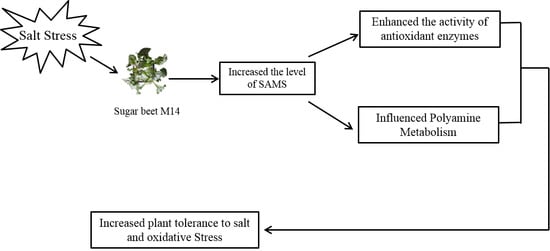
Overexpression of S-Adenosyl-l-Methionine Synthetase 2 from Sugar Beet M14 Increased Arabidopsis Tolerance to Salt and Oxidative Stress
Abstract
:1. Introduction
2. Results
2.1. Identification of Differentially Expressed M14 Genes under Salt Stress Using SSH
2.2. Cloning of a BvM14-SAMS2 Gene and Sequence Analysis
2.3. Analysis of BvM14-SAMS2 Response to Salt Stress
2.4. Overexpression of BvM14-SAMS2 Confers Enhanced Salt and H2O2 Tolerance in Arabidopsis
2.5. Overexpression of BvM14-SAMS2 Increased Antioxidative Activities in Arabidopsis
2.6. Overexpression of BvM14-SAMS2 Greatly Influenced Polyamine Metabolism
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Construction of a Subtractive cDNA Library
4.3. Molecular Cloning of BvM14-SAMS2 Gene and Sequence Analysis
4.4. Real-Time Quantitative PCR
4.5. Isolation of T-DNA Insertion Mutants in AtSAMS3
4.6. Constitutive Expression of BvM14-SAMS2 in Arabidopsis and Stress Tolerance Analysis
4.7. Determination of Antioxidant Enzyme Activities, Lipid Peroxidation, H2O2 Concentration, SAM Concentration and Polyamine Metabolism
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Zhu, J.K. Genetic analysis of plant salt tolerance using Arabidopsis. Plant Physiol. 2000, 124, 941–948. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.K. Plant salt tolerance. Trends Plant Sci. 2001, 6, 66–71. [Google Scholar] [CrossRef]
- Munns, R.; Tester, M. Mechanisms of salinity tolerance. Annu. Rev. Plant Biol. 2008, 59, 651–681. [Google Scholar] [CrossRef] [PubMed]
- Turan, M.A.; Elkarim, A.H.A.; Taban, N.; Taban, S. Effect of salt stress on growth, stomatal resistance, proline and chlorophyll concentrations on maize plant. Afr. J. Agric. Res. 2009, 4, 893–897. [Google Scholar]
- Barkla, B.J.; Castellanos-Cervantes, T.; de León, J.L.; Matros, A.; Mock, H.P.; Perez-Alfocea, F.; Salekdeh, G.H.; Witzel, K.; Zörb, C. Elucidation of salt stress defense and tolerance mechanisms of crop plants using proteomics—Current achievements and perspectives. Proteomics 2013, 13, 1885–1900. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Ma, C.; Wang, L.; Chen, S.; Li, H. Salt stress induced proteome and transcriptome changes in sugar beet monosomic addition line M14. J. Plant Physiol. 2012, 169, 839–850. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Zhang, Y.; Zhu, N.; Koh, J.; Ma, C.; Pan, Y.; Yu, B.; Chen, S.; Li, H. Proteomic analysis of salt tolerance in sugar beet monosomic addition line M14. J. Proteome Res. 2013, 12, 4931–4950. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Lai, J.; Sun, S.; Li, Y.; Liu, Y.; Liang, L.; Chen, M.; Xie, Q. Comparison analysis of transcripts from the halophyte Thellungiella halophila. J. Integr. Plant Biol. 2008, 50, 1327–1335. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Ma, X.L.; Zhang, Q.; Ma, C.L.; Wang, P.P.; Sun, Y.F.; Zhao, Y.X.; Zhang, H. Expressed sequence tags from a NaCl-treated Suaeda salsa cDNA library. Gene 2001, 267, 193–200. [Google Scholar] [CrossRef]
