Sperm RNA Payload: Implications for Intergenerational Epigenetic Inheritance
Abstract
:1. Introduction
2. Small RNA Classes and Their Functions
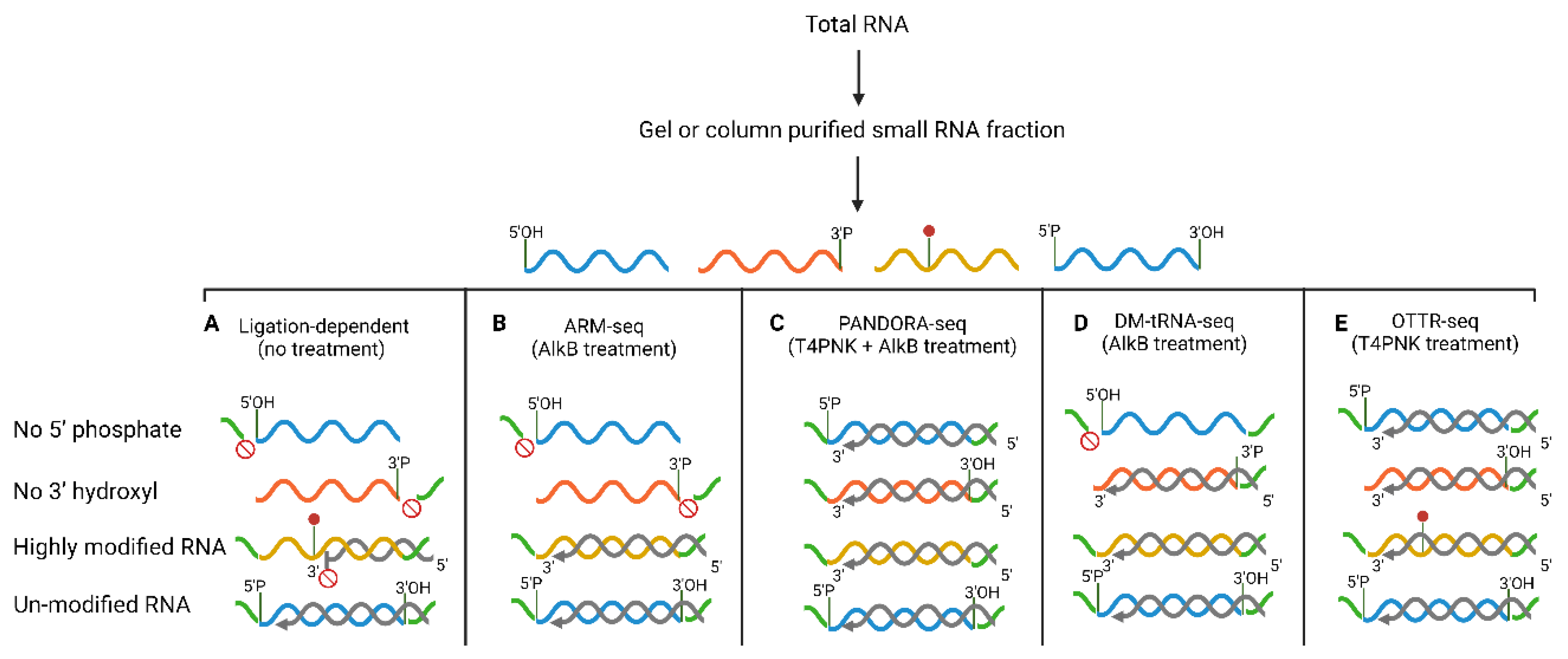
3. Current Methods for Sequencing sRNAs
4. The Mammalian Sperm RNA Payload
The Epididymis Shapes the sRNA Payload of Mature Sperm
5. Paternal Environmental Effects on Offspring Phenotypes
6. Role of Sperm Non-Coding RNAs in Intergenerational Inheritance
6.1. tRNA Fragments
6.2. Micro-RNAs
| Environmental Factors | Tissue (Species) | Altered sRNA Subtype | Primary Detection Method | Citation |
|---|---|---|---|---|
| High-fat diet | F0 Sperm (rat) | 15 miRNAs (let-7c, miR-293, miR-880, etc.), 41 tRFs (tRFGluCTC etc.) and 1092 piRNAs (piR-025883, piR-015935, piR-036085) | NEBNext® Multiplex Small RNA Library Prep Set for Illumina (New England Biolabs, Ipswich, MA, USA) | [129] |
| High-fat diet | F0 Sperm (mouse) | 11.53% tRFs (tRFGluCTC, tRFGlyGCC etc.), and 2.28% miRNAs (miR-3107, miR-142, miR-7210, etc.) | TruSeq Small RNA Sample Prep Kit (Illumina, San Diego, CA, USA) | [16] |
| High-sugar diet | F0 Sperm (mouse) | miR-19 | - | [86] |
| Western diet | RsRNAs (16S, 18S, 28S), tRFs (tRFSerGCT, tRFGluCTC, tRFiMetCAT, miRNAs (miR-10b, miR-10a, miR-125a) | TruSeq Small RNA Sample Pre Kit (Illumina) | [144] | |
| Low-protein diet | F0 Sperm (rat) | tRFGlyGCC, tRFGlyCCC, tRFLys CTT, tRFHisGTG and miRNA Let7 | 3′ and 5′ ligation-dependent protocol | [15] |
| vinclozolin | F3 Sperm (rat) | 13 miRNAs (miR-101, miR-1247 etc.), 16 piRNAs (piR_000216, piR_000333 etc.), 19 Mitochondrial RNAs, 59 tRFs (tRFProAGG, tRF AlaAGC etc.) | Ion RNA Kit v2 (Life Technologies, Carslbad, CA, USA) | [130] |
| DDT | F1, F2, and F3, sperm (rat) | tRFs and piRNAs | NEBNext Multiplex Small RNA Library Prep Set for Illumina | [131] |
| chronic mild stress (CMS) | F0 Sperm (mouse) | miRNAs (miR-199a, miR-103, miR-146a etc.) | TruSeq Small RNA Sample Pre Kit (Illumina) | [105] |
| chronic variable stress (CVS) | F0 Sperm (mouse) | miR-29c, miR-30a, miR-30a, miR-32, miR-193-5p, miR-204, miR-375, miR-532-3p, miR-698 | qRT-PCR | [101] |
| chronic social instability (CSI) | F0 Sperm (mouse) | miR-449, miR-34 | qRT-PCR | [145] |
| unpredictable maternal separation combined with unpredictable maternal stress (MSUS) | F1 Sperm (mouse) | miRNAs (miR-375-3p, miR-375-5p, miR-200b-3p etc.) | modified Illumina v1.5 (3′ and 5′ ligation-dependent protocol) | [133] |
| hypothalamic–pituitary–adrenal (HPA) axis dysregulation by feeding corticosterone (CORT) for 4 weeks | F1 Sperm (mouse) | miRNAs (miR-144, miR-190B, miR-192 etc.) | NA | [134] |
| long-term exercise | F0 Sperm (mouse) | miR-483, miR-431, miR-221, miR-21 | qRT-PCR | [135] |
| short-term exercise | F0 Sperm (mouse) | miR-19b, miR-455 and miR-133a, tRFGly and tRF-Pro | NA | [136] |
