Raman Multi-Omic Snapshot and Statistical Validation of Structural Differences between Herpes Simplex Type I and Epstein–Barr Viruses
Abstract
:1. Introduction
2. Results
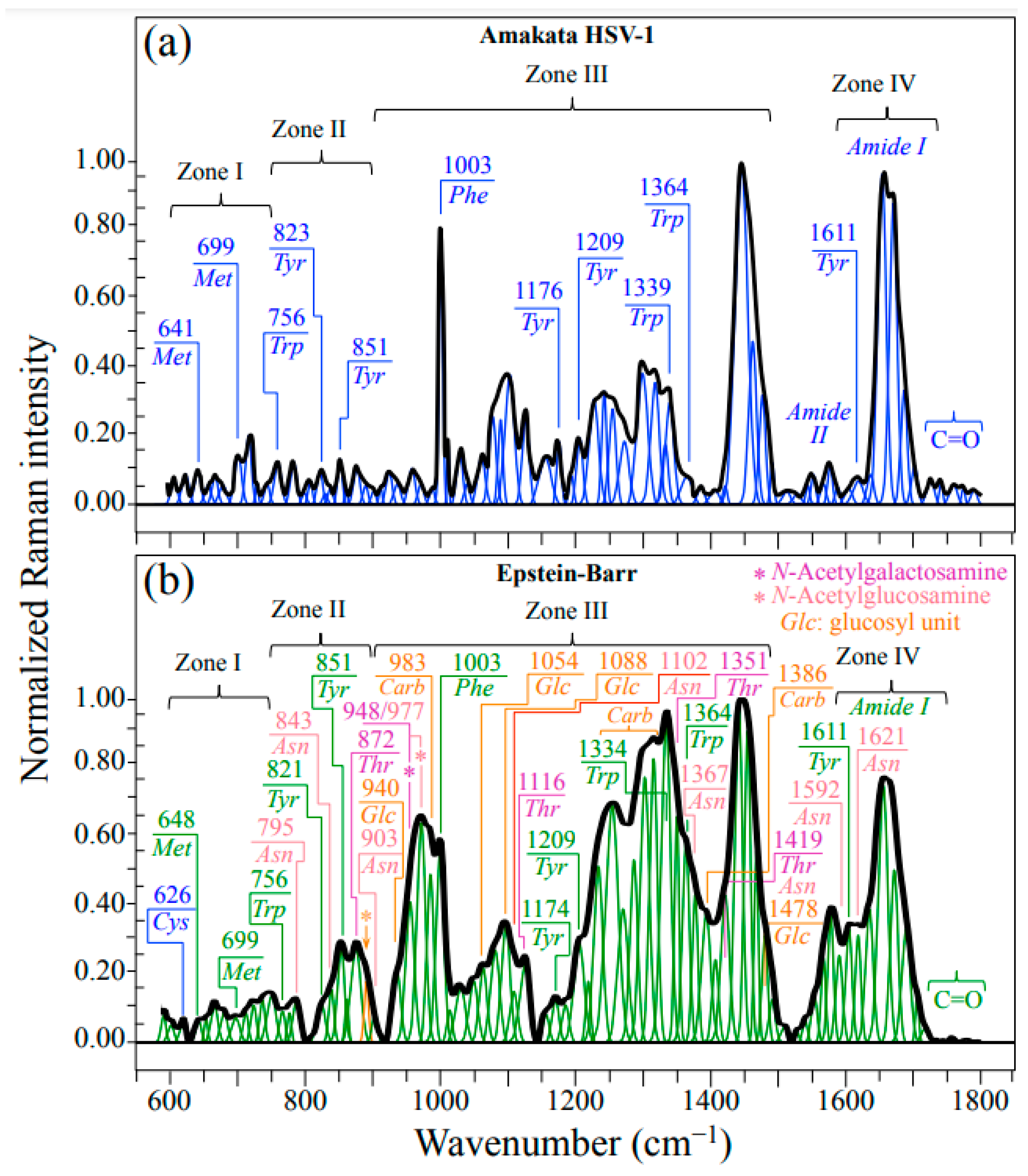
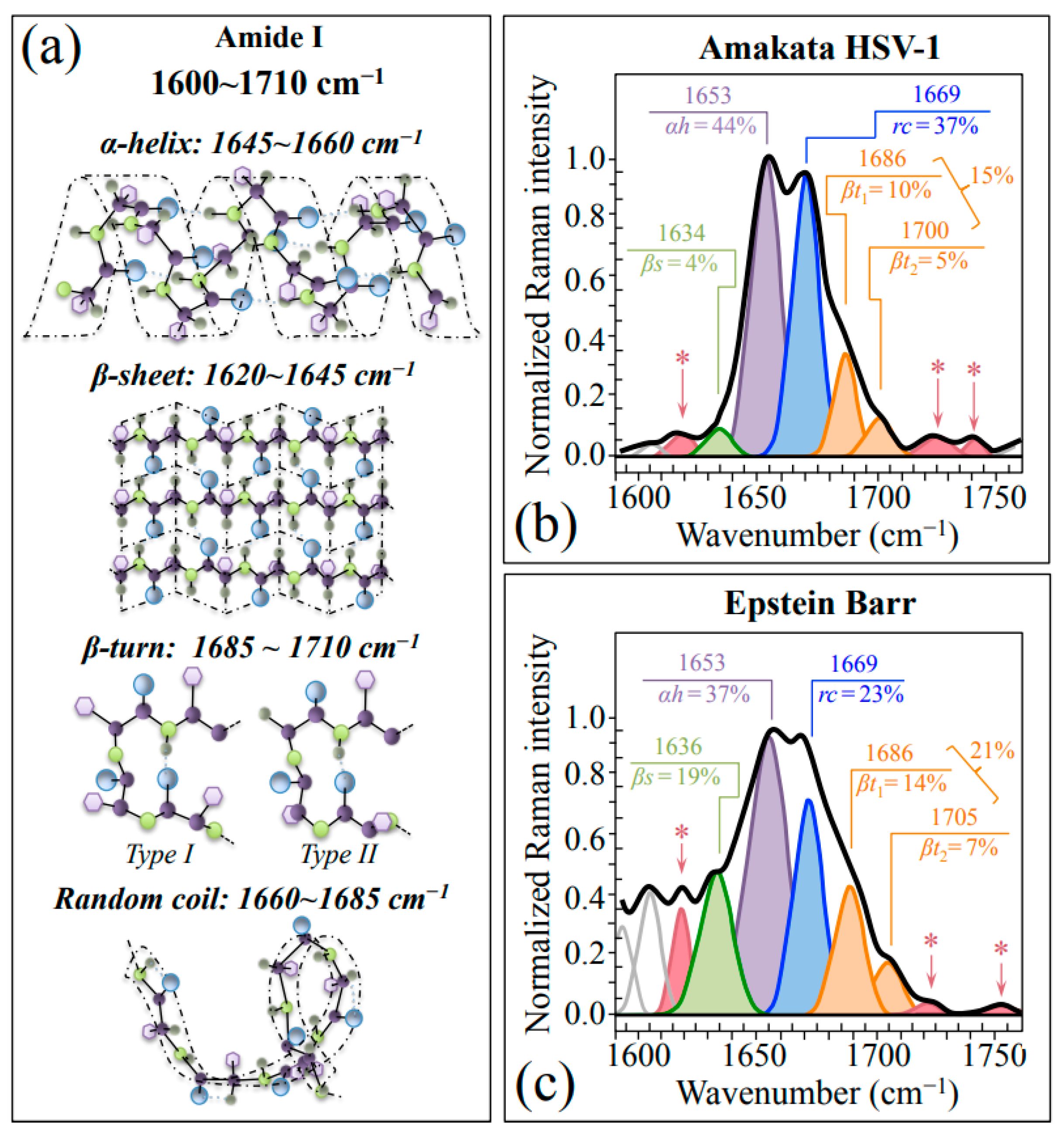
2.1. Average Raman Spectra and Assignments of Deconvoluted Sub-Bands
2.2. PCA Statistical Analysis and Raman Barcoding
3. Discussion
3.1. Raman Fingerprints of Structural Differences between HSV-1 and EBV
3.2. Raman Evidences for O-Glycosylation in EBV
3.3. Comparison and Validation through the Literature Data from Raman and Other Analytical Methods
4. Materials and Methods
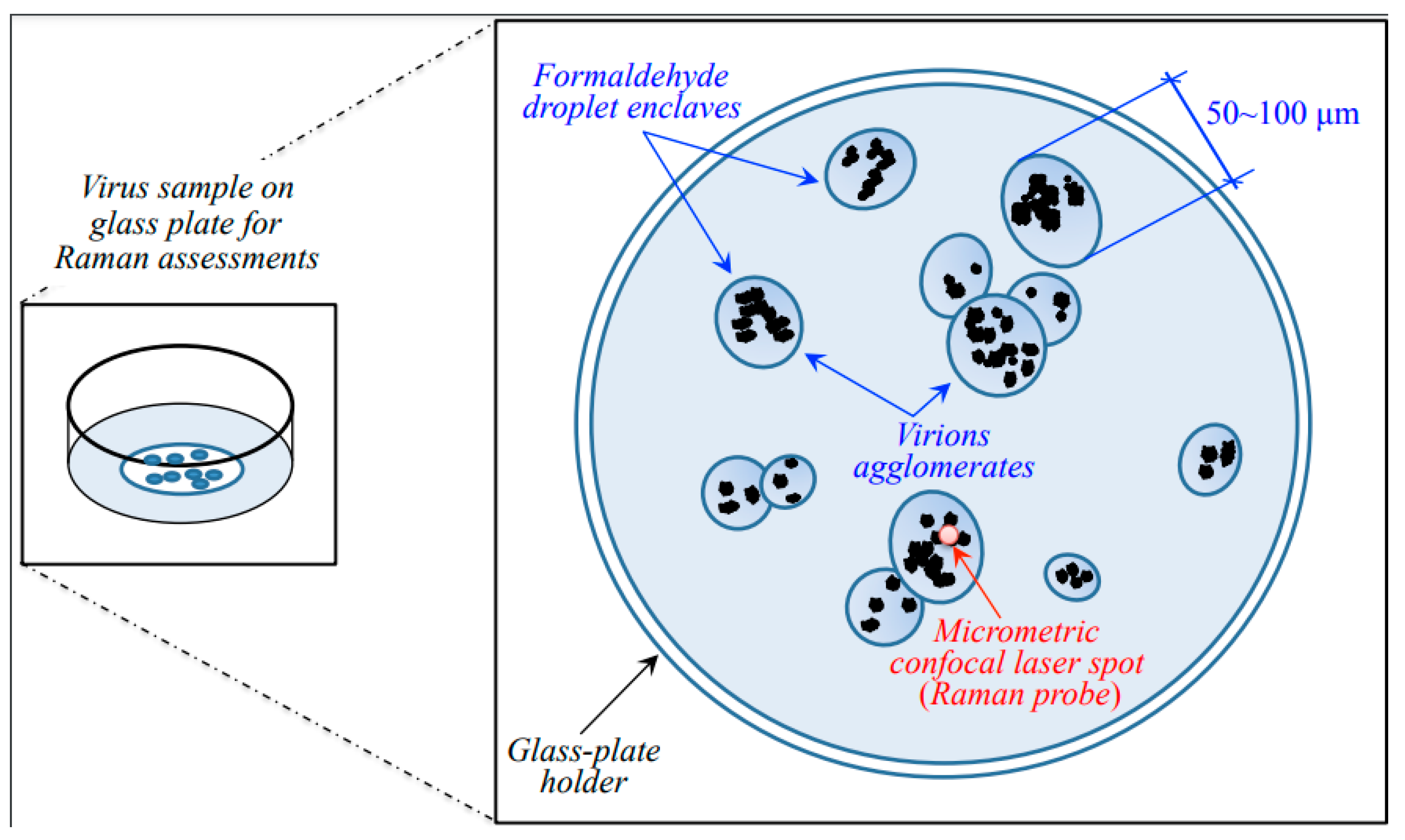
4.1. Virus Samples
4.2. Raman Samples and Spectroscopic Procedures
4.3. Statistical Analyses and Buildup of Raman Barcodes
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Whitley, R.J. Herpesviruses. In Medical Microbiology, 4th ed.; Baron, S., Ed.; The University of Texas Medical Branch: Galveston, TX, USA, 1996; Chapter 68. [Google Scholar]
- Sandri-Goldin, R.M. Alpha Herpesviruses: Molecular and Cellular Biology; Caister Academic Press: Norwich, UK, 2006. [Google Scholar]
- McGeoch, D.J.; Rixon, F.J.; Davison, A.J. Topics in herpesvirus genomics and evolution. Virus Res. 2006, 117, 90–104. [Google Scholar] [CrossRef] [PubMed]
- Balasubramaniam, R.; Kuperstein, A.S.; Stoopler, E.T. Update on oral herpes virus infections. Dent. Clin. N. Am. 2014, 58, 265–280. [Google Scholar] [CrossRef] [PubMed]
- McGeoch, D.J.; Cook, S.; Dolan, A.; Jamieson, F.E.; Telford, E.A. Molecular phylogeny and evolutionary timescale for the family of mammalian herpesviruses. J. Mol. Biol. 1995, 247, 443–458. [Google Scholar] [CrossRef]
- Burrell, C.J.; Howard, C.R.; Murphy, F.A. Virion structure and composition. In Fenner and White’s Medical Virology; Academic Press: London, UK, 2017; pp. 27–37. [Google Scholar]
- Villanueva, R.A.; Rouillé, Y.; Dubuisson, J. Interactions between virus proteins and host cell membranes during the viral life cycle. Int. Rev. Cytol. 2005, 245, 171–244. [Google Scholar] [PubMed]
- Casjens, S. Principles of virion structure, function and assembly. In Structural Biology of Viruses; Chiu, W., Burnett, R.M., Garcia, R.L., Eds.; Oxford University Press: New York, NY, USA, 1997; pp. 3–37. [Google Scholar]
- Baker, M.L.; Jiang, W.; Rixon, F.J.; Chiu, W. Common ancestry of herpesviruses and tailed DNA bacteriophages. J. Virol. 2005, 79, 14967–14970. [Google Scholar] [CrossRef]
- Meents, A.; Wiedorn, M.O. Virus structures by X-ray free-electron lasers. Annu. Rev. Virol. 2019, 6, 161–176. [Google Scholar] [CrossRef]
- Hryc, C.F.; Chen, D.-H.; Afonine, P.V.; Jakana, J.; Wang, Z.; Haase-Pettingell, C.; Jiang, W.; Adams, P.D.; King, J.A.; Schmid, M.F.; et al. Accurate model annotation of a near-atomic resolution cryo-EM map. Proc. Natl. Acad. Sci. USA 2017, 114, 3103–3108. [Google Scholar] [CrossRef]
