Analysis of Putative Epigenetic Regulatory Elements in the FXN Genomic Locus
Abstract
:1. Background
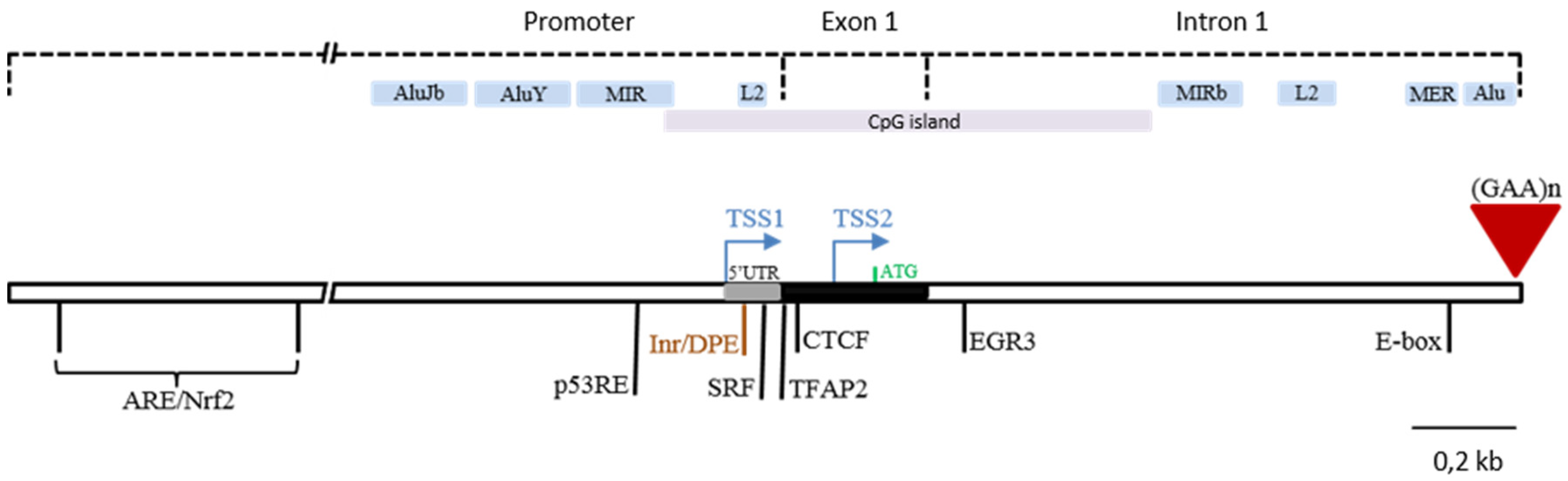
2. Frataxin Gene Regulation
3. Deciphering Regulatory Elements in the FXN Locus
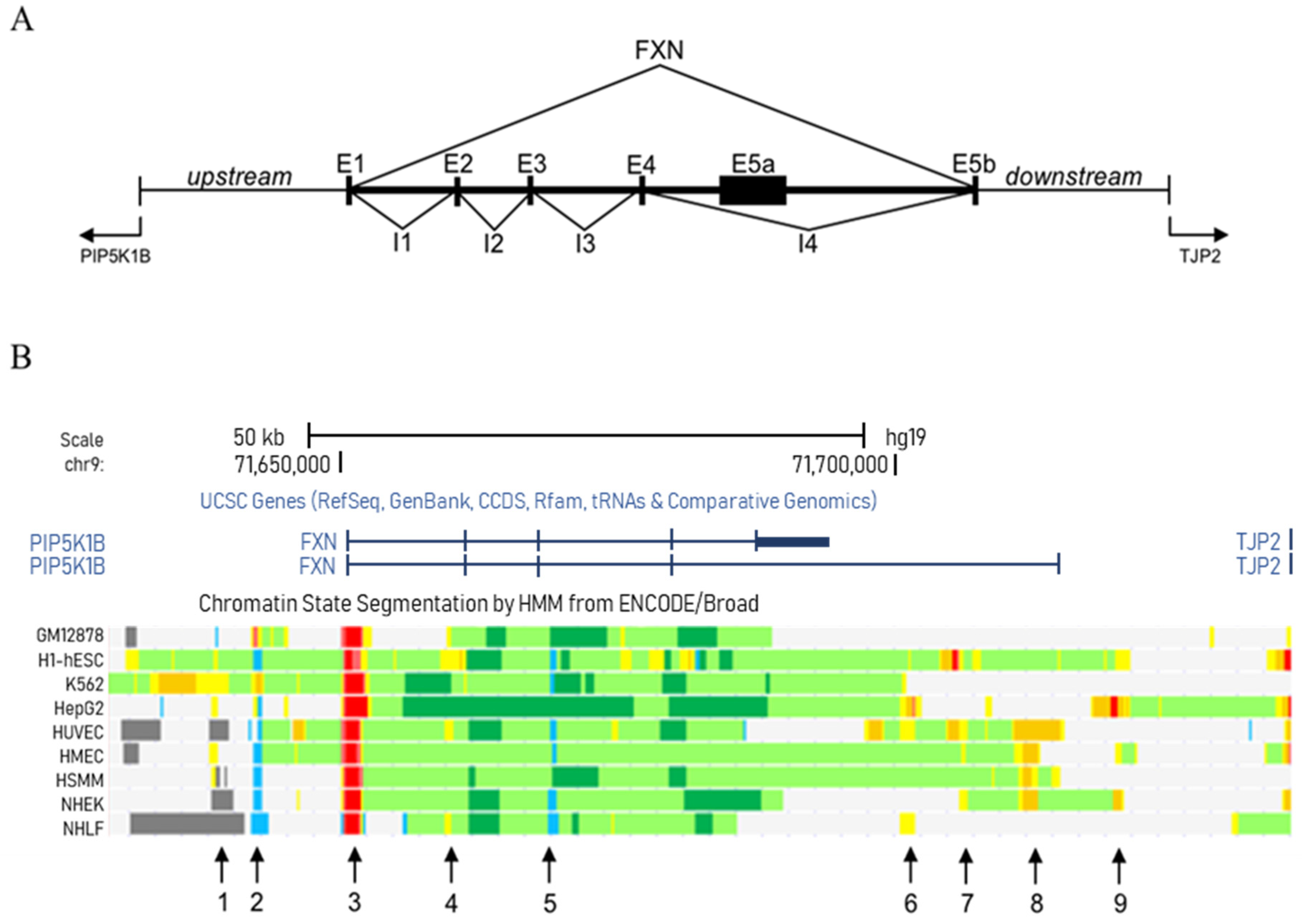
3.1. Study of the FXN Locus in the Cell Lines Provided by the ENCODE Consortium
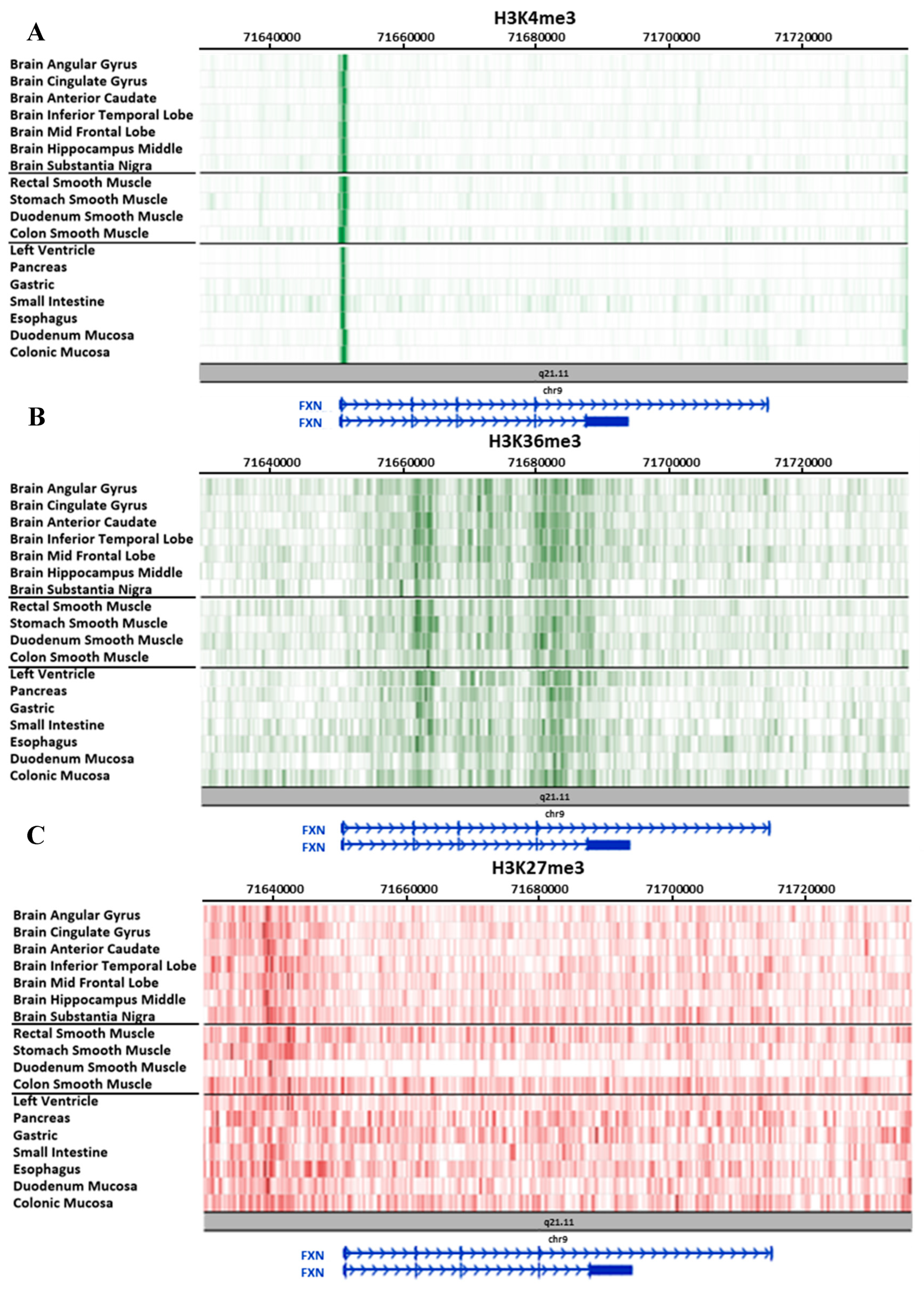
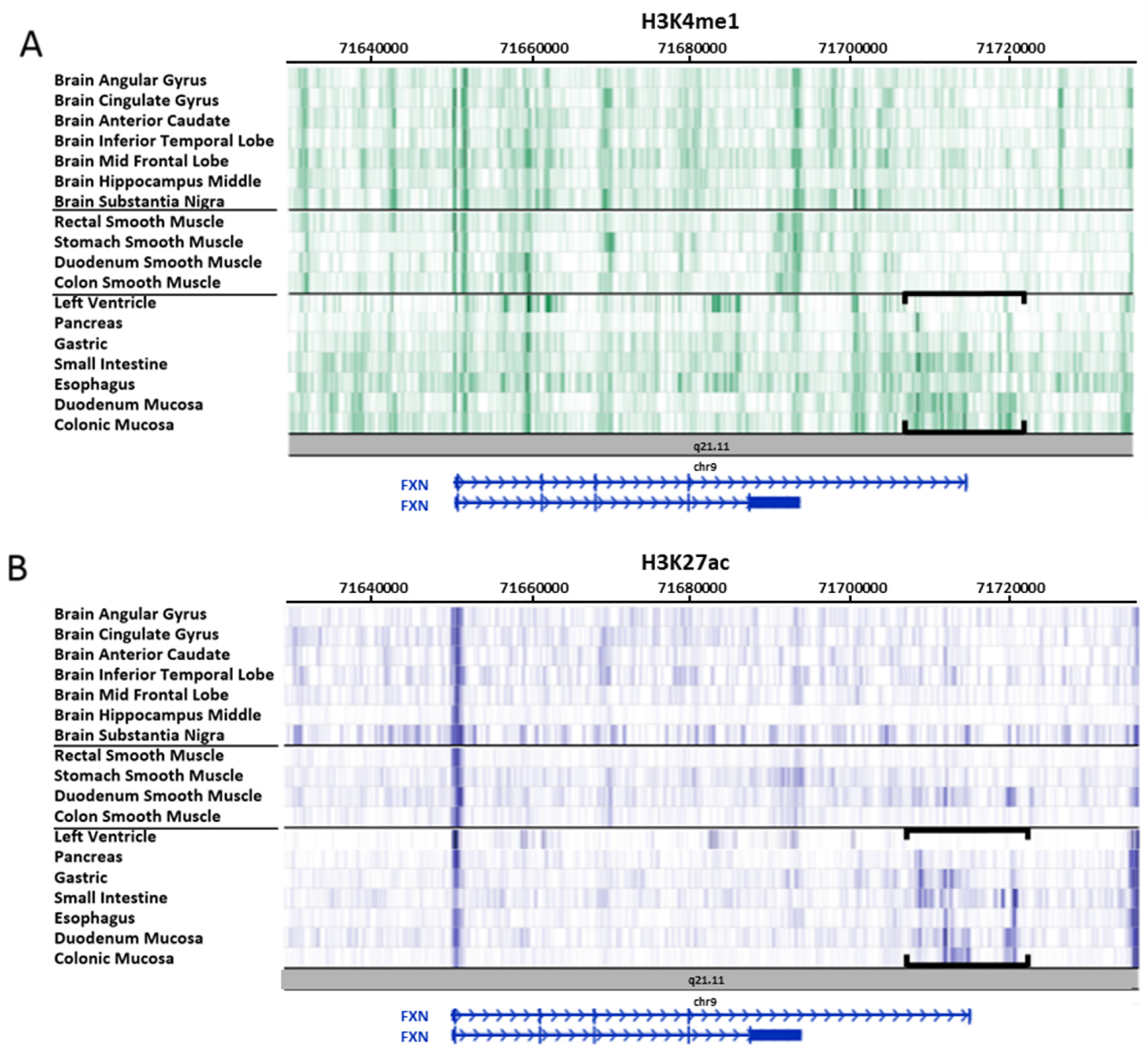
3.2. Study on Tissues and Primary Cultures Provided by the Roadmap Consortium
4. Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Schulz, J.B.; Boesch, S.; Bürk, K.; Durr, A.; Giunti, P.; Mariotti, C.; Pousset, F.; Schöls, L.; Vankan, P.; Pandolfo, M. Diagnosis and treatment of Friedreich ataxia: A European perspective. Nat. Rev. Neurol. 2009, 5, 222–234. [Google Scholar] [CrossRef] [PubMed]
- Pandolfo, M. The molecular basis of Friedreich ataxia. Adv. Exp. Med. Biol. 2002, 516, 99–118. [Google Scholar] [PubMed]
- Bürk, K. Friedreich Ataxia: Current status and future prospects. Cerebellum Ataxias 2017, 4, 4. [Google Scholar] [CrossRef] [Green Version]
- Cook, A.; Giunti, P. Friedreich’s ataxia: Clinical features, pathogenesis and management. Br. Med Bull. 2017, 124, 19–30. [Google Scholar] [CrossRef]
- Campuzano, V.; Montermini, L.; Moltò, M.D.; Pianese, L.; Cossée, M.; Cavalcanti, F.; Monros, E.; Rodius, F.; Duclos, F.; Monticelli, A.; et al. Friedreich’s ataxia: Autosomal recessive disease caused by an intronic GAA triplet repeat expansion. Science 1996, 271, 1423–1427. [Google Scholar] [CrossRef]
- Al-Mahdawi, S.; Ging, H.; Bayot, A.; Cavalcanti, F.B.; La Cognata, V.; Cavallaro, S.; Giunti, P.; Pook, M.A. Large Interruptions of GAA Repeat Expansion Mutations in Friedreich Ataxia Are Very Rare. Front. Cell. Neurosci. 2018, 12. [Google Scholar] [CrossRef] [Green Version]
- Pastore, A.; Puccio, H. Frataxin: A protein in search for a function. J. Neurochem. 2013, 126, 43–52. [Google Scholar] [CrossRef]
- Clark, E.; Butler, J.S.; Isaacs, C.J.; Napierala, M.; Lynch, D.R. Selected missense mutations impair frataxin processing in Friedreich ataxia. Ann. Clin. Transl. Neurol. 2017, 4, 575–584. [Google Scholar] [CrossRef] [Green Version]
- Navarro, J.A.; Llorens, J.V.; Soriano, S.; Botella, J.A.; Schneuwly, S.; Martínez-Sebastián, M.J.; Moltó, M.D. Overexpression of Human and Fly Frataxins in Drosophila Provokes Deleterious Effects at Biochemical, Physiological and Developmental Levels. PLoS ONE 2011, 6, e21017. [Google Scholar] [CrossRef] [Green Version]
