Whole-Genome Sequences of 13 Chinese Indigenous Pinewood Nematodes, Bursaphelenchus xylophilus
Abstract
:1. Introduction
2. Results
2.1. SNP Distribution in the Chinese Indigenous Pinewood Nematodes

| IDs | Species | Host Tree | Insect Vector | Climate Conditions | Geographic Region (District, Province) | No. of Samples |
|---|---|---|---|---|---|---|
| A1 | Bursaphelenchus xylophilus | Pinus sylvestris var. mongholica | Monochamus | Cold temperate | Hunnan, Shenyang, Liaoning | 1 |
| A2 | Bursaphelenchus xylophilus | Pinus tabuliformis | Monochamus | Cold temperate | Qingyuan, Fushun, Liaoning | 1 |
| A3 | Bursaphelenchus xylophilus | Pinus tabuliformis | Monochamus | Cold temperate | Dongzhou, Fushun, Liaoning | 1 |
| A4 | Bursaphelenchus xylophilus | Pinus tabuliformis | Monochamus | Cold temperate | Xihu, Benxi, Liaoning | 1 |
| A5 | Bursaphelenchus xylophilus | Pinus tabuliformis | Monochamus | Cold temperate | Mingshan, Benxi, Liaoning | 1 |
| A6 | Bursaphelenchus xylophilus | Pinus koraiensis | Monochamus | Cold temperate | Kaiyuan, Tieling, Liaoning | 1 |
| A7 | Bursaphelenchus xylophilus | Pinus tabuliformis | Monochamus | Cold temperate | Tieling, Tieling, Liaoning | 1 |
| A8 | Bursaphelenchus xylophilus | Pinus tabuliformis | Monochamus | Cold temperate | Dengta, Liaoning, Liaoning | 1 |
| A9 | Bursaphelenchus xylophilus | Pinus tabuliformis | Monochamus | Cold temperate | Liaoyang, Liaoning | 1 |
| A10 | Bursaphelenchus xylophilus | Pinus koraiensis | Monochamus | Cold temperate | Fengcheng, Dandong, Liaoning | 1 |
| A11 | Bursaphelenchus xylophilus | Pinus thunbergii | Monochamus | Cold temperate | Changhai, Dalian, Liaoning | 1 |
| B1 | Bursaphelenchus xylophilus | Pinus massoniana | Monochamus | Subtropic | Yingtan, Jianxi | 1 |
| B2 | Bursaphelenchus xylophilus | Pinus massoniana | Monochamus | Subtropic | Anshan, Anhui | 1 |
| Total | 13 |
2.2. Genetic Relationship among Chinese Indigenous Pinewood Nematodes
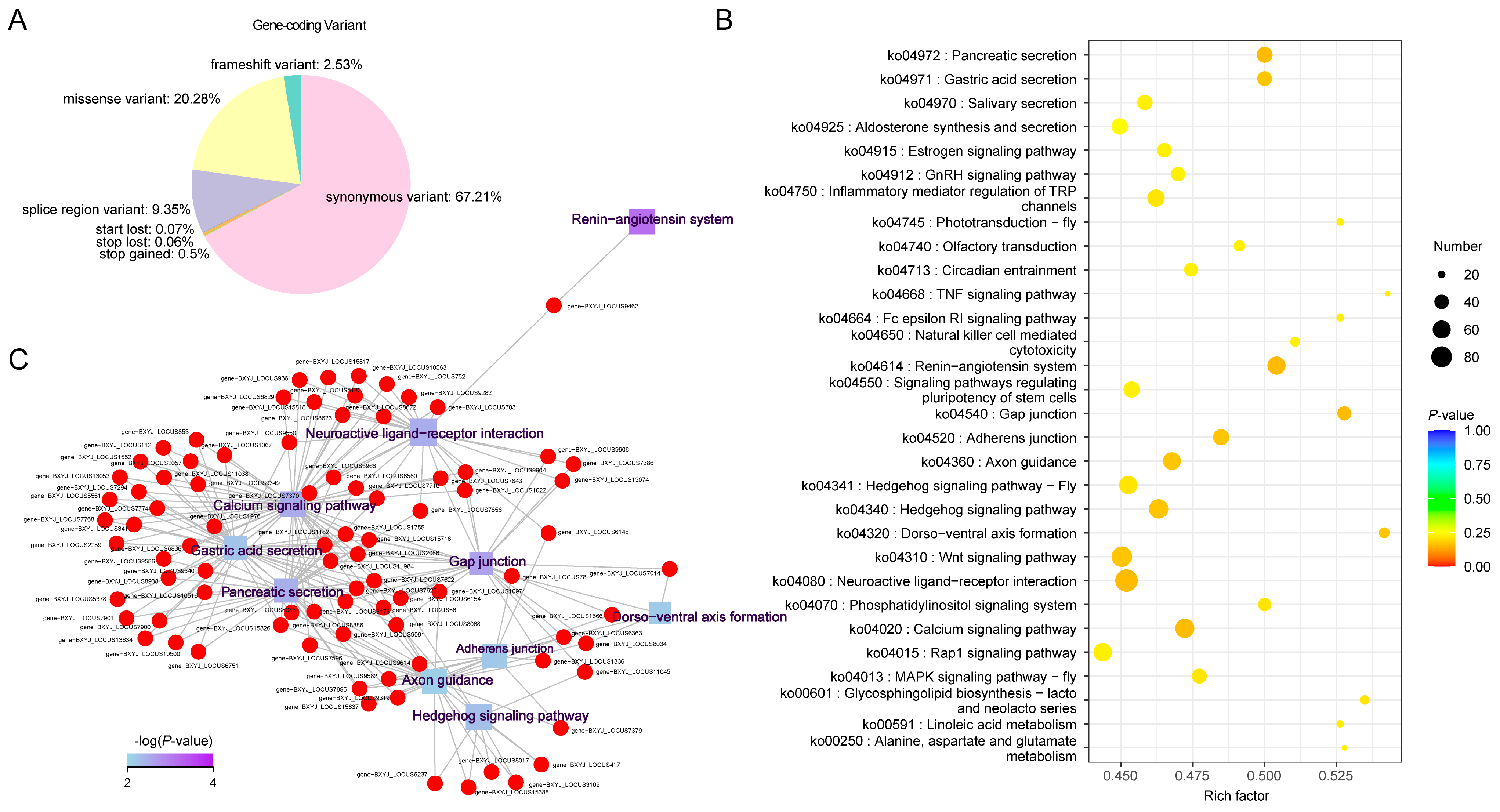
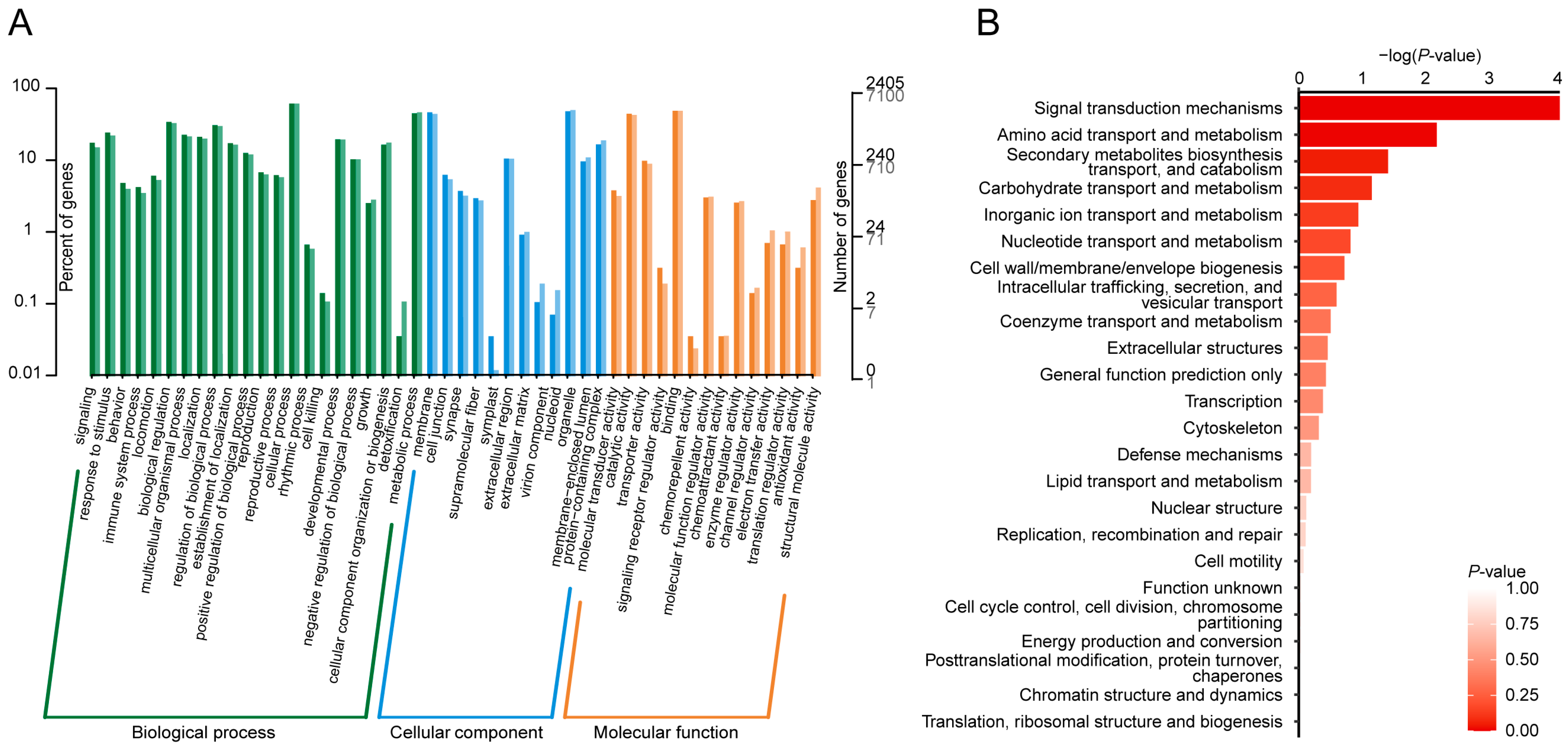
2.3. Annotation of Genetic Variants and Associated Genes
2.4. Specific Genetic Variants in Cold-Temperate Regions
3. Discussion
4. Materials and Methods
4.1. Sampling
4.2. Genomic DNA Isolation, Sequence Library Preparation, and Quality Control
4.3. Short-Read Preprocessing, Variant Calling, and Filtering
4.4. Population Structure Analysis with Principal Component Analysis and Phylogenetic Tree