6.3. PIWI-Interacting RNAs
6.4. Circular RNAs
6.5. Long RNAs
7. Interplay among Different Epigenetic Markers
8. Role of Sperm sRNAs in Intergenerational Inheritance: Evidence from Human Studies?
9. Conclusions and Future Directions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bartolomei, M.S.; Ferguson-Smith, A.C. Mammalian genomic imprinting. Cold Spring Harb. Perspect. Biol. 2011, 3, a002592. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sharma, U.; Rando, O.J. Metabolic Inputs into the Epigenome. Cell Metab. 2017, 25, 544–558. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Heard, E.; Martienssen, R.A. Transgenerational epigenetic inheritance: Myths and mechanisms. Cell 2014, 157, 95–109. [Google Scholar] [CrossRef] [Green Version]
- Rando, O.J. Daddy issues: Paternal effects on phenotype. Cell 2012, 151, 702–708. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Boskovic, A.; Rando, O.J. Transgenerational Epigenetic Inheritance. Annu. Rev. Genet. 2018, 52, 21–41. [Google Scholar] [CrossRef]
- Miska, E.A.; Ferguson-Smith, A.C. Transgenerational inheritance: Models and mechanisms of non-DNA sequence-based inheritance. Science 2016, 354, 59–63. [Google Scholar] [CrossRef]
- Burton, N.O.; Greer, E.L. Multigenerational epigenetic inheritance: Transmitting information across generations. Semin. Cell Dev. Biol. 2022, 127, 121–132. [Google Scholar] [CrossRef]
- Shi, J.; Zhang, Y.; Tan, D.; Zhang, X.; Yan, M.; Zhang, Y.; Franklin, R.; Shahbazi, M.; Mackinlay, K.; Liu, S.; et al. PANDORA-seq expands the repertoire of regulatory small RNAs by overcoming RNA modifications. Nat. Cell Biol. 2021, 23, 424–436. [Google Scholar] [CrossRef]
- Aravin, A.; Gaidatzis, D.; Pfeffer, S.; Lagos-Quintana, M.; Landgraf, P.; Iovino, N.; Morris, P.; Brownstein, M.J.; Kuramochi-Miyagawa, S.; Nakano, T.; et al. A novel class of small RNAs bind to MILI protein in mouse testes. Nature 2006, 442, 203–207. [Google Scholar] [CrossRef]
- Keam, S.P.; Hutvagner, G. tRNA-Derived Fragments (tRFs): Emerging New Roles for an Ancient RNA in the Regulation of Gene Expression. Life 2015, 5, 1638–1651. [Google Scholar] [CrossRef] [Green Version]
- Shi, J.; Ko, E.A.; Sanders, K.M.; Chen, Q.; Zhou, T. SPORTS1.0: A Tool for Annotating and Profiling Non-coding RNAs Optimized for rRNA- and tRNA-derived Small RNAs. Genom. Proteom. Bioinform. 2018, 16, 144–151. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chan, S.W.; Zilberman, D.; Xie, Z.; Johansen, L.K.; Carrington, J.C.; Jacobsen, S.E. RNA silencing genes control de novo DNA methylation. Science 2004, 303, 1336. [Google Scholar] [CrossRef] [PubMed]
- Volpe, T.A.; Kidner, C.; Hall, I.M.; Teng, G.; Grewal, S.I.; Martienssen, R.A. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 2002, 297, 1833–1837. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sharma, U.; Conine, C.C.; Shea, J.M.; Boskovic, A.; Derr, A.G.; Bing, X.Y.; Belleannee, C.; Kucukural, A.; Serra, R.W.; Sun, F.; et al. Biogenesis and function of tRNA fragments during sperm maturation and fertilization in mammals. Science 2016, 351, 391–396. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, Q.; Yan, M.; Cao, Z.; Li, X.; Zhang, Y.; Shi, J.; Feng, G.H.; Peng, H.; Zhang, X.; Zhang, Y.; et al. Sperm tsRNAs contribute to intergenerational inheritance of an acquired metabolic disorder. Science 2016, 351, 397–400. [Google Scholar] [CrossRef] [Green Version]
- Peng, H.; Shi, J.; Zhang, Y.; Zhang, H.; Liao, S.; Li, W.; Lei, L.; Han, C.; Ning, L.; Cao, Y.; et al. A novel class of tRNA-derived small RNAs extremely enriched in mature mouse sperm. Cell Res. 2012, 22, 1609–1612. [Google Scholar] [CrossRef]
- Aravin, A.A.; Sachidanandam, R.; Girard, A.; Fejes-Toth, K.; Hannon, G.J. Developmentally regulated piRNA clusters implicate MILI in transposon control. Science 2007, 316, 744–747. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Xu, C. Role of microRNAs in mammalian spermatogenesis and testicular germ cell tumors. Reproduction 2015, 149, R127–R137. [Google Scholar] [CrossRef] [Green Version]
- Chen, X.; Li, X.; Guo, J.; Zhang, P.; Zeng, W. The roles of microRNAs in regulation of mammalian spermatogenesis. J. Anim. Sci. Biotechnol. 2017, 8, 35. [Google Scholar] [CrossRef]
- Yuan, S.; Schuster, A.; Tang, C.; Yu, T.; Ortogero, N.; Bao, J.; Zheng, H.; Yan, W. Sperm-borne miRNAs and endo-siRNAs are important for fertilization and preimplantation embryonic development. Development 2015, 143, 635–647. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grivna, S.T.; Beyret, E.; Wang, Z.; Lin, H. A novel class of small RNAs in mouse spermatogenic cells. Genes Dev. 2006, 20, 1709–1714. [Google Scholar] [CrossRef] [Green Version]
- Girard, A.; Sachidanandam, R.; Hannon, G.J.; Carmell, M.A. A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 2006, 442, 199–202. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, T.; Takeda, A.; Tsukiyama, T.; Mise, K.; Okuno, T.; Sasaki, H.; Minami, N.; Imai, H. Identification and characterization of two novel classes of small RNAs in the mouse germline: Retrotransposon-derived siRNAs in oocytes and germline small RNAs in testes. Genes Dev. 2006, 20, 1732–1743. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lau, N.C.; Seto, A.G.; Kim, J.; Kuramochi-Miyagawa, S.; Nakano, T.; Bartel, D.P.; Kingston, R.E. Characterization of the piRNA complex from rat testes. Science 2006, 313, 363–367. [Google Scholar] [CrossRef] [Green Version]
- Li, X.Z.; Roy, C.K.; Dong, X.; Bolcun-Filas, E.; Wang, J.; Han, B.W.; Xu, J.; Moore, M.J.; Schimenti, J.C.; Weng, Z.; et al. An ancient transcription factor initiates the burst of piRNA production during early meiosis in mouse testes. Mol. Cell 2013, 50, 67–81. [Google Scholar] [CrossRef] [Green Version]
- Aravin, A.A.; Sachidanandam, R.; Bourc’his, D.; Schaefer, C.; Pezic, D.; Toth, K.F.; Bestor, T.; Hannon, G.J. A piRNA pathway primed by individual transposons is linked to de novo DNA methylation in mice. Mol. Cell 2008, 31, 785–799. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reuter, M.; Berninger, P.; Chuma, S.; Shah, H.; Hosokawa, M.; Funaya, C.; Antony, C.; Sachidanandam, R.; Pillai, R.S. Miwi catalysis is required for piRNA amplification-independent LINE1 transposon silencing. Nature 2011, 480, 264–267. [Google Scholar] [CrossRef]
