Characterization of the Chloroplast Genome of Trentepohlia odorata (Trentepohliales, Chlorophyta), and Discussion of its Taxonomy
Abstract
:1. Introduction
2. Results
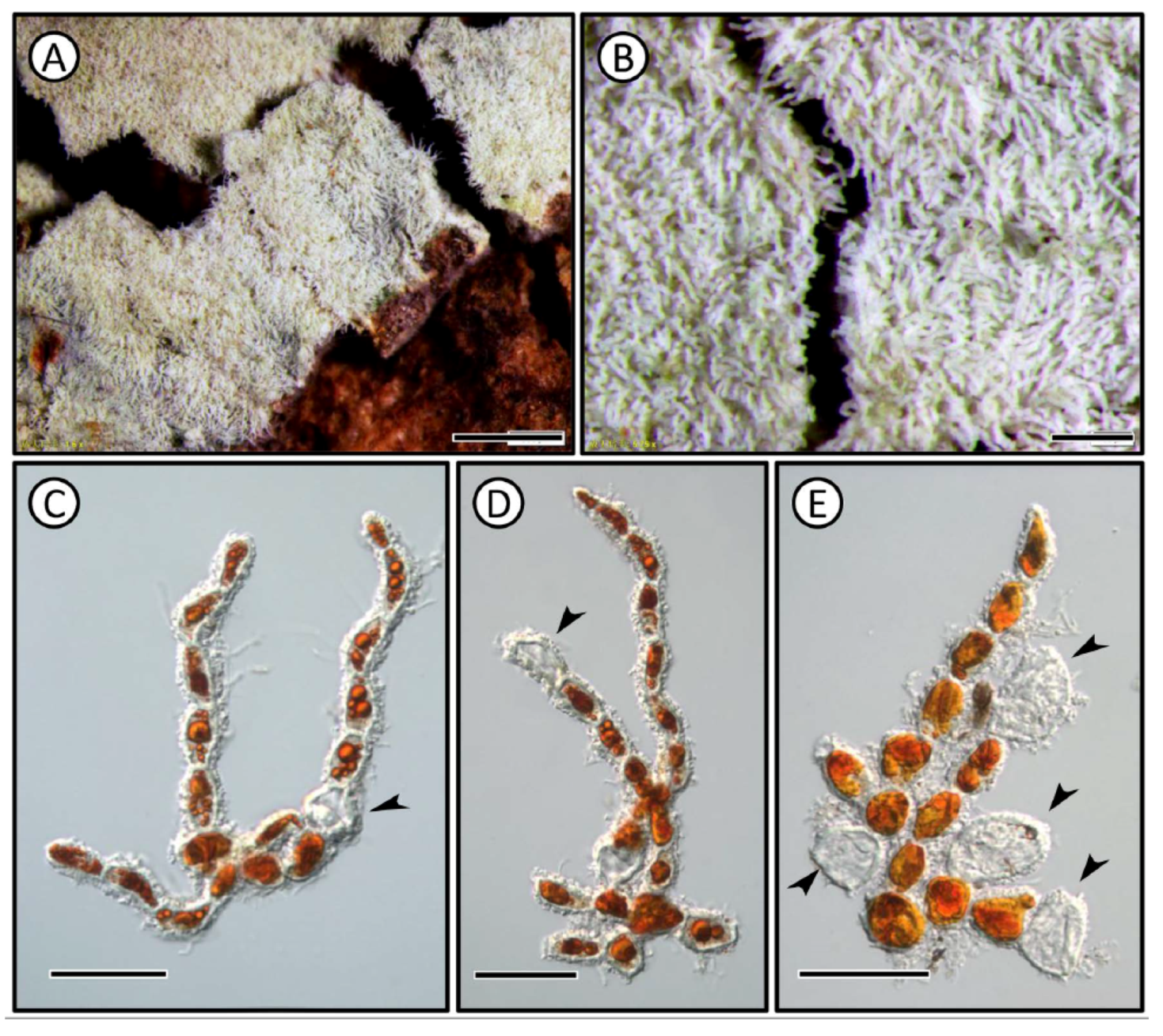
2.1. Morphological Observation
2.2. Phylogenetic Analysis
2.3. Chloroplast Genome Analysis
3. Discussion
4. Materials and Methods
4.1. Morphology and Cultivation
4.2. DNA Extraction, PCR, and Phylogenetic Reconstruction
4.3. Chloroplast Genome Assembly and Annotation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Guiry, M.D.; Guiry, G.M. AlgaeBase. World-Wide Electronic Publication, National University of Ireland, Galway. 2019. Available online: http://www.algaebase.org (accessed on 5 March 2019).
