrossRef]
- Alexiev, U.; Rimke, I.; Pöhlmann, T. Elucidation of the nature of the conformational changes of the EF-interhelical loop in bacteriorhodopsin and of the helix VIII on the cytoplasmic surface of bovine rhodopsin: A time-resolved fluorescence depolarization study. J. Mol. Biol. 2003, 328, 705–719. [Google Scholar] [CrossRef]
- Granier, S.; Kim, S.; Shafer, A.M.; Ratnala, V.R.P.; Fung, J.J.; Zare, R.N.; Kobilka, B. Structure and conformational changes in the C-terminal domain of the beta2-adrenoceptor: Insights from fluorescence resonance energy transfer studies. J. Biol. Chem. 2007, 282, 13895–13905. [Google Scholar] [CrossRef] [Green Version]
- Selvin, P.R. The renaissance of fluorescence resonance energy transfer. Nat. Struct. Biol. 2000, 7, 730–734. [Google Scholar] [CrossRef]
- Rahmeh, R.; Damian, M.; Cottet, M.; Orcel, H.; Mendre, C.; Durroux, T.; Sharma, K.S.; Durand, G.; Pucci, B.; Trinquet, E.; et al. Structural insights into biased G protein-coupled receptor signaling revealed by fluorescence spectroscopy. Proc. Natl. Acad. Sci. USA 2012, 109, 6733–6738. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cha, A.; Snyder, G.E.; Selvin, P.R.; Bezanilla, F. Atomic scale movement of the voltage-sensing region in a potassium channel measured via spectroscopy. Nature 1999, 402, 809–813. [Google Scholar] [CrossRef]
- Posson, D.J.; Ge, P.; Miller, C.; Bezanilla, F.; Selvin, P.R. Small vertical movement of a K+ channel voltage sensor measured with luminescence energy transfer. Nature 2005, 436, 848–851. [Google Scholar] [CrossRef] [PubMed]
- Lohse, M.J.; Nuber, S.; Hoffmann, C. Fluorescence/bioluminescence resonance energy transfer techniques to study G-protein-coupled receptor activation and signaling. Pharmacol. Rev. 2012, 64, 299–336. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lohse, M.J.; Maiellaro, I.; Calebiro, D. Kinetics and mechanism of G protein-coupled receptor activation. Curr. Opin. Cell Biol. 2014, 27, 87–93. [Google Scholar] [CrossRef]
- Kauk, M.; Hoffmann, C. Intramolecular and Intermolecular FRET Sensors for GPCRs—Monitoring Conformational Changes and Beyond. Trends Pharmacol. Sci. 2018, 39, 123–135. [Google Scholar] [CrossRef] [PubMed]
- Sandhu, M.; Touma, A.M.; Dysthe, M.; Sadler, F.; Sivaramakrishnan, S.; Vaidehi, N. Conformational plasticity of the intracellular cavity of GPCR-G-protein complexes leads to G-protein promiscuity and selectivity. Proc. Natl. Acad. Sci. USA 2019, 116, 11956–11965. [Google Scholar] [CrossRef] [Green Version]
- Namkung, Y.; LeGouill, C.; Kumar, S.; Cao, Y.; Teixeira, L.B.; Lukasheva, V.; Giubilaro, J.; Simões, S.C.; Longpré, J.-M.; Devost, D.; et al. Functional selectivity profiling of the angiotensin II type 1 receptor using pathway-wide BRET signaling sensors. Sci. Signal. 2018, 11, eaat1631. [Google Scholar] [CrossRef] [Green Version]