4.5. Annotation Analysis and Functional Enrichment Analyses
4.6. Annotation Analysis of Specific Genetic Variants in Cold-Temperate Regions
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Tóth, Á. Bursaphelenchus xylophilus, the pinewood nematode: Its significance and a historical review. Acta Biol. Szeged. 2011, 55, 213–217. [Google Scholar]
- Yang, A.; Ding, X.; Feng, Y.; Chen, T.; Ye, J. Genetic Diversity and Population Structure of Bursaphelenchus xylophilus in Central China Based on SNP Markers. Forests 2023, 14, 1443. [Google Scholar] [CrossRef]
- Estoup, A.; Guillemaud, T. Reconstructing routes of invasion using genetic data: Why, how and so what? Mol. Ecol. 2010, 19, 4113–4130. [Google Scholar] [CrossRef]
- Wang, B.; Ma, L.; Wang, F.; Wang, B.; Hao, X.; Xu, J.; Ma, Y. Low temperature extends the lifespan of Bursaphelenchus xylophilus through the cGMP pathway. Int. J. Mol. Sci. 2017, 18, 2320. [Google Scholar] [CrossRef]
- Yang, A.; Ding, X.; Feng, Y.; Zhao, R.; Ye, J. Genetic diversity and genome-wide association analysis of pine wood nematode populations in different regions of China. Front. Plant Sci. 2023, 14, 1183772. [Google Scholar]
- Feng, Y.; Jian, W.; Ding, X.; Ye, J. Genetic Diversity and Population Structure of Bursaphelenchus xylophilus in Guangdong, Guangxi, and Jiangsu Provinces in China. Forests 2024, 15, 934. [Google Scholar] [CrossRef]
- Ding, X.; Guo, Y.; Ye, J.; Wu, X.; Lin, S.; Chen, F.; Zhu, L.; Huang, L.; Song, X.; Zhang, Y. Population differentiation and epidemic tracking of Bursaphelenchus xylophilus in China based on chromosome-level assembly and whole-genome sequencing data. Pest Manag. Sci. 2022, 78, 1213–1226. [Google Scholar] [CrossRef]
- Kikuchi, T.; Cotton, J.A.; Dalzell, J.J.; Hasegawa, K.; Kanzaki, N.; McVeigh, P.; Takanashi, T.; Tsai, I.J.; Assefa, S.A.; Cock, P.J. Genomic insights into the origin of parasitism in the emerging plant pathogen Bursaphelenchus xylophilus. PLoS Pathog. 2011, 7, e1002219. [Google Scholar] [CrossRef] [PubMed]
- Figueiredo, J.; Simoes, M.J.; Gomes, P.; Barroso, C.; Pinho, D.; Conceicao, L.; Fonseca, L.; Abrantes, I.; Pinheiro, M.; Egas, C. Assessment of the geographic origins of pinewood nematode isolates via single nucleotide polymorphism in effector genes. PLoS ONE 2013, 8, e83542. [Google Scholar] [CrossRef]