- Harrison, S.C. Principles of virus structure. In Fields Virology, 4th ed.; Knipe, D.M., Howley, P.M., Eds.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2011; pp. 53–85. [Google Scholar]
- Khaykelson, D.; Raviv, U. Studying viruses using solution X-ray scattering. Biophys. Rev. 2020, 24, 41–48. [Google Scholar] [CrossRef]
- Thomas, G.J., Jr. Viruses and nucleoproteins. In Biological Applications of Raman Spectroscopy; Spiro, T.G., Ed.; John Wiley & Sons: New York, NY, USA, 1987; Volume 1, pp. 135–201. [Google Scholar]
- Thomas, G.J., Jr. Raman spectroscopy of protein and nucleic acid assemblies. Annu. Rev. Biophys. Biomol. Struct. 1999, 28, 1–27. [Google Scholar] [CrossRef]
- Tsuboi, M.; Overman, S.A.; Thomas, G.J., Jr. Orientation of tryptophan-26 in coat protein subunits of the filamentous virus Ff by polarized Raman microspectroscopy. Biochemistry 1996, 35, 10403–10410. [Google Scholar] [CrossRef]
- Lambert, P.J.; Whitman, A.G.; Dyson, O.F.; Akula, S.M. Raman spectroscopy: The gateway into tomorrow’s virology. Virol. J. 2006, 3, 51. [Google Scholar] [CrossRef] [PubMed]
- Pezzotti, G.; Ohgitani, E.; Shin-Ya, M.; Adachi, T.; Marin, E.; Boschetto, F.; Zhu, W.; Mazda, O. Instantaneous “catch-and-kill” inactivation of SARS-CoV-2 by nitride ceramics. Clin. Transl. Med. 2020, 10, e212. [Google Scholar] [CrossRef]
- Pezzotti, G.; Boschetto, F.; Ohgitani, E.; Fujita, Y.; Shin-Ya, M.; Adachi, T.; Yamamoto, T.; Kanamura, N.; Marin, E.; Zhu, W.; et al. Mechanisms of instantaneous inactivation of SARS-CoV-2 by silicon nitride bioceramic. Mater. Today Bio. 2021, 12, 100144. [Google Scholar] [CrossRef] [PubMed]
- Pezzotti, G.; Ohgitani, E.; Fujita, Y.; Imamura, H.; Shin-Ya, M.; Adachi, T.; Yamamoto, T.; Kanamura, N.; Marin, E.; Zhu, W.; et al. Raman fingerprints of the SARS-CoV-2 Delta variant and mechanisms of its instantaneous inactivation by silicon nitride bioceramics. ACS Infect. Dis. 2022, 8, 1563–1581. [Google Scholar] [CrossRef] [PubMed]
- Pezzotti, G.; Boschetto, F.; Ohgitani, E.; Fujita, Y.; Zhu, W.; Marin, E.; McEntire, B.J.; Bal, B.S.; Mazda, O. Silicon nitride: A potent solid-state bioceramic inactivator of ssRNA viruses. Sci. Rep. 2021, 11, 2977. [Google Scholar] [CrossRef]
- Pezzotti, G.; Boschetto, F.; Ohgitani, E.; Fujita, Y.; Shin-Ya, M.; Adachi, T.; Yamamoto, T.; Kanamura, N.; Marin, E.; Zhu, W.; et al. Raman molecular fingerprints of SARS-CoV-2 British variant and the concept of Raman barcode. Adv. Sci. 2021, 2021, 2103287. [Google Scholar] [CrossRef]
- Pezzotti, G.; Zhu, W.; Adachi, T.; Horiguchi, S.; Marin, E.; Boschetto, F.; Ogitani, E.; Mazda, O. Metabolic machinery encrypted in the Raman spectrum of influenza A virus-inoculated mammalian cells. J. Cell. Physiol. 2019, 235, 5146–5170. [Google Scholar] [CrossRef] [PubMed]
- Qiu, S.; Weng, Y.; Li, Y.; Chen, Y.; Pan, Y.; Liu, J.; Lin, W.; Chen, X.; Li, M.; Lin, T.; et al. Raman profile alterations of irradiated human nasopharyngeal cancer cells detected with laser tweezer Raman spectroscopy. RCS Adv. 2020, 10, 14368–14373. [Google Scholar] [CrossRef] [PubMed]
