Germline Mutation Enrichment in Pathways Controlling Endothelial Cell Homeostasis in Patients with Brain Arteriovenous Malformation: Implication for Molecular Diagnosis
Abstract
:1. Introduction
2. Results
2.1. WES, Bioinformatic Analysis, and Filtering
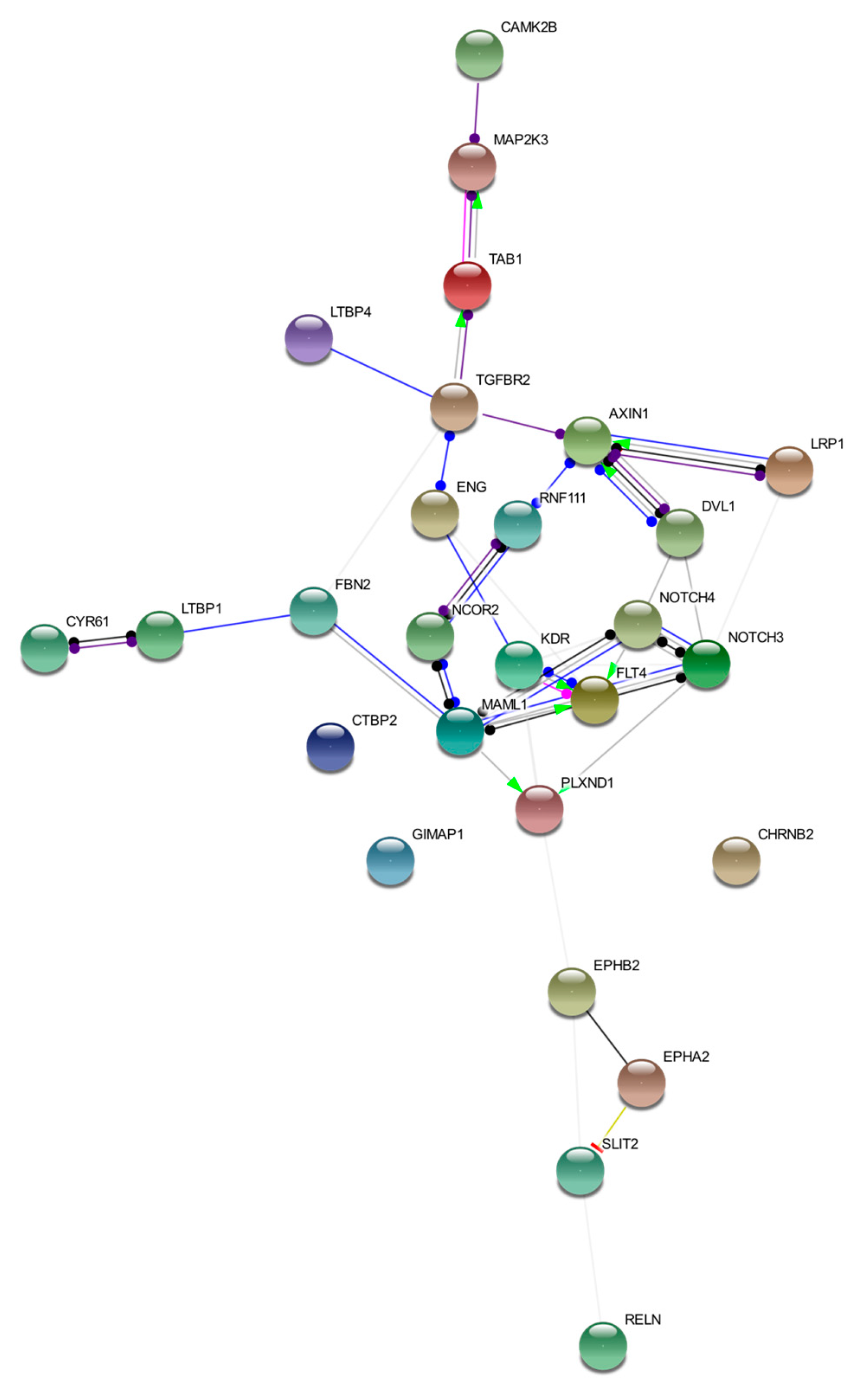
2.2. Gene Clustering and Prioritization
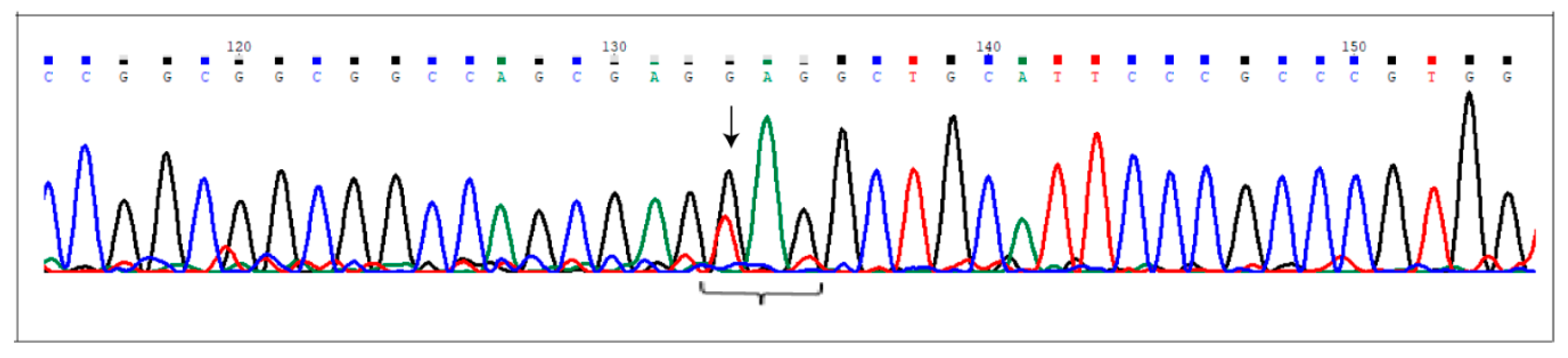
2.3. Sanger Validation
3. Discussion
3.1. AVM1
3.2. AVM2
3.3. AVM3
3.4. AVM4
3.5. AVM5
3.6. Novel Insights
3.7. Final Considerations
4. Materials and Methods
4.1. Patient Recruitment and WES Analysis
4.2. Bioinformatic Analysis
4.3. Variant Filtering Criteria
4.4. Gene Clustering and Prioritization
4.5. Sanger Validation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Winkler, E.A.; Birk, H.; Burkhardt, J.K.; Chen, X.; Yue, J.K.; Guo, D.; Rutledge, W.C.; Lasker, G.F.; Partow, C.; Tihan, T.; et al. Reductions in Brain Pericytes Are Associated With Arteriovenous Malformation Vascular Instability. J. Neurosurg. 2018, 129, 1464–1474. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ozpinar, A.; Mendez, G.; Abla, A.A. Epidemiology, genetics, pathophysiology, and prognostic classifications of cerebral arteriovenous malformations. Handb. Clin. Neurol. 2017, 143, 5–13. [Google Scholar] [PubMed]
- Zhang, D.; Zhou, F.; Zhao, X.; Bao, B.; Chen, J.; Yang, J. Endoglin is a conserved regulator of vasculogenesis in zebrafish—Implications for hereditary haemorrhagic telangiectasia. Biosci. Rep. 2019, 39. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Crist, A.M.; Lee, A.R.; Patel, N.R.; Westhoff, D.E.; Meadows, S.M. Vascular Deficiency of Smad4 Causes Arteriovenous Malformations: A Mouse Model of Hereditary Hemorrhagic Telangiectasia. Angiogenesis 2018, 21, 363–380. [Google Scholar] [CrossRef] [Green Version]
- Wooderchak-Donahue, W.L.; McDonald, J.; O’Fallon, B.; Upton, P.D.; Li, W.; Roman, B.L.; Young, S.; Plant, P.; Fülöp, G.T.; Langa, C.; et al. BMP9 Mutations Cause a Vascular-Anomaly Syndrome With Phenotypic Overlap With Hereditary Hemorrhagic Telangiectasia. Am. J. Hum. Genet. 2013, 93, 530–537. [Google Scholar] [CrossRef] [Green Version]
- Sasahara, A.; Kasuya, H.; Akagawa, H.; Ujiie, H.; Kubo, O.; Sasaki, T.; Onda, H.; Sakamoto, Y.; Krischek, B.; Hori, T.; et al. Increased expression of ephrin A1 in brain arteriovenous malformation: DNA microarray analysis. Neurosurg. Rev. 2007, 30, 299–305. [Google Scholar] [CrossRef] [Green Version]
- Bai, J.; Wang, Y.J.; Liu, L.; Zhao, Y.L. Ephrin B2 and EphB4 selectively mark arterial and venous vessels in cerebral arteriovenous malformation. J. Int. Med. Res. 2014, 42, 405–415. [Google Scholar] [CrossRef]
- Fang, J.; Hirschi, K. Molecular regulation of arteriovenous endothelial cell specification. F1000Research 2019, 8, F1000 Faculty Rev-1208. [Google Scholar] [CrossRef] [Green Version]
- Vaser, R.; Adusumalli, S.; Leng, S.N.; Sikic, M.; Ng, P.C. SIFT missense predictions for genomes. Nat. Protoc. 2016, 11, 1–9. [Google Scholar] [CrossRef]
- Schwarz, J.M.; Cooper, D.N.; Schuelke, M.; Seelow, D. MutationTaster2: Mutation prediction for the deep-sequencing age. Nat. Methods 2014, 11, 361–362. [Google Scholar] [CrossRef]
- Chen, X.; Liu, Y.; Zhou, S.; Nie, S.; Lin, Z.; Zhou, C.; Sun, J.; Gao, X.; Huang, Y. Methylation of the CDKN2A Gene Increases the Risk of Brain Arteriovenous Malformations. J. Mol. Neurosci. 2019, 69, 316–323. [Google Scholar] [CrossRef] [PubMed]
- Thomas, J.M.; Surendran, S.; Abraham, M.; Rajavelu, A.; Kartha, C.C. Genetic and epigenetic mechanisms in the development of arteriovenous malformations in the brain. Clin. Epigenetics 2016, 8, 78. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tual-Chalot, S.; Garcia-Collado, M.; Redgrave, R.E.; Singh, E.; Davison, B.; Park, C.; Lin, H.; Luli, S.; Jin, Y.; Wang, Y.; et al. Loss of Endothelial Endoglin Promotes High-Output Heart Failure Through Peripheral Arteriovenous Shunting Driven by VEGF Signaling. Circ. Res. 2020, 126, 243–257. [Google Scholar] [CrossRef] [PubMed]
- Delev, D.; Pavlova, A.; Grote, A.; Boström, A.; Höllig, A.; Schramm, J.; Fimmers, R.; Oldenburg, J.; Simon, M. NOTCH4 gene polymorphisms as potential risk factors for brain arteriovenous malformation development and hemorrhagic presentation. J. Neurosurg. 2017, 126, 1552–1559. [Google Scholar] [CrossRef]
- Goss, J.A.; Huang, A.Y.; Smith, E.; Konczyk, D.J.; Smits, P.J.; Sudduth, C.L.; Stapleton, C.; Patel, A.; Alexandrescu, S.; Warman, M.L.; et al. Somatic mutations in intracranial arteriovenous malformations. PLoS ONE 2019, 14, e0226852. [Google Scholar] [CrossRef] [Green Version]
- Al-Olabi, L.; Polubothu, S.; Dowsett, K.; Andrews, K.A.; Stadnik, P.; Joseph, A.P.; Knox, R.; Pittman, A.; Clark, G.; Baird, W.; et al. Mosaic RAS/MAPK Variants Cause Sporadic Vascular Malformations Which Respond to Targeted Therapy. J. Clin. Investig. 2018, 128, 1496–1508. [Google Scholar] [CrossRef]
- Priemer, D.S.; Vortmeyer, A.O.; Zhang, S.; Chang, H.Y.; Curless, K.L.; Cheng, L. Activating KRAS mutations in arteriovenous malformations of the brain: Frequency and clinicopathologic correlation. Hum. Pathol. 2019, 89, 33–39. [Google Scholar] [CrossRef]
