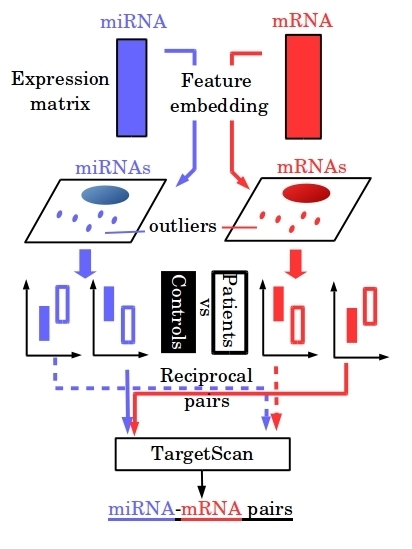
Identification of More Feasible MicroRNA–mRNA Interactions within Multiple Cancers Using Principal Component Analysis Based Unsupervised Feature Extraction
Abstract
:1. Introduction
2. Results and Discussion
2.1. Hepatocellular Carcinoma (HCC)
2.2. Non-Small Cell Lung Cancer (NSCLC)
2.3. Esophageal Squamous Cell Cancer (ESCC)
2.4. Prostate Cancer
2.5. Colorectal/Colon Cancer
2.6. Breast Cancer
2.7. Confirmation of Significance of the FDR Criterion
2.8. Discrimination Performance between Patients and Healthy Controls
2.9. Confident Candidate Selection by PCA-Based Unsupervised FE
2.10. Usefulness of Unmatched Data and Number of False Negatives
3. Materials and Methods
3.1. Gene Expression Profiles
3.1.1. HCC
3.1.2. NSCLC
3.1.3. ESCC
3.1.4. Prostate Cancer
3.1.5. Colorectal/Colon Cancer
3.1.6. Breast Tumors
3.2. PCA-Based Unsupervised FE
3.3. Identification of Significant miRNA–mRNA Pairs
3.4. Validation Using Starbase
3.5. Discrimination between Patients and Healthy Controls
3.6. Validation Using FDR
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Ha, M.; Kim, V.N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 2014, 15, 509–524. [Google Scholar] [CrossRef] [PubMed]
- Cloonan, N. Re-thinking miRNA–mRNA interactions: Intertwining issues confound target discovery. Bioessays 2015, 37, 379–388. [Google Scholar] [CrossRef] [PubMed]
- Taguchi, Y.H.; Iwadate, M.; Umeyama, H. Principal component analysis-based unsupervised feature extraction applied to in silico drug discovery for posttraumatic stress disorder-mediated heart disease. BMC Bioinform. 2015, 16, 139. [Google Scholar] [CrossRef] [PubMed]
- Li, J.H.; Liu, S.; Zhou, H.; Qu, L.H.; Yang, J.H. starBase v2.0: Decoding miRNA–ceRNA, miRNA–ncRNA and protein–RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014, 42, D92–D97. [Google Scholar] [CrossRef] [PubMed]
- Jiang, B.Y.; Zhang, X.C.; Su, J.; Meng, W.; Yang, X.N.; Yang, J.J.; Zhou, Q.; Chen, Z.Y.; Chen, Z.H.; Xie, Z.; et al. BCL11A overexpression predicts survival and relapse in non-small cell lung cancer and is modulated by microRNA-30a and gene amplification. Mol. Cancer 2013, 12, 61. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Chen, X.; Liang, H.; Deng, T.; Chen, W.; Zhang, S.; Liu, M.; Gao, X.; Liu, Y.; Zhao, C.; et al. miR-143 and miR-145 synergistically regulate ERBB3 to suppress cell proliferation and invasion in breast cancer. Mol. Cancer 2014, 13, 220. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B (Methodol.) 1995, 57, 289–300. [Google Scholar]
- Ding, M.; Li, J.; Yu, Y.; Liu, H.; Yan, Z.; Wang, J.; Qian, Q. Integrated analysis of miRNA, gene, and pathway regulatory networks in hepatic cancer stem cells. J. Transl. Med. 2015, 13, 259. [Google Scholar] [CrossRef] [PubMed]
- Ma, R.; Wang, C.; Wang, J.; Wang, D.; Xu, J. miRNA–mRNA interaction network in non-small-cell lung cancer. Interdiscip. Sci. 2015. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Zhang, Q.; Zhang, M.; Zhang, Y.; Li, F.; Lei, P. Analysis for the mechanism between the small cell lung cancer and non-small cell lung cancer combing the miRNA and mRNA expression profiles. Thorac. Cancer 2015, 6, 70–79. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Huang, Y.; Zhu, W.; Zhou, S.; Zhou, J.; Zeng, F.; Liu, X.; Zhang, Y.; Yu, J. An integrated analysis of miRNA and mRNA expressions in non-small cell lung cancers. PLoS ONE 2011, 6, e26502. [Google Scholar] [CrossRef] [PubMed]