- Pezzotti, G.; Ohgitani, E.; Ikegami, S.; Shin-Ya, M.; Adachi, T.; Yamamoto, T.; Kanamura, N.; Marin, E.; Zhu, W.; Okuma, K.; et al. Instantaneous inactivation of herpes simplex virus by silicon nitride bioceramics. Int. J. Mol. Struct. 2023, 24, 12657. [Google Scholar] [CrossRef]
- Reyes-Goddard, J.M.; Barr, H.; Stone, N. Surface enhanced Raman scattering of herpes simplex virus in tear film. Photodiagn. Photodyn. Ther. 2008, 5, 42–49. [Google Scholar] [CrossRef]
- Huleihel, M.; Shufan, E.; Zeiri, L.; Salman, A. Detection of Vero cells infected with Herpes simplex Types 1 and 2 and Varicella zoster viruses using Raman spectroscopy and advanced statistical methods. PLoS ONE 2016, 11, e0153599. [Google Scholar] [CrossRef]
- Wiercigroch, E.; Szafraniec, E.; Czamara, K.; Pacia, M.Z.; Majzner, K.; Kochan, K.; Kaczor, A.; Baranska, M.; Malek, K. Raman and infrared spectroscopy of carbohydrates: A review, Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2017, 185, 317–335. [Google Scholar] [CrossRef] [PubMed]
- Aubrey, K.L.; Thomas, G.J., Jr. Raman spectroscopy of filamentous bacteriophage Ff (fd, M13, f1) incorporating specifically-deuterated alanine and tryptophan side chains: Assignments and structural interpretation. Biophys. J. 1991, 60, 1337–1349. [Google Scholar] [CrossRef] [PubMed]
- Pezzotti, G.; Kobara, M.; Nakaya, T.; Imamura, H.; Miyamoto, N.; Adachi, T.; Yamamoto, T.; Kanamura, N.; Ohgitani, E.; Marin, E.; et al. Raman spectroscopy of oral Candida species: Molecular-scale analyses, chemometrics, and barcode identification. Int. J. Mol. Sci. 2022, 23, 5359. [Google Scholar] [CrossRef]
- Pezzotti, G. Raman spectroscopy in cell biology and microbiology. J. Raman Spectrosc. 2021, 52, 2348–2443. [Google Scholar] [CrossRef]
- Zhu, G.; Zhu, X.; Fan, Q.; Wan, X. Raman spectra of amino acids and their aqueous solutions. Spectrochim. Acta Part A 2011, 78, 1187–1195. [Google Scholar] [CrossRef]
- Golichenko, B.O.; Naseka, V.M.; Strelchuk, V.V.; Kolomys, O.F. Raman study of L-Asparagine and L-Glutamine molecules adsorbed on aluminum films in a wide frequency range. Semicond. Phys. Quantum Electron. Optoelectron. 2017, 20, 297–304. [Google Scholar] [CrossRef]
- Hildebrandt, P.G.; Copeland, R.A.; Spiro, T.G.; Otlewski, J.; Laskowski, M., Jr.; Prendergast, F.G. Tyrosine hydrogen-bonding and environmental effects in proteins probed by ultraviolet resonance Raman spectroscopy. Biochemistry 1988, 27, 5426–5433. [Google Scholar] [CrossRef]
- Hernandez, B.; Coic, Y.-M.; Pluger, F.; Kruglik, S.G.; Ghomi, M. All characteristic Raman markers of tyrosine and tyrosinate originate from phenol ring fundamental vibrations. J. Raman Spectrosc. 2016, 47, 210–220. [Google Scholar] [CrossRef]
- Madzharova, F.; Heiner, Z.; Guehlke, M.; Kneipp, J. Surface-enhanced hyper-Raman spectra of adenine, guanine, cytosine, thymine, and uracil. J. Phys. Chem. C 2016, 120, 15415–15423. [Google Scholar] [CrossRef]