- Zouari, N.; Saad, R.B.; Legavre, T.; Azaza, J.; Sabau, X.; Jaoua, M.; Masmoudi, K.; Hassairi, A. Identification and sequencing of ESTs from the halophyte grass Aeluropus littoralis. Gene 2007, 404, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Jha, B.; Agarwal, P.K.; Reddy, P.S.; Lal, S.; Sopory, S.K.; Reddy, M.K. Identification of salt-induced genes from Salicornia brachiata,an extreme halophyte through expressed sequence tags analysis. Genes Genet. Syst. 2009, 84, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Askari, H.; Edqvist, J.; Hajheidari, M.; Kafi, M.; Salekdeh, G.H. Effects of Salinity levels on proteome of Suaeda aegyptiaca leaves. Proteomics 2006, 6, 2542–2554. [Google Scholar] [CrossRef] [PubMed]
- Pang, Q.; Chen, S.; Dai, S.; Chen, Y.; Wang, Y.; Yan, X. Comparative proteomics of salt tolerance in Arabidopsis thaliana and Thellungiella halophila. J. Proteome Res. 2010, 9, 2584–2599. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Meng, Y.; Li, B.; Ma, X.; Lai, Y.; Si, E.; Yang, K.; Xu, X.; Shang, X.; Wang, H.; Wang, D. Physiological and proteomic analyses of salt stress response in the halophyte Halogeton glomeratus. Plant Cell Environ. 2015, 38, 655–669. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Fan, P.; Song, H.; Chen, X.; Li, X.; Li, Y. Comparative proteomic analysis of differentially expressed proteins in shoots of Salicornia europaeaunder different salinity. J. Proteome Res. 2009, 8, 3331–3345. [Google Scholar] [CrossRef] [PubMed]
- Roje, S. S-Adenosyl-l-methionine: Beyond the universal methyl group donor. Phytochemistry 2006, 67, 1686–1698. [Google Scholar] [CrossRef] [PubMed]
- Nagel, J.; Culley, L.K.; Lu, Y.; Liu, E.; Matthews, P.D.; Stevens, J.F.; Page, J.E. EST analysis of hop glandular trichomes identifies an O-methyltransferase that catalyzes the biosynthesis of xanthohumol. Plant Cell 2008, 20, 186–200. [Google Scholar] [CrossRef] [PubMed]
- Köllner, T.G.; Lenk, C.; Zhao, N.; Seidl-Adams, I.; Gershenzon, J.; Chen, F.; Degenhardt, J. Herbivore-induced SABATH methyltransferases of maize that methylate anthranilic acid using S-adenosyl-l-methionine. Plant Physiol. 2010, 153, 1795–1807. [Google Scholar] [CrossRef] [PubMed]
- Jang, S.J.; Wi, S.J.; Choi, Y.J.; An, G.; Park, K.Y. Increased polyamine biosynthesis enhances stress tolerance by preventing the accumulation of reactiveoxygen species:T-DNA mutational analysis of Oryza sativa lysine decarboxylase-like protein 1. Mol. Cells 2012, 34, 251–262. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Zhang, Y.; Shi, Y.; Zhang, Z.; Zou, Z.; Zhang, H.; Zhao, J. Effect of exogenous spermidine on polyamine content and metabolism in tomato exposed to salinity-alkalinity mixed stress. Plant Physiol. Biochem. 2012, 57, 200–209. [Google Scholar] [CrossRef] [PubMed]
- Gong, B.; Li, X.; VandenLangenberg, K.M.; Wen, D.; Sun, S.; Wei, M.; Li, Y.; Yang, F.; Shi, Q.; Wang, X. Overexpression of S-adenosyl-l-methionine synthetase increased tomato tolerance to alkali stress through polyamine metabolism. Plant Biotechnol. J. 2014, 12, 694–708. [Google Scholar] [CrossRef] [PubMed]