- Li, K.; Besse, E.K.; Ha, D.; Kovtunovych, G.; Rouault, T.A. Iron-dependent regulation of frataxin expression: Implications for treatment of Friedreich ataxia. Hum. Mol. Genet. 2008, 17, 2265–2273. [Google Scholar] [CrossRef] [Green Version]
- Pianese, L.; Tammaro, A.; Turano, M.; De Biase, I.; Monticelli, A.; Cocozza, S. Identification of a novel transcript of X25, the human gene involved in Friedreich ataxia. Neurosci. Lett. 2002, 320, 137–140. [Google Scholar] [CrossRef]
- Xia, H.; Cao, Y.; Dai, X.; Marelja, Z.; Zhou, D.; Mo, R.; Al-Mahdawi, S.; Pook, M.A.; Leimkühler, S.; Rouault, T.A.; et al. Novel Frataxin Isoforms May Contribute to the Pathological Mechanism of Friedreich Ataxia. PLoS ONE 2012, 7, e47847. [Google Scholar] [CrossRef] [Green Version]
- Kumari, D.; Biacsi, R.E.; Usdin, K. Repeat Expansion Affects Both Transcription Initiation and Elongation in Friedreich Ataxia Cells. J. Biol. Chem. 2010, 286, 4209–4215. [Google Scholar] [CrossRef] [Green Version]
- Greene, E.; Entezam, A.; Kumari, D.; Usdin, K. Ancient repeated DNA elements and the regulation of the human frataxin promoter. Genomics 2005, 85, 221–230. [Google Scholar] [CrossRef]
- Ågren, J.A.; Clark, A.G. Selfish genetic elements. PLoS Genet. 2018, 14, e1007700. [Google Scholar] [CrossRef]
- Nishibuchi, G.; Déjardin, J. The molecular basis of the organization of repetitive DNA-containing constitutive heterochromatin in mammals. Chromosom. Res. 2017, 25, 77–87. [Google Scholar] [CrossRef]
- Greene, E.; Mahishi, L.; Entezam, A.; Kumari, D.; Usdin, K. Repeat-induced epigenetic changes in intron 1 of the frataxin gene and its consequences in Friedreich ataxia. Nucleic Acids Res. 2007, 35, 3383–3390. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, K.; Singh, A.; Crooks, D.R.; Dai, X.; Cong, Z.; Pan, L.; Ha, D.; Rouault, T.A. Expression of Human Frataxin Is Regulated by Transcription Factors SRF and TFAP2. PLoS ONE 2010, 5, e12286. [Google Scholar] [CrossRef] [PubMed]
- De Biase, I.; Chutake, Y.K.; Rindler, P.M.; Bidichandani, S.I. Epigenetic Silencing in Friedreich Ataxia Is Associated with Depletion of CTCF (CCCTC-Binding Factor) and Antisense Transcription. PLoS ONE 2009, 4, e7914. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bushey, A.M.; Dorman, E.R.; Corces, V.G. Chromatin Insulators: Regulatory Mechanisms and Epigenetic Inheritance. Mol. Cell 2008, 32, 1–9. [Google Scholar] [CrossRef] [Green Version]
