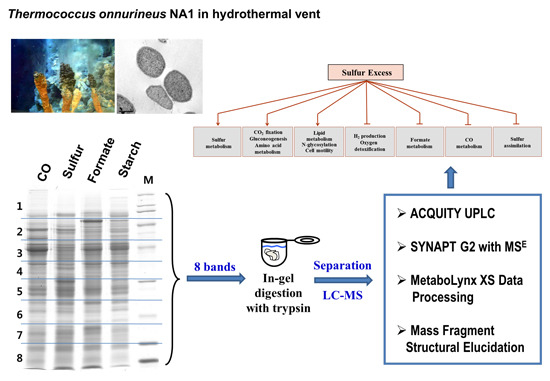
Proteomic Insights into Sulfur Metabolism in the Hydrogen-Producing Hyperthermophilic Archaeon Thermococcus onnurineus NA1
Abstract
:1. Introduction
2. Results and Discussion
2.1. The Growth of T. onnurineus NA1 under Sulfur Culture Conditions
2.2. Identification of Proteins Expressed under Sulfur Culture Conditions
| Substrate Ratio (A/B) | Sulfur/CO | Sulfur/Formate | Sulfur/Starch |
|---|---|---|---|
| p Value < 0.05 a | 330 | 336 | 359 |
| A/B ≥ 1.5 | 169 | 159 | 143 |
| A/B ≤ 0.67 | 130 | 141 | 172 |
| 1.5 > A/B > 0.67 | 31 | 36 | 44 |
| p Value > 0.05 | 114 | 109 | 112 |
| Subtotal | 444 | 445 | 471 |
| Total | – | 589 | – |
2.3. General Trends in the Proteome of T. onnurineus NA1 Grown with S°

| Protein Name | Gene Identification | Ratio a | p Value b | ||||
|---|---|---|---|---|---|---|---|
| Sulfur/CO | Sulfur/Formate | Sulfur/Starch | Sulfur/CO | Sulfur/Formate | Sulfur/Starch | ||
| gltB-1 Glutamate synthase small chain (SuDH I, SudA) | TON_0057 | Y | Y | Y | Y | Y | Y |
| NADH:polysulfide oxidoreductase (NPSOR) | TON_0129 | 1.36 | 1.17 | 1.70 | 0.89 | 0.75 | 0.99 |
| NADH oxidase (NSR) | TON_0305 | Y | 3.00 | 2.39 | Y | 1.00 | 1.00 |
| Protein disulfide oxidoreductase (PDO) | TON_0319 | 2.53 | 3.06 | 1.40 | 1.00 | 1.00 | 1.00 |
| ATPase C-terminus (SipB) | TON_0916 | Y | Y | Y | Y | Y | Y |
| Iron-molybdenum cofactor-binding protein (SipA) | TON_0919 | Y | Y | Y | Y | Y | Y |
| Putative oxidoreductase (SuDH II, SudX) | TON_1336 | Y | Y | Y | Y | Y | Y |
| Ferredoxin-NADP+ reductase subunit α (SuDH II, SudY) | TON_1337 | Y | Y | Y | Y | Y | Y |
| NADH dehydrogenase subunit D | TON_0487 | 2.48 | Y | 1.75 | 1.00 | Y | 0.97 |
| V-type ATP synthase subunit E | TON_1749 | Y | 1.30 | 1.35 | Y | 0.65 | 0.77 |
| V-type ATP synthase subunit C | TON_1750 | Y | Y | 1.36 | Y | Y | 0.69 |
| V-type ATP synthase subunit F | TON_1751 | Y | Y | Y | Y | Y | Y |
| V-type ATP synthase subunit A | TON_1752 | 2.12 | 1.57 | 1.79 | 1.00 | 1.00 | 1.00 |
| V-type ATP synthase subunit B | TON_1753 | 2.66 | 1.58 | 1.88 | 1.00 | 1.00 | 1.00 |
| V-type ATP synthase subunit D | TON_1754 | Y | Y | Y | Y | Y | Y |
| ABC-type transport system involved in Fe–S cluster assembly, ATPase component (SufC) | TON_0530 | Y | 0.92 | 0.85 | Y | 0.40 | 0.41 |
| ABC-type transport system involved in Fe–S cluster assembly (SufBD) | TON_0531 | 0.74 | 0.84 | Y | 0.27 | 0.35 | Y |
| Hypothetical protein TON_0849 (SufBD-domain containing protein) | TON_0849 | 4.26 | 2.29 | 0.37 | 1.00 | 1.00 | 0.00 |
| Hypothetical protein TON_0850 (SufBD-domain containing protein) | TON_0850 | Y | Y | 0.38 | Y | Y | 0.00 |
| ATPase (MrP/Nbp35 family ATP-binding protein) | TON_1483 | 4.90 | 4.06 | 1.14 | 1.00 | 1.00 | 0.96 |
| Phosphoenolpyruvate carboxykinase (PCK) | TON_0192 | 0.90 | 1.13 | 2.16 | 0.26 | 0.80 | 1.00 |
| 2-Oxoglutarate ferredoxin oxidoreductase subunit alpha (KGOR_α) | TON_0584 | 0.97 | Y | Y | 0.41 | Y | Y |
| 2-Oxoglutarate ferredoxin oxidoreductase subunit beta (KGOR_β) | TON_0586 | Y | Y | 1.58 | Y | Y | 0.69 |
| Ribulose bisphosphate carboxylase (RuBisCO) type III | TON_1234 | 1.12 | 1.28 | 1.32 | 0.96 | 1.00 | 1.00 |
| Archaeal succinyl-CoA synthetase forming), large subunit (SCS) | TON_1665 | 2.51 | 2.10 | 1.84 | 1.00 | 1.00 | 1.00 |
| Phosphoenolpyruvate synthase (PPS) | TON_0311 | 1.77 | 1.68 | 0.96 | 1.00 | 1.00 | 0.11 |
| Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) | TON_0639 | 1.30 | 0.52 | Y | 0.89 | 0.00 | Y |
| Thermophile-specific fructose-1,6-bisphosphatase (FBPase) | TON_1497 | 2.77 | 1.60 | Y | 1.00 | 1.00 | Y |
| Mevalonate kinase (MVK) | TON_0133 | Y | Y | Y | Y | Y | Y |
| Geranylgeranyl hydrogenase | TON_0316 | 1.51 | 1.92 | 0.81 | 1.00 | 1.00 | 0.03 |
| UDP-N-acetylglucosamine 2-epimerase (WecB) | TON_0501 | Y | Y | 1.17 | Y | Y | 0.58 |
| UDP-N-acetyl-d-mannosaminuronate dehydrogenase (WecC) | TON_0502 | 1.90 | 1.93 | 1.45 | 0.98 | 1.00 | 1.00 |
| Archaeal flagella-related protein C (FlaC) | TON_1184 | Y | Y | Y | Y | Y | Y |
| Bifunctional carboxypeptidase/aminoacylase | TON_0348 | 4.26 | 2.20 | Y | 1.00 | 0.93 | Y |
| Methionine aminopeptidase | TON_0362 | Y | Y | Y | Y | Y | Y |
| Deblocking aminopeptidase (DAP) | TON_0369 | 1.55 | 1.52 | 1.65 | 1.00 | 1.00 | 1.00 |
| PepQ-1 X-pro dipeptidase | TON_0481 | 4.39 | 3.56 | 0.76 | 1.00 | 1.00 | 0.00 |
| ATP-dependent protease Lon | TON_0529 | Y | Y | Y | Y | Y | Y |
| Hypothetical endoglucanase | TON_0570 | 1.38 | 1.17 | 1.15 | 0.93 | 0.72 | 0.72 |
| Prolyl endopeptidase | TON_0611 | Y | Y | 1.60 | Y | Y | 0.88 |
| Xaa-Pro aminopeptidase | TON_0651 | 2.86 | 2.41 | 1.68 | 0.95 | 0.85 | 0.81 |
| Zinc-dependent protease | TON_0804 | Y | Y | 0.58 | Y | Y | 0.23 |