- Carmell, M.A.; Girard, A.; van de Kant, H.J.G.; Bourc’his, D.; Bestor, T.H.; de Rooij, D.G.; Hannon, G.J. MIWI2 is essential for spermatogenesis and repression of transposons in the mouse male germline. Dev. Cell 2007, 12, 503–514. [Google Scholar] [CrossRef] [Green Version]
- Wu, P.-H.; Fu, Y.; Cecchini, K.; Özata, D.M.; Arif, A.; Yu, T.; Colpan, C.; Gainetdinov, I.; Weng, Z.; Zamore, P.D. The evolutionarily conserved piRNA-producing locus pi6 is required for male mouse fertility. Nat. Genet. 2020, 52, 728–739. [Google Scholar] [CrossRef]
- Anderson, P.; Ivanov, P. tRNA fragments in human health and disease. FEBS Lett. 2014, 588, 4297–4304. [Google Scholar] [CrossRef] [Green Version]
- Kumar, P.; Kuscu, C.; Dutta, A. Biogenesis and Function of Transfer RNA-Related Fragments (tRFs). Trends Biochem. Sci. 2016, 41, 679–689. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, S.R.; Collins, K. Starvation-induced cleavage of the tRNA anticodon loop in Tetrahymena thermophila. J. Biol. Chem. 2005, 280, 42744–42749. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sobala, A.; Hutvagner, G. Transfer RNA-derived fragments: Origins, processing, and functions. Wiley Interdiscip. Rev. RNA 2011, 2, 853–862. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wilson, B.; Dutta, A. Function and Therapeutic Implications of tRNA Derived Small RNAs. Front. Mol. Biosci. 2022, 9, 888424. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.S.; Shibata, Y.; Malhotra, A.; Dutta, A. A novel class of small RNAs: tRNA-derived RNA fragments (tRFs). Genes Dev. 2009, 23, 2639–2649. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fu, H.; Feng, J.; Liu, Q.; Sun, F.; Tie, Y.; Zhu, J.; Xing, R.; Sun, Z.; Zheng, X. Stress induces tRNA cleavage by angiogenin in mammalian cells. FEBS Lett. 2009, 583, 437–442. [Google Scholar] [CrossRef] [Green Version]
- Thompson, D.M.; Parker, R. The RNase Rny1p cleaves tRNAs and promotes cell death during oxidative stress in Saccharomyces cerevisiae. J. Cell Biol. 2009, 185, 43–50. [Google Scholar] [CrossRef] [Green Version]
- Yamasaki, S.; Ivanov, P.; Hu, G.F.; Anderson, P. Angiogenin cleaves tRNA and promotes stress-induced translational repression. J. Cell Biol. 2009, 185, 35–42. [Google Scholar] [CrossRef] [Green Version]
- Andersen, K.L.; Collins, K. Several RNase T2 enzymes function in induced tRNA and rRNA turnover in the ciliate Tetrahymena. Mol. Biol. Cell 2012, 23, 36–44. [Google Scholar] [CrossRef]
- Zhang, W.; Foo, M.; Eren, A.M.; Pan, T. tRNA modification dynamics from individual organisms to metaepitranscriptomics of microbiomes. Mol. Cell 2022, 82, 891–906. [Google Scholar] [CrossRef]
- Tuorto, F.; Liebers, R.; Musch, T.; Schaefer, M.; Hofmann, S.; Kellner, S.; Frye, M.; Helm, M.; Stoecklin, G.; Lyko, F. RNA cytosine methylation by Dnmt2 and NSun2 promotes tRNA stability and protein synthesis. Nat. Struct. Mol. Biol. 2012, 19, 900–905. [Google Scholar] [CrossRef] [PubMed]
- Blanco, S.; Dietmann, S.; Flores, J.V.; Hussain, S.; Kutter, C.; Humphreys, P.; Lukk, M.; Lombard, P.; Treps, L.; Popis, M.; et al. Aberrant methylation of tRNAs links cellular stress to neuro-developmental disorders. EMBO J. 2014, 33, 2020–2039. [Google Scholar] [CrossRef] [PubMed]
- Schaefer, M.; Pollex, T.; Hanna, K.; Tuorto, F.; Meusburger, M.; Helm, M.; Lyko, F. RNA methylation by Dnmt2 protects transfer RNAs against stress-induced cleavage. Genes Dev. 2010, 24, 1590–1595. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ivanov, P.; Emara, M.M.; Villen, J.; Gygi, S.P.; Anderson, P. Angiogenin-induced tRNA fragments inhibit translation initiation. Mol. Cell 2011, 43, 613–623. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goodarzi, H.; Liu, X.; Nguyen, H.C.; Zhang, S.; Fish, L.; Tavazoie, S.F. Endogenous tRNA-Derived Fragments Suppress Breast Cancer Progression via YBX1 Displacement. Cell 2015, 161, 790–802. [Google Scholar] [CrossRef] [Green Version]
- Sobala, A.; Hutvagner, G. Small RNAs derived from the 5 end of tRNA can inhibit protein translation in human cells. RNA Biol. 2013, 10, 553–563. [Google Scholar] [CrossRef] [Green Version]
- Kim, H.K.; Fuchs, G.; Wang, S.; Wei, W.; Zhang, Y.; Park, H.; Roy-Chaudhuri, B.; Li, P.; Xu, J.; Chu, K.; et al. A transfer-RNA-derived small RNA regulates ribosome biogenesis. Nature 2017, 552, 57–62. [Google Scholar] [CrossRef]
- Elbarbary, R.A.; Takaku, H.; Uchiumi, N.; Tamiya, H.; Abe, M.; Takahashi, M.; Nishida, H.; Nashimoto, M. Modulation of gene expression by human cytosolic tRNase Z(L) through 5′-half-tRNA. PLoS ONE 2009, 4, e5908. [Google Scholar] [CrossRef]
- Saikia, M.; Jobava, R.; Parisien, M.; Putnam, A.; Krokowski, D.; Gao, X.H.; Guan, B.J.; Yuan, Y.; Jankowsky, E.; Feng, Z.; et al. Angiogenin-cleaved tRNA halves interact with cytochrome c, protecting cells from apoptosis during osmotic stress. Mol. Cell. Biol. 2014, 34, 2450–2463. [Google Scholar] [CrossRef] [Green Version]
- Schorn, A.J.; Gutbrod, M.J.; LeBlanc, C.; Martienssen, R. LTR-Retrotransposon Control by tRNA-Derived Small RNAs. Cell 2017, 170, 61–71.e11. [Google Scholar] [CrossRef] [Green Version]
- Chen, X.; Zheng, Y.; Lei, A.; Zhang, H.; Niu, H.; Li, X.; Zhang, P.; Liao, M.; Lv, Y.; Zhu, Z.; et al. Early cleavage of preimplantation embryos is regulated by tRNA(Gln-TTG)-derived small RNAs present in mature spermatozoa. J. Biol. Chem. 2020, 295, 10885–10900. [Google Scholar] [CrossRef] [PubMed]
- Crocker, O.J.; Trigg, N.A.; Conine, C.C. Cloning and Sequencing Eukaryotic Small RNAs. Curr. Protoc. 2022, 2, e495. [Google Scholar] [CrossRef] [PubMed]
- Hafner, M.; Landgraf, P.; Ludwig, J.; Rice, A.; Ojo, T.; Lin, C.; Holoch, D.; Lim, C.; Tuschl, T. Identification of microRNAs and other small regulatory RNAs using cDNA library sequencing. Methods 2008, 44, 3–12. [Google Scholar] [CrossRef] [Green Version]