- López-Bautista, J.M.; Rindi, F.; Guiry, M.D. Molecular systematics of the subaerial green algal order Trentepohliales: An assessment based on morphological and molecular data. Int. J. Syst. Evol. Microbiol. 2006, 56, 1709–1715. [Google Scholar] [CrossRef]
- Rindi, F.; Lam, D.W.; López-Bautista, J.M. Phylogenetic relationships and species circumscription in Trentepohlia and Printzina (Trentepohliales, Chlorophyta). Mol. Phylogenetics Evol. 2009, 52, 329–339. [Google Scholar] [CrossRef]
- Nelsen, M.P.; Plata, E.R.; Andrew, C.J.; Lücking, R.; Lumbsch, H.T. Phylogenetic diversity of Trentepohlialean algae Associated with lichen-forming fungi. J. Phycol. 2011, 47, 282–290. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Hu, Z.; Liu, G. Morphology and molecular phylogeny of Trentepohliales (Chlorophyta) from China. Eur. J. Phycol. 2017, 52, 330–341. [Google Scholar] [CrossRef]
- Klimešová, M.; Rindi, F.; Škaloud, P. DNA cloning demonstrates high genetic heterogeneity in populations of the subaerial green alga Trentepohlia (Trentepohliales, Chlorophyta). J. Phycol. 2018, 55, 224–235. [Google Scholar] [CrossRef] [PubMed]
- Hariot, M.P. Notes sur le genre Trentepohlia Martius (Suite). J. Bot. 1889, 3, 345–405. [Google Scholar]
- Cribb, A.B. A revision of some species of Trentepohlia especially from Queensland. Proc. R. Soc. Qld. 1958, 82, 17–34. [Google Scholar]
- Sarma, P. The freshwater Chaetophorales of New Zealand. Nova Hedwigia 1986, 58, 1–169. [Google Scholar]
- Printz, H. Vorarbeiten zu einer Monographie der Trentepohliaceae. Nytt Magazin für Naturvidenskaberne 1939, 80, 137–210. [Google Scholar]
- Fučíková, K.; Leliaert, F.; Cooper, E.D.; Škaloud, P.; Clerck, O.D.; Gurgel, C.F.D.; Lewis, L.A.; Lewis, P.O.; López-Bautistaet, J.M.; Delwiche, C.F.; et al. New phylogenetic hypotheses for the core Chlorophyta based on chloroplast sequence data. Front. Ecol. Evol. 2014, 2, 63. [Google Scholar]
- Turmel, M.; Lemieux, C. Evolution of the plastid genome in green algae. Adv. Bot. Res. 2017, 85, 157–193. [Google Scholar]
- Fang, L.; Leliaert, F.; Novis, P.M.; Zhang, Z.; Zhu, H.; Liu, G.; Penny, D.; Zhong, B. Improving phylogenetic inference of core Chlorophyta using chloroplast sequences with strong phylogenetic signals and heterogeneous models. Mol. Phylogenetics Evol. 2018, 127, 248–255. [Google Scholar] [CrossRef]
- Pombert, J.F.; Lemieux, C.; Turmel, M. The complete chloroplast DNA sequence of the green alga Oltmannsiellopsis viridis reveals a distinctive quadripartite architecture in the chloroplast genome of early diverging ulvophytes. BMC Biol. 2006, 4, 3–4. [Google Scholar] [CrossRef]
- Melton, J.T., III; Leliaert, F.; Tronholm, A.; López-Bautistaet, J.M. The complete chloroplast and mitochondrial genomes of the green macroalga Ulva sp. UNA00071828 (Ulvophyceae, Chlorophyta). PLoS ONE 2015, 10, e0121020. [Google Scholar] [CrossRef]
- Turmel, M.; Otis, C.; Lemieux, C. Mitochondrion-to-chloroplast DNA transfers and intragenomic proliferation of chloroplast group II introns in Gloeotilopsis green algae (Ulotrichales, Ulvophyceae). Genome Biol. Evol. 2016, 8, 2789–2805. [Google Scholar] [CrossRef]
- Cortona, A.D.; Leliaert, F.; Bogaert, K.A.; Turmel, M.; Boedeker, C.; Janouškovec, J.; López-Bautistaet, J.M.; Verbruggen, H.; Vandepoele, K.; Clerck, O.D. The plastid genome in Cladophorales green algae is encoded by hairpin chromosomes. Curr. Biol. 2017, 27, 3771–3782. [Google Scholar] [CrossRef]