| Zinc-dependent protease | TON_0805 | Y | Y | 1.30 | Y | Y | 0.57 |
| Acylamino acid-releasing enzyme (acylaminoacyl-peptidase) | TON_0969 | Y | Y | Y | Y | Y | Y |
| Deblocking aminopeptidase (DAP) | TON_1032 | 2.83 | 2.23 | 1.75 | 1.00 | 1.00 | 1.00 |
| d-aminopeptidase | TON_1067 | Y | Y | Y | Y | Y | Y |
| Intracellular protease I (Pfp1) | TON_1285 | Y | Y | 1.55 | Y | Y | 0.99 |
| Proteasome-activating nucleotidase (PAN) | TON_1385 | Y | Y | Y | Y | Y | Y |
| Acylamino acid-releasing enzyme (acylaminoacyl-peptidase) | TON_1543 | 5.87 | Y | Y | 1.00 | Y | Y |
| Cobalt-activating carboxypeptidase | TON_1687 | 26.05 | 8.33 | 3.22 | 1.00 | 1.00 | 1.00 |
| d-aminopeptidase | TON_1960 | Y | Y | Y | Y | Y | Y |
| Signal recognition particle protein Srp54 | TON_0123 | Y | Y | Y | Y | Y | Y |
| Signal recognition particle GTPase | TON_0592 | Y | Y | 1.42 | Y | Y | 0.77 |
| Preprotein translocase subunit SecD | TON_1744 | Y | Y | Y | Y | Y | Y |
| Prefoldin subunit α | TON_0914 | Y | Y | Y | Y | Y | Y |
| Tryptophan synthase subunit β | TON_0147 | Y | Y | Y | Y | Y | Y |
| Imidazolonepropionase-like amidohydrolase | TON_0704 | 2.77 | 4.22 | 1.97 | 1.00 | 1.00 | 1.00 |
| Hypothetical aspartate racemase | TON_0801 | Y | Y | Y | Y | Y | Y |
| l-asparaginase | TON_1392 | Y | Y | Y | Y | Y | Y |
| Putative glutamate synthase subunit β | TON_0702 | Y | Y | 1.46 | Y | Y | 0.68 |
| 4-Aminobutyrate aminotransferase | TON_1605 | Y | 4.35 | 14.15 | Y | 1.00 | 1.00 |
| Serine hydroxymethyltransferase | TON_0821 | 2.08 | 2.92 | 2.23 | 1.00 | 1.00 | 1.00 |
| Sarcosine oxidase, α subunit | TON_1282 | Y | Y | Y | Y | Y | Y |
| Glycine cleavage system protein H | TON_1334 | 2.23 | Y | Y | 0.97 | Y | Y |
| Glycine dehydrogenase subunit 1 | TON_0213 | 1.43 | 1.93 | 1.43 | 0.94 | 1.00 | 0.95 |
| Glycine dehydrogenase subunit 2 | TON_0214 | 1.63 | 1.86 | 1.82 | 1.00 | 1.00 | 1.00 |
| l-Threonine 3-dehydrogenase | TON_0397 | 1.20 | 1.34 | 1.30 | 1.00 | 1.00 | 1.00 |
| d-isomer specific 2-hydroxyacid dehydrogenase | TON_0569 | 1.77 | 1.72 | 1.34 | 0.99 | 1.00 | 0.92 |
| S-adenosyl-l-homocysteine hydrolase | TON_1212 | Y | 1.40 | Y | Y | 0.80 | Y |
| Diaminopimelate aminotransferase | TON_1785 | 2.2 | 1.32 | Y | 0.97 | 0.87 | Y |
| 50S ribosomal protein L4P | TON_0067 | 1.23 | 1.72 | 1.28 | 1.00 | 1.00 | 1.00 |
| 50S ribosomal protein L19e | TON_0084 | 1.12 | Y | Y | 0.60 | Y | Y |
| 50S ribosomal protein L18P | TON_0085 | 1.28 | 1.21 | 1.31 | 0.99 | 0.95 | 1.00 |
| 50S ribosomal protein L30P | TON_0087 | 1.46 | 1.62 | 1.51 | 1.00 | 1.00 | 0.98 |
| 50S ribosomal protein L15P | TON_0088 | 1.32 | 2.16 | 1.68 | 0.90 | 1.00 | 0.99 |
| 50S ribosomal protein L14e | TON_0094 | 1.43 | 1.92 | 1.75 | 1.00 | 0.99 | 0.99 |
| Acidic ribosomal protein P0 | TON_0181 | 1.15 | 1.55 | 1.42 | 0.98 | 1.00 | 1.00 |
| 50S ribosomal protein L21e | TON_0406 | 1.39 | Y | 2.23 | 0.85 | Y | 0.97 |
| 30S ribosomal protein S19P | TON_0070 | Y | Y | Y | Y | Y | Y |
| 30S ribosomal protein S4e | TON_0078 | 1.09 | 1.46 | 1.23 | 0.84 | 1.00 | 0.97 |
| 30S ribosomal protein S5P | TON_0086 | 1.11 | 1.34 | 1.26 | 0.82 | 0.98 | 0.99 |
| 30S ribosomal protein S13P | TON_0102 | 1.23 | 1.32 | 1.45 | 0.94 | 0.80 | 0.98 |
| 30S ribosomal protein S4 | TON_0103 | 1.21 | 1.60 | 1.21 | 0.90 | 1.00 | 0.92 |
| 30S ribosomal protein S12P | TON_0222 | 1.02 | 1.67 | 1.27 | 0.52 | 0.98 | 0.96 |
| Asparagine synthetase A | TON_0058 | Y | Y | Y | Y | Y | Y |
| Leucyl-tRNA synthetase | TON_0141 | Y | Y | 1.26 | Y | Y | 0.78 |
| Alanyl-tRNA synthetase-related protein | TON_0899 | Y | Y | Y | Y | Y | Y |
| Isoleucyl-tRNA synthetase | TON_1803 | 1.82 | 1.51 | 2.80 | 1.00 | 0.99 | 1.00 |
| Elongation factor 1-β | TON_1866 | Y | Y | Y | Y | Y | Y |
| Translation initiation factor IF-2 subunit γ | TON_1944 | 1.31 | 1.09 | 1.31 | 0.99 | 0.79 | 1.00 |
| Large helicase-related protein | TON_0613 | Y | Y | Y | Y | Y | Y |
| Hypothetical phosphate transport system regulator PhoU | TON_1464 | 1.34 | 1.20 | 1.46 | 0.97 | 0.88 | 0.99 |
| ABC transporter tungsten-binding protein (Tungsten) | TON_0014 | Y | 1.51 | 1.22 | Y | 0.88 | 0.79 |
| Small-conductance mechanosensitive channel | TON_0799 | Y | Y | Y | Y | Y | Y |
| ABC-type dipeptide/oligopeptide transport system (AppABC/OppBCDF) | TON_1764 | 8.50 | 5.21 | 3.32 | 1.00 | 1.00 | 1.00 |
| ABC-type dipeptide/oligopeptide transport system, ATPase component | TON_1767 | Y | Y | 2.29 | Y | Y | 0.92 |
| ABC-type dipeptide/oligopeptide transport system, ATPase component (AppABC/OppBCDF) | TON_1768 | 4.06 | 2.64 | 3.71 | 0.99 | 0.99 | 1.00 |
| ABC transporter related ATPase component | TON_1874 | Y | Y | Y | Y | Y | Y |