- Wu, B.; Li, C.; Zhang, P.; Yao, Q.; Wu, J.; Han, J.; Liao, L.; Xu, Y.; Lin, R.; Xiao, D.; et al. Dissection of miRNA–miRNA interaction in esophageal squamous cell carcinoma. PLoS ONE 2013, 8, e73191. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Li, D.; Yang, Y.; Jiang, G. An integrated analysis of the effects of microRNA and mRNA on esophageal squamous cell carcinoma. Mol. Med. Rep. 2015, 12, 945–952. [Google Scholar] [CrossRef] [PubMed]
- Meng, X.R.; Lu, P.; Mei, J.Z.; Liu, G.J.; Fan, Q.X. Expression analysis of miRNA and target mRNAs in esophageal cancer. Braz. J. Med. Biol. Res. 2014, 47, 811–817. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Edwards, A.; Fan, W.; Flemington, E.K.; Zhang, K. miRNA–mRNA correlation-network modules in human prostate cancer and the differences between primary and metastatic tumor subtypes. PLoS ONE 2012, 7, e40130. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.; Tang, W.; Du, P.; Wang, G.; Chen, W.; Li, J.; Zhu, Y.; Gao, J.; Cui, L. Identifying microRNA–mRNA regulatory network in colorectal cancer by a combination of expression profile and bioinformatics analysis. BMC Syst. Biol. 2012, 6, 68. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Gill, R.; Cooper, N.G.; Yoo, J.K.; Datta, S. Modeling microRNA–mRNA interactions using PLS regression in human colon cancer. BMC Med. Genom. 2011, 4, 44. [Google Scholar] [CrossRef] [PubMed]
- Bleckmann, A.; Leha, A.; Artmann, S.; Menck, K.; Salinas-Riester, G.; Binder, C.; Pukrop, T.; Beissbarth, T.; Klemm, F. Integrated miRNA and mRNA profiling of tumor-educated macrophages identifies prognostic subgroups in estrogen receptor-positive breast cancer. Mol. Oncol. 2015, 9, 155–166. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.F.; Jiang, W.H.; Han, Y.T.; He, L.F.; Zhang, H.L.; Ren, H. Integrated microRNA–mRNA analysis of pancreatic ductal adenocarcinoma. Genet. Mol. Res. 2015, 14, 10288–10297. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, X.; Li, Z.; Lin, H.; Gu, L.; Lin, Q.; Lu, Z.; Tzeng, C.M. Integrated miRNA and mRNA expression profiling to identify mRNA targets of dysregulated miRNAs in non-obstructive azoospermia. Sci. Rep. 2015, 5, 7922. [Google Scholar] [CrossRef] [PubMed]
- Naderi, E.; Mostafaei, M.; Pourshams, A.; Mohamadkhani, A. Network of microRNAs–mRNAs interactions in pancreatic cancer. Biomed. Res. Int. 2014, 2014, 534821. [Google Scholar] [CrossRef] [PubMed]
- Wei, L.; Lian, B.; Zhang, Y.; Li, W.; Gu, J.; He, X.; Xie, L. Application of microRNA and mRNA expression profiling on prognostic biomarker discovery for hepatocellular carcinoma. BMC Genom. 2014, 15 (Suppl. 1), S13. [Google Scholar] [CrossRef] [PubMed]
- Shih, T.C.; Tien, Y.J.; Wen, C.J.; Yeh, T.S.; Yu, M.C.; Huang, C.H.; Lee, Y.S.; Yen, T.C.; Hsieh, S.Y. MicroRNA-214 downregulation contributes to tumor angiogenesis by inducing secretion of the hepatoma-derived growth factor in human hepatoma. J. Hepatol. 2012, 57, 584–591. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Palencia, A.; Gomez-Morales, M.; Gomez-Capilla, J.A.; Pedraza, V.; Boyero, L.; Rosell, R.; Farez-Vidal, M.E. Gene expression profiling reveals novel biomarkers in nonsmall cell lung cancer. Int. J. Cancer 2011, 129, 355–364. [Google Scholar] [CrossRef] [PubMed]
- Tan, X.; Qin, W.; Zhang, L.; Hang, J.; Li, B.; Zhang, C.; Wan, J.; Zhou, F.; Shao, K.; Sun, Y.; et al. A 5-microRNA signature for lung squamous cell carcinoma diagnosis and hsa-miR-31 for prognosis. Clin. Cancer Res. 2011, 17, 6802–6811. [Google Scholar] [CrossRef] [PubMed]