- Lopes, R.P.; Marques, M.P.M.; Valero, R.; Tomkinson, J.; de Carvalho, L.A.E.B. Guanine: A combined study using vibrational spectroscopy and theoretical methods. Spectrosc. Int. J. 2012, 27, 273–292. [Google Scholar] [CrossRef]
- Sanchez-Cortes, S.; Garcia-Ramos, J.V. SERS of cytosine and its methylated derivatives on metal colloids. J. Raman Spectrosc. 1992, 23, 61–66. [Google Scholar] [CrossRef]
- Mathlouthi, M.; Seuvre, A.M.; Koenig, J.L.F.t.-i.r. and laser-Raman spectra of cytosine and cytidine. Carbohydr. Res. 1986, 146, 1–13. [Google Scholar] [CrossRef]
- Lopes, R.P.; Valero, R.; Tomkinson, J.; Marques, M.P.M.; de Carvalho, L.A.E.B. Applying vibrational spectroscopy to the study of nucleobases—Adenine as a case study. N. J. Chem. 2013, 37, 2691–2699. [Google Scholar] [CrossRef]
- Notingher, I.; Green, C.; Dyer, C. Discrimination between ricin and sulphur mustard toxicity in vitro using Raman spectroscopy. J. R. Soc. Interface 2004, 1, 79–90. [Google Scholar] [CrossRef]
- Rygula, A.; Majzner, K.; Marzec, K.M.; Kaczor, A.; Pilarczyk, M.; Baranska, M. Raman spectroscopy of proteins: A review. J. Raman Spectrosc. 2013, 44, 1061–1076. [Google Scholar] [CrossRef]
- Pezzotti, G.; Kobara, M.; Nakaya, T.; Imamura, H.; Fujii, T.; Miyamoto, N.; Adachi, T.; Yamamoto, T.; Kanamura, N.; Ohgitani, E.; et al. Raman metabolomics of Candida auris clades: Profiling and barcode identification. Int. J. Mol. Sci. 2022, 23, 11736. [Google Scholar] [CrossRef]
- Pezzotti, G.; Zhu, W.; Chikaguchi, H.; Marin, E.; Boschetto, F.; Masumura, T.; Sato, Y.-I.; Nakazaki, T. Raman molecular fingerprints of rice nutritional quality and the concept of Raman barcode. Front. Nutr. 2021, 8, 663569. [Google Scholar] [CrossRef]
- Pereira, L. Function of glycoprotein B homologues of the family Herpesviridae. Infect. Agents Dis. 1994, 3, 9–28. [Google Scholar]
- Xiao, J.; Palefsky, J.M.; Herrera, R.; Berline, J.; Tugizov, S.M. The Epstein-Barr virus BMRF-2 protein facilitates virus attachment to oral epithelial cells. Virology 2008, 370, 430–442. [Google Scholar] [CrossRef]
- Chowdhary, S.; Deka, R.; Panda, K.; Kumar, R.; Solomon, A.D.; Das, J.; Kanoujiya, S.; Gupta, A.K.; Sinha, S.; Ruokolainen, J.; et al. Recent updates on viral Oncogenesis: Available preventive and therapeutic entities. Mol. Pharm. 2023, In press. [CrossRef] [PubMed]
- DiMaio, F.; Song, Y.; Li, X.; Brunner, M.J.; Xu, C.; Conticello, V.; Egelman, E.; Marlovits, T.C.; Cheng, Y.; Baker, D. Atomic-accuracy models from 4.5-A cryo-electron microscopy data with density-guided iterative local refinement. Nat. Methods 2015, 12, 361–365. [Google Scholar] [CrossRef] [PubMed]
- Snijder, J.; Ortego, M.S.; Weidle, C.; Stuart, A.B.; Gray, M.D.; McElrath, M.J.; Pancera, M.; Veesler, D.; McGuire, A.T. An antibody targeting the fusion machinery neutralizes dual-tropic infection and defines a site of vulnerability on Epstein-Barr virus. Immunity 2018, 48, 799–811. [Google Scholar] [CrossRef] [PubMed]