| Hydrogenase 4, component G or formate hydrogen lyase, subunit 5 (Mhy I) | TON_0276 | 1.73 | Y | 2.94 | 0.94 | Y | 0.94 |
| Oxidoreductase iron-sulfur protein (Fdh1B) | TON_0280 | Y | Y | Y | Y | Y | Y |
| fdhA formate dehydrogenase, α subunit (FdhA) | TON_0281 | Y | Y | Y | Y | Y | Y |
| Protein Name | Gene Identification | Ratio a | p Value b | ||||
|---|---|---|---|---|---|---|---|
| Sulfur/CO | Sulfur/Formate | Sulfur/Starch | Sulfur/CO | Sulfur/Formate | Sulfur/Starch | ||
| Hypothetical transcription regulator (SurR) | TON_0318 | - | - | S | - | - | S |
| Membrane bound hydrogenase, NiFe-hydrogenase large subunit 2 | TON_1593 | C | - | S | C | - | S |
| Cytochrome-c3 hydrogenase subunit gamma (SH II, Sulf-II) | TON_0054 | - | - | S | - | - | S |
| Cytosolic NiFe-hydrogenase, α subunit (SH I, Sulf-I) | TON_0534 | 0.32 | - | 0.57 | 0.00 | - | 0.11 |
| Cytosolic NiFe-hydrogenase, δ subunit (SH I, Sulf-I) | TON_0535 | C | - | - | C | - | - |
| Cytochrome-c3 hydrogenase subunit gamma (SH I, Sulf-I) | TON_0536 | C | - | - | C | - | - |
| Sulfhydrogenase beta subunit (SH Iβ) | TON_0537 | C | - | - | C | - | - |
| ATPase involved in chromosome partitioning | TON_0262 | 0.78 | 0.53 | 0.47 | 0.07 | 0.00 | 0.00 |
| Hydrogenase maturation protease HycI | TON_0263 | - | F | - | - | F | - |
| Formate hydrogen lyase, subunit 7 (Mhy I) | TON_0274 | - | - | S | - | - | S |
| Hydrogenase maturation protein HypF | TON_0286 | 0.26 | 0.41 | 0.18 | 0.01 | 0.06 | 0.00 |
| Hydrogenase expression/formation protein HypE | TON_0287 | 0.69 | 0.86 | 0.45 | 0.16 | 0.41 | 0.00 |
| Coenzyme F420 hydrogenase alpha subunit (Frh_α) | TON_1559 | - | F | S | - | F | S |
| CoenzymeF420 hydrogenase/dehydrogenase beta subunit (Frh_β) | TON_1561 | - | F | S | - | F | S |
| TonB-dependent receptor protein:Formate dehydrogenase, subunit FdhD | TON_1562 | - | F | - | - | F | - |
| Hypothetical formate dehydrogenase, α subunit (Fdh2) | TON_1563 | C | F | S | C | F | S |
| 4Fe-4S cluster-binding protein | TON_1564 | - | F | - | - | F | - |
| Hydrogenase 4, component G or formate hydrogen lyase, subunit 5 (Mhy II, Mfh2) | TON_1569 | - | F | - | - | F | - |
| Formate hydrogen lyase subunit 6 (Mhy II) | TON_1570 | - | F | - | - | F | - |
| Hydrogenase 4, component I or formate hydrogen lyase, subunit 7 | TON_1571 | - | F | - | - | F | - |
| Hypothetical protein TON_1572 | TON_1572 | - | F | - | - | F | - |
| Hypothetical Multisubunit Na+/H+ antiporter MnhB subunit (MnhB) | TON_1577 | - | F | - | - | F | - |
| NADH dehydrogenase subunit C (MBX) | TON_0488 | C | - | - | C | - | - |
| 4Fe-4S ferredoxin, iron-sulfur binding domain protein (CooF) | TON_1017 | C | - | - | C | - | - |
| Hypothetical ATP-binding protein (CooC) | TON_1019 | C | - | - | C | - | - |
| Hydrogenase 4, subunit 5 (Mch) | TON_1023 | C | - | - | C | - | - |
| NADH dehydrogenase (ubiquinone), 20 kDa subunit (Mch) | TON_1024 | C | - | - | C | - | - |
| Transcription regulator, PadR family | TON_0114 | - | F | - | - | F | - |
| Hypothetical transcription regulator (TrmB) | TON_0332 | 0.76 | 0.81 | 0.54 | 0.00 | 0.00 | 0.00 |
| Hypothetical transcription regulator (Lrp/AsnC family transcriptional regulator) | TON_0662 | - | F | - | - | F | - |
| Transcription regulator (Lrp/AsnC family transcriptional regulator) | TON_1284 | - | F | S | - | F | S |
| Transcription factor (TBP) | TON_1309 | - | F | S | - | F | S |
| Transcription regulator (Phosphate uptake regulator, phoU) | TON_1393 | - | - | S | - | - | S |
| Hypothetical transcription regulator | TON_1436 | - | - | S | - | - | S |
| Transcription regulator (Lrp/AsnC family Transcription regulator) | TON_1510 | 0.39 | 0.29 | 0.54 | 0.00 | 0.00 | 0.02 |
| Transcription regulator, ArsR family | TON_1663 | - | - | S | - | - | S |
| Hypothetical transcription regulator (TrmB-like protein) | TON_1797 | 0.82 | 0.73 | 0.44 | 0.35 | 0.19 | 0.00 |
| Manganese-dependent transcription regulator | TON_1956 | - | - | S | - | - | S |
| Peroxiredoxin | TON_0829 | 0.17 | 0.08 | 0.16 | 0.00 | 0.00 | 0.00 |
| Thioredoxin peroxidase | TON_0862 | C | - | S | C | - | S |
| Type A flavoprotein (FdpA, flavodiiron protein) | TON_0863 | 0.36 | 0.41 | 0.46 | 0.00 | 0.00 | 0.00 |
| NAD(P)H:rubredoxin oxidoreductase (NROR) | TON_0865 | C | - | S | C | - | S |
| Rubrerythrin (Rr) | TON_0866 | 0.34 | 0.48 | 0.70 | 0.00 | 0.00 | 0.00 |
| Sor superoxide reductase (SOR) | TON_0868 | - | 0.32 | - | - | 0.00 | - |
| Ferredoxin-NADP+ reductase subunit α (FNOR) | TON_0056 | - | - | S | - | - | S |
| Cysteine desulfurase | TON_0289 | C | F | S | C | F | S |
| Cysteine synthase (OASS) | TON_1004 | - | - | S | - | - | S |
| Hypothetical protein TON_1360 (SAT, serine acetyltransferase) | TON_1360 | - | - | 0.88 | - | - | 0.44 |