- Shimizu, R.; Lan, N.N.; Tai, T.T.; Adachi, Y.; Kawazoe, A.; Mu, A.; Taketani, S. p53 directly regulates the transcription of the human frataxin gene and its lack of regulation in tumor cells decreases the utilization of mitochondrial iron. Gene 2014, 551, 79–85. [Google Scholar] [CrossRef] [PubMed]
- Sahdeo, S.; Scott, B.D.; McMackin, M.Z.; Jasoliya, M.; Brown, B.; Wulff, H.; Perlman, S.L.; Pook, M.A.; Cortopassi, G.A. Dyclonine rescues frataxin deficiency in animal models and buccal cells of patients with Friedreich’s ataxia. Hum. Mol. Genet. 2014, 23, 6848–6862. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, B.; Fritsche, K.L.; Beversdorf, D.Q.; Gu, Z.; Lee, J.C.; Folk, W.R.; Greenlief, C.M.; Sun, G.Y. Yin-Yang Mechanisms Regulating Lipid Peroxidation of Docosahexaenoic Acid and Arachidonic Acid in the Central Nervous System. Front. Neurol. 2019, 10, 642. [Google Scholar] [CrossRef]
- Yang, B.; Li, R.; Greenlief, C.M.; Fritsche, K.; Gu, Z.; Cui, J.; Lee, J.C.; Beversdorf, D.; Sun, G.Y. Unveiling anti-oxidative and anti-inflammatory effects of docosahexaenoic acid and its lipid peroxidation product on lipopolysaccharide-stimulated BV-2 microglial cells. J. Neuroinflamm. 2018, 15, 202. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Inbar-Feigenberg, M.; Choufani, S.; Butcher, D.T.; Roifman, M.; Weksberg, R. Basic concepts of epigenetics. Fertil. Steril. 2013, 99, 607–615. [Google Scholar] [CrossRef] [PubMed]
- Molina-Serrano, D.; Schiza, V.; Kirmizis, A. Cross-talk among epigenetic modifications: Lessons from histone arginine methylation. Biochem. Soc. Trans. 2013, 41, 751–759. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Luger, K.; Mäder, A.W.; Richmond, R.K.; Sargent, D.F.; Richmond, T.J. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 1997, 389, 251–260. [Google Scholar] [CrossRef]
- Murr, R. Interplay between different epigenetic modifications and mechanisms. Adv. Genet. 2010, 70, 101–141. [Google Scholar]
- Musselman, C.A.; LaLonde, M.-E.; Côté, J.; Kutateladze, T. Perceiving the epigenetic landscape through histone readers. Nat. Struct. Mol. Biol. 2012, 19, 1218–1227. [Google Scholar] [CrossRef] [Green Version]
- Chutake, Y.K.; Costello, W.; Lam, C.; Bidichandani, S.I. Altered nucleosome positioning at the transcription start site and deficient transcriptional initiation in Friedreich ataxia. J. Biol. Chem. 2014, 289, 15194–15202. [Google Scholar] [CrossRef] [Green Version]