- Valery, C.; Deville-Foillard, S.; Lefebvre, C.; Taberner, N.; Legrand, P.; Meneau, F.; Meriadec, C.; Delvaux, C.; Bizien, T.; Kasotakis, E.; et al. Atomic view of the histidine environment stabilizing higher-pH conformations of pH-dependent proteins. Nat. Commun. 2015, 6, 7771. [Google Scholar] [CrossRef]
- Backovic, M.; Longnecker, R.; Jardetzky, T.S. Structure of a trimeric variant of the Epstein-Barr virus glycoprotein B. Proc. Natl. Acad. Sci. USA 2009, 106, 2880–2885. [Google Scholar] [CrossRef]
- Backovic, M.; Leser, G.P.; Lamb, R.A.; Longnecker, R.; Jardetzky, T.S. Characterization of EBV gB indicates properties of both class I and class II viral fusion proteins. Virology 2007, 368, 102–113. [Google Scholar] [CrossRef]
- Neuhierl, B.; Feederle, R.; Hammerschmidt, W.; Delecluse, H.J. Glycoprotein gp110 of Epstein-Barr virus determines viral tropism and efficiency of infection. Proc. Natl. Acad. Sci. USA 2002, 99, 15036–15041. [Google Scholar] [CrossRef]
- Miura, T.; Takeuchi, H.; Harada, I. Tryptophan Raman bands sensitive to hydrogen bonding and side-chain conformation. J. Raman Spectrosc. 1989, 20, 667–671. [Google Scholar] [CrossRef]
- Krafft, C.; Neudert, L.; Simat, T.; Salzer, R. Near infrared Raman spectra of human brain lipids. Spectrochim. Acta Part A 2005, 61, 1529–1535. [Google Scholar] [CrossRef]
- Thomas, G.J., Jr. New structural insights from Raman spectroscopy of proteins and their assemblies. Biopolymers 2002, 67, 214–225. [Google Scholar] [CrossRef]
- Bormashenko, E.; Fedorets, A.A.; Dombrovsky, L.A.; Nosonovsky, M. Survival of virus particles in water droplets: Hydrophobic forces and Landauer’s principle. Entropy 2021, 23, 181. [Google Scholar] [CrossRef]
- Weidner-Glunde, M.; Kruminis-Kaszkiel, E.; Savanagouder, M. Herpesviral latency—Common themes. Pathogens 2020, 9, 125. [Google Scholar] [CrossRef] [PubMed]
- Longnecker, R.; Kieff, E.; Cohen, J.I. Epstein–Barr Virus. In Fields Virology, 6th ed.; Knipe, D.M., Howley, P.M., Eds.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2013; Volume 2. [Google Scholar]
- Hernandez, B.; Pflueger, F.; Kruglik, S.G.; Ghomi, M. Characteristic Raman lines of phenylalanine analyzed by a multiconformational approach. J. Raman Spectrosc. 2013, 44, 827–833. [Google Scholar] [CrossRef]
- D’Amico, F.; Cammisuli, F.; Addobbati, R.; Rizzardi, C.; Gessini, A.; Masciovecchio, C.; Rossi, B.; Pascolo, L. Oxidative damage in DNA bases revealed by UV resonant Raman spectroscopy. Analyst 2015, 140, 1477–1485. [Google Scholar] [CrossRef] [PubMed]
- Gill, M.B.; Kutok, J.L.; Fingeroth, J.D. Epstein-Barr virus thymidine kinase is a centrosomal resident precisely localized to the periphery of centrioles. J. Virol. 2007, 81, 6523–6535. [Google Scholar] [CrossRef] [PubMed]
- Rasile, L.; Ghosh, K.; Raviprakash, K.; Ghosh, H.P. Effects of deletions in the carboxy-terminal hydrophobic region of herpes simplex virus glycoprotein gB on intracellular transport and membrane anchoring. J. Virol. 1993, 67, 4856–4866. [Google Scholar] [CrossRef]