- Leliaert, F.; López-Bautistaet, J.M. The chloroplast genomes of Bryopsis plumosa and Tydemania expeditiones (Bryopsidales, Chlorophyta): Compact genomes and genes of bacterial origin. BMC Genom. 2015, 16, 204. [Google Scholar] [CrossRef]
- Mcm, C.; Leliaert, F.; Marcelino, V.R.; Verbruggen, H. Large diversity of nonstandard genes and dynamic evolution of chloroplast genomes in Siphonous green algae (Bryopsidales, Chlorophyta). Genome Biol. Evol. 2018, 10, 1048–1061. [Google Scholar]
- Rindi, F.; Guiry, M.D. Diversity, life history, and ecology of Trentepohlia and Printzina (Trentepohliales, Chlorophyta) in urban habitats in Western Ireland. J. Phycol. 2002, 38, 39–54. [Google Scholar] [CrossRef]
- Ong, B.L.; Lim, M.; Wee, Y.C. Effects of desiccation and illumination on photosynthesis and pigmentation of an edaphic population of Trentepohlia odorata (Chlorophyta). J. Phycol. 1992, 28, 768–772. [Google Scholar] [CrossRef]
- Liu, G.X.; Zhang, Q.; Zhu, H.; Hu, Z.Y. Massive Trentepohlia-Bloom in a glacier valley of Mt. Gongga, China, and a new variety of Trentepohlia (Chlorophyta). PLoS ONE 2012, 7, e37725. [Google Scholar] [CrossRef]
- Brouard, J.S.; Otis, C.; Lemieux, C.; Turmel, M. The exceptionally large chloroplast genome of the green alga Floydiella terrestris illuminates the evolutionary history of the Chlorophyceae. Genome Biol. Evol. 2010, 2, 240–256. [Google Scholar] [CrossRef]
- Brouard, J.S.; Otis, C.; Lemieux, C.; Turmel, M. The Chloroplast genome of the green alga Schizomeris leibleinii (Chlorophyceae) provides evidence for bidirectional DNA replication from a single origin in the Chaetophorales. Genome Biol. Evol. 2011, 3, 505–515. [Google Scholar] [CrossRef]
- Chen, L.; Zhang, L.; Zhang, W.; Liu, T. Comparative analysis of growth and carotenoid accumulation of Trentepohlia arborum in aerial, subaerial, and aquatic cultivation. J. Appl. Phycol. 2015, 27, 1079–1087. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Gouy, M.; Guindon, S.; Gascuel, O. SeaView version 4: A multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol. Biol. Evol. 2010, 27, 221–224. [Google Scholar] [CrossRef]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.F.; Haeseler, A.V.; Jermiin, L.S. ModelFinder: Fast model selection for accurate phylogenetic estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef]
- Lam-Tung, N.; Schmidt, H.A.; Arndt, V.H.; Quang, M.B. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar]
- Ronquist, F.; Teslenko, M.; Paul, V.D.M.; Ayres, D.L.; Darling, A.; Höhna, S.; LArget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Luo, R.; Liu, B.; Xie, Y.; Li, Z.; Huang, W.; Yuan, J.; He, G.; Chen, Y.; Pan, Q.; Liu, Y.; et al. SOAPdenovo2: An empirically improved memory-efficient short-read de novo assembler. Gigascience 2012, 1, 18. [Google Scholar] [CrossRef]
- Koren, S.; Walenz, B.P.; Berlin, K.; Miller, J.R.; Bergman, N.H.; Phillippy, A.M. Canu: Scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017, 27, 722–736. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schaffer, A.A.; Zhang, J.H.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef] [Green Version]