- Guo, Z.; Tan, J.; Zhuo, C.; Wang, C.; Xiang, B.; Wang, Z. Abscisic acid, H2O2 and nitricoxide interactions mediated cold-induced S-adenosylmethionine synthetase in Medicag osativa subsp. falcata that confers cold tolerance through up-regulating polyamine oxidation. Plant Biotechnol. J. 2014, 12, 601–612. [Google Scholar] [CrossRef] [PubMed]
- Qi, Y.C.; Wang, F.F.; Zhang, H.; Liu, W.Q. Overexpression of Suadea salsa S-adenosylmethionine synthetase gene promotes salt tolerance in transgenic tobacco. Acta Physiol. Plant 2010, 32, 263–269. [Google Scholar] [CrossRef]
- Li, H.; Cao, H.; Wang, Y.; Pang, Q.; Ma, C.; Chen, S. Proteomic analysis of sugar beet apomictic monosomic addition line M14. J. Proteom. 2009, 73, 297–308. [Google Scholar] [CrossRef] [PubMed]
- Ge, Y.; He, G.; Wang, Z.; Guo, D.; Qin, R.; Li, R. GISH and BAC-FISH study of apomicitic Beta M14. Sci. China Ser. C Life Sci. 2007, 37, 209–216. [Google Scholar]
- Guo, D.; Kang, C.; Liu, L.; Li, Y. Study of apomixis in theallotriploid beet (VVC). Agric. Sci. China 1999, 32, 1–5. [Google Scholar]
- Wang, Y.; Zhan, Y.; Wu, C.; Gong, S.; Zhu, N.; Chen, S.; Li, H. Cloning of a cystatin gene from sugar beet M14 that can enhance plant salt tolerance. Plant Sci. 2012, 191, 93–99. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Han, Y.; Tao, F.; Chong, K. Knockdown of SAMS genes encoding S-adenosyl-l-methionine synthetases causes methylation alterations of DNAs and histones and leads to late flowering in rice. J. Plant Physiol. 2011, 168, 1837–1843. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.H.; Chae, H.S.; Lee, J.H.; Hwang, B.; Hahn, K.W.; Kang, B.G.; Kim, W.T. Structure and expression of two cDNAs encoding S-adenosyl-l-methionine synthetase of rice (Oryza sativa L.). Biochim. Biophys. Acta 1997, 1354, 13–18. [Google Scholar] [CrossRef]
- Hazarika, P.; Rajam, M.V. Biotic and abiotic stress tolerance in transgenic tomatoes by constitutive expression of S-adenosylmethionine decarboxylase gene. Physiol. Mol. Biol. Plants 2011, 17, 115–128. [Google Scholar] [CrossRef] [PubMed]
- Mao, D.; Yu, F.; Li, J.; Van de Poel, B.; Tan, D.; Li, J.; Liu, Y.; Li, X.; Dong, M.; Chen, L.; et al. FERONIA receptor kinase interacts with S-adenosylmethionine synthetase and suppresses S-adenosylmethionine production and ethylene biosynthesis in Arabidopsis. Plant Cell Environ. 2015, 38, 2566–2574. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Zou, T.; McCormick, S. S-Adenosylmethionine Synthetase 3 is important for pollen tube growth. Plant Physiol. 2016, 172, 244–253. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.H.; Kim, S.H.; Palaniyandi, S.A.; Yang, S.H.; Suh, J.W. Expression of potato S-adenosyl-l-methionine synthase (SbSAMS) gene altered developmental characteristics and stress responses in transgenic Arabidopsis plants. Plant Physiol. Biochem. 2015, 87, 84–91. [Google Scholar] [CrossRef] [PubMed]