| Molybdenum cofactor biosynthesis protein A (MoaA) | TON_1410 | - | F | S | - | F | S |
| Hypothetical protein TON_1504 (PSP, O-phosphoserine phosphatase) | TON_1504 | 0.66 | 0.41 | 0.53 | 0.18 | 0.03 | 0.09 |
| Hypothetical protein TON_1706 (PAP phosphatase) | TON_1706 | - | F | - | - | F | - |
2.4. Proteins Up-Regulated in Response to Growth on Sulfur
2.4.1. CO2 Fixation
2.4.2. Lipid Biosynthesis
2.4.3. Protein Glycosylation and Motility
2.4.4. Proteolytic Pathways and Protein Translocation
2.4.5. Formate Oxidation to CO2
2.4.6. Reductive Sulfur Metabolism
2.4.7. Fe–S Cluster Biogenesis
2.5. Antagonistic Switch between H2-Metabolism and Reductive Sulfur Metabolism
2.6. Proteins Down-Regulated in Response to Growth on Sulfur
2.6.1. H2-Metabolism

2.6.2. Oxygen Detoxification
2.6.3. Transcription Regulation
2.6.4. Sulfur Assimilation Pathway
3. Experimental Section
3.1. Strain and Culture Conditions
3.2. Protein Preparation and Enzyme Digestion
3.3. Analysis by Nano-UPLC-MSE Tandem Mass Spectrometry and Quantitative Analysis
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Bertoldo, C.; Antranikian, G. The order thermococcales. In The Prokaryotes, 3rd ed.; Dworkin, M., Falkow, S., Rosenberg, E., Schleifer, K.H., Stackebrandt, E., Eds.; Springer: New York, NY, USA, 2006; Volume 3, pp. 69–81. [Google Scholar]
- Liu, Y.; Beer, L.L.; Whitman, W.B. Sulfur metabolism in archaea reveals novel processes. Environ. Microbiol. 2012, 14, 2632–2644. [Google Scholar] [CrossRef] [PubMed]
- Amend, J.P.; Shock, E.L. Energetics of overall metabolic reactions of thermophilic and hyperthermophilic Archaea and bacteria. FEMS Microbiol. Rev. 2001, 25, 175–243. [Google Scholar] [CrossRef] [PubMed]
- Huber, R.; Stetter, K.O. Discovery of hyperthermophilic microorganisms. Methods Enzymol. 2001, 330, 11–24. [Google Scholar] [PubMed]
- Santangelo, T.J.; Cubonova, L.; Reeve, J.N. Deletion of alternative pathways for reductant recycling in Thermococcus kodakarensis increases hydrogen production. Mol. Microbiol. 2011, 81, 897–911. [Google Scholar] [CrossRef] [PubMed]
- Kanai, T.; Matsuoka, R.; Beppu, H.; Nakajima, A.; Okada, Y.; Atomi, H.; Imanaka, T. Distinct physiological roles of the three [NiFe]-hydrogenase orthologs in the hyperthermophilic archaeon Thermococcus kodakarensis. J. Bacteriol. 2011, 193, 3109–3116. [Google Scholar] [CrossRef] [PubMed]
- Chou, C.J.; Shockley, K.R.; Conners, S.B.; Lewis, D.L.; Comfort, D.A.; Adams, M.W.; Kelly, R.M. Impact of substrate glycoside linkage and elemental sulfur on bioenergetics of and hydrogen production by the hyperthermophilic archaeon Pyrococcus furiosus. Appl. Environ. Microbiol. 2007, 73, 6842–6853. [Google Scholar] [CrossRef] [PubMed]
- Schut, G.J.; Nixon, W.J.; Lipscomb, G.L.; Scott, R.A.; Adams, M.W. Mutational Analyses of the enzymes involved in the metabolism of hydrogen by the hyperthermophilic archaeon Pyrococcus furiosus. Front. Microbiol. 2012, 3, 163. [Google Scholar] [CrossRef] [PubMed]
- Schut, G.J.; Boyd, E.S.; Peters, J.W.; Adams, M.W. The modular respiratory complexes involved in hydrogen and sulfur metabolism by heterotrophic hyperthermophilic archaea and their evolutionary implications. FEMS Microbiol. Rev. 2013, 37, 182–203. [Google Scholar] [PubMed]
- Schut, G.J.; Zhou, J.; Adams, M.W. DNA microarray analysis of the hyperthermophilic archaeon Pyrococcus furiosus: Evidence for an new type of sulfur-reducing enzyme complex. J. Bacteriol 2001, 183, 7027–7036. [Google Scholar] [CrossRef] [PubMed]
- Schut, G.J.; Bridger, S.L.; Adams, M.W. Insights into the metabolism of elemental sulfur by the hyperthermophilic archaeon Pyrococcus furiosus: Characterization of a coenzyme A- dependent NAD(P)H sulfur oxidoreductase. J. Bacteriol. 2007, 189, 4431–4441. [Google Scholar] [CrossRef] [PubMed]
- Bridger, S.L.; Clarkson, S.M.; Stirrett, K.; DeBarry, M.B.; Lipscomb, G.L.; Schut, G.J.; Westpheling, J.; Scott, R.A.; Adams, M.W. Deletion strains reveal metabolic roles for key elemental sulfur-responsive proteins in Pyrococcus furiosus. J. Bacteriol. 2011, 193, 6498–6504. [Google Scholar] [CrossRef] [PubMed]
- Bae, S.S.; Kim, Y.J.; Yang, S.H.; Lim, J.K.; Jeon, J.H.; Lee, H.S.; Kang, S.G.; Kim, S.-J.; Lee, J.-H. Thermoccoccus onnurineus sp. nov., a hyperthermophilic archaeon isolated from a deep-sea hydrothermal vent area at the PACMANUS field. J. Microbiol. Biotechnol. 2006, 16, 1826–1831. [Google Scholar]
- Bae, S.S.; Kim, T.W.; Lee, H.S.; Kwon, K.K.; Kim, Y.J.; Kim, M.S.; Lee, J.H.; Kang, S.G. H2 production from CO, formate or starch using the hyperthermophilic archaeon, Thermococcus onnurineus. Biotechnol. Lett. 2012, 34, 75–79. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.S.; Kang, S.G.; Bae, S.S.; Lim, J.K.; Cho, Y.; Kim, Y.J.; Jeon, J.H.; Cha, S.S.; Kwon, K.K.; Kim, H.T.; et al. The complete genome sequence of Thermococcus onnurineus NA1 reveals a mixed heterotrophic and carboxydotrophic metabolism. J. Bacteriol. 2008, 190, 7491–7499. [Google Scholar]