- Davis, M.R.; Andersson, R.; Severin, J.; de Hoon, M.; Bertin, N.; Baillie, J.K.; Kawaji, H.; Sandelin, A.; Forrest, A.R.; Summers, K.M. FANTOM Consortium: Transcriptional profiling of the human fibrillin/LTBP gene family, key regulators of mesenchymal cell functions. Mol. Genet. Metab. 2014, 112, 73–83. [Google Scholar] [CrossRef]
- Doetschman, T.; Barnett, J.V.; Runyan, R.B.; Camenisch, T.D.; Heimark, R.L.; Granzier, H.L.; Conway, S.J.; Azhar, M. Transforming growth factor beta signaling in adult cardiovascular diseases and repair. Cell Tissue Res. 2012, 347, 203–223. [Google Scholar] [CrossRef] [Green Version]
- Vézina, A.; Charfi, C.; Zgheib, A.; Annabi, B. Cerebrovascular Angiogenic Reprogramming upon LRP1 Repression: Impact on Sphingosine-1-Phosphate-Mediated Signaling in Brain Endothelial Cell Chemotactism. Mol. Neurobiol. 2018, 55, 3551–3563. [Google Scholar] [CrossRef]
- Nakajima, C.; Haffner, P.; Goerke, S.M.; Zurhove, K.; Adelmann, G.; Frotscher, M.; Herz, J.; Bock, H.H.; May, P. The lipoprotein receptor LRP1 modulates sphingosine-1-phosphate signaling and is essential for vascular development. Development 2014, 141, 4513–4525. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grimsey, N.J.; Lin, Y.; Narala, R.; Rada, C.C.; Mejia-Pena, H.; Trejo, J. G protein-coupled receptors activate p38 MAPK via a non-canonical TAB1-TAB2- and TAB1-TAB3-dependent pathway in endothelial cells. J. Biol. Chem. 2019, 294, 5867–5878. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sylvain-Prévost, S.; Ear, T.; Simard, F.A.; Fortin, C.F.; Dubois, C.M.; Flamand, N.; McDonald, P.P. Activation of TAK1 by Chemotactic and Growth Factors, and Its Impact on Human Neutrophil Signaling and Functional Responses. J. Immunol. 2015, 195, 5393–5403. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Di, Y.; Li, S.; Wang, L.; Zhang, Y.; Dorf, M.E. Homeostatic interactions between MEKK3 and TAK1 involved in NF-kappaB signaling. Cell. Signal. 2008, 20, 705–713. [Google Scholar] [CrossRef] [Green Version]
- Noshiro, S.; Mikami, T.; Kataoka-Sasaki, Y.; Sasaki, M.; Hashi, K.; Ohtaki, S.; Wanibuchi, M.; Mikuni, N.; Kocsis, J.D.; Honmou, O. Biological relevance of tissue factor and IL-6 in arteriovenous malformations. Neurosurg. Rev. 2017, 40, 359–367. [Google Scholar] [CrossRef]
- Shoemaker, L.D.; Fuentes, L.F.; Santiago, S.M.; Allen, B.M.; Cook, D.J.; Steinberg, G.K.; Chang, S.D. Human brain arteriovenous malformations express lymphatic-associated genes. Ann. Clin. Transl. Neurol. 2014, 1, 982–995. [Google Scholar] [CrossRef]
- Murphy, P.A.; Kim, T.N.; Huang, L.; Nielsen, C.M.; Lawton, M.T.; Adams, R.H.; Schaffer, C.B.; Wang, R.A. Constitutively active Notch4 receptor elicits brain arteriovenous malformations through enlargement of capillary-like vessels. Proc. Natl. Acad. Sci. USA 2014, 111, 18007–18012. [Google Scholar] [CrossRef] [Green Version]
- Hou, Y.; Shin, Y.J.; Han, E.J.; Choi, J.S.; Park, J.M.; Cha, J.H.; Choi, J.Y.; Lee, M.Y. Distribution of vascular endothelial growth factor receptor-3/Flt4 mRNA in adult rat central nervous system. J. Chem. Neuroanat. 2011, 42, 56–64. [Google Scholar] [CrossRef]
- Chaqour, B. Caught between a “Rho” and a hard place: Are CCN1/CYR61 and CCN2/CTGF the arbiters of microvascular stiffness? J. Cell Commun. Signal. 2019. [Google Scholar] [CrossRef]
- Adams, R.H.; Wilkinson, G.A.; Weiss, C.; Diella, F.; Gale, N.W.; Deutsch, U.; Risau, W.; Klein, R. Roles of ephrinB ligands and EphB receptors in cardiovascular development: Demarcation of arterial/venous domains, vascular morphogenesis, and sprouting angiogenesis. Genes Dev. 1999, 13, 295–306. [Google Scholar] [CrossRef]
- Liu, H.; Devraj, K.; Möller, K.; Liebner, S.; Hecker, M.; Korff, T. EphrinB-mediated reverse signalling controls junctional integrity and pro-inflammatory differentiation of endothelial cells. Thromb. Haemost. 2014, 112, 151–163. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, N.; Zhao, W.D.; Liu, D.X.; Liang, Y.; Fang, W.G.; Li, B.; Chen, Y.H. Inactivation of EphA2 promotes tight junction formation and impairs angiogenesis in brain endothelial cells. Microvasc. Res. 2011, 82, 113–121. [Google Scholar] [CrossRef] [PubMed]
- Doronzo, G.; Astanina, E.; Corà, D.; Chiabotto, G.; Comunanza, V.; Noghero, A.; Neri, F.; Puliafito, A.; Primo, L.; Spampanato, C.; et al. TFEB controls vascular development by regulating the proliferation of endothelial cells. EMBO J. 2019, 38, e98250. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Oh, W.J.; Gaiano, N.; Yoshida, Y.; Gu, C. Semaphorin 3E-Plexin-D1 signaling regulates VEGF function in developmental angiogenesis via a feedback mechanism. Genes Dev. 2011, 25, 1399–1411. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gu, C.; Yoshida, Y.; Livet, J.; Reimert, D.V.; Mann, F.; Merte, J.; Henderson, C.E.; Jessell, T.M.; Kolodkin, A.L.; Ginty, D.D. Semaphorin 3E and plexin-D1 control vascular pattern independently of neuropilins. Science 2005, 307, 265–268. [Google Scholar] [CrossRef]
- Li, G.J.; Yang, Y.; Yang, G.K.; Wan, J.; Cui, D.L.; Ma, Z.H.; Du, L.J.; Zhang, G.M. Slit2 suppresses endothelial cell proliferation and migration by inhibiting the VEGF-Notch signaling pathway. Mol. Med. Rep. 2017, 15, 1981–1988. [Google Scholar] [CrossRef] [Green Version]