- Rajeswari, M.R. DNA triplex structures in neurodegenerative disorder, Friedreich’s ataxia. J. Biosci. 2012, 37, 519–532. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, N.; Chastain, P.D.; Parniewski, P.; Ohshima, K.; Pandolfo, M.; Griffith, J.D.; Wells, R.D. Sticky DNA: Self-association properties of long GAA.TTC repeats in R.R.Y triplex structures from Friedreich’s ataxia. Mol. Cell 1999, 3, 465–475. [Google Scholar] [CrossRef]
- Grabczyk, E.; Mancuso, M.; Sammarco, M. A persistent RNA.DNA hybrid formed by transcription of the Friedreich ataxia triplet repeat in live bacteria, and by T7 RNAP in vitro. Nucleic Acids Res. 2007, 35, 5351–5359. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Groh, M.; Lufino, M.M.P.; Wade-Martins, R.; Gromak, N. R-loops Associated with Triplet Repeat Expansions Promote Gene Silencing in Friedreich Ataxia and Fragile X Syndrome. PLoS Genet. 2014, 10, e1004318. [Google Scholar] [CrossRef] [Green Version]
- Saveliev, A.; Everett, C.; Sharpe, T.; Webster, Z.; Festenstein, R. DNA triplet repeats mediate heterochromatin-protein-1-sensitive variegated gene silencing. Nature 2003, 422, 909–913. [Google Scholar] [CrossRef]
- Wells, R.D. DNA triplexes and Friedreich ataxia. FASEB J. 2008, 22, 1625–1634. [Google Scholar] [CrossRef] [Green Version]
- Ohshima, K.; Montermini, L.; Wells, R.D.; Pandolfo, M. Inhibitory Effects of Expanded GAA·TTC Triplet Repeats from Intron I of the Friedreich Ataxia Gene on Transcription and Replicationin Vivo. J. Biol. Chem. 1998, 273, 14588–14595. [Google Scholar] [CrossRef] [Green Version]
- Marmolino, D.; Acquaviva, F. Friedreich’s Ataxia: From the (GAA) n Repeat Mediated Silencing to New Promising Molecules for Therapy. Cerebellum 2009, 8, 245–259. [Google Scholar] [CrossRef]
- Punga, T.; Bühler, M. Long intronic GAA repeats causing Friedreich ataxia impede transcription elongation. EMBO Mol. Med. 2010, 2, 120–129. [Google Scholar] [CrossRef]
- Evans-Galea, M.V.; Carrodus, N.; Rowley, S.; Corben, L.A.; Tai, G.; Saffery, R.; Galati, J.C.; Wong, N.C.; Craig, J.M.; Lynch, D.R.; et al. FXN methylation predicts expression and clinical outcome in Friedreich ataxia. Ann. Neurol. 2012, 71, 487–497. [Google Scholar] [CrossRef]