- Kim, Y.J.; Lee, H.S.; Kim, E.S.; Bae, S.S.; Lim, J.K.; Matsumi, R.; Lebedinsky, A.V.; Sokolova, T.G.; Kozhevnikova, D.A.; Cha, S.S.; et al. Formate-driven growth coupled with H2 production. Nature 2010, 467, 352–355. [Google Scholar]
- Lim, J.K.; Kang, S.G.; Lebedinsky, A.V.; Lee, J.H.; Lee, H.S. Identification of a novel class of membrane-bound [NiFe]-hydrogenases in Thermococcus onnurineus NA1 by in silico analysis. Appl. Environ. Microbiol. 2010, 76, 6286–6289. [Google Scholar] [CrossRef] [PubMed]
- Lim, J.K.; Bae, S.S.; Kim, T.W.; Lee, J.H.; Lee, H.S.; Kang, S.G. Thermodynamics of formate-oxidizing metabolism and implications for H2 production. Appl. Environ. Microbiol. 2012, 78, 7393–7397. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.S.; Bae, S.S.; Kim, Y.J.; Kim, T.W.; Lim, J.K.; Lee, S.H.; Choi, A.R.; Jeon, J.H.; Lee, J.H.; Lee, H.S.; et al. CO-dependent H2 production by genetically engineered Thermococcus onnurineus NA1. Appl. Environ. Microbiol. 2013, 79, 2048–2053. [Google Scholar]
- Kawarabayasi, Y.; Sawada, M.; Horikawa, H.; Haikawa, Y.; Hino, Y.; Yamamoto, S.; Sekine, M.; Baba, S.; Kosugi, H.; Hosoyama, A.; et al. Complete sequence and gene organization of the genome of a hyper-thermophilic archaebacterium, Pyrococcus horikoshii OT3. DNA Res. 1998, 5, 55–76. [Google Scholar]
- Robb, F.T.; Maeder, D.L.; Brown, J.R.; DiRuggiero, J.; Stump, M.D.; Yeh, R.K.; Weiss, R.B.; Dunn, D.M. Genomic sequence of hyperthermophile, Pyrococcus furiosus: Implications for physiology and enzymology. Methods Enzymol. 2001, 330, 134–157. [Google Scholar] [PubMed]
- Cohen, G.N.; Barbe, V.; Flament, D.; Galperin, M.; Heilig, R.; Lecompte, O.; Poch, O.; Prieur, D.; Querellou, J.; Ripp, R.; et al. An integrated analysis of the genome of the hyperthermophilic archaeon Pyrococcus abyssi. Mol. Microbiol. 2003, 47, 1495–1512. [Google Scholar]
- Fukui, T.; Atomi, H.; Kanai, T.; Matsumi, R.; Fujiwara, S.; Imanaka, T. Complete genome sequence of the hyperthermophilic archaeon Thermococcus kodakaraensis KOD1 and comparison with Pyrococcus genomes. Genome Res. 2005, 15, 352–363. [Google Scholar] [CrossRef] [PubMed]
- Zivanovic, Y.; Armengaud, J.; Lagorce, A.; Leplat, C.; Guerin, P.; Dutertre, M.; Anthouard, V.; Forterre, P.; Wincker, P.; Confalonieri, F. Genome analysis and genome-wide proteomics of Thermococcus gammatolerans, the most radioresistant organism known amongst the Archaea. Genome Biol. 2009, 10, R70. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mardanov, A.V.; Ravin, N.V.; Svetlitchnyi, V.A.; Beletsky, A.V.; Miroshnichenko, M.L.; Bonch-Osmolovskaya, E.A.; Skryabin, K.G. Metabolic versatility and indigenous origin of the archaeon Thermococcus sibiricus, isolated from a siberian oil reservoir, as revealed by genome analysis. Appl. Environ. Microbiol. 2009, 75, 4580–4588. [Google Scholar] [CrossRef] [PubMed]
- Vannier, P.; Marteinsson, V.T.; Fridjonsson, O.H.; Oger, P.; Jebbar, M. Complete genome sequence of the hyperthermophilic, piezophilic, heterotrophic, and carboxydotrophic archaeon Thermococcus barophilus MP. J. Bacteriol. 2011, 193, 1481–1482. [Google Scholar] [CrossRef] [PubMed]
- Oger, P.; Sokolova, T.G.; Kozhevnikova, D.A.; Chernyh, N.A.; Bartlett, D.H.; Bonch-Osmolovskaya, E.A.; Lebedinsky, A.V. Complete genome sequence of the hyperthermophilic archaeon Thermococcus sp. strain AM4, capable of organotrophic growth and growth at the expense of hydrogenogenic or sulfidogenic oxidation of carbon monoxide. J. Bacteriol. 2011, 193, 7019–7020. [Google Scholar] [PubMed]
- Kwon, S.O.; Kang, S.G.; Park, S.H.; Kim, Y.H.; Choi, J.S.; Lee, J.H.; Kim, S.I. Proteomic characterization of the sulfur-reducing hyperthermophilic archaeon Thermococcus onnurineus NA1 by 2-DE/MS-MS. Extremophiles 2009, 13, 379–387. [Google Scholar] [CrossRef] [PubMed]
- Yun, S.H.; Kwon, S.O.; Park, G.W.; Kim, J.Y.; Kang, S.G.; Lee, J.H.; Chung, Y.H.; Kim, S.; Choi, J.S.; Kim, S.I. Proteome analysis of Thermococcus onnurineus NA1 reveals the expression of hydrogen gene cluster under carboxydotrophic growth. J. Proteomics 2011, 74, 1926–1933. [Google Scholar] [CrossRef] [PubMed]
- Moon, Y.J.; Kwon, J.; Yun, S.H.; Lim, H.L.; Kim, M.S.; Kang, S.G.; Lee, J.H.; Choi, J.S.; Kim, S.I.; Chung, Y.H. Proteome analyses of hydrogen-producing hyperthermophilic archaeon Thermococcus onnurineus NA1 in different one-carbon substrate culture conditions. Mol. Cell. Proteomics 2012, 11, M111015420. [Google Scholar] [CrossRef]