- Castro, M.; Nikolaev, V.O.; Palm, D.; Lohse, M.J.; Vilardaga, J.-P. Turn-on switch in parathyroid hormone receptor by a two-step parathyroid hormone binding mechanism. Proc. Natl. Acad. Sci. USA 2005, 102, 16084–16089. [Google Scholar] [CrossRef] [Green Version]
- Audet, N.; Gales, C.; Archer-Lahlou, E.; Vallieres, M.; Schiller, P.W.; Bouvier, M.; Pineyro, G. Bioluminescence Resonance Energy Transfer Assays Reveal Ligand-specific Conformational Changes within Preformed Signaling Complexes Containing -Opioid Receptors and Heterotrimeric G Proteins. J. Biol. Chem. 2008, 283, 15078–15088. [Google Scholar] [CrossRef] [Green Version]
- Picard, L.-P.; Schönegge, A.M.; Lohse, M.J.; Bouvier, M. Bioluminescence resonance energy transfer-based biosensors allow monitoring of ligand- and transducer-mediated GPCR conformational changes. Commun. Biol. 2018, 1, 1–7. [Google Scholar] [CrossRef]
- Galés, C.; Van Durm, J.J.J.; Schaak, S.; Pontier, S.; Percherancier, Y.; Audet, M.; Paris, H.; Bouvier, M. Probing the activation-promoted structural rearrangements in preassembled receptor–G protein complexes. Nat. Struct. Mol. Biol. 2006, 13, 778–786. [Google Scholar] [CrossRef]
- Quast, R.B.; Margeat, E. Studying GPCR conformational dynamics by single molecule fluorescence. Mol. Cell. Endocrinol. 2019, 493, 110469. [Google Scholar] [CrossRef] [PubMed]
- Calebiro, D.; Sungkaworn, T. Single-Molecule Imaging of GPCR Interactions. Trends Pharmacol. Sci. 2018, 39, 109–122. [Google Scholar] [CrossRef] [PubMed]
- Mazal, H.; Haran, G. Single-molecule FRET methods to study the dynamics of proteins at work. Curr. Opin. Biomed. Eng. 2019, 12, 8–17. [Google Scholar] [CrossRef] [PubMed]
- Hellenkamp, B.; Schmid, S.; Doroshenko, O.; Opanasyuk, O.; Kühnemuth, R.; Rezaei Adariani, S.; Ambrose, B.; Aznauryan, M.; Barth, A.; Birkedal, V.; et al. Precision and accuracy of single-molecule FRET measurements—A multi-laboratory benchmark study. Nat. Methods 2018, 15, 669–676. [Google Scholar] [CrossRef]
- Vafabakhsh, R.; Levitz, J.; Isacoff, E.Y. Conformational dynamics of a class C G protein-coupled receptor. Nature 2015, 524, 497–501. [Google Scholar] [CrossRef]
- Olofsson, L.; Felekyan, S.; Doumazane, E.; Scholler, P.; Fabre, L.; Zwier, J.M.; Rondard, P.; Seidel, C.A.M.; Pin, J.-P.; Margeat, E. Fine tuning of sub-millisecond conformational dynamics controls metabotropic glutamate receptors agonist efficacy. Nat. Commun. 2014, 5, 5206. [Google Scholar] [CrossRef]
- Gutzeit, V.A.; Thibado, J.; Stor, D.S.; Zhou, Z.; Blanchard, S.C.; Andersen, O.S.; Levitz, J. Conformational dynamics between transmembrane domains and allosteric modulation of a metabotropic glutamate receptor. eLife 2019, 8. [Google Scholar] [CrossRef]
- Habrian, C.H.; Levitz, J.; Vyklicky, V.; Fu, Z.; Hoagland, A.; McCort-Tranchepain, I.; Acher, F.; Isacoff, E.Y. Conformational pathway provides unique sensitivity to a synaptic mGluR. Nat. Commun. 2019, 10. [Google Scholar] [CrossRef] [Green Version]