- Shigematsu, M.; Kawamura, T.; Kirino, Y. Generation of 2′,3′-Cyclic Phosphate-Containing RNAs as a Hidden Layer of the Transcriptome. Front. Genet. 2018, 9, 562. [Google Scholar] [CrossRef] [Green Version]
- Motorin, Y.; Muller, S.; Behm-Ansmant, I.; Branlant, C. Identification of modified residues in RNAs by reverse transcription-based methods. Methods Enzym. 2007, 425, 21–53. [Google Scholar]
- Upton, H.E.; Ferguson, L.; Temoche-Diaz, M.M.; Liu, X.-M.; Pimentel, S.C.; Ingolia, N.T.; Schekman, R.; Collins, K. Low-bias ncRNA libraries using ordered two-template relay: Serial template jumping by a modified retroelement reverse transcriptase. Proc. Natl. Acad. Sci. USA 2021, 118, e2107900118. [Google Scholar] [CrossRef] [PubMed]
- Cozen, A.E.; Quartley, E.; Holmes, A.D.; Hrabeta-Robinson, E.; Phizicky, E.M.; Lowe, T.M. ARM-seq: AlkB-facilitated RNA methylation sequencing reveals a complex landscape of modified tRNA fragments. Nat. Methods 2015, 12, 879–884. [Google Scholar] [CrossRef] [Green Version]
- Zheng, G.; Qin, Y.; Clark, W.C.; Dai, Q.; Yi, C.; He, C.; Lambowitz, A.M.; Pan, T. Efficient and quantitative high-throughput tRNA sequencing. Nat. Methods 2015, 12, 835–837. [Google Scholar] [CrossRef]
- Sharma, U.; Sun, F.; Conine, C.C.; Reichholf, B.; Kukreja, S.; Herzog, V.A.; Ameres, S.L.; Rando, O.J. Small RNAs Are Trafficked from the Epididymis to Developing Mammalian Sperm. Dev. Cell 2018, 46, 481–494.e6. [Google Scholar] [CrossRef]
- Behrens, A.; Rodschinka, G.; Nedialkova, D.D. High-resolution quantitative profiling of tRNA abundance and modification status in eukaryotes by mim-tRNAseq. Mol. Cell 2021, 81, 1802–1815.e7. [Google Scholar] [CrossRef]
- Gustafsson, H.T.; Galan, C.; Yu, T.; Upton, H.E.; Ferguson, L.; Kaymak, E.; Weng, Z.; Collins, K.; Rando, O.J. Deep sequencing of yeast and mouse tRNAs and tRNA fragments using OTTR. bioRxiv 2022. [Google Scholar] [CrossRef]
- Ferguson, L.; Upton, H.E.; Pimentel, S.C.; Mok, A.; Lareau, L.F.; Collins, K.; Ingolia, N.T. Streamlined and sensitive mono- and diribosome profiling in yeast and human cells. bioRxiv 2023. [Google Scholar] [CrossRef]
- Thomas, N.K.; Poodari, V.C.; Jain, M.; Olsen, H.E.; Akeson, M.; Abu-Shumays, R.L. Direct Nanopore Sequencing of Individual Full Length tRNA Strands. ACS Nano 2021, 15, 16642–16653. [Google Scholar] [CrossRef]
- Hao, S.L.; Ni, F.D.; Yang, W.X. The dynamics and regulation of chromatin remodeling during spermiogenesis. Gene 2019, 706, 201–210. [Google Scholar] [CrossRef]
- Gòdia, M.; Castelló, A.; Rocco, M.; Cabrera, B.; Rodríguez-Gil, J.E.; Balasch, S.; Lewis, C.; Sánchez, A.; Clop, A. Identification of circular RNAs in porcine sperm and evaluation of their relation to sperm motility. Sci. Rep. 2020, 10, 7985. [Google Scholar] [CrossRef] [PubMed]
- Chioccarelli, T.; Pierantoni, R.; Manfrevola, F.; Porreca, V.; Fasano, S.; Chianese, R.; Cobellis, G. Histone Post-Translational Modifications and CircRNAs in Mouse and Human Spermatozoa: Potential Epigenetic Marks to Assess Human Sperm Quality. J. Clin. Med. 2020, 9, 640. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, Y.H.; Wang, A.; Song, C.; Shankar, G.; Srivastava, R.K.; Au, K.F.; Li, X.Z. Single-molecule long-read sequencing reveals a conserved intact long RNA profile in sperm. Nat. Commun. 2021, 12, 1361. [Google Scholar] [CrossRef] [PubMed]
- James, E.R.; Carrell, D.T.; Aston, K.I.; Jenkins, T.G.; Yeste, M.; Salas-Huetos, A. The Role of the Epididymis and the Contribution of Epididymosomes to Mammalian Reproduction. Int. J. Mol. Sci. 2020, 21, 5377. [Google Scholar] [CrossRef]
- Bourc’his, D.; Voinnet, O. A Small-RNA Perspective on Gametogenesis, Fertilization, and Early Zygotic Development. Science 2010, 330, 617–622. [Google Scholar] [CrossRef] [Green Version]
- Martinez, G.; Panda, K.; Kohler, C.; Slotkin, R.K. Silencing in sperm cells is directed by RNA movement from the surrounding nurse cell. Nat. Plants 2016, 2, 16030. [Google Scholar] [CrossRef] [PubMed]
- Trigg, N.A.; Eamens, A.L.; Nixon, B. The contribution of epididymosomes to the sperm small RNA profile. Reproduction 2019, 157, R209–R223. [Google Scholar] [CrossRef] [Green Version]
- Sharma, U. Paternal Contributions to Offspring Health: Role of Sperm Small RNAs in Intergenerational Transmission of Epigenetic Information. Front. Cell Dev. Biol. 2019, 7, 215. [Google Scholar] [CrossRef] [PubMed]
- Reilly, J.N.; McLaughlin, E.A.; Stanger, S.J.; Anderson, A.L.; Hutcheon, K.; Church, K.; Mihalas, B.P.; Tyagi, S.; Holt, J.E.; Eamens, A.L.; et al. Characterisation of mouse epididymosomes reveals a complex profile of microRNAs and a potential mechanism for modification of the sperm epigenome. Sci. Rep. 2016, 6, 31794. [Google Scholar] [CrossRef] [Green Version]
- Nixon, B.; Stanger, S.J.; Mihalas, B.P.; Reilly, J.N.; Anderson, A.L.; Tyagi, S.; Holt, J.E.; McLaughlin, E.A. The microRNA signature of mouse spermatozoa is substantially modified during epididymal maturation. Biol. Reprod. 2015, 93, 91. [Google Scholar] [CrossRef]
- Belleannee, C.; Calvo, E.; Caballero, J.; Sullivan, R. Epididymosomes convey different repertoires of microRNAs throughout the bovine epididymis. Biol. Reprod. 2013, 89, 30. [Google Scholar] [CrossRef]
- Twenter, H.; Klohonatz, K.; Davis, K.; Bass, L.; Coleman, S.J.; Bouma, G.J.; Bruemmer, J.E. Transfer of MicroRNAs From Epididymal Epithelium to Equine Spermatozoa. J. Equine Vet. Sci. 2020, 87, 102841. [Google Scholar] [CrossRef]
- Rompala, G.R.; Ferguson, C.; Homanics, G.E. Coincubation of sperm with epididymal extracellular vesicle preparations from chronic intermittent ethanol-treated mice is sufficient to impart anxiety-like and ethanol-induced behaviors to adult progeny. Alcohol 2020, 87, 111–120. [Google Scholar] [CrossRef]
- Chan, J.C.; Morgan, C.P.; Adrian Leu, N.; Shetty, A.; Cisse, Y.M.; Nugent, B.M.; Morrison, K.E.; Jasarevic, E.; Huang, W.; Kanyuch, N.; et al. Reproductive tract extracellular vesicles are sufficient to transmit intergenerational stress and program neurodevelopment. Nat. Commun. 2020, 11, 1499. [Google Scholar] [CrossRef] [Green Version]