- Godina, G.; Kirsch, C.; Dörfler, V.; Barg, M.; Singh, P.R.; Vandenbossche, B.; Strauch, O.; Ehlers, R.-U.; Molina, C. Single nucleotide polymorphism markers in Heterorhabditis bacteriophora associated with virulence at low temperature. Nematology 2022, 24, 925–938. [Google Scholar] [CrossRef]
- Smith, H.M.; Smith, B.P.; Morales, N.B.; Moskwa, S.; Clingeleffer, P.R.; Thomas, M.R. SNP markers tightly linked to root knot nematode resistance in grapevine (Vitis cinerea) identified by a genotyping-by-sequencing approach followed by Sequenom MassARRAY validation. PLoS ONE 2018, 13, e0193121. [Google Scholar] [CrossRef] [PubMed]
- Pu, J.; Wang, Z.; Chung, H. Climate change and the genetics of insecticide resistance. Pest Manag. Sci. 2020, 76, 846–852. [Google Scholar] [CrossRef] [PubMed]
- Kirk, H.; Dorn, S.; Mazzi, D. Molecular genetics and genomics generate new insights into invertebrate pest invasions. Evol. Appl. 2013, 6, 842–856. [Google Scholar] [CrossRef]
- Pélissié, B.; Crossley, M.S.; Cohen, Z.P.; Schoville, S.D. Rapid evolution in insect pests: The importance of space and time in population genomics studies. Curr. Opin. Insect Sci. 2018, 26, 8–16. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Sun, J. Pinewood nematode Bursaphelenchus xylophilus (Steiner and buhrer) nickle. In Biological Invasions and Its Management in China; Springer: Singapore, 2017; Volume 2, pp. 3–21. [Google Scholar]
- Pereira, F.; Moreira, C.; Fonseca, L.; van Asch, B.; Mota, M.; Abrantes, I.; Amorim, A. New insights into the phylogeny and worldwide dispersion of two closely related nematode species, Bursaphelenchus xylophilus and Bursaphelenchus mucronatus. PLoS ONE 2013, 8, e56288. [Google Scholar] [CrossRef]
- Akbulut, S.; Stamps, W. Insect vectors of the pinewood nematode: A review of the biology and ecology of Monochamus species. For. Pathol. 2012, 42, 89–99. [Google Scholar] [CrossRef]
- Xiao, Y.; Guo, Q.; Xie, N.; Yuan, G.; Liao, M.; Gui, Q.; Ding, G. Predicting the global potential distribution of Bursaphelenchus xylophilus using an ecological niche model: Expansion trend and the main driving factors. BMC Ecol. Evol. 2024, 24, 48. [Google Scholar] [CrossRef]
- Xu, Q.; Zhang, X.; Li, J.; Ren, J.; Ren, L.; Luo, Y. Pine wilt disease in Northeast and Northwest China: A comprehensive risk review. Forests 2023, 14, 174. [Google Scholar] [CrossRef]
- Mendes, A.V. Characterization of MicroRNAs Implicated in Maritime Pine Resistance to Pinewood Nematode. Master’s Thesis, Instituto Superior de Agronomia, Universidade de Lisboa, Lisbon, Portugal, 2022. [Google Scholar]