- Castaldo, I.; Pinelli, M.; Monticelli, A.; Acquaviva, F.; Giacchetti, M.; Filla, A.; Sacchetti, S.; Keller, S.; Avvedimento, V.E.; Chiariotti, L.; et al. DNA methylation in intron 1 of the frataxin gene is related to GAA repeat length and age of onset in Friedreich ataxia patients. J. Med Genet. 2008, 45, 808–812. [Google Scholar] [CrossRef] [PubMed]
- Al-Mahdawi, S.; Pinto, R.M.; Ismail, O.; Varshney, D.; Lymperi, S.; Sandi, C.; Trabzuni, D.; Pook, M.A. The Friedreich ataxia GAA repeat expansion mutation induces comparable epigenetic changes in human and transgenic mouse brain and heart tissues. Hum. Mol. Genet. 2007, 17, 735–746. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rivera, C.M.; Ren, B. Mapping human epigenomes. Cell 2013, 155, 39–55. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nageshwaran, S.; Festenstein, R. Epigenetics and Triplet-Repeat Neurological Diseases. Front. Neurol. 2015, 6, 1010. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sandi, C.; Sandi, M.; Virmouni, S.A.; Al-Mahdawi, S.; Pook, M.A. Epigenetic-based therapies for Friedreich ataxia. Front. Genet. 2014, 5, 165. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Consortium, E.P. An integrated encyclopedia of DNA elements in the human genome. Nature 2012, 489, 57–74. [Google Scholar] [CrossRef]
- Kundaje, A.; Roadmap Epigenomics Consortium; Meuleman, W.; Ernst, J.; Bilenky, M.; Yen, A.; Heravi-Moussavi, A.; Kheradpour, P.; Zhang, Z.; Wang, J.; et al. Integrative analysis of 111 reference human epigenomes. Nature 2015, 518, 317–330. [Google Scholar] [CrossRef] [Green Version]
- The ENCODE Project Consortium The ENCODE (ENCyclopedia Of DNA Elements) Project. Science 2004, 306, 636–640. [CrossRef] [Green Version]
- Luo, Y.; Hitz, B.C.; Gabdank, I.; A Hilton, J.; Kagda, M.S.; Lam, B.; Myers, Z.; Sud, P.; Jou, J.; Lin, K.; et al. New developments on the Encyclopedia of DNA Elements (ENCODE) data portal. Nucleic Acids Res. 2020, 48, D882–D889. [Google Scholar] [CrossRef]
- Thomas, D.J.; Rosenbloom, K.R.; Clawson, H.; Hinrichs, A.S.; Trumbower, H.; Raney, B.J.; Karolchik, N.; Barber, G.P.; Harte, R.A.; Hillman-Jackson, J.; et al. The ENCODE Project at UC Santa Cruz. Nucleic Acids Res. 2006, 35, D663–D667. [Google Scholar] [CrossRef] [Green Version]
- Sloan, C.A.; Chan, E.; Davidson, J.; Malladi, V.; Strattan, J.S.; Hitz, B.C.; Gabdank, I.; Narayanan, A.; Ho, M.; Ho, M.C.; et al. ENCODE data at the ENCODE portal. Nucleic Acids Res. 2015, 44, D726–D732. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Geiman, T.M.; Robertson, K.D. Chromatin remodeling, histone modifications, and DNA methylation?how does it all fit together? J. Cell. Biochem. 2002, 87, 117–125. [Google Scholar] [CrossRef] [PubMed]