- Yang, H.Y.; Kwon, J.; Cho, E.J.; Choi, H.I.; Park, C.; Park, H.R.; Park, S.H.; Chung, K.J.; Ryoo, Z.Y.; Cho, K.O.; et al. Proteomic analysis of protein expression affected by peroxiredoxin V knock-down in hypoxic kidney. J. Proteome Res. 2010, 9, 4003–4015. [Google Scholar]
- Gao, B.B.; Stuart, L.; Feener, E.P. Label-free quantitative analysis of one-dimensional PAGE LC/MS/MS proteome: Application on angiotensin II-stimulated smooth muscle cells secretome. Mol. Cell. Proteomics 2008, 7, 2399–2409. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, S.; Inui, M.; Yukawa, H.; Kanao, T.; Tomizawa, K.; Atomi, H.; Imanaka, T. Phototrophic growth of a Rubisco-deficient mesophilic purple nonsulfur bacterium harboring a Type III Rubisco from a hyperthermophilic archaeon. J. Biotechnol. 2006, 124, 532–544. [Google Scholar] [CrossRef] [PubMed]
- Sato, T.; Atomi, H.; Imanaka, T. Archaeal type III RuBisCOs function in a pathway for AMP metabolism. Science 2007, 315, 1003–1006. [Google Scholar] [CrossRef] [PubMed]
- Adams, M.W.; Holden, J.F.; Menon, A.L.; Schut, G.J.; Grunden, A.M.; Hou, C.; Hutchins, A.M.; Jenney, F.E., Jr.; Kim, C.; Ma, K.; et al. Key role for sulfur in peptide metabolism and in regulation of three hydrogenases in the hyperthermophilic archaeon Pyrococcus furiosus. J. Bacteriol. 2001, 183, 716–724. [Google Scholar]
- Yang, D.; Shipman, L.W.; Roessner, C.A.; Scott, A.I.; Sacchettini, J.C. Structure of the Methanococcus jannaschii mevalonate kinase, a member of the GHMP kinase superfamily. J. Biol. Chem. 2002, 277, 9462–9467. [Google Scholar] [CrossRef] [PubMed]
- Primak, Y.A.; Du, M.; Miller, M.C.; Wells, D.H.; Nielsen, A.T.; Weyler, W.; Beck, Z.Q. Characterization of a feedback-resistant mevalonate kinase from the archaeon Methanosarcina mazei. Appl. Environ. Microbiol. 2011, 77, 7772–7778. [Google Scholar] [CrossRef] [PubMed]
- Nishimura, Y.; Eguchi, T. Biosynthesis of archaeal membrane lipids: Digeranylgeranylglycero-phospholipid reductase of the thermoacidophilic archaeon Thermoplasma acidophilum. J. Biochem. 2006, 139, 1073–1081. [Google Scholar] [CrossRef] [PubMed]
- Xu, Q.; Eguchi, T.; Mathews, I.I.; Rife, C.L.; Chiu, H.J.; Farr, C.L.; Feuerhelm, J.; Jaroszewski, L.; Klock, H.E.; Knuth, M.W.; et al. Insights into substrate specificity of geranylgeranyl reductases revealed by the structure of digeranylgeranylglycerophospholipid reductase, an essential enzyme in the biosynthesis of archaeal membrane lipids. J. Mol. Biol. 2010, 404, 403–417. [Google Scholar]
- Murakami, M.; Shibuya, K.; Nakayama, T.; Nishino, T.; Yoshimura, T.; Hemmi, H. Geranylgeranyl reductase involved in the biosynthesis of archaeal membrane lipids in the hyperthermophilic archaeon Archaeoglobus fulgidus. FEBS J. 2007, 274, 805–814. [Google Scholar] [CrossRef] [PubMed]
- Sato, S.; Murakami, M.; Yoshimura, T.; Hemmi, H. Specific partial reduction of geranylgeranyl diphosphate by an enzyme from the thermoacidophilic archaeon Sulfolobus acidocaldarius yields a reactive prenyl donor, not a dead-end product. J. Bacteriol. 2008, 190, 3923–3929. [Google Scholar] [CrossRef] [PubMed]
- Van de Vossenberg, J.L.; Driessen, A.J.; Konings, W.N. The essence of being extremophilic: The role of the unique archaeal membrane lipids. Extremophiles 1998, 2, 163–170. [Google Scholar] [CrossRef] [PubMed]
- Namboori, S.C.; Graham, D.E. Acetamido sugar biosynthesis in the Euryarchaea. J. Bacteriol. 2008, 190, 2987–2996. [Google Scholar] [CrossRef] [PubMed]
- Abu-Qarn, M.; Eichler, J. Protein N-glycosylation in Archaea: Defining Haloferax volcanii genes involved in S-layer glycoprotein glycosylation. Mol. Microbiol. 2006, 61, 511–525. [Google Scholar] [CrossRef] [PubMed]
- Jarrell, K.F.; Jones, G.M.; Nair, D.B. Biosynthesis and role of N-linked glycosylation in cell surface structures of archaea with a focus on flagella and S layers. Int. J. Microbiol. 2010, 2010, 470138. [Google Scholar] [CrossRef] [PubMed]
- White, R.H. Structures of the modified folates in the extremely thermophilic archaebacterium Thermococcus litoralis. J. Bacteriol. 1993, 175, 3661–3663. [Google Scholar] [PubMed]
- White, R.H. Structures of the modified folates in the thermophilic archaebacteria Pyrococcus furiosus. Biochemistry 1993, 32, 745–753. [Google Scholar] [CrossRef] [PubMed]
- Sauer, F.D.; Blackwell, B.A.; Kramer, J.K.; Marsden, B.J. Structure of a novel cofactor containing N-(7-mercaptoheptanoyl)-O-3-phosphothreonine. Biochemistry 1990, 29, 7593–7600. [Google Scholar] [CrossRef] [PubMed]
- Eichler, J.; Adams, M.W. Posttranslational protein modification in Archaea. Microbiol. Mol. Biol. Rev. 2005, 69, 393–425. [Google Scholar] [CrossRef] [PubMed]