- Garcia, N.J.; Chen, J.; Longnecker, R. Modulation of Epstein-Barr virus glycoprotein B (gB) fusion activity by the gB cytoplasmic tail domain. mBio 2013, 4, e00571-12. [Google Scholar] [CrossRef]
- Bagdonaite, I.; Norden, R.; Joshi, H.J.; King, S.L.; Vakhrushev, S.Y.; Olofsson, S.; Wandall, H.H. Global mapping of O-glycosylation of Varicella Zoster virus, Human Cytomegalovirus, and Epstein-Barr virus. J. Biol. Chem. 2016, 292, 12014–12028. [Google Scholar] [CrossRef]
- Sitompul, L.S.; Widodo, N.; Djati, M.S.; Utomo, D.H. Epitope mapping of gp350/220 conserved domain of Epstein-Barr virus to develop nasopharyngeal carcinoma (npc) vaccine. Bioinformation 2012, 8, 479–482. [Google Scholar] [CrossRef]
- Wang, Y.-C.; Lin, V.; Loring, J.F.; Peterson, S.E. The ‘sweet’ spot of cellular pluripotency: Protein glycosylation in human pluripotent stem cells and its applications in regenerative medicine. Exp. Opin. Biol. Ther. 2015, 15, 679–687. [Google Scholar] [CrossRef]
- Arboleda, P.H.; Loppnow, G.R. Raman spectroscopy as a discovery tool in carbohydrate chemistry. Anal. Chem. 2000, 72, 2093–2098. [Google Scholar] [CrossRef] [PubMed]
- Serafini-Cessi, F.; Malagolini, N.; Nanni, M.; Dall’Olio, F.; Fiume, G.C.; Tanner, J.; Kieff, E. Characterization of N- and O-linked oligosaccharides of glycoprotein 350 from Epstein-Barr virus. Virology 1989, 170, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Dennis, J.W.; Laferte’, S.; Waghorne, C.; Breitman, M.L.; Kerbel, R.S. Branching of Asn-linked oligosaccharides is directly associated with metastasis. Science 1987, 236, 582–585. [Google Scholar] [CrossRef]
- Minami, M.; Kita, M.; Yan, X.-Q.; Yamamoto, T.; Iida, T.; Sekigawa, K.; Iwakura, Y.; Imanishi, J. Role of IFN-γ and tumor necrosis Factor-α in Herpes simplex virus Type 1 infection. J. Interferon Cytokine Res. 2002, 22, 671–676. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Ichimura, H.; Iida, T.; Minami, M.; Kobayashi, K.; Kita, M.; Sotozono, C.; Tagawa, Y.-I.; Iwakura, Y.; Imanishi, J. Kinetics of cytokine production in the cornea and trigeminal ganglion of C57BL/6 mice after corneal HSV-a infection. J. Interferon Cytokine Res. 1999, 19, 609–615. [Google Scholar] [CrossRef]
- Salman, A.; Shufan, E.; Zeiri, L.; Huleihel, M. Characterization and detection of Vero cells infected with herpes simplex virus type 1 using Raman spectroscopy and advanced statistical methods. Methods 2014, 68, 364–370. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Ren, L.; Wang, Q.; Wen, Z.; Liu, C.; Ding, Y. Raman spectroscopy: A potential diagnostic tool for oral diseases. Front. Cell. Infect. Microbiol. 2022, 12, 775236. [Google Scholar] [CrossRef] [PubMed]