- Zhang, S.; Kim, J.Y.; Xu, S.; Liu, H.; Yin, M.; Koroleva, M.; Guo, J.; Pei, X.; Jin, Z.G. Endothelial-specific YY1 governs sprouting angiogenesis through directly interacting with RBPJ. Proc. Natl. Acad. Sci. USA 2020, 117, 4792–4801. [Google Scholar] [CrossRef]
- Koinuma, D.; Shinozaki, M.; Komuro, A.; Goto, K.; Saitoh, M.; Hanyu, A.; Ebina, M.; Nukiwa, T.; Miyazawa, K.; Imamura, T.; et al. Arkadia amplifies TGF-beta superfamily signalling through degradation of Smad7. EMBO J. 2003, 22, 6458–6470. [Google Scholar] [CrossRef] [Green Version]
- Zhou, P.; Wan, X.; Zou, Y.; Chen, Z.; Zhong, A. Transforming growth factor beta (TGF-β) is activated by the CtBP2-p300-AP1 transcriptional complex in chronic renal failure. Int. J. Biol. Sci. 2020, 16, 204–215. [Google Scholar] [CrossRef] [Green Version]
- Villa, C.; Colombo, G.; Meneghini, S.; Gotti, C.; Moretti, M.; Ferini-Strambi, L.; Chisci, E.; Giovannoni, R.; Becchetti, A.; Combi, R. CHRNA2 and Nocturnal Frontal Lobe Epilepsy: Identification and Characterization of a Novel Loss of Function Mutation. Front. Mol. Neurosci. 2019, 12, 17. [Google Scholar] [CrossRef] [Green Version]
- Heeschen, C.; Weis, M.; Aicher, A.; Dimmeler, S.; Cooke, J.P. A novel angiogenic pathway mediated by non-neuronal nicotinic acetylcholine receptors. J. Clin. Investig. 2002, 110, 527–536. [Google Scholar] [CrossRef]
- Peghaire, C.; Bats, M.L.; Sewduth, R.; Jeanningros, S.; Jaspard, B.; Couffinhal, T.; Duplàa, C.; Dufourcq, P. Fzd7 (Frizzled-7) Expressed by Endothelial Cells Controls Blood Vessel Formation Through Wnt/β-Catenin Canonical Signaling. Arterioscler. Thromb. Vasc. Biol. 2016, 36, 2369–2380. [Google Scholar] [CrossRef] [Green Version]
- Liu, W.; Rui, H.; Wang, J.; Lin, S.; He, Y.; Chen, M.; Li, Q.; Ye, Z.; Zhang, S.; Chan, S.C.; et al. Axin is a scaffold protein in TGF-beta signaling that promotes degradation of Smad7 by Arkadia. EMBO J. 2006, 25, 1646–1658. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jensen, L.D.; Hot, B.; Ramsköld, D.; Germano, R.F.V.; Yokota, C.; Giatrellis, S.; Lauschke, V.M.; Hubmacher, D.; Li, M.X.; Hupe, M.; et al. Disruption of the Extracellular Matrix Progressively Impairs Central Nervous System Vascular Maturation Downstream of β-Catenin Signaling. Arterioscler. Thromb. Vasc. Biol. 2019, 39, 1432–1447. [Google Scholar] [CrossRef] [PubMed]
- Eisa-Beygi, S.; Benslimane, F.M.; El-Rass, S.; Prabhudesai, S.; Abdelrasoul, M.K.A.; Simpson, P.M.; Yalcin, H.C.; Burrows, P.E.; Ramchandran, R. Characterization of Endothelial Cilia Distribution During Cerebral-Vascular Development in Zebrafish (Danio rerio). Arterioscler. Thromb. Vasc. Biol. 2018, 38, 2806–2818. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Noren, D.P.; Chou, W.H.; Lee, S.H.; Qutub, A.A.; Warmflash, A.; Wagner, D.S.; Popel, A.S.; Levchenko, A. Endothelial cells decode VEGF-mediated Ca2+ signaling patterns to produce distinct functional responses. Sci. Signal. 2016, 9, ra20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yamamura, H.; Suzuki, Y.; Yamamura, H.; Asai, K.; Imaizumi, Y. Hypoxic stress up-regulates Kir2.1 expression and facilitates cell proliferation in brain capillary endothelial cells. Biochem. Biophys. Res. Commun. 2016, 476, 386–392. [Google Scholar] [CrossRef] [PubMed]
- Yamazaki, D.; Kito, H.; Yamamoto, S.; Ohya, S.; Yamamura, H.; Asai, K.; Imaizumi, Y. Contribution of K(ir)2 potassium channels to ATP-induced cell death in brain capillary endothelial cells and reconstructed HEK293 cell model. Am. J. Physiol. Cell. Physiol. 2011, 300, C75–C86. [Google Scholar] [CrossRef]
- Hauer, A.J.; Kleinloog, R.; Giuliani, F.; Rinkel, G.J.E.; de Kort, G.A.; Berkelbach van der Sprenkel, J.W.; van der Zwan, A.; Gosselaar, P.H.; van Rijen, P.C.; de Boer-Bergsma, J.J.; et al. RNA-Sequencing Highlights Inflammation and Impaired Integrity of the Vascular Wall in Brain Arteriovenous Malformations. Stroke 2020, 51, 268–274. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [Green Version]
- Wang, K.; Zhao, S.; Liu, B.; Zhang, Q.; Li, Y.; Liu, J.; Shen, Y.; Ding, X.; Lin, J.; Wu, Y.; et al. Perturbations of BMP/TGF-β and VEGF/VEGFR signalling pathways in non-syndromic sporadic brain arteriovenous malformations (BAVM). J. Med. Genet. 2018, 55, 675–684. [Google Scholar] [CrossRef] [PubMed]
- Scimone, C.; Donato, L.; Alafaci, C.; Granata, F.; Rinaldi, C.; Longo, M.; D’Angelo, R.; Sidoti, A. High-Throughput Sequencing to Detect Novel Likely Gene-Disrupting Variants in Pathogenesis of Sporadic Brain Arteriovenous Malformations. Front. Genet. 2020, 11, 146. [Google Scholar] [CrossRef] [PubMed]
- Guijarro-Muñoz, I.; Cuesta, A.M.; Alvarez-Cienfuegos, A.; Geng, J.G.; Alvarez-Vallina, L.; Sanz, L. The axonal repellent Slit2 inhibits pericyte migration: Potential implications in angiogenesis. Exp. Cell Res. 2012, 318, 371–378. [Google Scholar] [CrossRef]
- Harde, E.; Nicholson, L.; Furones Cuadrado, B.; Bissen, D.; Wigge, S.; Urban, S.; Segarra, M.; Ruiz de Almodóvar, C.; Acker-Palmer, A. EphrinB2 regulates VEGFR2 during dendritogenesis and hippocampal circuitry development. Elife 2019, 8, e49819. [Google Scholar] [CrossRef]
- Spetzler, R.F.; Martin, N.A. A Proposed Grading System for Arteriovenous Malformations. J. Neurosurg. 1986, 65, 476–483. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [Green Version]