- Kaati, G.; Bygren, L.; Edvinsson, S. Cardiovascular and diabetes mortality determined by nutrition during parents’ and grandparents’ slow growth period. Eur. J. Hum. Genet. 2002, 10, 682–688. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bygren, L.O.; Kaati, G.; Edvinsson, S. Longevity determined by paternal ancestors’ nutrition during their slow growth period. Acta Biotheor. 2001, 49, 53–59. [Google Scholar] [CrossRef] [PubMed]
- Pembrey, M.E.; Bygren, L.O.; Kaati, G.; Edvinsson, S.; Northstone, K.; Sjostrom, M.; Golding, J.; Team, A.S. Sex-specific, male-line transgenerational responses in humans. Eur. J. Hum. Genet. 2006, 14, 159–166. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lumey, L.; Stein, A.D.; Kahn, H.S.; van der Pal-de Bruin, K.M.; Blauw, G.; Zybert, P.A.; Susser, E.S. Cohort Profile: The Dutch Hunger Winter Families Study. Int. J. Epidemiol. 2007, 36, 1196–1204. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huypens, P.; Sass, S.; Wu, M.; Dyckhoff, D.; Tschöp, M.; Theis, F.; Marschall, S.; de Angelis, M.H.; Beckers, J. Epigenetic germline inheritance of diet-induced obesity and insulin resistance. Nat. Genet. 2016, 48, 497–499. [Google Scholar] [CrossRef]
- Carone, B.R.; Fauquier, L.; Habib, N.; Shea, J.M.; Hart, C.E.; Li, R.; Bock, C.; Li, C.; Gu, H.; Zamore, P.D.; et al. Paternally induced transgenerational environmental reprogramming of metabolic gene expression in mammals. Cell 2010, 143, 1084–1096. [Google Scholar] [CrossRef] [Green Version]
- Grandjean, V.; Fourre, S.; De Abreu, D.A.; Derieppe, M.A.; Remy, J.J.; Rassoulzadegan, M. RNA-mediated paternal heredity of diet-induced obesity and metabolic disorders. Sci. Rep. 2015, 5, 18193. [Google Scholar] [CrossRef] [Green Version]
- Fullston, T.; Ohlsson Teague, E.M.; Palmer, N.O.; DeBlasio, M.J.; Mitchell, M.; Corbett, M.; Print, C.G.; Owens, J.A.; Lane, M. Paternal obesity initiates metabolic disturbances in two generations of mice with incomplete penetrance to the F2 generation and alters the transcriptional profile of testis and sperm microRNA content. FASEB J. 2013, 27, 4226–4243. [Google Scholar] [CrossRef]
- Watkins, A.J.; Sinclair, K.D. Paternal low protein diet affects adult offspring cardiovascular and metabolic function in mice. Am. J. Physiol. Heart Circ. Physiol. 2014, 306, H1444–H1452. [Google Scholar] [CrossRef] [Green Version]
- Ly, N.H.; Maekawa, T.; Yoshida, K.; Liu, Y.; Muratani, M.; Ishii, S. RNA-Sequencing Analysis of Paternal Low-Protein Diet-Induced Gene Expression Change in Mouse Offspring Adipocytes. G3 Genes Genomes Genet. 2019, 9, 2161–2170. [Google Scholar] [CrossRef] [Green Version]
- Holland, M.L.; Lowe, R.; Caton, P.W.; Gemma, C.; Carbajosa, G.; Danson, A.F.; Carpenter, A.A.; Loche, E.; Ozanne, S.E.; Rakyan, V.K. Early-life nutrition modulates the epigenetic state of specific rDNA genetic variants in mice. Science 2016, 353, 495–498. [Google Scholar] [CrossRef] [Green Version]
- Ng, S.F.; Lin, R.C.; Laybutt, D.R.; Barres, R.; Owens, J.A.; Morris, M.J. Chronic high-fat diet in fathers programs beta-cell dysfunction in female rat offspring. Nature 2010, 467, 963–966. [Google Scholar] [CrossRef]
- Kim, P.; Choi, C.S.; Park, J.H.; Joo, S.H.; Kim, S.Y.; Ko, H.M.; Kim, K.C.; Jeon, S.J.; Park, S.H.; Han, S.-H.; et al. Chronic exposure to ethanol of male mice before mating produces attention deficit hyperactivity disorder-like phenotype along with epigenetic dysregulation of dopamine transporter expression in mouse offspring: ADHD-Like Phenotype Induced by EtOH Exposure to Sire Mice. J. Neurosci. Res. 2014, 92, 658–670. [Google Scholar] [PubMed]
- Vallaster, M.P.; Kukreja, S.; Bing, X.Y.; Ngolab, J.; Zhao-Shea, R.; Gardner, P.D.; Tapper, A.R.; Rando, O.J. Paternal nicotine exposure alters hepatic xenobiotic metabolism in offspring. eLife 2017, 6, e24771. [Google Scholar] [CrossRef]
- Liang, F.; Diao, L.; Liu, J.; Jiang, N.; Zhang, J.; Wang, H.; Zhou, W.; Huang, G.; Ma, D. Paternal ethanol exposure and behavioral abnormities in offspring: Associated alterations in imprinted gene methylation. Neuropharmacology 2014, 81, 126–133. [Google Scholar] [CrossRef] [PubMed]
- Wimmer, M.E.; Briand, L.A.; Fant, B.; Guercio, L.A.; Arreola, A.C.; Schmidt, H.D.; Sidoli, S.; Han, Y.; Garcia, B.A.; Pierce, R.C. Paternal cocaine taking elicits epigenetic remodeling and memory deficits in male progeny. Mol. Psychiatry 2017, 22, 1641–1650. [Google Scholar] [CrossRef] [Green Version]
- McCarthy, D.M.; Morgan, T.J., Jr.; Lowe, S.E.; Williamson, M.J.; Spencer, T.J.; Biederman, J.; Bhide, P.G. Nicotine exposure of male mice produces behavioral impairment in multiple generations of descendants. PLoS Biol. 2018, 16, e2006497. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hawkey, A.B.; White, H.; Pippen, E.; Greengrove, E.; Rezvani, A.H.; Murphy, S.K.; Levin, E.D. Paternal nicotine exposure in rats produces long-lasting neurobehavioral effects in the offspring. Neurotoxicol. Teratol. 2019, 74, 106808. [Google Scholar] [CrossRef]
- Dias, B.G.; Ressler, K.J. Parental olfactory experience influences behavior and neural structure in subsequent generations. Nat. Neurosci. 2014, 17, 89–96. [Google Scholar] [CrossRef] [Green Version]
- Mao, Z.; Xia, W.; Chang, H.; Huo, W.; Li, Y.; Xu, S. Paternal BPA exposure in early life alters Igf2 epigenetic status in sperm and induces pancreatic impairment in rat offspring. Toxicol. Lett. 2015, 238, 30–38. [Google Scholar] [CrossRef]
- Anway, M.D.; Cupp, A.S.; Uzumcu, M.; Skinner, M.K. Epigenetic transgenerational actions of endocrine disruptors and male fertility. Science 2005, 308, 1466–1469. [Google Scholar] [CrossRef] [Green Version]
- Rodgers, A.B.; Morgan, C.P.; Bronson, S.L.; Revello, S.; Bale, T.L. Paternal stress exposure alters sperm microRNA content and reprograms offspring HPA stress axis regulation. J. Neurosci. 2013, 33, 9003–9012. [Google Scholar] [CrossRef] [Green Version]