- Gregory, T.R. Understanding natural selection: Essential concepts and common misconceptions. Evol. Educ. Outreach 2009, 2, 156–175. [Google Scholar] [CrossRef]
- Toruño, T.Y.; Stergiopoulos, I.; Coaker, G. Plant-pathogen effectors: Cellular probes interfering with plant defenses in spatial and temporal manners. Annu. Rev. Phytopathol. 2016, 54, 419–441. [Google Scholar] [CrossRef]
- Wang, H.; Liu, J.; He, J.; Huang, D.; Xi, Y.; Xiao, T.; Ouyang, Q.; Zhang, S.; Wan, S.; Chen, X. Potential mechanisms underlying the therapeutic roles of sinisan formula in depression: Based on network pharmacology and molecular docking study. Front. Psychiatry 2022, 13, 1063489. [Google Scholar] [CrossRef] [PubMed]
- Bisht, A.; Tewari, D.; Kumar, S.; Chandra, S. Network pharmacology, molecular docking, and molecular dynamics simulation to elucidate the mechanism of anti-aging action of Tinospora cordifolia. Mol. Divers. 2024, 28, 1743–1763. [Google Scholar] [CrossRef] [PubMed]
- Shinya, R.; Morisaka, H.; Kikuchi, T.; Takeuchi, Y.; Ueda, M.; Futai, K. Secretome analysis of the pine wood nematode Bursaphelenchus xylophilus reveals the tangled roots of parasitism and its potential for molecular mimicry. PLoS ONE 2013, 8, e67377. [Google Scholar] [CrossRef]
- Chen, X.; Song, X.; Li, J.; Wang, J.; Yan, Y.; Yang, F. Integrated proteomic, phosphoproteomic, and N-glycoproteomic analyses of small extracellular vesicles from C2C12 myoblasts identify specific PTM patterns in ligand-receptor interactions. Cell Commun. Signal. 2024, 22, 273. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Feng, Y.; Wang, X.; Cui, J.; Deng, X.; Zhang, X. Adaptation of pine wood nematode Bursaphelenchus xylophilus to β-pinene stress. BMC Genom. 2020, 21, 478. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Borneman, J.; Ruegger, P.; Liang, L.; Zhang, K.-Q. Molecular Mechanisms of the Interactions between Nematodes and Nematophagous Microorganisms. In Plant Defence: Biological Control; Springer: Cham, Switzerland, 2020; pp. 421–441. [Google Scholar]
- Ma, J.-E.; Li, L.-M.; Jiang, H.-Y.; Zhang, X.-J.; Li, J.; Li, G.-Y.; Yuan, L.-H.; Wu, J.; Chen, J.-P. Transcriptomic analysis identifies genes and pathways related to myrmecophagy in the Malayan pangolin (Manis javanica). PeerJ 2017, 5, e4140. [Google Scholar] [CrossRef]