- Pezzotti, G.; Adachi, T.; Gasparutti, I.; Vincini, G.; Zhu, W.; Boffelli, M.; Rondinella, A.; Marin, E.; Ichioka, H.; Yamamoto, T.; et al. Vibrational monitor of early demineralization in tooth enamel after in vitro exposure to phosphoridic liquid. Spectrochim. Acta A Mol. Biomol. Spectrosc. 2017, 173, 19–33. [Google Scholar] [CrossRef]
- Pezzotti, G.; Adachi, T.; Imamura, H.; Bristol, D.R.; Adachi, K.; Yamamoto, T.; Kanamura, N.; Marin, E.; Zhu, W.; Kawai, T.; et al. In situ Raman study of neurodegenerated human neuroblastoma cells exposed to outer-membrane vesicles isolated from Porphyromonas gingivalis. Int. J. Mol. Sci. 2023, 24, 13351. [Google Scholar] [CrossRef]
- Chaudhary, I.; Jackson, N.; Denning, D.; O’Neill, L.; Byrne, H.J. Contributions of vibrational spectroscopy to virology: A review. Clin. Spectrosc. 2022, 4, 100022. [Google Scholar] [CrossRef]
- Salehi, H.; Ramoji, A.; Mougari, S.; Merida, P.; Neyret, A.; Popp, J.; Muriaux, D.; Horvat, B.; Cuisinier, F. Specific intracellular signature of SARS-CoV-2 infection using confocal Raman microscopy. Commun. Chem. 2022, 5, 85. [Google Scholar] [CrossRef] [PubMed]
- Desai, S.; Mishra, S.V.; Joshi, A.; Sarkar, D.; Hole, A.; Mishra, R.; Dutt, S.; Chilakapati, M.K.; Gupta, S.; Dutt, A. Raman spectroscopy-based detection of RNA viruses in saliva: A preliminary report. J. Biophotonics 2020, 13, e202000189. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, D.; Jakhmola, S.; Pathak, D.K.; Kumar, R.; Jha, H.C. Temporal in vitro Raman spectroscopy for monitoring replication kinetics of Epstein-Barr virus infection in glial cells. ACS Omega 2020, 5, 29547–29560. [Google Scholar] [CrossRef]
- Krishna, C.M.; Sockalingum, G.D.; Kurien, J.; Rao, L.; Venteo, L.; Pluot, M.; Manfait, M.; Kartha, V.B. Micro-Raman spectroscopy for optical pathology of oral squamous cell carcinoma. Appl. Spectrosc. 2004, 58, 1128–1135. [Google Scholar] [CrossRef]
- Mian, S.A.; Yorucu, C.; Ullah, M.S.; Rehman, I.U.; Colley, H.E. Raman spectroscopy can discriminate between normal, dysplastic and cancerous oral mucosa: A tissue-engineering approach. J. Tissue Eng. Regener. Med. 2017, 11, 3253–3262. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Keshavarz, M.; Kassanos, P.; Kidy, Z.; Roddan, A.; Yeatman, E.; Thompson, A.J. SERS detection of breast cancer-derived exosomes using a nanostructured Pt-black template. Adv. Sensor Res. 2023, 2, 2200039. [Google Scholar] [CrossRef]
- Abdali, S.; Refstrup, P.; Nielsen, O.F.; Bohr, H. Enkephalins: Raman spectral analysis and comparison as a function of pH 1-13. Biopolymers 2003, 72, 318–328. [Google Scholar] [CrossRef]
- Michen, B.; Graule, T. Isoelectric points of viruses. J. Appl. Microbiol. 2010, 109, 388–397. [Google Scholar] [CrossRef]
- Hutt-Fletcher, L.M. Epstein-Barr virus entry. J. Virol. 2007, 81, 7825–7832. [Google Scholar] [CrossRef]
- Miller, N.; Hutt-Fletcher, L.M. Epstein-Barr virus enters B cells and epithelial cells by different routes. J. Virol. 1992, 66, 3409–3414. [Google Scholar] [CrossRef]