- Weiss, I.C.; Franklin, T.B.; Vizi, S.; Mansuy, I.M. Inheritable effect of unpredictable maternal separation on behavioral responses in mice. Front. Behav. Neurosci. 2011, 5, 3. [Google Scholar] [CrossRef] [Green Version]
- Dietz, D.M.; Laplant, Q.; Watts, E.L.; Hodes, G.E.; Russo, S.J.; Feng, J.; Oosting, R.S.; Vialou, V.; Nestler, E.J. Paternal transmission of stress-induced pathologies. Biol. Psychiatry 2011, 70, 408–414. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rompala, G.R.; Simons, A.; Kihle, B.; Homanics, G.E. Paternal Preconception Chronic Variable Stress Confers Attenuated Ethanol Drinking Behavior Selectively to Male Offspring in a Pre-Stress Environment Dependent Manner. Front. Behav. Neurosci. 2018, 12, 257. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Chen, Z.-P.; Hu, H.; Lei, J.; Zhou, Z.; Yao, B.; Chen, L.; Liang, G.; Zhan, S.; Zhu, X.; et al. Sperm microRNAs confer depression susceptibility to offspring. Sci. Adv. 2021, 7, eabd7605. [Google Scholar] [CrossRef]
- Morgan, C.P.; Bale, T.L. Early prenatal stress epigenetically programs dysmasculinization in second-generation offspring via the paternal lineage. J. Neurosci. 2011, 31, 11748–11755. [Google Scholar] [CrossRef] [Green Version]
- Wu, L.; Lu, Y.; Jiao, Y.; Liu, B.; Li, S.; Li, Y.; Xing, F.; Chen, D.; Liu, X.; Zhao, J.; et al. Paternal Psychological Stress Reprograms Hepatic Gluconeogenesis in Offspring. Cell Metab. 2016, 23, 735–743. [Google Scholar] [CrossRef] [Green Version]
- Banks, S.; King, S.A.; Irvine, D.S.; Saunders, P.T.K. Impact of a mild scrotal heat stress on DNA integrity in murine spermatozoa. Reproduction 2005, 129, 505–514. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wan, X.; He, X.; Liu, Q.; Wang, X.; Ding, X.; Li, H. Frequent and mild scrotal heat stress in mice epigenetically alters glucose metabolism in the male offspring. Am. J. Physiol. Endocrinol. Metab. 2020, 319, E291–E304. [Google Scholar] [CrossRef]
- Sun, W.; Dong, H.; Becker, A.S.; Dapito, D.H.; Modica, S.; Grandl, G.; Opitz, L.; Efthymiou, V.; Straub, L.G.; Sarker, G.; et al. Cold-induced epigenetic programming of the sperm enhances brown adipose tissue activity in the offspring. Nat. Med. 2018, 24, 1372–1383. [Google Scholar] [CrossRef] [Green Version]
- Lismer, A.; Dumeaux, V.; Lafleur, C.; Lambrot, R.; Brind’Amour, J.; Lorincz, M.C.; Kimmins, S. Histone H3 lysine 4 trimethylation in sperm is transmitted to the embryo and associated with diet-induced phenotypes in the offspring. Dev. Cell 2021, 56, 671–686.e6. [Google Scholar] [CrossRef]
- Xue, J.; Schoenrock, S.A.; Valdar, W.; Tarantino, L.M.; Ideraabdullah, F.Y. Maternal vitamin D depletion alters DNA methylation at imprinted loci in multiple generations. Clin. Epigenet. 2016, 8, 107. [Google Scholar] [CrossRef] [Green Version]
- McPherson, N.O.; Fullston, T.; Kang, W.X.; Sandeman, L.Y.; Corbett, M.A.; Owens, J.A.; Lane, M. Paternal under-nutrition programs metabolic syndrome in offspring which can be reversed by antioxidant/vitamin food fortification in fathers. Sci. Rep. 2016, 6, 27010. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Anderson, L.M.; Riffle, L.; Wilson, R.; Travlos, G.S.; Lubomirski, M.S.; Alvord, W.G. Preconceptional fasting of fathers alters serum glucose in offspring of mice. Nutrition 2006, 22, 327–331. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sellem, E.; Marthey, S.; Rau, A.; Jouneau, L.; Bonnet, A.; Perrier, J.-P.; Fritz, S.; Le Danvic, C.; Boussaha, M.; Kiefer, H.; et al. A comprehensive overview of bull sperm-borne small non-coding RNAs and their diversity across breeds. Epigenet. Chromatin 2020, 13, 19. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McCarrey, J.R.; Lehle, J.D.; Raju, S.S.; Wang, Y.F.; Nilsson, E.E.; Skinner, M.K. Tertiary Epimutations—A Novel Aspect of Epigenetic Transgenerational Inheritance Promoting Genome Instability. PLoS ONE 2016, 11, e0168038. [Google Scholar] [CrossRef] [Green Version]
- Watkins, A.J.; Sirovica, S.; Stokes, B.; Isaacs, M.; Addison, O.; Martin, R.A. Paternal low protein diet programs preimplantation embryo gene expression, fetal growth and skeletal development in mice. Biochim. Biophys. Acta Mol. Basis Dis. 2017, 1863, 1371–1381. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, N.M.; de Oliveira Andrade, F.; Jin, L.; Zhang, X.; Macon, M.; Cruz, M.I.; Benitez, C.; Wehrenberg, B.; Yin, C.; Wang, X.; et al. Maternal intake of high n-6 polyunsaturated fatty acid diet during pregnancy causes transgenerational increase in mammary cancer risk in mice. Breast Cancer Res. 2017, 19, 77. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Assis, S.; Warri, A.; Cruz, M.I.; Laja, O.; Tian, Y.; Zhang, B.; Wang, Y.; Huang, T.H.; Hilakivi-Clarke, L. High-fat or ethinyl-oestradiol intake during pregnancy increases mammary cancer risk in several generations of offspring. Nat. Commun. 2012, 3, 1053. [Google Scholar] [CrossRef] [Green Version]
- Da Cruz, R.S.; Chen, E.; Smith, M.; Bates, J.; de Assis, S. Diet and Transgenerational Epigenetic Inheritance of Breast Cancer: The Role of the Paternal Germline. Front. Nutr. 2020, 7, 93. [Google Scholar] [CrossRef]
- Da Cruz, R.S.; Carney, E.J.; Clarke, J.; Cao, H.; Cruz, M.I.; Benitez, C.; Jin, L.; Fu, Y.; Cheng, Z.; Wang, Y.; et al. Paternal malnutrition programs breast cancer risk and tumor metabolism in offspring. Breast Cancer Res. 2018, 20, 99. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fontelles, C.C.; Carney, E.; Clarke, J.; Nguyen, N.M.; Yin, C.; Jin, L.; Cruz, M.I.; Ong, T.P.; Hilakivi-Clarke, L.; de Assis, S. Paternal overweight is associated with increased breast cancer risk in daughters in a mouse model. Sci. Rep. 2016, 6, 28602. [Google Scholar] [CrossRef] [Green Version]
- Binder, N.K.; Mitchell, M.; Gardner, D.K. Parental diet-induced obesity leads to retarded early mouse embryo development and altered carbohydrate utilisation by the blastocyst. Reprod. Fertil. Dev. 2012, 24, 804–812. [Google Scholar] [CrossRef]