- Hu, N.; Wang, C.; Clifford, R.J.; Yang, H.H.; Su, H.; Wang, L.; Wang, Y.; Xu, Y.; Tang, Z.Z.; Ding, T.; et al. Integrative genomics analysis of genes with biallelic loss and its relation to the expression of mRNA and micro-RNA in esophageal squamous cell carcinoma. BMC Genom. 2015, 16, 732. [Google Scholar] [CrossRef] [PubMed]
- Mathe, E.A.; Nguyen, G.H.; Bowman, E.D.; Zhao, Y.; Budhu, A.; Schetter, A.J.; Braun, R.; Reimers, M.; Kumamoto, K.; Hughes, D.; et al. MicroRNA expression in squamous cell carcinoma and adenocarcinoma of the esophagus: Associations with survival. Clin. Cancer Res. 2009, 15, 6192–6200. [Google Scholar] [CrossRef] [PubMed]
- Taylor, B.S.; Schultz, N.; Hieronymus, H.; Gopalan, A.; Xiao, Y.; Carver, B.S.; Arora, V.K.; Kaushik, P.; Cerami, E.; Reva, B.; et al. Integrative genomic profiling of human prostate cancer. Cancer Cell 2010, 18, 11–22. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.D.; Ceniccola, K.; Yang, Q.; Andrawis, R.; Patel, V.; Ji, Y.; Rhim, J.; Olender, J.; Popratiloff, A.; Latham, P.; et al. Identification and Functional Validation of Reciprocal microRNA-mRNA Pairings in African American Prostate Cancer Disparities. Clin. Cancer Res. 2015, 21, 4970–4984. [Google Scholar] [CrossRef] [PubMed]
- Sheffer, M.; Bacolod, M.D.; Zuk, O.; Giardina, S.F.; Pincas, H.; Barany, F.; Paty, P.B.; Gerald, W.L.; Notterman, D.A.; Domany, E. Association of survival and disease progression with chromosomal instability: A genomic exploration of colorectal cancer. Proc. Natl. Acad. Sci. USA 2009, 106, 7131–7136. [Google Scholar] [CrossRef] [PubMed]
- Li, E.; Ji, P.; Ouyang, N.; Zhang, Y.; Wang, X.Y.; Rubin, D.C.; Davidson, N.O.; Bergamaschi, R.; Shroyer, K.R.; Burke, S.; et al. Differential expression of miRNAs in colon cancer between African and Caucasian Americans: Implications for cancer racial health disparities. Int. J. Oncol. 2014, 45, 587–594. [Google Scholar] [CrossRef] [PubMed]
- Farazi, T.A.; Horlings, H.M.; Ten Hoeve, J.J.; Mihailovic, A.; Halfwerk, H.; Morozov, P.; Brown, M.; Hafner, M.; Reyal, F.; van Kouwenhove, M.; et al. MicroRNA sequence and expression analysis in breast tumors by deep sequencing. Cancer Res. 2011, 71, 4443–4453. [Google Scholar] [CrossRef] [PubMed]
- Taguchi, Y.H. Identification of aberrant gene expression associated with aberrant promoter methylation in primordial germ cells between E13 and E16 rat F3 generation vinclozolin lineage. BMC Bioinform. 2015, 16 (Suppl. 18), S16. [Google Scholar] [CrossRef] [PubMed]
- Taguchi, Y.h. Integrative Analysis of Gene Expression and Promoter Methylation during Reprogramming of a Non-Small-Cell Lung Cancer Cell Line Using Principal Component Analysis-Based Unsupervised Feature Extraction. In Intelligent Computing in Bioinformatics; Huang, D.S., Han, K., Gromiha, M., Eds.; Springer International Publishing: Heidelberg, Germany, 2014; Volume 8590, LNCS; pp. 445–455. [Google Scholar]
- Taguchi, Y.h.; Iwadate, M.; Umeyama, H.; Murakami, Y.; Okamoto, A. Heuristic principal component analysis-aased unsupervised feature extraction and its application to bioinformatics. In Big Data Analytics in Bioinformatics and Healthcare; Wang, B., Li, R., Perrizo, W., Eds.; IGI Global: Hershey, PA, USA, 2015; pp. 138–162. [Google Scholar]
- Taguchi, Y.H.; Iwadate, M.; Umeyama, H. Heuristic principal component analysis-based unsupervised feature extraction and its application to gene expression analysis of amyotrophic lateral sclerosis data sets. In Proceedings of the 2015 IEEE Conference on Computational Intelligence in Bioinformatics and Computational Biology (CIBCB), Niagara Falls, ON, Canada, 12–15 August 2015; pp. 1–10.