- Garrison, E.; Marth, G. Haplotype-based variant detection from short-read sequencing. arXiv 2012, arXiv:1207.3907v2. [Google Scholar]
- Wang, K.; Li, M.; Hakonarson, H. ANNOVAR: Functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010, 38, e164. [Google Scholar] [CrossRef]
- Karczewski, K.J.; Francioli, L.C.; Tiao, G.; Cummings, B.B.; Alföldi, J.; Wang, Q.; Collins, R.L.; Laricchia, K.M.; Ganna, A.; Birnbaum, D.P.; et al. Variation across 141,456 human exomes and genomes reveals the spectrum of loss-of-function intolerance across human protein-coding genes. BioRxiv 2019, 531210. [Google Scholar] [CrossRef] [Green Version]
- 1000 Genomes Project Consortium; Auton, A.; Brooks, L.D.; Durbin, R.M.; Garrison, E.P.; Kang, H.M.; Korbel, J.O.; Marchini, J.L.; McCarthy, S.; McVean, G.A.; et al. A global reference for human genetic variation. Nature 2015, 526, 68–74. [Google Scholar]
- Bindea, G.; Mlecnik, B.; Hackl, H.; Charoentong, P.; Tosolini, M.; Kirilovsky, A.; Fridman, W.H.; Pagès, F.; Trajanoski, Z.; Galon, J. ClueGO: A cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 2009, 25, 1091–1093. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, J.; Bardes, E.E.; Aronow, B.J.; Jegga, A.G. ToppGene Suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res. 2009, 37, W305–W311. [Google Scholar] [CrossRef] [PubMed]
| Sample | GO ID | GO Term | Ontology Source | Term p-Value Corrected with Bonferroni Step Down | Group p-Value Corrected with Bonferroni Step Down | Associated Genes Found |
|---|---|---|---|---|---|---|
| AVM1 | GO:0003341 | cilium movement | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.00 | 0.00 | (ASPM, CFAP206, DNAH1, DNAH5, HYDIN, TEKT5) |
| GO:0035082 | axoneme assembly | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.01 | 0.00 | (CFAP206, DNAH1, DNAH5, HYDIN, RP1L1) | |
| R-HSA:2129379 | molecules associated with elastic fibers | REACTOME_Pathways_27.02.2019 | 0.01 | 0.05 | (FBN2, FBN3, LTBP1, LTBP4) | |
| GO:2000105 | positive regulation of DNA-dependent DNA replication | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.01 | 0.14 | (CDC7, FLG, SYTL2) | |
| GO:0097722 | sperm motility | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.02 | 0.00 | (ASPM, CACNA1I, DNAH1, DNAH5, SLC22A16, TEKT5) | |
| R-HSA:1566948 | elastic fiber formation | REACTOME_Pathways_27.02.2019 | 0.02 | 0.05 | (FBN2, FBN3, LTBP1, LTBP4) | |
| GO:0030317 | flagellated sperm motility | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.02 | 0.00 | (ASPM, CACNA1I, DNAH1, DNAH5, SLC22A16, TEKT5) | |
| GO:0018410 | C-terminal protein amino acid modification | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.03 | 0.02 | (ASPM, LCMT2, SH2B1) | |
| GO:0005044 | scavenger receptor activity | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.03 | 0.03 | (LRP1, MEGF10, STAB1) | |
| GO:1903078 | positive regulation of protein localization to plasma membrane | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.03 | 0.03 | (CARD14, LRP1, NKD2) | |
| WP:3668 | hypothesized pathways in pathogenesis of cardiovascular disease | WikiPathways_27.02.2019 | 0.04 | 0.05 | (FBN2, FBN3, LTBP1) | |
| GO:0001578 | microtubule bundle formation | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.05 | 0.00 | (CFAP206, DNAH1, DNAH5, HYDIN, RP1L1) | |
| GO:1903392 | negative regulation of adherens junction organization | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.05 | 0.07 | (DLEC1, LRP1, TEP1) | |
| GO:0051895 | negative regulation of focal adhesion assembly | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.05 | 0.07 | (DLEC1, LRP1, TEP1) | |
| AVM2 | GO:0031122 | cytoplasmic microtubule organization | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.03 | 0.02 | (GOLGA4, KIF19, MBD1, PCM1, TUBGCP6) |
| GO:0042267 | natural killer cell mediated cytotoxicity | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.03 | 0.08 | (LGALS9, LILRB1, PIK3R6) | |
| GO:0070228 | regulation of lymphocyte apoptotic process | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.03 | 0.09 | (IRS2, LGALS9, SLC39A10) | |
| R-HSA:450346 | activated human TAK1 phosphorylates MKK3/MKK6 | REACTOME_Reactions_27.02.2019 | 0.04 | 0.00 | (MAP2K3, NOD1, TAB1) | |
| GO:0061098 | positive regulation of protein tyrosine kinase activity | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.05 | 0.02 | (AGXT, DGKQ, DOK7, ERCC6, RELN) | |
| R-HSA:450302 | activated TAK1 mediates p38 MAPK activation | REACTOME_Pathways_27.02.2019 | 0.06 | 0.00 | (MAP2K3, NOD1, TAB1) | |
| R-HSA:448424 | interleukin-17 signaling | REACTOME_Pathways_27.02.2019 | 0.06 | 0.00 | (MAP2K3, NOD1, TAB1) | |
| R-HSA:382054 | PDGF binds to extracellular matrix proteins | REACTOME_Reactions_27.02.2019 | 0.07 | 0.04 | (COL6A3, COL6A6, SPP1) | |
| WP:231 | TNF alpha signaling pathway | WikiPathways_27.02.2019 | 0.08 | 0.00 | (KSR1, MAP2K3, NFKBIE, NSMAF, TAB1) | |
| R-HSA:186797 | signaling by PDGF | REACTOME_Pathways_27.02.2019 | 0.08 | 0.04 | (COL6A3, COL6A6, GRB7, SPP1) | |
| GO:0036498 | IRE1-mediated unfolded protein response | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.08 | 0.04 | (ARFGAP3, EXTL1, NR1H4, VAPB, WFS1) | |
| GO:0035176 | social behavior | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.09 | 0.02 | (HTT, MBD1, PCM1) | |