- Krumsiek, J.; Arnold, R.; Rattei, T. Gepard: A rapid and sensitive tool for creating dotplots on genome scale. Bioinformatics 2007, 23, 1026–1028. [Google Scholar] [CrossRef]
- Wyman, S.K.; Jansen, R.K.; Boore, J.L. Automatic annotation of organellar genomes with DOGMA. Bioinformatics 2004, 20, 3252–3255. [Google Scholar] [CrossRef] [Green Version]
- Cantarel, B.L.; Korf, I.; Robb, S.M.C.; Parra, G.; Ross, E.; Moore, B.; Holt, C.; Alvarado, A.S.; Yandell, M. MAKER: An easy-to-use annotation pipeline designed for emerging model organism genomes. Genome Res. 2008, 18, 188–196. [Google Scholar] [CrossRef]
- Lowe, T.M.; Eddy, S.R. tRNAscan-SE: A program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 1997, 25, 955–964. [Google Scholar] [CrossRef]
- Lagesen, K.; Hallin, P.; Rødland, E.A.; Stærfeldt, H.H.; Rognes, T.; Ussery, D.W. RNAmmer: Consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res. 2007, 35, 3100–3108. [Google Scholar] [CrossRef]
- Michel, F.; Umesono, K.; Ozeki, H. Comparative and functional anatomy of group II catalytic introns—A review. Gene 1989, 82, 5–30. [Google Scholar] [CrossRef]
- Michel, F.; Westhof, E. Modelling of the three-dimensional architecture of group I catalytic introns based on comparative sequence analysis. J. Mol. Biol. 1990, 216, 585–610. [Google Scholar] [CrossRef]
- Lohse, M.; Drechsel, O.; Bock, R. OrganellarGenomeDRAW (OGDRAW): A tool for the easy generation of high-quality custom graphical maps of plastid and mitochondrial genomes. Curr. Genet. 2007, 52, 267–274. [Google Scholar] [CrossRef]







| Gene Products | Genes |
|---|---|
| ATP synthase | atpA, B, F, H, I |
| Chlorophyll biosynthesis | chlB, I, L, N |
| Cytochrome b6/f | petA, B, D, G |
| Photosystem I | psaA, B, C, I, J |
| Photosystem II | psbA, B, C, D, E, F, H, I, J, K, L, M, N, T, Z |
| Large subunit ribosomal proteins | rpl2, 5, 12, 14, 16, 20, 23 |
| RNA polymerase | rpoA, B, C1, C2 |
| Small subunit ribosomal proteins | rps2, 3, 4, 7, 8, 9, 11, 12, 14, 18, 19 |
| Unknown function proteins | ycf1, 3 |
| Other proteins | clpp (2 copies), ccsA |
| Rubisco | rbcL |
| Translation factors | tufA, infA |
| Ribosomal RNAs | rrl (2 copies), rrs |
| Transfer RNAs | trnA(TGC), C(GCA), D(GTC), E(TTC), F(GAA), G(TCC), H(GTG), I(AAT), K(TTT), L(CAA), L(TAA), L(TAG), M(CAT) (5 copies), N(GTT), P(TGG), Q(TTG), R(ACG), R(CCT), R(TCG), R(TCT), S(GCT), S(TGA), T(TGT), V(TAC), W(CCA), Y(GTA) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, H.; Hu, Y.; Liu, F.; Hu, Z.; Liu, G. Characterization of the Chloroplast Genome of Trentepohlia odorata (Trentepohliales, Chlorophyta), and Discussion of its Taxonomy. Int. J. Mol. Sci. 2019, 20, 1774. https://doi.org/10.3390/ijms20071774
Zhu H, Hu Y, Liu F, Hu Z, Liu G. Characterization of the Chloroplast Genome of Trentepohlia odorata (Trentepohliales, Chlorophyta), and Discussion of its Taxonomy. International Journal of Molecular Sciences. 2019; 20(7):1774. https://doi.org/10.3390/ijms20071774
Chicago/Turabian StyleZhu, Huan, Yuxin Hu, Feng Liu, Zhengyu Hu, and Guoxiang Liu. 2019. "Characterization of the Chloroplast Genome of Trentepohlia odorata (Trentepohliales, Chlorophyta), and Discussion of its Taxonomy" International Journal of Molecular Sciences 20, no. 7: 1774. https://doi.org/10.3390/ijms20071774