- Umeyama, H.; Iwadate, M.; Taguchi, Y.H. TINAGL1 and B3GALNT1 are potential therapy target genes to suppress metastasis in non-small cell lung cancer. BMC Genom. 2014, 15 (Suppl. 9), S2. [Google Scholar] [CrossRef] [PubMed]
- Murakami, Y.; Kubo, S.; Tamori, A.; Itami, S.; Kawamura, E.; Iwaisako, K.; Ikeda, K.; Kawada, N.; Ochiya, T.; Taguchi, Y.H. Comprehensive analysis of transcriptome and metabolome analysis in Intrahepatic Cholangiocarcinoma and Hepatocellular Carcinoma. Sci. Rep. 2015, 5, 16294. [Google Scholar] [CrossRef] [PubMed]
- Murakami, Y.; Tanahashi, T.; Okada, R.; Toyoda, H.; Kumada, T.; Enomoto, M.; Tamori, A.; Kawada, N.; Taguchi, Y.H.; Azuma, T. Comparison of Hepatocellular Carcinoma miRNA Expression Profiling as Evaluated by Next Generation Sequencing and Microarray. PLoS ONE 2014, 9, e106314. [Google Scholar] [CrossRef] [PubMed]
- Murakami, Y.; Toyoda, H.; Tanahashi, T.; Tanaka, J.; Kumada, T.; Yoshioka, Y.; Kosaka, N.; Ochiya, T.; Taguchi, Y.H. Comprehensive miRNA expression analysis in peripheral blood can diagnose liver disease. PLoS ONE 2012, 7, e48366. [Google Scholar] [CrossRef] [PubMed]
- Taguchi, Y.H.; Murakami, Y. Universal disease biomarker: Can a fixed set of blood microRNAs diagnose multiple diseases? BMC Res. Notes 2014, 7, 581. [Google Scholar] [CrossRef] [PubMed]
- Taguchi, Y.H.; Murakami, Y. Principal component analysis based feature extraction approach to identify circulating microRNA biomarkers. PLoS ONE 2013, 8, e66714. [Google Scholar] [CrossRef] [PubMed]
- Kinoshita, R.; Iwadate, M.; Umeyama, H.; Taguchi, Y.H. Genes associated with genotype-specific DNA methylation in squamous cell carcinoma as candidate drug targets. BMC Syst. Biol. 2014, 8 (Suppl. 1), S4. [Google Scholar] [CrossRef] [PubMed]
- Ishida, S.; Umeyama, H.; Iwadate, M.; Taguchi, Y.H. Bioinformatic Screening of Autoimmune Disease Genes and Protein Structure Prediction with FAMS for Drug Discovery. Protein Pept. Lett. 2014, 21, 828–839. [Google Scholar] [CrossRef] [PubMed]
- Taguchi, Y.h.; Okamoto, A. Principal Component Analysis for Bacterial Proteomic Analysis. In Pattern Recognition in Bioinformatics; Shibuya, T., Kashima, H., Sese, J., Ahmad, S., Eds.; Springer International Publishing: Heidelberg, Germany, 2012; Volume 7632, LNCS; pp. 141–152. [Google Scholar]
- Agarwal, V.; Bell, G.W.; Nam, J.W.; Bartel, D.P. Predicting effective microRNA target sites in mammalian mRNAs. Elife 2015, 4. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2015. [Google Scholar]
- Klaus, B.; Strimmer, K. fdrtool: Estimation of (Local) False Discovery Rates and Higher Criticism. Available online: https://cran.r-project.org/web/packages/fdrtool/index.html (accessed on 10 March 2016).