| R-HSA:373739 | ankyrins link voltage-gated sodium and potassium channels to spectrin and L1 | REACTOME_Reactions_27.02.2019 | 0.10 | 0.05 | (ANK2, SCN7A, SPTA1) | |
| R-HSA:445095 | interaction between L1 and ankyrins | REACTOME_Pathways_27.02.2019 | 0.11 | 0.05 | (ANK2, SCN7A, SPTA1) | |
| GO:1901618 | organic hydroxy compound transmembrane transporter activity | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.11 | 0.03 | (AQP7, HTT, SLC10A6) | |
| GO:0006890 | retrograde vesicle-mediated transport, Golgi to endoplasmic reticulum | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.13 | 0.05 | (ARFGAP3, CENPE, ERGIC1, HTT, TAPBP) | |
| GO:0035036 | sperm-egg recognition | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.13 | 0.05 | (CATSPER2, ZAN, ZP1) | |
| GO:0021846 | cell proliferation in forebrain | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.13 | 0.02 | (KIF14, MBD1, PCM1) | |
| GO:0018195 | peptidyl-arginine modification | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.13 | 0.05 | (NR1H4, PADI2, PRMT7) | |
| GO:0002753 | cytoplasmic pattern recognition receptor signaling pathway | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.15 | 0.00 | (ALPK1, DHX58, NOD1, TAB1) | |
| R-HSA:6807875 | ARFGAP, cargo, v-SNAREs, and p24 proteins bind nascent COPI complex | REACTOME_Reactions_27.02.2019 | 0.19 | 0.05 | (ANK2, ARFGAP3, SPTA1) | |
| R-HSA:6807877 | ARFGAPs stimulate ARF GTPase activity | REACTOME_Reactions_27.02.2019 | 0.19 | 0.05 | (ANK2, ARFGAP3, SPTA1) | |
| R-HSA:450294 | MAP kinase activation | REACTOME_Pathways_27.02.2019 | 0.20 | 0.00 | (MAP2K3, NOD1, TAB1) | |
| R-HSA:375165 | NCAM signaling for neurite out-growth | REACTOME_Pathways_27.02.2019 | 0.22 | 0.04 | (COL6A3, COL6A6, SPTA1) | |
| R-HSA:2022090 | assembly of collagen fibrils and other multimeric structures | REACTOME_Pathways_27.02.2019 | 0.22 | 0.04 | (COL6A3, COL6A6, LAMA3) | |
| R-HSA:168643 | nucleotide-binding domain, leucine rich repeat containing receptor (NLR) signaling pathways | REACTOME_Pathways_27.02.2019 | 0.23 | 0.00 | (NOD1, PSTPIP1, TAB1) | |
| GO:1903573 | negative regulation of response to endoplasmic reticulum stress | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.23 | 0.04 | (NR1H4, PRKN, WFS1) | |
| AVM3 | GO:0038089 | positive regulation of cell migration by vascular endothelial growth factor signaling pathway | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.01 | 0.17 | (KDR, MYO1C, PKD1) |
| GO:0007157 | heterophilic cell–cell adhesion via plasma membrane cell adhesion molecules | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.04 | 0.09 | (HMCN1, RP9, SPG7) | |
| GO:0120193 | tight junction organization | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.04 | 0.09 | (EPHA2, MYO1C, PDZD4) | |
| GO:0032688 | negative regulation of interferon-beta production | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.04 | 0.08 | (LRRFIP1, MYO1C, NLRC3) | |
| GO:0048739 | cardiac muscle fiber development | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.08 | 0.05 | (MYOM2, OBSL1, SPG7) | |
| AVM4 | GO:0001539 | cilium or flagellum-dependent cell motility | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.00 | 0.00 | (DNAH11, DNAH14, DNAH2, DNAH3, DNAH8, GAS8) |
| GO:0060285 | cilium-dependent cell motility | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.00 | 0.00 | (DNAH11, DNAH14, DNAH2, DNAH3, DNAH8, GAS8) | |
| GO:0007221 | positive regulation of transcription of Notch receptor target | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.00 | 0.00 | (MAML1, NOTCH3, OPN1LW, PLXND1, SPHKAP) | |
| WP:334 | GPCRs, class B secretin-like | WikiPathways_27.02.2019 | 0.01 | 0.10 | (ADCYAP1R1, ADGRG2, GLP2R, SCTR) | |
| GO:0003279 | cardiac septum development | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.01 | 0.00 | (CRELD1, DNAH11, LRP2, MAML1, PLXND1, SLIT2, SMO, TAB1) | |
| GO:0071503 | response to heparin | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.02 | 0.01 | (AOC1, GPIHBP1, SLIT2) | |
| R-HSA:9021450 | PLXND1 gene expression is stimulated by NOTCH1/NOTCH3 coactivator complexes | REACTOME_Reactions_27.02.2019 | 0.02 | 0.00 | (MAML1, NOTCH3, PLXND1) | |
| GO:0061476 | response to anticoagulant | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.03 | 0.01 | (AOC1, GPIHBP1, SLIT2) | |
| R-HSA:2162123 | synthesis of prostaglandin (PG) and thromboxane (TX) | REACTOME_Pathways_27.02.2019 | 0.03 | 0.19 | (AKR1C3, HPGD, TBXAS1) | |
| GO:0003205 | cardiac chamber development | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.04 | 0.00 | (CRELD1, DNAH11, LRP2, MAML1, PKD1, PLXND1, SLIT2, SMO, TAB1) | |
| GO:0048278 | vesicle docking | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.05 | 0.15 | (CTBP2, SCFD2, STX11) | |
| GO:0007157 | heterophilic cell–cell adhesion via plasma membrane cell adhesion molecules | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.05 | 0.15 | (FAT4, HMCN1, NUFIP2) | |
| GO:0021795 | cerebral cortex cell migration | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.05 | 0.15 | (SLIT2, SRGAP2, SUN1) | |
| GO:0002753 | cytoplasmic pattern recognition receptor signaling pathway | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.05 | 0.25 | (ALPK1, IRAK2, TAB1) | |