- Peleg, G.; Ghanouni, P.; Kobilka, B.K.; Zare, R.N. Single-molecule spectroscopy of the β2 adrenergic receptor: Observation of conformational substates in a membrane protein. Proc. Natl. Acad. Sci. USA 2001, 98, 8469–8474. [Google Scholar] [CrossRef] [Green Version]
- Bockenhauer, S.; Fürstenberg, A.; Yao, X.J.; Kobilka, B.K.; Moerner, W.E. Conformational dynamics of single G protein-coupled receptors in solution. J. Phys. Chem. B 2011, 115, 13328–13338. [Google Scholar] [CrossRef] [Green Version]
- Lamichhane, R.; Liu, J.J.; Pljevaljcic, G.; White, K.L.; van der Schans, E.; Katritch, V.; Stevens, R.C.; Wüthrich, K.; Millar, D.P. Single-molecule view of basal activity and activation mechanisms of the G protein-coupled receptor β2AR. Proc. Natl. Acad. Sci. USA 2015, 112, 14254–14259. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lamichhane, R.; Liu, J.J.; White, K.L.; Katritch, V.; Stevens, R.C.; Wüthrich, K.; Millar, D.P. Biased Signaling of the G-Protein-Coupled Receptor β2AR Is Governed by Conformational Exchange Kinetics. Structure 2020, 28, 371–377.e3. [Google Scholar] [CrossRef] [PubMed]
- Barth, A. Infrared spectroscopy of proteins. Biochim. Biophys. Acta (BBA)-Bioenerg. 2007, 1767, 1073–1101. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fahmy, K.; Jäger, F.; Beck, M.; Zvyaga, T.A.; Sakmar, T.P.; Siebert, F. Protonation states of membrane-embedded carboxylic acid groups in rhodopsin and metarhodopsin II: A Fourier-transform infrared spectroscopy study of site-directed mutants. Proc. Natl. Acad. Sci. USA 1993, 90, 10206–10210. [Google Scholar] [CrossRef] [Green Version]
- Lüdeke, S.; Beck, M.; Yan, E.C.Y.; Sakmar, T.P.; Siebert, F.; Vogel, R. The role of Glu181 in the photoactivation of rhodopsin. J. Mol. Biol. 2005, 353, 345–356. [Google Scholar] [CrossRef]
- Vogel, R.; Siebert, F.; Lüdeke, S.; Hirshfeld, A.; Sheves, M. Agonists and partial agonists of rhodopsin: Retinals with ring modifications. Biochemistry 2005, 44, 11684–11699. [Google Scholar] [CrossRef]
- Mahalingam, M.; Martínez-Mayorga, K.; Brown, M.F.; Vogel, R. Two protonation switches control rhodopsin activation in membranes. Proc. Natl. Acad. Sci. USA 2008, 105, 17795–17800. [Google Scholar] [CrossRef] [Green Version]
- Madathil, S.; Fahmy, K. Lipid Protein Interactions Couple Protonation to Conformation in a Conserved Cytosolic Domain of G Protein-coupled Receptors. J. Biol. Chem. 2009, 284, 28801–28809. [Google Scholar] [CrossRef] [Green Version]
- Ye, S.; Zaitseva, E.; Caltabiano, G.; Schertler, G.F.X.; Sakmar, T.P.; Deupi, X.; Vogel, R. Tracking G-protein-coupled receptor activation using genetically encoded infrared probes. Nature 2010, 464, 1386–1389. [Google Scholar] [CrossRef]
- Zaitseva, E.; Brown, M.F.; Vogel, R. Sequential Rearrangement of Interhelical Networks upon Rhodopsin Activation in Membranes: The Meta IIa Conformational Substate. J. Am. Chem. Soc. 2010, 132, 4815–4821. [Google Scholar] [CrossRef] [Green Version]