- Zago, E.; Morsa, S.; Dat, J.F.; Alard, P.; Ferrarini, A.; Inzé, D.; Delledonne, M.; van Breusegem, F. Nitric oxide-and hydrogen peroxide-responsive gene regulation during cell death induction in tobacco. Plant Physiol. 2006, 141, 404–411. [Google Scholar] [CrossRef] [PubMed]
- Asada, K. The water-water cyclein in chloroplasts: Scavenging of active oxygens and dissipation of excess photons. Annu. Rev. Plant Physiol. Plant. Mol. Biol. 1999, 50, 601–639. [Google Scholar] [CrossRef] [PubMed]
- Diao, Q.N.; Song, Y.J.; Shi, D.M.; Qi, H.Y. Nitric oxide induced by polyamines involves antioxidant systems against chilling stress intomato (Lycopersicon esculentum Mill.) seedling. J. Zhejiang Univ Sci. B 2016, 17, 916–930. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Zeng, J.; Li, Y.; Rong, X.; Sun, J.; Sun, T.; Li, M.; Wang, L.; Feng, Y.; Chai, R.; et al. Expression of TaWRKY44, a wheat WRKY gene, in transgenic tobacco confers multiple abiotic stress tolerances. Front. Plant Sci. 2015, 6, 615. [Google Scholar] [CrossRef] [PubMed]
- Diao, Q.; Song, Y.; Shi, D.; Qi, H. Interaction of polyamines, abscisic acid, nitric oxide, and hydrogen peroxide under chilling stress in tomato (Lycopersicon esculentum Mill.) seedlings. Front. Plant Sci. 2017, 8, 203. [Google Scholar] [CrossRef] [PubMed]
- Ha, H.C.; Sirisoma, N.S.; Kuppusamy, P.; Zweier, J.L.; Woster, P.M.; Casero, R.A., Jr. The natural polyamine spermine functions directly as a free radical scavenger. Proc. Natl. Acad. Sci. USA 1998, 95, 11140–11145. [Google Scholar] [CrossRef] [PubMed]
- Kusano, T.; Berberich, T.; Tateda, C.; Takahashi, Y. Polyamines: Essential factors for growth and survival. Planta 2008, 228, 367–381. [Google Scholar] [CrossRef] [PubMed]
- Botella, M.A.; Xu, Y.; Prabha, T.N.; Zhao, Y.; Narasimhan, M.L.; Wilson, K.A.; Nielsen, S.S.; Bressan, R.A.; Hasegawa, P.M. Differential expression of soybean cysteine proteinase inhibitor genes during development and in response to wounding and methyl jasmonate. Plant Physiol. 1996, 112, 1201–1210. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.D.; Gibson, T.J.; Plewniak, F.; Jeanmougin, F.; Higgins, D.G. The CLUSTAL_X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997, 25, 4876–4882. [Google Scholar] [CrossRef] [PubMed]
- Lichtenthaler, H.K. Chlorophylls and carotenoids: Pigments of photosynthetic biomembranes. Methods Enzymol. 1987, 148, 350–382. [Google Scholar]
- Roeder, S.; Dreschler, K.; Wirtz, M.; Cristescu, S.M.; van Harren, F.J.; Hell, R.; Piechulla, B. SAM levels, gene expression of SAM synthetase, methionine synthase and ACC oxidase, and ethylene emission from N. suaveolens flowers. Plant Mol. Biol. 2009, 70, 535–546. [Google Scholar] [CrossRef] [PubMed]







| Unigene Number | Length (bp) | Annotation | Score | E-Value |
|---|---|---|---|---|
| L1 | 354 | Glutathione S-transferase | 159 | 6 × 10−33 |
| L2 | 231 | Late embryogenesis abundant protein | 73 | 1 × 10−5 |
| L3 | 174 | Chloroplastic chlorophyll a-b binding 8 | 154 | 2 × 10−27 |
| L4 | 150 | ATP-binding cassette transporter C family member 14 | 70 | 3 × 10−11 |