| GO:0007616 | long-term memory | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.09 | 0.04 | (CTNS, GRIA1, PJA2, PRNP) | |
| GO:0007613 | memory | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.11 | 0.04 | (ADGRF1, CHRNB2, CIC, CTNS, GRIA1, PJA2, PRNP) | |
| R-HSA:9013508 | NOTCH3 intracellular domain regulates transcription | REACTOME_Pathways_27.02.2019 | 0.14 | 0.00 | (MAML1, NOTCH3, PLXND1) | |
| GO:0035904 | aorta development | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.25 | 0.00 | (LRP2, PLXND1, TAB1) | |
| GO:0060840 | artery development | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.25 | 0.00 | (HPGD, LRP2, NOTCH3, PLXND1, TAB1) | |
| GO:0045747 | positive regulation of Notch signaling pathway | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.27 | 0.00 | (MAML1, NEPRO, OPN1LW, SPHKAP) | |
| GO:0035082 | axoneme assembly | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.33 | 0.00 | (DNAH8, GAS8, RSPH1) | |
| WP:268 | Notch signaling | WikiPathways_27.02.2019 | 0.39 | 0.00 | (CTBP2, MAML1, NOTCH3) | |
| GO:1903846 | positive regulation of cellular response to transforming growth factor beta stimulus | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.40 | 0.00 | (OPN1LW, RNF111, SPHKAP) | |
| GO:0030511 | positive regulation of transforming growth factor beta receptor signaling pathway | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.40 | 0.00 | (OPN1LW, RNF111, SPHKAP) | |
| R-HSA:9012852 | signalling by NOTCH3 | REACTOME_Pathways_27.02.2019 | 0.41 | 0.00 | (MAML1, NOTCH3, PLXND1) | |
| GO:0060976 | coronary vasculature development | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.41 | 0.00 | (LRP2, PLXND1, TAB1) | |
| GO:0060411 | cardiac septum morphogenesis | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.42 | 0.00 | (DNAH11, LRP2, SLIT2, SMO) | |
| AVM5 | WP:2572 | primary focal segmental glomerulosclerosis (FSGS) | WikiPathways_27.02.2019 | 0.03 | 0.05 | (AGRN, CAMK2B, LAMB2, LMX1B, MKI67) |
| GO:0006027 | glycosaminoglycan catabolic process | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.04 | 0.08 | (AGRN, CD44, GPX4) | |
| GO:0031952 | regulation of protein autophosphorylation | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.08 | 0.04 | (CA5A, CAMK2B, ENG) | |
| GO:1904886 | beta-catenin destruction complex disassembly | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.08 | 0.05 | (AXIN1, CA5A, DVL1) | |
| GO:0097150 | neuronal stem cell population maintenance | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.09 | 0.05 | (BRCA2, FANCC, MBD1) | |
| R-HSA:2142753 | arachidonic acid metabolism | REACTOME_Pathways_27.02.2019 | 0.09 | 0.05 | (CYP4B1, CYP4F3, GPX4, PRXL2B) | |
| GO:1904063 | negative regulation of cation transmembrane transport | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.10 | 0.04 | (ARHGEF40, CA5A, CAMK2B, EPHB2, KEL) | |
| GO:0007528 | neuromuscular junction development | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.15 | 0.05 | (AGRN, DVL1, LAMB2) | |
| GO:0034766 | negative regulation of ion transmembrane transport | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.18 | 0.04 | (ARHGEF40, CA5A, CAMK2B, EPHB2, KEL) | |
| GO:0043267 | negative regulation of potassium ion transport | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.22 | 0.04 | (CA5A, CPAMD8, KEL) | |
| GO:0044091 | membrane biogenesis | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.25 | 0.04 | (CA5A, EPHB2, PTPRH) | |
| GO:2001258 | negative regulation of cation channel activity | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.26 | 0.04 | (CA5A, CAMK2B, EPHB2) | |
| GO:0071709 | membrane assembly | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.28 | 0.04 | (CA5A, EPHB2, PTPRH) | |
| GO:0016248 | channel inhibitor activity | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.28 | 0.04 | (CA5A, CAMK2B, PHPT1) | |
| GO:0008200 | ion channel inhibitor activity | GO_BiologicalProcess-EBI-UniProt-GOA_27.02.2019_00h00 | 0.28 | 0.04 | (CA5A, CAMK2B, PHPT1) |
| Ontology | Feature | ID | Name | Genes |
|---|---|---|---|---|
| GO: Biological Process | Vessel development | GO:0001525 | Angiogenesis | ACVRL1 ENG EPHA2 EPHB2 GDF2 KDR NOTCH3 PLXND1 SLIT2 TGFBR2 |
| GO:0001568 | Blood vessel development | ACVRL1 ENG EPHA2 EPHB2 GDF2 KDR PKD1 LRP1 LTBP1 TAB1 HPGD LRP2 NOTCH3 PLXND1 SLIT2 SMO TGFBR2 | ||
| GO:0001569 | Branching involved in blood vessel morphogenesis | ENG GDF2 PLXND1 TGFBR2 | ||
| GO:0001570 | Vasculogenesis | ENG EPHA2 GDF2 SMO KDR TGFBR2 | ||
| GO:0001944 | Vasculature development | ACVRL1 ENG EPHA2 GDF2 KDR PKD1 LRP1 LTBP1 HPGD LRP2 NOTCH3 PLXND1 SLIT2 SMO TAB1 TGFBR2 | ||
| GO:0048514 | Blood vessel morphogenesis | ACVRL1 ENG EPHA2 EPHB2 GDF2 LRP1 KDR HPGD LRP2 NOTCH3 PLXND1 SLIT2 SMO TGFBR2 | ||
| GO:1901342 | Regulation of vasculature development | ACVRL1 ENG EPHA2 GDF2 KDR PLXND1 TGFBR2 | ||
| TGFBR signaling | GO:0007179 | Transforming growth factor beta receptor signaling pathway | ACVRL1 AXIN1 ENG FBN2 GDF2 LTBP1 LTBP4 HPGD RNF111 SMAD4 TAB1 TGFBR2 | |
| GO:0017015 | Regulation of transforming growth factor beta receptor signaling pathway | AXIN1 ENG FBN2 GDF2 LTBP1 LTBP4 RNF111 SMAD4 TGFBR2 | ||
| GO:0071559 | Response to transforming growth factor beta | ACVRL1 AXIN1 ENG FBN2 GDF2 HPGD RNF111 LTBP1 LTBP4 SMAD4 TAB1 TGFBR2 | ||