- White, J.M. Viral and cellular membrane fusion proteins. Annu. Rev. Physiol. 1990, 52, 675–697. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Yeh, Y.-T.; Xue, Y.; Wang, Z.; Zhang, N.; Ricker, R.; Yu, Z.; Roder, A.; Lopez, N.P.; Organtini, L.; et al. Accurate virus identification with interpretable Raman signatures by machine learning. Proc. Natl. Acad. Sci. USA 2022, 119, e2118836119. [Google Scholar] [CrossRef] [PubMed]




| Structural Difference | Raman Fingerprints | Implications |
|---|---|---|
| Carbohydrate signals: stronger in EBV than HSV-1 | 940, 983, 1054, and 1088 cm−1 (C−C & C−O stretching in Glc ring) 1250~1360 cm−1 (CH2 twisting & wagging) | These signals reveal the prominent presence of glycoproteins in EBV |
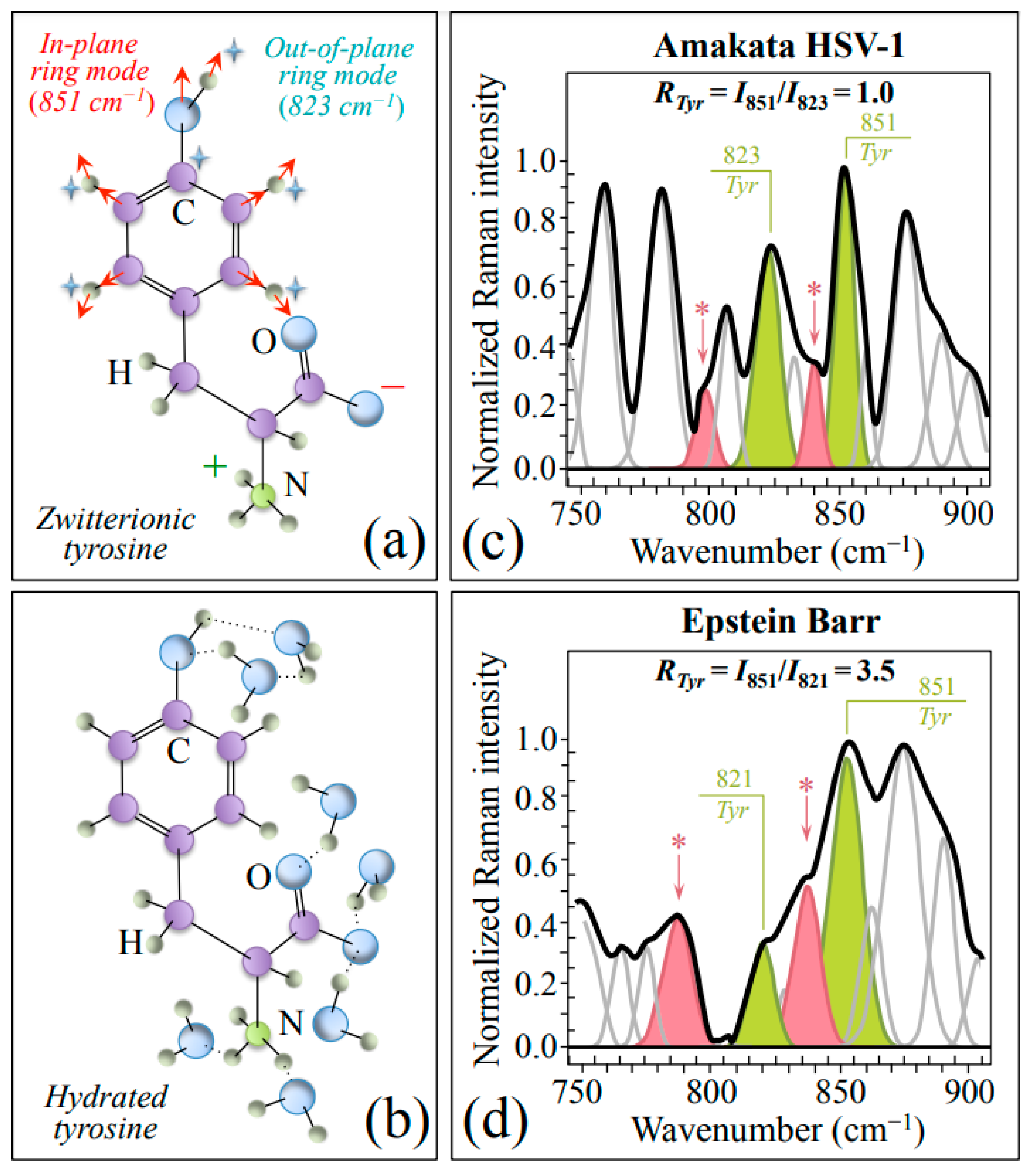
| Tyrosine ratio, RTyr = I851/I823, higher in EBV than HSV-1 | 851/823 cm−1 tyrosine doublet (in-plane & out-of-plane ring vibrations) | Significantly more acidic pH at EBV virion/environment interface |
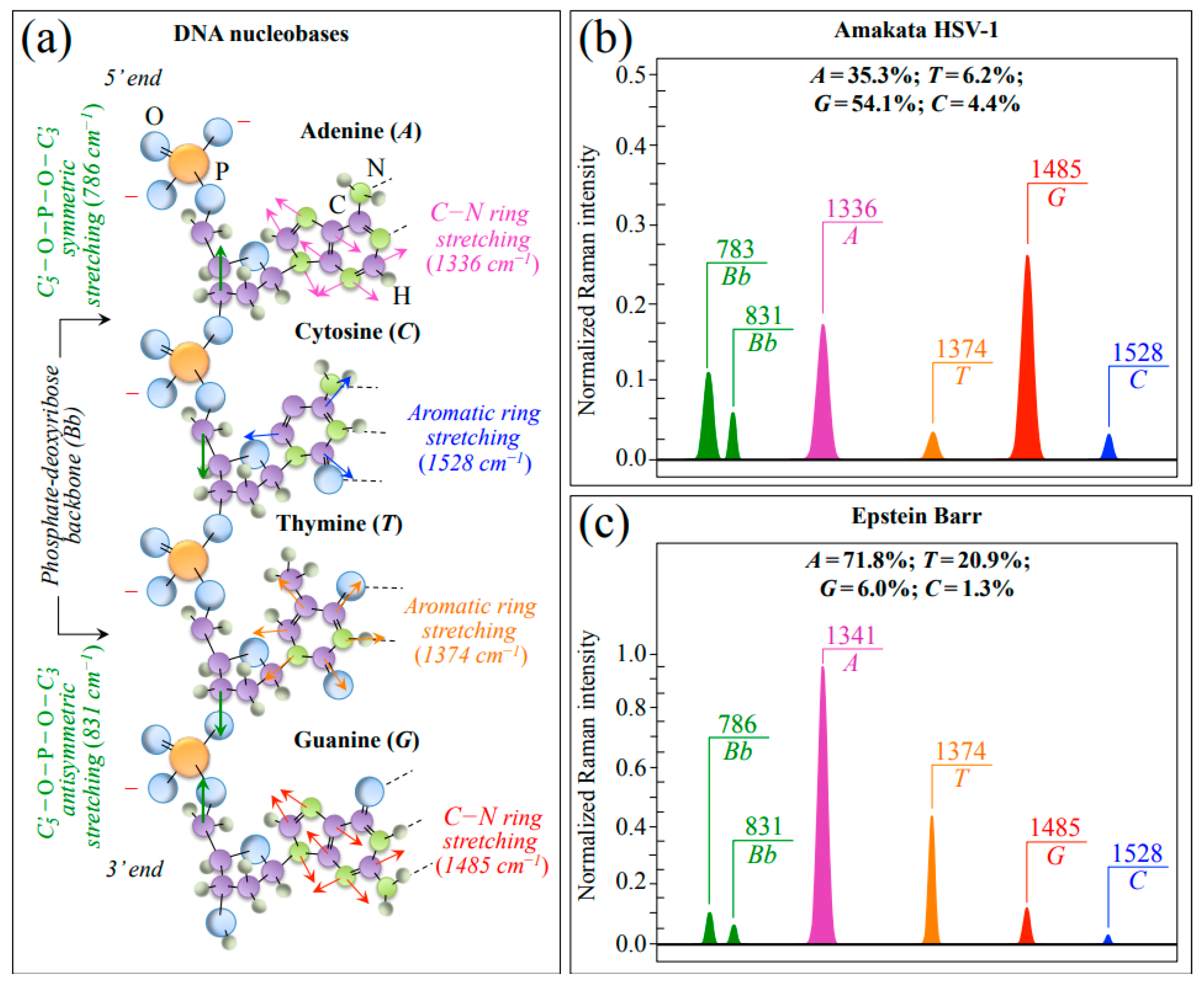
| Differences in Raman genomic structures between EBV and HSV-1 | 1528 (C) &1374 (T) cm−1 (ring stretching in pyrimidines) 1336 (A) & 1485 (G) cm−1 (C−N ring stretching in purines) | Different fractions of purines and pyrimidines as expected from different genome structures |
| Differences in protein secondary structure: β-sheet/β-turn fractions higher in EBV than HSV-1 | Amide I bands: 1634~1636 cm−1 → β-sheet 1653 cm−1 → α-sheet 1669 cm−1 → random coil 1686/1705 cm−1 → β-turn | More rigid protein structure in EBV; it also justifies the stronger threonine signals found in EBV |
| Increased tryptophan torsional angle, χ2,1, in EBV as compared to HSV-1 | 1557 → 1551 cm−1 (C=C stretching in tryptophan pyrrole ring) | Higher hydrophilicity in EBV helps to retain temperature and pH during latency |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pezzotti, G.; Ohgitani, E.; Imamura, H.; Ikegami, S.; Shin-Ya, M.; Adachi, T.; Adachi, K.; Yamamoto, T.; Kanamura, N.; Marin, E.; et al. Raman Multi-Omic Snapshot and Statistical Validation of Structural Differences between Herpes Simplex Type I and Epstein–Barr Viruses. Int. J. Mol. Sci. 2023, 24, 15567. https://doi.org/10.3390/ijms242115567
Pezzotti G, Ohgitani E, Imamura H, Ikegami S, Shin-Ya M, Adachi T, Adachi K, Yamamoto T, Kanamura N, Marin E, et al. Raman Multi-Omic Snapshot and Statistical Validation of Structural Differences between Herpes Simplex Type I and Epstein–Barr Viruses. International Journal of Molecular Sciences. 2023; 24(21):15567. https://doi.org/10.3390/ijms242115567
Chicago/Turabian StylePezzotti, Giuseppe, Eriko Ohgitani, Hayata Imamura, Saki Ikegami, Masaharu Shin-Ya, Tetsuya Adachi, Keiji Adachi, Toshiro Yamamoto, Narisato Kanamura, Elia Marin, and et al. 2023. "Raman Multi-Omic Snapshot and Statistical Validation of Structural Differences between Herpes Simplex Type I and Epstein–Barr Viruses" International Journal of Molecular Sciences 24, no. 21: 15567. https://doi.org/10.3390/ijms242115567