- Elgeti, M.; Kazmin, R.; Rose, A.S.; Szczepek, M.; Hildebrand, P.W.; Bartl, F.J.; Scheerer, P.; Hofmann, K.P. The arrestin-1 finger loop interacts with two distinct conformations of active rhodopsin. J. Biol. Chem. 2018, 293, 4403–4410. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pope, A.L.; Sanchez-Reyes, O.B.; South, K.; Zaitseva, E.; Ziliox, M.; Vogel, R.; Reeves, P.J.; Smith, S.O. A Conserved Proline Hinge Mediates Helix Dynamics and Activation of Rhodopsin. Structure 2020, 28, 1004–1013.e4. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Chance, M.R. Protein Footprinting Comes of Age: Mass Spectrometry for Biophysical Structure Assessment. Mol. Cell. Proteom. 2017, 16, 706–716. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, X.; Chien, E.Y.T.; Chalmers, M.J.; Pascal, B.D.; Gatchalian, J.; Stevens, R.C.; Griffin, P.R. Dynamics of the beta2-adrenergic G-protein coupled receptor revealed by hydrogen-deuterium exchange. Anal. Chem. 2010, 82, 1100–1108. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Orban, T.; Jastrzebska, B.; Gupta, S.; Wang, B.; Miyagi, M.; Chance, M.R.; Palczewski, K. Conformational dynamics of activation for the pentameric complex of dimeric G protein-coupled receptor and heterotrimeric G protein. Structure 2012, 20, 826–840. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- West, G.M.; Chien, E.Y.T.; Katritch, V.; Gatchalian, J.; Chalmers, M.J.; Stevens, R.C.; Griffin, P.R. Ligand-dependent perturbation of the conformational ensemble for the GPCR β2 adrenergic receptor revealed by HDX. Structure 2011, 19, 1424–1432. [Google Scholar] [CrossRef] [Green Version]
- Kahsai, A.W.; Xiao, K.; Rajagopal, S.; Ahn, S.; Shukla, A.K.; Sun, J.; Oas, T.G.; Lefkowitz, R.J. Multiple ligand-specific conformations of the β2-adrenergic receptor. Nat. Chem. Biol. 2011, 7, 692–700. [Google Scholar] [CrossRef] [Green Version]
- Dror, R.O.; Pan, A.C.; Arlow, D.H.; Borhani, D.W.; Maragakis, P.; Shan, Y.; Xu, H.; Shaw, D.E. Pathway and mechanism of drug binding to G-protein-coupled receptors. Proc. Natl. Acad. Sci. USA 2011, 108, 13118–13123. [Google Scholar] [CrossRef] [Green Version]
- Saleh, N.; Kleinau, G.; Heyder, N.; Clark, T.; Hildebrand, P.W.; Scheerer, P. Binding, Thermodynamics, and Selectivity of a Non-peptide Antagonist to the Melanocortin-4 Receptor. Front. Pharmacol. 2018, 9, 560. [Google Scholar] [CrossRef] [Green Version]
- Milanos, L.; Saleh, N.; Kling, R.C.; Kaindl, J.; Tschammer, N.; Clark, T. Identification of Two Distinct Sites for Antagonist and Biased Agonist Binding to the Human Chemokine Receptor CXCR3. Angew. Chem. 2016, 55, 15277–15281. [Google Scholar] [CrossRef]
- Bock, A.; Merten, N.; Schrage, R.; Dallanoce, C.; Bätz, J.; Klöckner, J.; Schmitz, J.; Matera, C.; Simon, K.; Kebig, A.; et al. The allosteric vestibule of a seven transmembrane helical receptor controls G-protein coupling. Nat. Commun. 2012, 3, 1044. [Google Scholar] [CrossRef] [PubMed]
- Saleh, N.; Saladino, G.; Gervasio, F.L.; Haensele, E.; Banting, L.; Whitley, D.C.; Sopkova-de Oliveira Santos, J.; Bureau, R.; Clark, T. A Three-Site Mechanism for Agonist/Antagonist Selective Binding to Vasopressin Receptors. Angew. Chem. 2016, 55, 8008–8012. [Google Scholar] [CrossRef] [PubMed]