- Yurist-Doutsch, S.; Chaban, B.; VanDyke, D.J.; Jarrell, K.F.; Eichler, J. Sweet to the extreme: Protein glycosylation in Archaea. Mol. Microbiol. 2008, 68, 1079–1084. [Google Scholar] [CrossRef] [PubMed]
- Ohmura, N.; Tsugita, K.; Koizumi, J.I.; Saika, H. Sulfur-binding protein of flagella of Thiobacillus ferrooxidans. J. Bacteriol. 1996, 178, 5776–5780. [Google Scholar] [PubMed]
- Lee, Y.G.; Leem, S.H.; Chung, Y.H.; Kim, S.I. Characterization of thermostable deblocking aminopeptidases of archaeon Thermococcus onnurineus NA1 by proteomic and biochemical approaches. J. Microbiol. 2012, 50, 792–797. [Google Scholar] [CrossRef] [PubMed]
- Fukui, T.; Eguchi, T.; Atomi, H.; Imanaka, T. A membrane-bound archaeal Lon protease displays ATP-independent proteolytic activity towards unfolded proteins and ATP-dependent activity for folded proteins. J. Bacteriol. 2002, 184, 3689–3698. [Google Scholar] [CrossRef] [PubMed]
- Cha, S.S.; An, Y.J.; Lee, C.R.; Lee, H.S.; Kim, Y.G.; Kim, S.J.; Kwon, K.K.; de Donatis, G.M.; Lee, J.H.; Maurizi, M.R.; et al. Crystal structure of Lon protease: Molecular architecture of gated entry to a sequestered degradation chamber. EMBO J. 2010, 29, 3520–3530. [Google Scholar]
- Horwitz, A.A.; Navon, A.; Groll, M.; Smith, D.M.; Reis, C.; Goldberg, A.L. ATP-induced structural transitions in PAN, the proteasome-regulatory ATPase complex in Archaea. J. Biol. Chem. 2007, 282, 22921–22929. [Google Scholar] [CrossRef] [PubMed]
- Takacs, M.; Toth, A.; Bogos, B.; Varga, A.; Rakhely, G.; Kovacs, K.L. Formate hydrogenlyase in the hyperthermophilic archaeon, Thermococcus litoralis. BMC Microbiol. 2008, 8, 88. [Google Scholar] [CrossRef] [PubMed]
- Silva, P.J.; van den Ban, E.C.; Wassink, H.; Haaker, H.; de Castro, B.; Robb, F.T.; Hagen, W.R. Enzymes of hydrogen metabolism in Pyrococcus furiosus. Eur. J. Biochem. 2000, 267, 6541–6551. [Google Scholar] [CrossRef] [PubMed]
- Kobori, H.; Ogino, M.; Orita, I.; Nakamura, S.; Imanaka, T.; Fukui, T. Characterization of NADH oxidase/NADPH polysulfide oxidoreductase and its unexpected participation in oxygen sensitivity in an anaerobic hyperthermophilic archaeon. J. Bacteriol. 2010, 192, 5192–5202. [Google Scholar] [CrossRef] [PubMed]
- Campbell, B.J.; Smith, J.L.; Hanson, T.E.; Klotz, M.G.; Stein, L.Y.; Lee, C.K.; Wu, D.; Robinson, J.M.; Khouri, H.M.; Eisen, J.A.; et al. Adaptations to submarine hydrothermal environments exemplified by the genome of Nautilia profundicola. PLoS Genet. 2009, 5, e1000362. [Google Scholar]
- Ladenstein, R.; Ren, B. Protein disulfides and protein disulfide oxidoreductases in hyperthermophiles. FEBS J. 2006, 273, 4170–4185. [Google Scholar] [CrossRef] [PubMed]
- Hagen, W.R.; Silva, P.J.; Amorim, M.A.; Hagedoorn, P.L.; Wassink, H.; Haaker, H.; Robb, F.T. Novel structure and redox chemistry of the prosthetic groups of the iron-sulfur flavoprotein sulfide dehydrogenase from Pyrococcus furiosus; evidence for a [2Fe–2S] cluster with Asp(Cys)3 ligands. J. Biol. Inorg. Chem. 2000, 5, 527–534. [Google Scholar] [CrossRef] [PubMed]
- Ma, K.; Adams, M.W. Sulfide dehydrogenase from the hyperthermophilic archaeon Pyrococcus furiosus: A new multifunctional enzyme involved in the reduction of elemental sulfur. J. Bacteriol. 1994, 176, 6509–6517. [Google Scholar] [PubMed]
- Schut, G.J.; Brehm, S.D.; Datta, S.; Adams, M.W. Whole-genome DNA microarray analysis of A hyperthermophile and an archaeon: Pyrococcus furiosus grown on carbohydrates or peptides. J. Bacteriol. 2003, 185, 3935–3947. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Sieprawska-Lupa, M.; Whitman, W.B.; White, R.H. Cysteine is not the sulfur source for iron-sulfur cluster and methionine biosynthesis in the methanogenic archaeon Methanococcus maripaludis. J. Biol. Chem. 2010, 285, 31923–31929. [Google Scholar] [CrossRef] [PubMed]
- Lipscomb, G.L.; Keese, A.M.; Cowart, D.M.; Schut, G.J.; Thomm, M.; Adams, M.W.; Scott, R.A. SurR: A transcriptional activator and repressor controlling hydrogen and elemental sulphur metabolism in Pyrococcus furiosus. Mol. Microbiol. 2009, 71, 332–349. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Lipscomb, G.L.; Keese, A.M.; Schut, G.J.; Thomm, M.; Adams, M.W.; Wang, B.C.; Scott, R.A. SurR regulates hydrogen production in Pyrococcus furiosus by a sulfur-dependent redox switch. Mol. Microbiol. 2010, 77, 1111–1122. [Google Scholar] [CrossRef] [PubMed]
- Sapra, R.; Bagramyan, K.; Adams, M.W. A simple energy-conserving system: Proton reduction coupled to proton translocation. Proc. Natl. Acad. Sci. USA 2003, 100, 7545–7550. [Google Scholar] [CrossRef] [PubMed]
- Ma, K.; Adams, M.W. Hydrogenases I and II from Pyrococcus furiosus. Methods Enzymol. 2001, 331, 208–216. [Google Scholar] [PubMed]