- Krawetz, S.A.; Kruger, A.; Lalancette, C.; Tagett, R.; Anton, E.; Draghici, S.; Diamond, M.P. A survey of small RNAs in human sperm. Hum. Reprod. 2011, 26, 3401–3412. [Google Scholar] [CrossRef] [Green Version]
- Rassoulzadegan, M.; Grandjean, V.; Gounon, P.; Vincent, S.; Gillot, I.; Cuzin, F. RNA-mediated non-mendelian inheritance of an epigenetic change in the mouse. Nature 2006, 441, 469–474. [Google Scholar] [CrossRef] [Green Version]
- Brink, R.A. A Genetic Change Associated with the R Locus in Maize Which Is Directed and Potentially Reversible. Genetics 1956, 41, 872–889. [Google Scholar] [CrossRef]
- Chandler, V.L. Paramutation: From maize to mice. Cell 2007, 128, 641–645. [Google Scholar] [CrossRef] [Green Version]
- Arteaga-Vazquez, M.A.; Chandler, V.L. Paramutation in maize: RNA mediated trans-generational gene silencing. Curr. Opin. Genet. Dev. 2010, 20, 156–163. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Castro Barbosa, T.; Ingerslev, L.R.; Alm, P.S.; Versteyhe, S.; Massart, J.; Rasmussen, M.; Donkin, I.; Sjogren, R.; Mudry, J.M.; Vetterli, L.; et al. High-fat diet reprograms the epigenome of rat spermatozoa and transgenerationally affects metabolism of the offspring. Mol. Metab. 2016, 5, 184–197. [Google Scholar] [CrossRef] [PubMed]
- Schuster, A.; Skinner, M.K.; Yan, W. Ancestral vinclozolin exposure alters the epigenetic transgenerational inheritance of sperm small noncoding RNAs. Env. Epigenet 2016, 2, dvw004. [Google Scholar] [CrossRef] [Green Version]
- Skinner, M.K.; Ben Maamar, M.; Sadler-Riggleman, I.; Beck, D.; Nilsson, E.; McBirney, M.; Klukovich, R.; Xie, Y.; Tang, C.; Yan, W. Alterations in sperm DNA methylation, non-coding RNA and histone retention associate with DDT-induced epigenetic transgenerational inheritance of disease. Epigenet. Chromatin 2018, 11, 8. [Google Scholar] [CrossRef] [Green Version]
- Rodgers, A.B.; Morgan, C.P.; Leu, N.A.; Bale, T.L. Transgenerational epigenetic programming via sperm microRNA recapitulates effects of paternal stress. Proc. Natl. Acad. Sci. USA 2015, 112, 13699–13704. [Google Scholar] [CrossRef] [Green Version]
- Gapp, K.; Jawaid, A.; Sarkies, P.; Bohacek, J.; Pelczar, P.; Prados, J.; Farinelli, L.; Miska, E.; Mansuy, I.M. Implication of sperm RNAs in transgenerational inheritance of the effects of early trauma in mice. Nat. Neurosci. 2014, 17, 667–669. [Google Scholar] [CrossRef] [Green Version]
- Short, A.K.; Fennell, K.A.; Perreau, V.M.; Fox, A.; O’Bryan, M.K.; Kim, J.H.; Bredy, T.W.; Pang, T.Y.; Hannan, A.J. Elevated paternal glucocorticoid exposure alters the small noncoding RNA profile in sperm and modifies anxiety and depressive phenotypes in the offspring. Transl. Psychiatry 2016, 6, e837. [Google Scholar] [CrossRef] [Green Version]
- Murashov, A.K.; Pak, E.S.; Koury, M.; Ajmera, A.; Jeyakumar, M.; Parker, M.; Williams, O.; Ding, J.; Walters, D.; Neufer, P.D. Paternal long-term exercise programs offspring for low energy expenditure and increased risk for obesity in mice. FASEB J. 2016, 30, 775–784. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Short, A.K.; Yeshurun, S.; Powell, R.; Perreau, V.M.; Fox, A.; Kim, J.H.; Pang, T.Y.; Hannan, A.J. Exercise alters mouse sperm small noncoding RNAs and induces a transgenerational modification of male offspring conditioned fear and anxiety. Transl. Psychiatry 2017, 7, e1114. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Boskovic, A.; Bing, X.Y.; Kaymak, E.; Rando, O.J. Control of noncoding RNA production and histone levels by a 5′ tRNA fragment. Genes Dev. 2020, 34, 118–131. [Google Scholar] [CrossRef] [PubMed]
- Macfarlan, T.S.; Gifford, W.D.; Driscoll, S.; Lettieri, K.; Rowe, H.M.; Bonanomi, D.; Firth, A.; Singer, O.; Trono, D.; Pfaff, S.L. Embryonic stem cell potency fluctuates with endogenous retrovirus activity. Nature 2012, 487, 57–63. [Google Scholar] [CrossRef] [Green Version]
- Kiani, J.; Grandjean, V.; Liebers, R.; Tuorto, F.; Ghanbarian, H.; Lyko, F.; Cuzin, F.; Rassoulzadegan, M. RNA–Mediated Epigenetic Heredity Requires the Cytosine Methyltransferase Dnmt2. PLoS Genet. 2013, 9, e1003498. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Zhang, X.; Shi, J.; Tuorto, F.; Li, X.; Liu, Y.; Liebers, R.; Zhang, L.; Qu, Y.; Qian, J.; et al. Dnmt2 mediates intergenerational transmission of paternally acquired metabolic disorders through sperm small non-coding RNAs. Nat. Cell Biol. 2018, 20, 535–540. [Google Scholar] [CrossRef] [Green Version]
- Lee, G.S.; Conine, C.C. The Transmission of Intergenerational Epigenetic Information by Sperm microRNAs. Epigenomes 2022, 6, 12. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S. A Regulator of Metabolic Reprogramming: MicroRNA Let-7. Transl. Oncol. 2019, 12, 1005–1013. [Google Scholar] [CrossRef] [PubMed]
- Fontelles, C.C.; Guido, L.N.; Rosim, M.P.; Andrade Fde, O.; Jin, L.; Inchauspe, J.; Pires, V.C.; de Castro, I.A.; Hilakivi-Clarke, L.; de Assis, S.; et al. Paternal programming of breast cancer risk in daughters in a rat model: Opposing effects of animal- and plant-based high-fat diets. Breast Cancer Res. 2016, 18, 71. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Raad, G.; Serra, F.; Martin, L.; Derieppe, M.A.; Gilleron, J.; Costa, V.L.; Pisani, D.F.; Amri, E.Z.; Trabucchi, M.; Grandjean, V. Paternal multigenerational exposure to an obesogenic diet drives epigenetic predisposition to metabolic diseases in mice. eLife 2021, 10, e61736. [Google Scholar] [CrossRef]
- Dickson, D.A.; Paulus, J.K.; Mensah, V.; Lem, J.; Saavedra-Rodriguez, L.; Gentry, A.; Pagidas, K.; Feig, L.A. Reduced levels of miRNAs 449 and 34 in sperm of mice and men exposed to early life stress. Transl. Psychiatry 2018, 8, 101. [Google Scholar] [CrossRef] [Green Version]
- Ashe, A.; Sapetschnig, A.; Weick, E.M.; Mitchell, J.; Bagijn, M.P.; Cording, A.C.; Doebley, A.L.; Goldstein, L.D.; Lehrbach, N.J.; Le Pen, J.; et al. piRNAs can trigger a multigenerational epigenetic memory in the germline of C. elegans. Cell 2012, 150, 88–99. [Google Scholar] [CrossRef] [Green Version]