- Saleh, N.; Hucke, O.; Kramer, G.; Schmidt, E.; Montel, F.; Lipinski, R.; Ferger, B.; Clark, T.; Hildebrand, P.W.; Tautermann, C.S. Multiple Binding Sites Contribute to the Mechanism of Mixed Agonistic and Positive Allosteric Modulators of the Cannabinoid CB1 Receptor. Angew. Chem. 2018, 57, 2580–2585. [Google Scholar] [CrossRef]
- Saleh, N.; Ibrahim, P.; Clark, T. Differences between G-Protein-Stabilized Agonist-GPCR Complexes and their Nanobody-Stabilized Equivalents. Angew. Chem. 2017, 56, 9008–9012. [Google Scholar] [CrossRef] [PubMed]
- Saleh, N.; Saladino, G.; Gervasio, F.L.; Clark, T. Investigating allosteric effects on the functional dynamics of β2-adrenergic ternary complexes with enhanced-sampling simulations. Chem. Sci. 2017, 8, 4019–4026. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lally, C.C.M.; Bauer, B.; Selent, J.; Sommer, M.E. C-edge loops of arrestin function as a membrane anchor. Nat. Commun. 2017, 8, 14258. [Google Scholar] [CrossRef] [PubMed]
- Shukla, A.K.; Westfield, G.H.; Xiao, K.; Reis, R.I.; Huang, L.-Y.; Tripathi-Shukla, P.; Qian, J.; Li, S.; Blanc, A.; Oleskie, A.N.; et al. Visualization of arrestin recruitment by a G-protein-coupled receptor. Nature 2014, 512, 218–222. [Google Scholar] [CrossRef] [PubMed]
- Goncalves, J.A.; South, K.; Ahuja, S.; Zaitseva, E.; Opefi, C.A.; Eilers, M.; Vogel, R.; Reeves, P.J.; Smith, S.O. Highly conserved tyrosine stabilizes the active state of rhodopsin. Proc. Natl. Acad. Sci. USA 2010, 107, 19861–19866. [Google Scholar] [CrossRef] [Green Version]
- Ahuja, S.; Hornak, V.; Yan, E.C.Y.; Syrett, N.; Goncalves, J.A.; Hirshfeld, A.; Ziliox, M.; Sakmar, T.P.; Sheves, M.; Reeves, P.J.; et al. Helix movement is coupled to displacement of the second extracellular loop in rhodopsin activation. Nat. Struct. Mol. Biol. 2009, 16, 168–175. [Google Scholar] [CrossRef] [Green Version]
- Bokoch, M.P.; Zou, Y.; Rasmussen, S.G.F.; Liu, C.W.; Nygaard, R.; Rosenbaum, D.M.; Fung, J.J.; Choi, H.-J.; Thian, F.S.; Kobilka, T.S.; et al. Ligand-specific regulation of the extracellular surface of a G-protein-coupled receptor. Nature 2010, 463, 108–112. [Google Scholar] [CrossRef] [Green Version]
- Getmanova, E.; Patel, A.B.; Klein-Seetharaman, J.; Loewen, M.C.; Reeves, P.J.; Friedman, N.; Sheves, M.; Smith, S.O.; Khorana, H.G. NMR spectroscopy of phosphorylated wild-type rhodopsin: Mobility of the phosphorylated C-terminus of rhodopsin in the dark and upon light activation. Biochemistry 2004, 43, 1126–1133. [Google Scholar] [CrossRef]
- Langen, R.; Cai, K.; Altenbach, C.; Khorana, H.G.; Hubbell, W.L. Structural features of the C-terminal domain of bovine rhodopsin: A site-directed spin-labeling study. Biochemistry 1999, 38, 7918–7924. [Google Scholar] [CrossRef] [PubMed]