| Number of Samples | Number of Probes | ||||
|---|---|---|---|---|---|
| Cancers | GEO ID | Tumors | Controls | Selected | Non-Selected |
| HCC | |||||
| mRNA | GSE45114 | 24 | 25 | 269 | 22,963 |
| miRNA | GSE36915 | 68 | 21 | 58 | 1087 |
| NSCLC | |||||
| mRNA | GSE18842 | 46 | 45 | 1098 | 53,504 |
| miRNA | GSE15008 | 187 | 174 | 268 | 3428 |
| ESCC | |||||
| mRNA | GSE38129 | 30 | 30 | 189 | 22,088 |
| miRNA | GSE19337 | 76 | 76 | 37 | 1217 |
| Prostate cancer | |||||
| mRNA | GSE21032 | 150 | 29 | 399 | 43,020 |
| miRNA | GSE84318 | 27 | 27 | 23 | 700 |
| Colon/colorectal cancer | |||||
| mRNA | GSE41258 | 186 | 54 | 309 | 21,974 |
| miRNA | GSE48267 | 30 | 30 | 12 | 839 |
| Breast cancer | |||||
| mRNA | GSE29174 | 110 | 11 | 980 | 33,600 |
| miRNA | GSE28884 | 173 | 16 | 18 | 2258 |
| HCC | NSCLC | ESCC | ||||
|---|---|---|---|---|---|---|
| mRNA | miRNA | mRNA | miRNA | mRNA | miRNA | |
| FDR | 262 | 38 | 978 | 0 | 189 | 0 |
| BH | 269 | 58 | 1091 | 268 | 189 | 37 |
| Prostate Cancer | Colon/Colorectal Cancer | Breast Cancer | ||||
| mRNA | miRNA | mRNA | miRNA | mRNA | miRNA | |
| FDR | 399 | 7 | 305 | 12 | 861 | 0 |
| BH | 399 | 23 | 309 | 12 | 908 | 18 |
| mRNA | miRNA | |||
|---|---|---|---|---|
| HCC | hc | HCC | hc | |
| HCC | 20 | 0 | 64 | 0 |
| hc | 4 | 25 | 4 | 21 |
| (L, p-value, odds ratio) | (4, , ∞) | (10, *, ∞) | ||
| NSCLC | hc | NSCLC | hc | |
| NSCLC | 46 | 0 | 171 | 12 |
| hc | 0 | 45 | 16 | 162 |
| (L, p-value, odds ratio) | (2,*, ∞) | (5, *, ) | ||
| ESCC | hc | ESCC | hc | |
| ESCC | 28 | 2 | 63 | 11 |
| hc | 2 | 28 | 13 | 65 |
| (L, p-value, odds ratio) | (2, , ) | (6, *, ) | ||
| Pancreatic cancer | hc | Pancreatic cancer | hc | |
| Pancreatic cancer | 139 | 4 | 22 | 3 |
| hc | 11 | 25 | 5 | 24 |
| (L, p-value, odds ratio) | (8, *, ) | (4, , ) | ||
| Colorectal cancer | hc | Colon cancer | hc | |
| Colon/Colorectal cancer | 178 | 5 | 27 | 3 |
| hc | 8 | 49 | 3 | 27 |
| (L, p-value, odds ratio) | (8, *, ) | (4, , ) | ||
| Breast cancer | hc | Breast cancer | hc | |
| Breast cancer | 110 | 0 | 169 | 5 |
| hc | 0 | 11 | 4 | 11 |
| (L, p-value, odds ratio) | (3, , ∞) | (18, , ) | ||
© 2016 by the author; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Taguchi, Y.-h. Identification of More Feasible MicroRNA–mRNA Interactions within Multiple Cancers Using Principal Component Analysis Based Unsupervised Feature Extraction. Int. J. Mol. Sci. 2016, 17, 696. https://doi.org/10.3390/ijms17050696
Taguchi Y-h. Identification of More Feasible MicroRNA–mRNA Interactions within Multiple Cancers Using Principal Component Analysis Based Unsupervised Feature Extraction. International Journal of Molecular Sciences. 2016; 17(5):696. https://doi.org/10.3390/ijms17050696
Chicago/Turabian StyleTaguchi, Y-h. 2016. "Identification of More Feasible MicroRNA–mRNA Interactions within Multiple Cancers Using Principal Component Analysis Based Unsupervised Feature Extraction" International Journal of Molecular Sciences 17, no. 5: 696. https://doi.org/10.3390/ijms17050696