| GO:0071560 | Cellular response to transforming growth factor beta stimulus | ACVRL1 AXIN1 ENG FBN2 GDF2 LTBP1 LTBP4 HPGD RNF111 SMAD4 TAB1 TGFBR2 | ||
| GO:1903844 | Regulation of cellular response to transforming growth factor beta stimulus | AXIN1 ENG FBN2 GDF2 LTBP1 LTBP4 SMAD4 TGFBR2 | ||
| Heart development | GO:0003007 | Heart morphogenesis | ACVRL1 DNAH11 ENG SMAD4 FAT4 LRP2 PLXND1 SLIT2 SMAD4 SMO TAB1 TGFBR2 | |
| GO:0003205 | Cardiac chamber development | DNAH11 ENG LRP2 MAML1 PLXND1 SLIT2 LRP1 LTBP1 SMAD4 SMO TAB1 TGFBR2 | ||
| GO:0003206 | Cardiac chamber morphogenesis | DNAH11 ENG LRP2 SLIT2 SMAD4 SMO TGFBR2 | ||
| GO:0003208 | Cardiac ventricle morphogenesis | ENG LRP2 SMAD4 TGFBR2 | ||
| GO:0003231 | Cardiac ventricle development | ENG LTBP1 LRP2 SLIT2 SMAD4 TGFBR2 | ||
| GO:0003279 | Cardiac septum development | DNAH11 ENG LRP1 LTBP1 LRP2 MAML1 PLXND1 SLIT2 SMAD4 SMO TAB1 TGFBR2 | ||
| GO:0007507 | Heart development | ACVRL1 DNAH5 DNAH11 DVL1 ENG KDR PKD1 LRP1 LTBP1 FAT4 LRP2 MAML1 PLXND1 SLIT2 SMAD4 SMO TAB1 TGFBR2 | ||
| GO:0060411 | Cardiac septum morphogenesis | DNAH11 ENG LRP2 SLIT2 SMAD4 SMO TGFBR2 | ||
| GO:0072358 | Cardiovascular system development | ACVRL1 ENG EPHA2 EPHB2 GDF2 KDR PKD1 LRP1 LTBP1 HPGD LRP2 NOTCH3 PLXND1 SLIT2 SMO TAB1 TGFBR2 | ||
| GO:0072359 | Circulatory system development | ACVRL1 DNAH5 DNAH11 DVL1 ENG EPHA2 EPHB2 FAT4 GDF2 LRP1 LTBP1 KDR PKD1 HPGD LRP2 MAML1 NOTCH3 PLXND1 SLIT2 SMAD4 SMO TAB1 TGFBR2 | ||
| GO:2000826 | Regulation of heart morphogenesis | ENG SMAD4 SMO TGFBR2 | ||
| BMP signaling | GO:0030509 | BMP signaling pathway | ACVRL1 ENG GDF2 KDR LRP2 SMAD4 | |
| GO:0030510 | Regulation of BMP signaling pathway | ACVRL1 ENG GDF2 KDR LRP2 SMAD4 | ||
| GO:0030513 | Positive regulation of BMP signaling pathway | ACVRL1 ENG GDF2 KDR SMAD4 | ||
| GO:0071772 | Response to BMP | ACVRL1 ENG GDF2 KDR LRP2 SMAD4 | ||
| GO:0071773 | Cellular response to BMP stimulus | ACVRL1 ENG GDF2 KDR LRP2 SMAD4 | ||
| Endothelial/mesenchymal differentiation | GO:0001935 | Endothelial cell proliferation | ACVRL1 ENG EPHA2 GDF2 KDR | |
| GO:0001936 | Regulation of endothelial cell proliferation | ACVRL1 ENG GDF2 KDR | ||
| GO:0003158 | Endothelium development | ACVRL1 ENG GDF2 KDR SMAD4 | ||
| GO:0045446 | Endothelial cell differentiation | ACVRL1 ENG GDF2 KDR SMAD4 | ||
| GO:0048762 | Mesenchymal cell differentiation | ACVRL1 ENG SMAD4 SMO TGFBR2 | ||
| GO:0060485 | Mesenchyme development | ACVRL1 ENG SMAD4 SMO TGFBR2 | ||
| Hypoxia response | GO:0001666 | Response to hypoxia | ACVRL1 CHRNB2 ENG KDR SMAD4 TGFBR2 | |
| GO:0036293 | Response to decreased oxygen levels | ACVRL1 CHRNB2 ENG KDR SMAD4 TGFBR2 | ||
| GO:0070482 | Response to oxygen levels | ACVRL1 CHRNB2 ENG KDR SMAD4 TGFBR2 | ||
| Human Phenotype | Cerebrovascular malformations | HP:0100026 | Arteriovenous malformation | ACVRL1 BRCA2 DVL1 ENG FANCC GDF2 SMAD4 |
| HP:0001009 | Telangiectasia | ACVRL1 BRCA2 ENG FANCC GDF2 SMAD4 | ||
| HP:0001048 | Cavernous hemangioma | ACVRL1 AXIN1 ENG GDF2 SMAD4 | ||
| Benign/malignant neoplasm | HP:0005306 | Capillary hemangioma | ACVRL1 DVL1 ENG GDF2 KDR SMAD4 | |
| HP:0001028 | Hemangioma | ACVRL1 AXIN1 DVL1 ENG GDF2 KDR PRKN SMAD4 | ||
| HP:0100742 | Vascular neoplasm | ACVRL1 AXIN1 DVL1 ENG GDF2 KDR PRKN SMAD4 | ||
| Vessel dilatation | HP:0002624 | Abnormal venous morphology | ACVRL1 AXIN1 BRCA2 ENG EPHB2 GDF2 NOTCH3 SMAD4 | |
| HP:0002619 | Varicose veins | ACVRL1 AXIN1 ENG GDF2 NOTCH3 SMAD4 | ||
| HP:0004414 | Abnormality of the pulmonary artery | ACVRL1 ENG FBN2 GDF2 LTBP4 SMAD4 TGFBR2 | ||
| HP:0004930 | Abnormality of the pulmonary vasculature | ACVRL1 AXIN1 ENG FBN2 GDF2 LTBP4 SMAD4 TGFBR2 | ||
| HP:0100659 | Abnormality of the cerebral vasculature | ACVRL1 BRCA2 ENG GDF2 NOTCH3 PKD1 PRNP SMAD4 SMO TGFBR2 WFS1 | ||
| HP:0009145 | Abnormal cerebral artery morphology | ACVRL1 BRCA2 ENG FANCC GDF2 NOTCH3 PKD1 SMAD4 TGFBR2 | ||
| HP:0011004 | Abnormal systemic arterial morphology | ACVRL1 DNAH11 ENG FBN2 GDF2 NOTCH3 PKD1 SMAD4 TGFBR2 | ||
| Mouse Phenotype | Cerebrovascular malformations | MP:0006093 | Arteriovenous malformation | ACVRL1 ENG GDF2 NOTCH3 |
| Vessel dilatation | MP:0000259 | Abnormal vascular development | ACVRL1 CD44 CTBP2 ENG EPHA2 KDR HPGD MAML1 NOTCH3 PLXND1 LTBP1 SMAD4 SMO SPP1 TGFBR2 | |
| MP:0001614 | Abnormal blood vessel morphology | ACVRL1 CD44 CTBP2 DNAH11 DNAH5 ENG EPHA2 FBN2 GDF2 KDR PKD1 LMX1B LRP1 LTBP1 LTBP4 HPGD LRP2 MAML1 NOTCH3 PLXND1 SLIT2 NR1H4 SMAD4 SPP1 TAB1 TGFBR2 | ||
| MP:0003410 | Abnormal artery development | ACVRL1 ENG NOTCH3 PLXND1 SMO KDR LTBP1 TGFBR2 | ||
| MP:0004787 | Abnormal dorsal aorta morphology | ACVRL1 ENG KDR TGFBR2 | ||
| MP:0000267 | Abnormal heart development | ACVRL1 AXIN1 CTBP2 DNAH5 ENG KDR MAML1 PKD1 SMAD4 SMO TGFBR2 | ||
| Defects in embryo vasculogenesis | MP:0001719 | Absent vitelline blood vessels | ACVRL1 CTBP2 ENG KDR SMO TGFBR2 | |
| MP:0003229 | Abnormal vitelline vasculature morphology | ACVRL1 CTBP2 ENG KDR SMO SMAD4 TGFBR2 | ||
| Disease | Cerebrovascular malformations | C0003857 | Congenital arteriovenous malformation | ACVRL1 ENG NOTCH3 SMAD4 |
| Vessel disorders | C0334533 | Arteriovenous hemangioma | ACVRL1 ENG NOTCH3 SMAD4 | |