- Cavalli, G.; Heard, E. Advances in epigenetics link genetics to the environment and disease. Nature 2019, 571, 489–499. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ernst, J.; Kellis, M. Discovery and characterization of chromatin states for systematic annotation of the human genome. Nat. Biotechnol. 2010, 28, 817–825. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ernst, J.; Kellis, M. ChromHMM: Automating chromatin-state discovery and characterization. Nat. Methods 2012, 9, 215–216. [Google Scholar] [CrossRef] [Green Version]
- Ernst, J.; Kellis, M. Chromatin-state discovery and genome annotation with ChromHMM. Nat. Protoc. 2017, 12, 2478–2492. [Google Scholar] [CrossRef]
- Chan, R.C.; Libbrecht, M.W.; Roberts, E.G.; Bilmes, J.A.; Noble, W.S.; Hoffman, M.M. Segway 2.0: Gaussian mixture models and minibatch training. Bioinformatics 2017, 34, 669–671. [Google Scholar] [CrossRef] [Green Version]
- Hoffman, M.M.; Ernst, J.; Wilder, S.; Kundaje, A.; Harris, R.S.; Libbrecht, M.; Giardine, B.; Ellenbogen, P.M.; Bilmes, J.A.; Birney, E.; et al. Integrative annotation of chromatin elements from ENCODE data. Nucleic Acids Res. 2012, 41, 827–841. [Google Scholar] [CrossRef] [Green Version]
- Ernst, J.; Kheradpour, P.; Mikkelsen, T.S.; Shoresh, N.; Ward, L.D.; Epstein, C.B.; Zhang, X.; Wang, L.; Issner, R.; Coyne, M.; et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 2011, 473, 43–49. [Google Scholar] [CrossRef]
- Osman, E.; Miller, M.R.; Robbins, K.L.; Lombardi, A.M.; Atkinson, A.K.; Brehm, A.J.; Lorson, C.L. Morpholino antisense oligonucleotides targeting intronic repressor Element1 improve phenotype in SMA mouse models. Hum. Mol. Genet. 2014, 23, 4832–4845. [Google Scholar] [CrossRef] [Green Version]
- Ott, C.J.; Blackledge, N.P.; Kerschner, J.L.; Leir, S.-H.; Crawford, G.E.; Cotton, C.U.; Harris, A. Intronic enhancers coordinate epithelial-specific looping of the active CFTR locus. Proc. Natl. Acad. Sci. USA 2009, 106, 19934–19939. [Google Scholar] [CrossRef] [Green Version]
- Belbellaa, B.; Reutenauer, L.; Messaddeq, N.; Monassier, L.; Puccio, H. High levels of frataxin overexpression leads to mitochondrial and cardiac toxicity in mouse models. BioRxiv 2020. [Google Scholar] [CrossRef] [Green Version]