- Altenbach, C.; Klein-Seetharaman, J.; Hwa, J.; Khorana, H.G.; Hubbell, W.L. Structural features and light-dependent changes in the sequence 59–75 connecting helices I and II in rhodopsin: A site-directed spin-labeling study. Biochemistry 1999, 38, 7945–7949. [Google Scholar] [CrossRef] [PubMed]
- Altenbach, C.; Cai, K.; Khorana, H.G.; Hubbell, W.L. Structural features and light-dependent changes in the sequence 306–322 extending from helix VII to the palmitoylation sites in rhodopsin: A site-directed spin-labeling study. Biochemistry 1999, 38, 7931–7937. [Google Scholar] [CrossRef] [PubMed]
- Farrens, D.L.; Altenbach, C.; Yang, K.; Hubbell, W.L.; Khorana, H.G. Requirement of rigid-body motion of transmembrane helices for light activation of rhodopsin. Science 1996, 274, 768–770. [Google Scholar] [CrossRef]
- Altenbach, C.; Cai, K.; Klein-Seetharaman, J.; Khorana, H.G.; Hubbell, W.L. Structure and function in rhodopsin: Mapping light-dependent changes in distance between residue 65 in helix TM1 and residues in the sequence 306-319 at the cytoplasmic end of helix TM7 and in helix H8. Biochemistry 2001, 40, 15483–15492. [Google Scholar] [CrossRef] [PubMed]
- Altenbach, C.; Klein-Seetharaman, J.; Cai, K.; Khorana, H.G.; Hubbell, W.L. Structure and function in rhodopsin: Mapping light-dependent changes in distance between residue 316 in helix 8 and residues in the sequence 60–75, covering the cytoplasmic end of helices TM1 and TM2 and their connection loop CL1. Biochemistry 2001, 40, 15493–15500. [Google Scholar] [CrossRef]
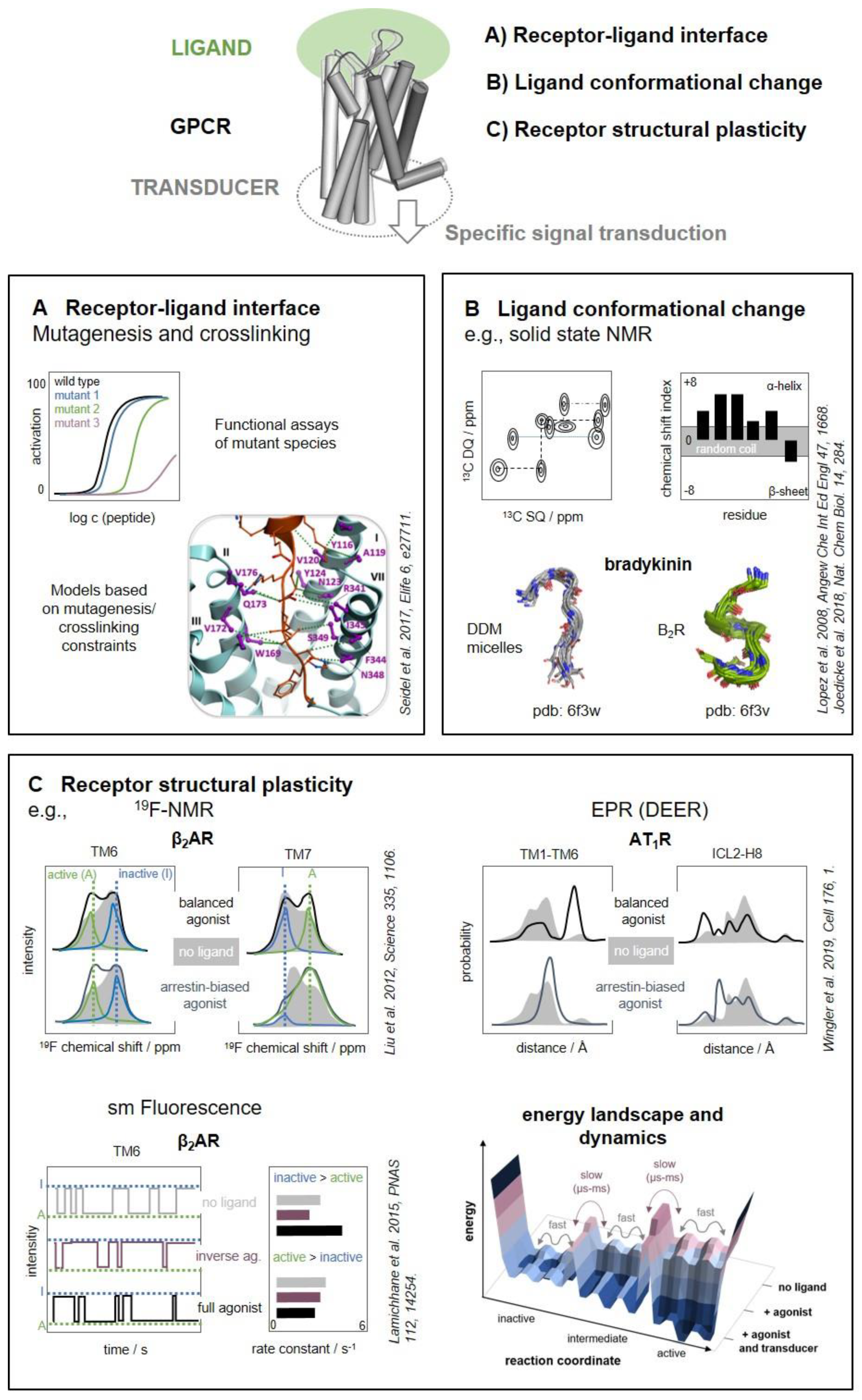


| Method | Label | Introduced by | Provides Information About | Comments | Ref. |
|---|---|---|---|---|---|
| NMR (solution and solid-state) | 13Cε-methionine or other specifically labeled aa: 13C/15N-tryptophane 13Cδ1 isoleucine 15N-valine 13Cζ-tyrosine; 13Cβ-cysteine etc. | residue-specific during biosynthesis (expression medium) | chemical environment/distinct conformational states, exchange rates between different states can be extracted short-range interactions (via cross-polarization in solid-state NMR) | 13Cε-methionine exploits favorable relaxation profile and flexibility of methyl-groups; deuterated background possible; 15N-valine: backbone assignment | [205,206,207,208,209,210] [217,233,234] [175] [236] [319,320] |
| 13C-(di)methyl-lysine | residue-specific; chemically: reductive methylation of primary amines with 13CH2O | chemical environment/distinct conformational states short-range interactions (participation in salt bridges) | exploits favorable relaxation profile and flexibility of methyl-groups; negative charge of side chain preserved | [209,321] | |
| 19F | site-specific (1 label) chemically: (single) reactive cysteine plus TET or BTFA | chemical environment/distinct conformational states exchange rates between different states can be extracted short-range interactions (19F-NOE; 19F-31P REDOR) | direct excitation and no background allow straightforward extraction of entropic and enthalpic parameters <8 Å; up to 12 Å; e.g., phosphorylation sites | [174,214,218,223,224,225,226,227,228,229,230] [226] [322] | |