- Shirayama, M.; Seth, M.; Lee, H.C.; Gu, W.; Ishidate, T.; Conte, D., Jr.; Mello, C.C. piRNAs initiate an epigenetic memory of nonself RNA in the C. elegans germline. Cell 2012, 150, 65–77. [Google Scholar] [CrossRef] [Green Version]
- Chen, L.L. The expanding regulatory mechanisms and cellular functions of circular RNAs. Nat. Rev. Mol. Cell Biol. 2020, 21, 475–490. [Google Scholar] [CrossRef]
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak, A.; Maier, L.; Mackowiak, S.D.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013, 495, 333–338. [Google Scholar] [CrossRef]
- Tang, C.; Xie, Y.; Yu, T.; Liu, N.; Wang, Z.; Woolsey, R.J.; Tang, Y.; Zhang, X.; Qin, W.; Zhang, Y.; et al. m(6)A-dependent biogenesis of circular RNAs in male germ cells. Cell Res. 2020, 30, 211–228. [Google Scholar] [CrossRef]
- Gapp, K.; Parada, G.E.; Gross, F.; Corcoba, A.; Kaur, J.; Grau, E.; Hemberg, M.; Bohacek, J.; Miska, E.A. Single paternal dexamethasone challenge programs offspring metabolism and reveals multiple candidates in RNA-mediated inheritance. iScience 2021, 24, 102870. [Google Scholar] [CrossRef] [PubMed]
- Chioccarelli, T.; Falco, G.; Cappetta, D.; De Angelis, A.; Roberto, L.; Addeo, M.; Ragusa, M.; Barbagallo, D.; Berrino, L.; Purrello, M.; et al. FUS driven circCNOT6L biogenesis in mouse and human spermatozoa supports zygote development. Cell. Mol. Life Sci. 2022, 79, 50. [Google Scholar] [CrossRef] [PubMed]
- Gapp, K.; van Steenwyk, G.; Germain, P.L.; Matsushima, W.; Rudolph, K.L.M.; Manuella, F.; Roszkowski, M.; Vernaz, G.; Ghosh, T.; Pelczar, P.; et al. Alterations in sperm long RNA contribute to the epigenetic inheritance of the effects of postnatal trauma. Mol. Psychiatry 2020, 25, 2162–2174. [Google Scholar] [CrossRef] [Green Version]
- Alleman, M.; Sidorenko, L.; McGinnis, K.; Seshadri, V.; Dorweiler, J.E.; White, J.; Sikkink, K.; Chandler, V.L. An RNA-dependent RNA polymerase is required for paramutation in maize. Nature 2006, 442, 295–298. [Google Scholar] [CrossRef] [PubMed]
- Maniar, J.M.; Fire, A.Z. EGO-1, a C. elegans RdRP, modulates gene expression via production of mRNA-templated short antisense RNAs. Curr. Biol. 2011, 21, 449–459. [Google Scholar] [CrossRef] [Green Version]
- Rechavi, O.; Lev, I. Principles of Transgenerational Small RNA Inheritance in Caenorhabditis elegans. Curr. Biol. 2017, 27, R720–R730. [Google Scholar] [CrossRef] [Green Version]
- Holoch, D.; Moazed, D. RNA-mediated epigenetic regulation of gene expression. Nat. Rev. Genet. 2015, 16, 71–84. [Google Scholar] [CrossRef] [Green Version]
- Benetti, R.; Gonzalo, S.; Jaco, I.; Muñoz, P.; Gonzalez, S.; Schoeftner, S.; Murchison, E.; Andl, T.; Chen, T.; Klatt, P.; et al. A mammalian microRNA cluster controls DNA methylation and telomere recombination via Rbl2-dependent regulation of DNA methyltransferases. Nat. Struct. Mol. Biol. 2008, 15, 268–279. [Google Scholar] [CrossRef]
- Lai, F.; Shiekhattar, R. Where long noncoding RNAs meet DNA methylation. Cell Res. 2014, 24, 263–264. [Google Scholar] [CrossRef] [Green Version]
- Li, L.-C. Chromatin remodeling by the small RNA machinery in mammalian cells. Epigenetics 2014, 9, 45–52. [Google Scholar] [CrossRef] [Green Version]
- Ingerslev, L.R.; Donkin, I.; Fabre, O.; Versteyhe, S.; Mechta, M.; Pattamaprapanont, P.; Mortensen, B.; Krarup, N.T.; Barrès, R. Endurance training remodels sperm-borne small RNA expression and methylation at neurological gene hotspots. Clin. Epigenet. 2018, 10, 12. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Natt, D.; Kugelberg, U.; Casas, E.; Nedstrand, E.; Zalavary, S.; Henriksson, P.; Nijm, C.; Jaderquist, J.; Sandborg, J.; Flinke, E.; et al. Human sperm displays rapid responses to diet. PLoS Biol. 2019, 17, e3000559. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Donkin, I.; Versteyhe, S.; Ingerslev, L.R.; Qian, K.; Mechta, M.; Nordkap, L.; Mortensen, B.; Appel, E.V.R.; Jørgensen, N.; Kristiansen, V.B.; et al. Obesity and Bariatric Surgery Drive Epigenetic Variation of Spermatozoa in Humans. Cell Metab. 2016, 23, 369–378. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marczylo, E.L.; Amoako, A.A.; Konje, J.C.; Gant, T.W.; Marczylo, T.H. Smoking induces differential miRNA expression in human spermatozoa: A potential transgenerational epigenetic concern? Epigenetics 2012, 7, 432–439. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Lin, J.; Liu, M.; Li, R.; Tian, B.; Zhang, X.; Xu, B.; Liu, M.; Zhang, X.; Li, Y.; et al. Highly sensitive sequencing reveals dynamic modifications and activities of small RNAs in mouse oocytes and early embryos. Sci. Adv. 2016, 2, e1501482. [Google Scholar] [CrossRef] [PubMed] [Green Version]

| Sequencing Methods | Pretreatment of RNA | Sequencing of RNA with 5′OH | Sequencing of RNA with 2′–3′ Cyclic Phosphate/3′ Phosphate | RT through Modified RNA Nucleotides | Small RNA Classes Preferentially Sequenced |
|---|---|---|---|---|---|
| OTTR-seq | −/+ PNK | Yes | Yes | Yes | tRFs, rRNA fragments, miRNAs, and other non-modified small RNA classes |
| DM-tRNA | AlkB (wild type and D135S mutant) | No | No | Yes | tRFs, miRNAs, and other non-modified small RNA classes |
| ARM-seq | AlkB | No | No | No | tRFs, miRNAs, and other non-modified small RNA classes |
| PANDORA-seq | T4 PNK, AlkB | Yes | Yes | No | tRFs, rRNA fragments, miRNAs, and other non-modified small RNA classes |
| Ligation-dependent | No | No | No | No | miRNAs, and other non-modified small RNA classes |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, S.; Sharma, U. Sperm RNA Payload: Implications for Intergenerational Epigenetic Inheritance. Int. J. Mol. Sci. 2023, 24, 5889. https://doi.org/10.3390/ijms24065889
Liu S, Sharma U. Sperm RNA Payload: Implications for Intergenerational Epigenetic Inheritance. International Journal of Molecular Sciences. 2023; 24(6):5889. https://doi.org/10.3390/ijms24065889
Chicago/Turabian StyleLiu, Simeiyun, and Upasna Sharma. 2023. "Sperm RNA Payload: Implications for Intergenerational Epigenetic Inheritance" International Journal of Molecular Sciences 24, no. 6: 5889. https://doi.org/10.3390/ijms24065889