- Vannocci, T.; Manzano, R.N.; Beccalli, O.; Bettegazzi, B.; Grohovaz, F.; Cinque, G.; De Riso, A.; Quaroni, L.; Codazzi, F.; Pastore, A. Adding a temporal dimension to the study of Friedreich’s ataxia: The effect of frataxin overexpression in a human cell model. Dis. Model. Mech. 2018, 11, dmm032706. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Taylor, T.N.; Potgieter, D.; Anwar, S.; Senior, S.L.; Janezic, S.; Threlfell, S.; Ryan, B.; Parkkinen, L.; Deltheil, T.; Cioroch, M.; et al. Region-specific deficits in dopamine, but not norepinephrine, signaling in a novel A30P α-synuclein BAC transgenic mouse. Neurobiol. Dis. 2013, 62, 193–207. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gomez-Sebastian, S.; Gimenez-Cassina, A.; Diaz-Nido, J.; Lim, F.; Wade-Martins, R. Infectious delivery and expression of a 135 kb human FRDA genomic DNA locus complements Friedreich’s ataxia deficiency in human cells. Mol. Ther. 2007, 15, 248–254. [Google Scholar] [CrossRef] [PubMed]
- Giménez-Cassina, A.; Wade-Martins, R.; Gómez-Sebastián, S.; Corona, J.C.; Lim, F.; Diaz-Nido, J. Infectious delivery and long-term persistence of transgene expression in the brain by a 135-kb iBAC-FXN genomic DNA expression vector. Gene Ther. 2011, 18, 1015–1019. [Google Scholar] [CrossRef] [Green Version]
- Gakh, O.; Bedekovics, T.; Duncan, S.F.; Smith, D.Y.; Berkholz, N.S.; Isaya, G. Normal and Friedreich Ataxia Cells Express Different Isoforms of Frataxin with Complementary Roles in Iron-Sulfur Cluster Assembly*. J. Biol. Chem. 2010, 285, 38486–38501. [Google Scholar] [CrossRef] [Green Version]
- Schmucker, S.; Argentini, M.; Carelle-Calmels, N.; Martelli, A.; Puccio, H. The in vivo mitochondrial two-step maturation of human frataxin. Hum. Mol. Genet. 2008, 17, 3521–3531. [Google Scholar] [CrossRef] [Green Version]
- Perez-Luz, S.; Giménez-Cassina, A.; Fernández-Frías, I.; Wade-Martins, R.; Diaz-Nido, J. Delivery of the 135 kb human frataxin genomic DNA locus gives rise to different frataxin isoforms. Genomics 2015, 106, 76–82. [Google Scholar] [CrossRef] [Green Version]
- Fernández-Frías, I.; Pérez-Luz, S.; Díaz-Nido, J. Enhanced Production of Herpes Simplex Virus 1 Amplicon Vectors by Gene Modification and Optimization of Packaging Cell Growth Medium. Mol. Ther. Methods Clin. Dev. 2020, 17, 491–496. [Google Scholar] [CrossRef]
| Chromosome 9 | Chromatin State | |||