| EPR continuous wave pulsed (DEER) | N-O· (nitroxide radical, typically stabilized by proximal dimethyl groups in a pyrrole ring) | site-specific (1 or 2 labels) chemically: reactive cysteines plus MTSL etc. Uaa labeling possible [201,202,203] | chemical environment, dynamics, accessibility (continuous wave) long-range distances (continuous wave, pulsed (DEER)) | nitroxide scanning (SDSL) allows mapping of secondary structure and membrane boundaries, as well as structural transitions spectral broadening in continuous wave EPR 8–25 Å, DEER 20–70 Å | [191,243,244,245,323,324,325] [326,327,328] [173,214,249] |
| Fluorescence Spectroscopy FRET/LRET single molecule microscopy/spectroscopy | fluorophores | site-specific (1 or 2 labels) chemically: (single) reactive cysteines bioorthogonal ligation possible, e.g., “click”-chemistry at genetically engineered Azi; FlAsH-tags etc. [198,273] | shift of environment polarity lifetime of (sub)states (fluorescence anisotropy and lifetime) distances (quenching, FRET, LRET) rate of structural transitions (slower than ms) visualization of transient states not significantly populated in equilibrium | nM protein concentration sufficient, applications in living cells possible distance ranges: tryptophane-induced quenching 5–15 Å 20–80 Å with high orientational dependence for FRET, which is overcome by LRET sm fluorescence with environmentally sensitive fluorophores or smFRET | [192,260,261,262] [193,265,266,267] [260,264] [268] [270] [290,291,292,293] [220] |
| FTIR | intrinsic C=O bond vibrations N3 (or CN etc.) bond vibrations | label-free by genetic engineering, e.g., p-azido-phenylalanine (Azi) | protonation switches, secondary structure chemical environment (electrostatics)/distinct conformational states | application of difference spectra to extract responsive C=O signals kinetic resolution up to 1 μs bond vibrations in a region free of endogenous signals | [35,295,296,297,298,299] [204,300] |
| HDX HRF | (label-free) | in situ exchange proton-deuterium (reversible) in situ reaction hydroxyl-radical (irreversible) | accessibility, secondary structure, temporal resolution up to seconds accessibility, temporal resolution up to ms | Label free, global view on structural alterations Accuracy depends on coverage and resolution of mass spectrometry | [41,305,306,307,308] [41] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kaiser, A.; Coin, I. Capturing Peptide–GPCR Interactions and Their Dynamics. Molecules 2020, 25, 4724. https://doi.org/10.3390/molecules25204724
Kaiser A, Coin I. Capturing Peptide–GPCR Interactions and Their Dynamics. Molecules. 2020; 25(20):4724. https://doi.org/10.3390/molecules25204724
Chicago/Turabian StyleKaiser, Anette, and Irene Coin. 2020. "Capturing Peptide–GPCR Interactions and Their Dynamics" Molecules 25, no. 20: 4724. https://doi.org/10.3390/molecules25204724