- Van Haaster, D.J.; Silva, P.J.; Hagedoorn, P.L.; Jongejan, J.A.; Hagen, W.R. Reinvestigation of the steady-state kinetics and physiological function of the soluble NiFe-hydrogenase I of Pyrococcus furiosus. J. Bacteriol. 2008, 190, 1584–1587. [Google Scholar] [CrossRef] [PubMed]
- Forzi, L.; Sawers, R.G. Maturation of [NiFe]-hydrogenases in Escherichia coli. Biometals 2007, 20, 565–758. [Google Scholar] [CrossRef] [PubMed]
- Jenney, F.E., Jr.; Adams, M.W. Hydrogenases of the model hyperthermophiles. Ann. N. Y. Acad. Sci. 2008, 1125, 252–266. [Google Scholar] [CrossRef] [PubMed]
- Thorgersen, M.P.; Stirrett, K.; Scott, R.A.; Adams, M.W. Mechanism of oxygen detoxification by the surprisingly oxygen-tolerant hyperthermophilic archaeon, Pyrococcus furiosus. Proc. Natl. Acad. Sci. USA 2012, 109, 18547–18552. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.J.; Engelmann, A.; Horlacher, R.; Qu, Q.; Vierke, G.; Hebbeln, C.; Thomm, M.; Boos, W. TrmB, a sugar-specific transcriptional regulator of the trehalose/maltose ABC transporter from the hyperthermophilic archaeon Thermococcus litoralis. J. Biol. Chem. 2003, 278, 983–990. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.J.; Moulakakis, C.; Koning, S.M.; Hausner, W.; Thomm, M.; Boos, W. TrmB, a sugar sensing regulator of ABC transporter genes in Pyrococcus furiosus exhibits dual promoter specificity and is controlled by different inducers. Mol. Microbiol. 2005, 57, 1797–1807. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.S.; Kim, Y.J.; Lee, J.H.; Kang, S.G. Identification and characterization of inorganic pyrophosphatase and PAP phosphatase from Thermococcus onnurineus NA1. J. Bacteriol. 2009, 191, 3415–3419. [Google Scholar] [CrossRef] [PubMed]
- Sevcenco, A.M.; Bevers, L.E.; Pinkse, M.W.; Krijger, G.C.; Wolterbeek, H.T.; Verhaert, P.D.; Hagen, W.R.; Hagedoorn, P.L. Molybdenum incorporation in tungsten aldehyde oxidoreductase enzymes from Pyrococcus furiosus. J. Bacteriol. 2010, 192, 4143–4152. [Google Scholar] [CrossRef] [PubMed]
- Borup, B.; Ferry, J.G. Cysteine biosynthesis in the Archaea: Methanosarcina thermophila utilizes O-acetylserine sulfhydrylase. FEMS Microbiol. Lett. 2000, 189, 205–210. [Google Scholar] [CrossRef] [PubMed]
- Raven, N.; Sharp, R.J. Development of defined and minimal media for the growth of the hyperthermophilic archaeon Pyrococcus furiosus Vc1. FEMS Microbiol. Lett. 1997, 146, 135–141. [Google Scholar] [CrossRef]
- Hidese, R.; Inoue, T.; Imanaka, T.; Fujiwara, S. Cysteine desulphurase plays an important role in environmental adaptation of the hyperthermophilic archaeon Thermococcus kodakarensis. Mol. Microbiol. 2014, 93, 331–345. [Google Scholar] [CrossRef] [PubMed]
- Holden, J.F.; Takai, K.; Summit, M.; Bolton, S.; Zyskowski, J.; Baross, J.A. Diversity among three novel groups of hyperthermophilic deep-sea Thermococcus species from three sites in the northeastern Pacific Ocean. FEMS Microbiol. Ecol. 2001, 36, 51–60. [Google Scholar] [CrossRef] [PubMed]
- Sokolova, T.G.; Jeanthon, C.; Kostrikina, N.A.; Chernyh, N.A.; Lebedinsky, A.V.; Stackebrandt, E.; Bonch-Osmolovskaya, E.A. The first evidence of anaerobic CO oxidation coupled with H2 production by a hyperthermophilic archaeon isolated from a deep-sea hydrothermal vent. Extremophiles 2004, 8, 317–323. [Google Scholar] [CrossRef] [PubMed]
- Balch, W.E.; Wolfe, R.S. New approach to the cultivation of methanogenic bacteria: 2-Mercaptoethanesulfonic acid (HS-CoM)-dependent growth of Methanobacterium ruminantium in a pressureized atmosphere. Appl. Environ. Microbiol. 1976, 32, 781–791. [Google Scholar] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Moon, Y.-J.; Kwon, J.; Yun, S.-H.; Lim, H.L.; Kim, J.; Kim, S.J.; Kang, S.G.; Lee, J.-H.; Kim, S.I.; Chung, Y.-H. Proteomic Insights into Sulfur Metabolism in the Hydrogen-Producing Hyperthermophilic Archaeon Thermococcus onnurineus NA1. Int. J. Mol. Sci. 2015, 16, 9167-9195. https://doi.org/10.3390/ijms16059167
Moon Y-J, Kwon J, Yun S-H, Lim HL, Kim J, Kim SJ, Kang SG, Lee J-H, Kim SI, Chung Y-H. Proteomic Insights into Sulfur Metabolism in the Hydrogen-Producing Hyperthermophilic Archaeon Thermococcus onnurineus NA1. International Journal of Molecular Sciences. 2015; 16(5):9167-9195. https://doi.org/10.3390/ijms16059167
Chicago/Turabian StyleMoon, Yoon-Jung, Joseph Kwon, Sung-Ho Yun, Hye Li Lim, Jonghyun Kim, Soo Jung Kim, Sung Gyun Kang, Jung-Hyun Lee, Seung Il Kim, and Young-Ho Chung. 2015. "Proteomic Insights into Sulfur Metabolism in the Hydrogen-Producing Hyperthermophilic Archaeon Thermococcus onnurineus NA1" International Journal of Molecular Sciences 16, no. 5: 9167-9195. https://doi.org/10.3390/ijms16059167