|---|---|---|---|---|
| Nº | Start | End | ||
| 1 | 71638200 | 71639000 | Enhancer | |
| 2 | 71642200 | 71643000 | Insulator | |
| 3 | 71650400 | 71652100 | Promoter | |
| 4 | 71659400 | 71660200 | Enhancer | |
| 5 | 71668900 | 71669600 | Insulator | |
| 6 | 71700500 | 71702000 | Enhancer | |
| 7 | 71704200 | 71706600 | Enhancer | |
| 8 | 71710700 | 71713200 | Strong enhancer | |
| 9 | 71719600 | 71721400 | Enhancer | |
| H3K4me1 | |||||||
|---|---|---|---|---|---|---|---|
| Chromosome 9 | Tissues | ||||||
| Region | Start | End | Cerebral | Muscular | Others | ||
| Up | 1 | 71630400 | 71632000 | 12.14 | 13.50 | 9.67 | |
| 2 | 71634700 | 71636000 | 5.71 | 3.75 | 10.17 | ||
| 3 | 71638300 | 71639600 | 8.43 | 5.50 | 10.50 | ||
| 4 | 71642100 | 71643100 | 13.29 | 9.25 | 12.17 | ||
| 5 | 71650000 | 71650700 | 19.71 | 20.25 | 14.67 | ||
| ORF | I1 | 6 | 71651000 | 71652200 | 20.14 | 17.25 | 16.17 |
| 7 | 71659300 | 71660200 | 12.14 | 17.25 | 17.83 | ||
| I2 | 8 | 71661500 | 71662800 | 10.43 | 4.00 | 4.67 | |
| I3 | 9 | 71668400 | 71670400 | 12.29 | 11.50 | 8.67 | |
| 10 | 71678600 | 71681700 | 9.71 | 8.00 | 9.33 | ||
| I4 | 11 | 71683300 | 71686300 | 5.00 | 4.00 | 12.67 | |
| 12 | 71690700 | 71691900 | 5.29 | 11.50 | 5.33 | ||
| 13 | 71693000 | 71694500 | 16.71 | 15.50 | 12.67 | ||
| 14 | 71697600 | 71698700 | 12.50 | 6.50 | 9.67 | ||
| 15 | 71700300 | 71702400 | 11.86 | 8.50 | 15.50 | ||
| 16 | 71703700 | 71705900 | 8.71 | 6.50 | 13.00 | ||
| 17 | 71708600 | 71709200 | 3.86 | 2.50 | 21.33 | ||
| 18 | 71713500 | 71715400 | 5.57 | 4.00 | 13.17 | ||
| Down | 19 | 71718800 | 71721000 | 4.71 | 4.00 | 14.00 | |
| 20 | 71726500 | 71727400 | 12.00 | 2.25 | 6.67 | ||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fernández-Frías, I.; Pérez-Luz, S.; Díaz-Nido, J. Analysis of Putative Epigenetic Regulatory Elements in the FXN Genomic Locus. Int. J. Mol. Sci. 2020, 21, 3410. https://doi.org/10.3390/ijms21103410
Fernández-Frías I, Pérez-Luz S, Díaz-Nido J. Analysis of Putative Epigenetic Regulatory Elements in the FXN Genomic Locus. International Journal of Molecular Sciences. 2020; 21(10):3410. https://doi.org/10.3390/ijms21103410
Chicago/Turabian StyleFernández-Frías, Iván, Sara Pérez-Luz, and Javier Díaz-Nido. 2020. "Analysis of Putative Epigenetic Regulatory Elements in the FXN Genomic Locus" International Journal of Molecular Sciences 21, no. 10: 3410. https://doi.org/10.3390/ijms21103410