| L5 | 392 | Ca2+ transporting ATPase | 260 | 4 × 10−78 |
| L6 | 237 | MYB (v-myb avian myeloblastosis viral oncogene homolog) transcription factor | 52 | 2 × 10−7 |
| L7 | 359 | Salt-induced hydrophilic protein | 70.3 | 2 × 10−9 |
| L8 | 467 | Aldehyde dehydrogenase family 7 A1 | 119 | 1 × 10−32 |
| L9 | 233 | Cysteine proteinase inhibitor | 134 | 1 × 10−8 |
| L10 | 221 | Aldehyde dehydrogenase | 123 | 3 × 10−13 |
| L11 | 274 | S-adenosylmethionine decarboxylase | 155 | 1 × 10−36 |
| L12 | 148 | Pyruvate kinase family protein | 89 | 4 × 10−18 |
| L13 | 274 | Cysteine protease | 63 | 1 × 10−7 |
| L14 | 297 | Short-chain dehydrogenase | 73 | 7 × 10−11 |
| L15 | 246 | Carboxyl-terminal-processing protease | 132 | 1 × 10−30 |
| L16 | 301 | MYB transcription factor | 94 | 2 × 10−7 |
| L17 | 144 | Short-chain dehydrogenases/reductases family 7C | 198 | 5 × 10−8 |
| L18 | 326 | Vacuole ATPase subunit A | 144 | 3 × 10−66 |
| L19 | 528 | Putative mitochondrial carrier protein | 163 | 5 × 10−77 |
| L20 | 416 | Nitrite reductase | 254 | 4 × 10−66 |
| L21 | 512 | Lipid transfer protein | 184 | 4 × 10−45 |
| L22 | 314 | S-adenosylmethionine synthase 2 | 239 | 1 × 10−61 |
| L23 | 763 | Phosphoglycerate kinase | 407 | 8 × 10−112 |
| L24 | 236 | Cellulose synthase-like protein E6 | 71.2 | 5 × 10−11 |
| L25 | 450 | High-mobility group B6 transcription factor | 69 | 3 × 10−2 |
| L26 | 544 | Heat- and acid-stable phosphoprotein | 95 | 3 × 10−16 |
| L27 | 236 | Thioredoxin domain 2 | 68 | 6 × 10−31 |
| L28 | 437 | Hypothetical protein | 109 | 5 × 10−4 |
| L29 | 197 | DNA replication licensing factor minichromosome maintenance 4 (MCM4) | 88 | 2 × 10−2 |
| R1 | 245 | Thioredoxin domain 1 | 96 | 3 × 10−13 |
| R2 | 364 | S-adenosylmethionine synthase 2 | 194 | 1 × 10−22 |
| R3 | 208 | WD (Trp-Asp) repeat phosphoinositide-interacting | 174 | 6 × 10−30 |
| R4 | 447 | AAA-type ATPase family protein | 143 | 4 × 10−12 |
| R5 | 182 | Putative senescence-associated protein | 121 | 6 × 10−2 |
| R6 | 394 | S-adenosylmethionine synthase 2 | 74 | 3 × 10−7 |
| R7 | 113 | 60S ribosomal protein L19-3 | 863 | 4 × 10−14 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ma, C.; Wang, Y.; Gu, D.; Nan, J.; Chen, S.; Li, H. Overexpression of S-Adenosyl-l-Methionine Synthetase 2 from Sugar Beet M14 Increased Arabidopsis Tolerance to Salt and Oxidative Stress. Int. J. Mol. Sci. 2017, 18, 847. https://doi.org/10.3390/ijms18040847
Ma C, Wang Y, Gu D, Nan J, Chen S, Li H. Overexpression of S-Adenosyl-l-Methionine Synthetase 2 from Sugar Beet M14 Increased Arabidopsis Tolerance to Salt and Oxidative Stress. International Journal of Molecular Sciences. 2017; 18(4):847. https://doi.org/10.3390/ijms18040847
Chicago/Turabian StyleMa, Chunquan, Yuguang Wang, Dan Gu, Jingdong Nan, Sixue Chen, and Haiying Li. 2017. "Overexpression of S-Adenosyl-l-Methionine Synthetase 2 from Sugar Beet M14 Increased Arabidopsis Tolerance to Salt and Oxidative Stress" International Journal of Molecular Sciences 18, no. 4: 847. https://doi.org/10.3390/ijms18040847