| C0007820 | Cerebrovascular Disorders | ACVRL1 ENG GDF2 NOTCH3 PKD1 RELN SMAD4 SPP1 TGFBR2 | ||
| C0042373 | Vascular Diseases | ACVRL1 ENG KDR NOTCH3 PRKN SMAD4 SPP1 TGFBR2 | ||
| C0002940 | Aneurysm | ACVRL1 ENG LRP1 LTBP4 PKD1 TGFBR2 |
| Locus | Gene | HGNC ID | SNP ID | Ensembl Transcript ID | Coding Sequence Change | Ensembl Protein ID | Protein Sequence Change |
|---|---|---|---|---|---|---|---|
| 1p22.3 | CCN1 * | 2654 | rs765069158 | ENST00000451137 | c.G463C | ENSP00000398736 | p.Gly155Arg |
| 1p36.12 | EPHB2 | 3393 | rs142113032 | ENST00000400191 | c.847G>C | ENSP00000383053 | p.Asp283His |
| 1p36.13 | EPHA2 | 3386 | rs35484156 | ENST00000358432 | c.830C>T | ENSP00000351209 | p.Ser277Leu |
| 1p36.33 | DVL1 | 3084 | rs61735963 | ENST00000378891 | c.469G>A | ENSP00000368169 | p.Ala157Thr |
| 1q21.3 | CHRNB2 | 1962 | rs55685423 | ENST00000368476 | c.1191G>C | ENSP00000357461 | p.Gln397His |
| 2p22.3 | LTBP1 | 6714 | rs80163321 rs149319598 | ENST00000404816 | c. 2248 G>A c. 2410C>T | ENSP00000386043 | p.Val750Ile p.Pro804Ser |
| 3p24.1 | TGFBR2 * | 11773 | rs35766612 | ENST00000295754 | c.1159G>T | ENSP00000295754 | p.Val387Leu |
| 3q22.1 | PLXND1 | 9107 | rs137955512 | ENST00000324093 | c.2275C>T | ENSP00000317128 | p.Pro759Ser |
| 4p15.31 | SLIT2 | 11086 | rs115629108 | ENST00000504154 | c.4049G>A | ENSP00000422591 | p.Ser1350Asn |
| 4q12 | KDR | 6307 | rs755067067 | ENST00000263923 | c.1990C>T | ENSP00000263923 | p.Arg664Cys |
| 5q23.3 | FBN2 | 3604 | rs28763954 | ENST00000262464 | c.976C>T | ENSP00000262464 | p.Pro326Ser |
| 5q35.3 | MAML1 | 13632 | rs113636707 | ENST00000292599 | c.569G>A | ENSP00000292599 | p.Arg190His |
| 5q35.3 | FLT4 * | 3767 | rs200763913 | ENST00000261937 | c.1133G>A | ENSP00000261937 | p.Arg378His |
| 6p21.32 | NOTCH4 * | 7884 | rs8192573 | ENST00000375023 | c.4037G>A | ENSP00000364163 | p.Arg1346Gln |
| 7p13 | CAMK2B | 1461 | rs528355050 | ENST00000395749 | c.1577C>T | ENSP00000379098 | p.Pro526Leu |
| 7q22.1 | RELN | 9957 | rs114684479 rs79499902 | ENST00000428762 | c.3477C>A c.5284G>A | ENSP00000392423 | p.Asn1159Lys p.Val1762Ile |
| 7q36.1 | GIMAP1 * | 23237 | rs1326399257 | ENST00000307194 | c.699G>A | ENSP00000302833 | p.W233X |
| 9q34.11 | ENG * | 3349 | rs139398993 | ENST00000373203 | c.392C>T | ENSP00000362299 | p.Pro131Leu |
| 10q26.13 | CTBP2 | 2595 | rs1058301 | ENST00000334808 | c.387C>G | ENSP00000357816 | p.Asp129Glu |
| 12q13.3 | LRP1 | 6692 | rs113379328 | ENST00000243077 | c.7636G>A | ENSP00000243077 | p.Gly2546Ser |
| 12q24.31 | NCoR2 * | 7673 | Novel | ENST00000405201 | c..2078G>T | ENSP00000384018 | p.Glu693Ter |
| 15q22.1-q22.2 | RNF111 | 17384 | rs142916216 | ENST00000557998 | c.888T>G | ENSP00000452732 | p.Ile296Met |
| 16p13.3 | AXIN1 | 903 | rs200741961 | ENST00000262320 | c.644C>T | ENSP00000262320 | p.Ser215Leu |
| 17p11.2 | MAP2K3 | 6843 | rs33911218 rs36047035 | ENST00000342679 | c.118C>G c.164G>C | ENSP00000345083 | p.Pro40Ala p.Arg55Thr |
| 19p13.12 | NOTCH3 | 7883 | rs141320511 | ENST00000263388 | c.4552C>A | ENSP00000263388 | p.Leu1518Met |
| 19q13.2 | LTBP4 | 6717 | rs35809725 | ENST00000308370 | c.4499A>T | ENSP00000311905 | p.Tyr1500Phe |
| 22q13.1 | TAB1 | 18157 | rs536084162 rs140879164 | ENST00000216160 | c.19A>C c.560G>A | ENSP00000216160 | p.Ser7Arg p.Arg187His |
| Patient | Sex | Age (Years) | Age of Onset (Years) | Symptoms | Lesion Number | Spetzler–Martin Grading |
|---|---|---|---|---|---|---|
| AVM1 | M | 14 | 12 | Intracerebral hemorrhage following AVM rupture | 1—parietal left area | 2 |
| AVM2 | M | 31 | 18 | Tremor of the left leg, diffuse tremor, seizures | 1—front-parietal left area (not bleeding) | 2 |
| AVM3 | F | 32 | 29 | Dizziness, tinnitus, seizures nausea right hemiparesis visus reduction | 1— parietal left area | 3 |
| AVM4 | F | 8 | At birth | Drowsy status, finalistic limb movement | 1—proliferative microangiopathy, central left area | 3 |
| AVM5 | M | 7 | 5 | Sudden headache, vertigo, seizures | 1—anterior-parietal paramedian right area | 2 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Scimone, C.; Granata, F.; Longo, M.; Mormina, E.; Turiaco, C.; Caragliano, A.A.; Donato, L.; Sidoti, A.; D’Angelo, R. Germline Mutation Enrichment in Pathways Controlling Endothelial Cell Homeostasis in Patients with Brain Arteriovenous Malformation: Implication for Molecular Diagnosis. Int. J. Mol. Sci. 2020, 21, 4321. https://doi.org/10.3390/ijms21124321
Scimone C, Granata F, Longo M, Mormina E, Turiaco C, Caragliano AA, Donato L, Sidoti A, D’Angelo R. Germline Mutation Enrichment in Pathways Controlling Endothelial Cell Homeostasis in Patients with Brain Arteriovenous Malformation: Implication for Molecular Diagnosis. International Journal of Molecular Sciences. 2020; 21(12):4321. https://doi.org/10.3390/ijms21124321
Chicago/Turabian StyleScimone, Concetta, Francesca Granata, Marcello Longo, Enricomaria Mormina, Cristina Turiaco, Antonio A. Caragliano, Luigi Donato, Antonina Sidoti, and Rosalia D’Angelo. 2020. "Germline Mutation Enrichment in Pathways Controlling Endothelial Cell Homeostasis in Patients with Brain Arteriovenous Malformation: Implication for Molecular Diagnosis" International Journal of Molecular Sciences 21, no. 12: 4321. https://doi.org/10.3390/ijms21124321