- Li, Z.; Shen, L.; Hou, Q.; Zhou, Z.; Mei, L.; Zhao, H.; Wen, X. Identification of genes and metabolic pathways involved in resin yield in masson pine by integrative analysis of transcriptome, proteome and biochemical characteristics. Int. J. Mol. Sci. 2022, 23, 11420. [Google Scholar] [CrossRef]
- Rodrigues, A.M.; Carrasquinho, I.; António, C. Primary metabolite adjustments associated with pinewood nematode resistance in Pinus pinaster. Front. Plant Sci. 2021, 12, 777681. [Google Scholar] [CrossRef]
- Zhang, W.; Zhao, L.; Zhou, J.; Yu, H.; Zhang, C.; Lv, Y.; Lin, Z.; Hu, S.; Zou, Z.; Sun, J. Enhancement of oxidative stress contributes to increased pathogenicity of the invasive pine wood nematode. Philos. Trans. R. Soc. B 2019, 374, 20180323. [Google Scholar] [CrossRef]
- Li, Y.; Meng, F.; Deng, X.; Wang, X.; Feng, Y.; Zhang, W.; Pan, L.; Zhang, X. Comparative transcriptome analysis of the pinewood nematode Bursaphelenchus xylophilus reveals the molecular mechanism underlying its defense response to host-derived α-pinene. Int. J. Mol. Sci. 2019, 20, 911. [Google Scholar] [CrossRef]
- Overgaard, J.; MacMillan, H.A. The integrative physiology of insect chill tolerance. Annu. Rev. Physiol. 2017, 79, 187–208. [Google Scholar] [CrossRef] [PubMed]
- Clark, M.S.; Worland, M.R. How insects survive the cold: Molecular mechanisms—A review. J. Comp. Physiol. B 2008, 178, 917–933. [Google Scholar] [CrossRef]
- Babraham Bioinformatics. FastQC. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 1 August 2024).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Vasimuddin, M.; Misra, S.; Li, H.; Aluru, S. Efficient Architecture-Aware Acceleration of BWA-MEM for Multicore Systems. In Proceedings of the 2019 IEEE International Parallel and Distributed Processing Symposium (IPDPS), Rio de Janeiro, Brazil, 20–24 May 2019; pp. 314–324. [Google Scholar]
- Danecek, P.; Bonfield, J.K.; Liddle, J.; Marshall, J.; Ohan, V.; Pollard, M.O.; Whitwham, A.; Keane, T.; McCarthy, S.A.; Davies, R.M.; et al. Twelve years of SAMtools and BCFtools. GigaScience 2021, 10, giab008. [Google Scholar] [CrossRef] [PubMed]
- Van der Auwera, G.A.; Carneiro, M.O.; Hartl, C.; Poplin, R.; Del Angel, G.; Levy-Moonshine, A.; Jordan, T.; Shakir, K.; Roazen, D.; Thibault, J.; et al. From FastQ data to high confidence variant calls: The Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinform. 2013, 43, 11.10.1–11.10.33. [Google Scholar] [CrossRef] [PubMed]
- Broad Institute. Picard Toolkit. 2019. Available online: https://broadinstitute.github.io/picard/ (accessed on 1 August 2024).
- Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 2011, 27, 2987–2993. [Google Scholar] [CrossRef]
- Gu, Z.; Gu, L.; Eils, R.; Schlesner, M.; Brors, B. circlize implements and enhances circular visualization in R. Bioinformatics 2014, 30, 2811–2812. [Google Scholar] [CrossRef]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.I.; Daly, M.J.; et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef]
- Villanueva, R.A.M.; Chen, Z.J. ggplot2: Elegant Graphics for Data Analysis; Taylor & Francis: London, UK, 2019. [Google Scholar]
- Paradis, E.; Claude, J.; Strimmer, K. APE: Analyses of phylogenetics and evolution in R language. Bioinformatics 2004, 20, 289–290. [Google Scholar] [CrossRef]
- Cingolani, P.; Platts, A.; Wang, L.L.; Coon, M.; Nguyen, T.; Wang, L.; Land, S.J.; Lu, X.; Ruden, D.M. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly 2012, 6, 80–92. [Google Scholar] [CrossRef]
- Sayers, E.W.; Bolton, E.E.; Brister, J.R.; Canese, K.; Chan, J.; Comeau, D.C.; Connor, R.; Funk, K.; Kelly, C.; Kim, S. Database resources of the national center for biotechnology information. Nucleic Acids Res. 2022, 50, D20. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.; Hu, E.; Xu, S.; Chen, M.; Guo, P.; Dai, Z.; Feng, T.; Zhou, L.; Tang, W.; Zhan, L. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation 2021, 2, 100141. [Google Scholar] [CrossRef] [PubMed]

| Gene Symbol | Description | Annotation | Variant Type |
|---|---|---|---|
| Bm1-29045 | TKR78190.1 hypothetical protein | Genetic information processing; Translation | Stop gained, frame shifted |
| F59C6.8 | TKR58295.1 hypothetical protein | − * | Frameshift variant |
| Tag-151 | KAE9554954.1 hypothetical protein | − | Frameshift variant |
| Vha-12 | KHJ82218.1 hypothetical protein | Metabolism; Environmental information Processing; cellular processes | Frameshift variant |
| At1g67520 | VDN83815.1 unnamed protein product | − | Frameshift variant |
| Ugt-58 | KAE9553687.1 hypothetical protein | Metabolism | Frameshift variant |
| OatA | XP_003093275.1 hypothetical protein | − | Frameshift variant |
| Lys-3 | XP_024506972.1 Glycoside hydrolase | − | Frameshift variant |
| SETMAR | AAZ67092.1 transposase | Metabolism | Frameshift variant |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dong, B.; Wu, H.; Li, D.; Luo, Z.; He, S.; Hao, X.; Gao, J. Whole-Genome Sequences of 13 Chinese Indigenous Pinewood Nematodes, Bursaphelenchus xylophilus. Int. J. Mol. Sci. 2024, 25, 10492. https://doi.org/10.3390/ijms251910492
Dong B, Wu H, Li D, Luo Z, He S, Hao X, Gao J. Whole-Genome Sequences of 13 Chinese Indigenous Pinewood Nematodes, Bursaphelenchus xylophilus. International Journal of Molecular Sciences. 2024; 25(19):10492. https://doi.org/10.3390/ijms251910492
Chicago/Turabian StyleDong, Bo, Hao Wu, Debin Li, Zaiquan Luo, Shan He, Xin Hao, and Junxin Gao. 2024. "Whole-Genome Sequences of 13 Chinese Indigenous Pinewood Nematodes, Bursaphelenchus xylophilus" International Journal of Molecular Sciences 25, no. 19: 10492. https://doi.org/10.3390/ijms251910492






