Machine Learning-Based Classification of Transcriptome Signatures of Non-Ulcerative Bladder Pain Syndrome
Abstract
1. Introduction
2. Results
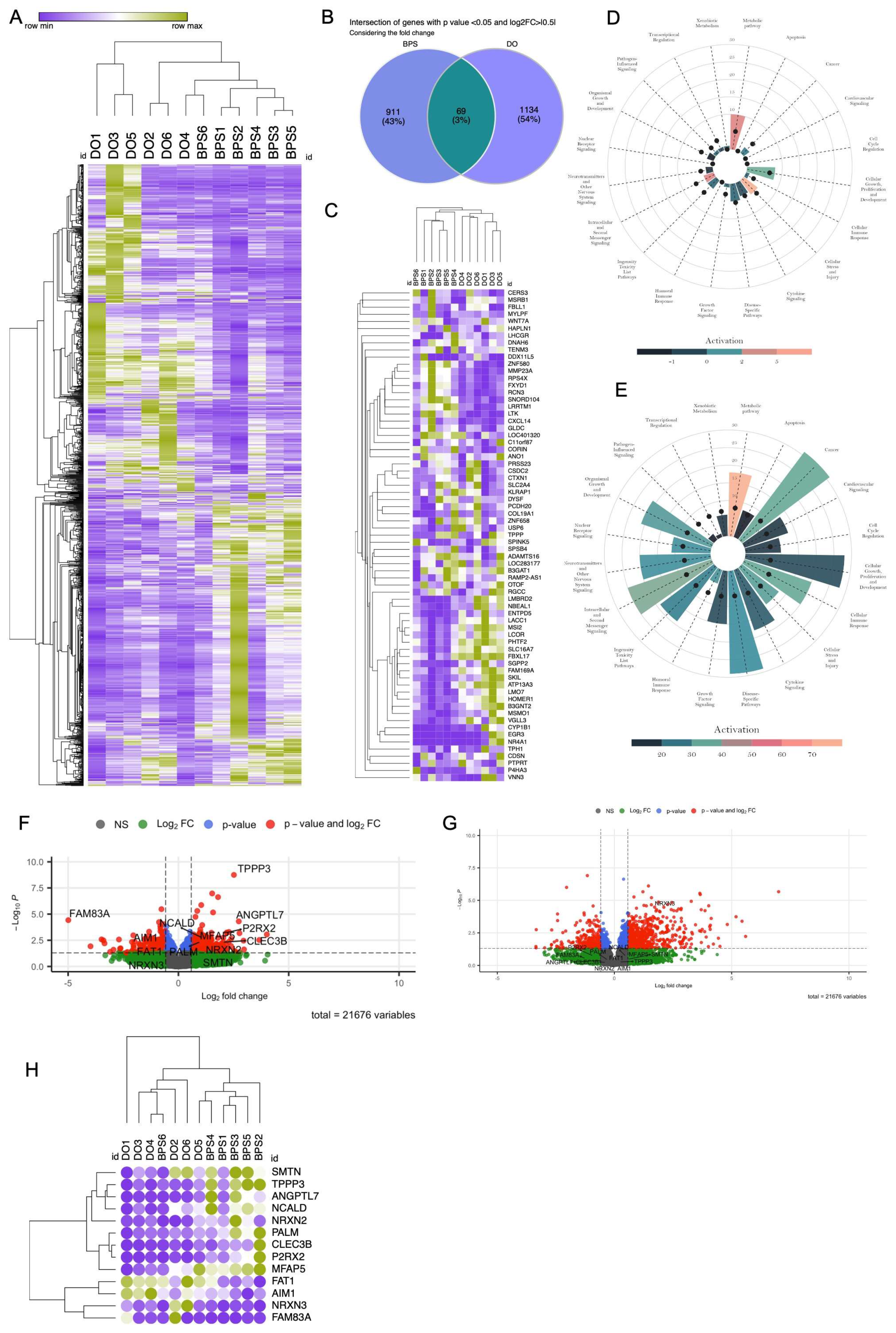
2.1. Transcriptomic Difference between BPS and DO
2.1.1. Differential Expression Analysis
2.1.2. Functional Enrichment Analysis
2.2. Validation of RNA-Seq Data Using QPCR
2.2.1. QPCR Gene Panel Selection Criteria
2.2.2. Statistical Examination of Selected Genes
- Normality Test on RNA Seq Data: According to the central limit theorem, the sampling distribution tends to be normal if the sample is large enough (n > 30). However, our sample size for RNAseq is smaller (n = 6); therefore, normality was checked by visual inspection [histogram plots, Q–Q plot (quantile-quantile plot)] and by significance tests. Using normalized read counts in histogram plots, Q–Q plots, and the Shapiro–Wilk normality test, we established that the distribution of the data was significantly different from normal (Figure S1B). Based on read counts in all groups (BPS, DO, and control), we visualized regulation for each of the selected genes in the NGS dataset (Figure 2). After performing the Kruskal–Wallis test, we showed that TPPP3, SMTN, ANDPTL7, NCALD, and P2RX2 are up-regulated in BPS compared to control and DO; FAT1, AIM1, and FAM83A are down-regulated; CLEC3B and PALM are higher in BPS than in DO; and NRXN3 is up-regulated in DO compared to BPS and control (Figure 2).
- Empirical Cumulative Distribution Function (ECDF) Analysis: ECDF is closely related to cumulative frequency and provides an alternative visualization of distribution. It reports for any given number the percentage of individuals that are below a set threshold. We applied this function to NGS read count data for the 13 selected marker genes and report an excellent separation in distribution of reads between BPS, DO, and controls for some genes (TPPP3, FAT1, ANGPTL7, AIM1, PALM, NCALD, P2RX2), but not others (SMTN, NRXN3, NRXN2, FAM83A, MFAP5) (Figure 3).
- Z-score-based Patient Grouping: To assess the potential utility of the 13 chosen genes for categorizing patients based on their LUTD type, we computed a patient z-score for each gene. This score represents the deviation of an expression value from the mean expression of that gene across all patients. Patients were then categorized into groups based on their calculated z-scores. A patient with a z-score >0 was classified as High (indicated by red bars in Figure S3), while a z-score <0 designated the patient as Low (indicated by green bars in Figure S3). Upon comparing z-score for patients with DO and BPS, it becomes evident that the selected genes effectively segregate the samples based on the type of LUTD. High z-scores are observed in BPS patients for TPPP3, ANGPTL7, CLEC3B, PALM, NCALD, and P2RX2. Conversely, low z-scores are observed in BPS patients for FAT1, AIM1, and NRXN3. Z-scores do not effectively distinguish between BPS and DO groups based on SMTN, FAM83A, and MFAP5 genes (Figure S3).
- Correlation Analysis: We used a correlogram to determine the relationship between different attributes (genes). Figure S4A shows correlation with significance values added. FAM83A, FAT1, and AM1 showed an opposite relationship or non-significant correlation to other genes, whereas the rest of the selected genes had a positive correlation to other genes. In particular, MFAP5 showed strong correlation to CLEC3B, PALM, SMTN, TPPP3, and NCALD (Figure S4A).
- Principal Component Analysis (PCA): We conducted principal component analysis (PCA) on 18 patients, divided into three groups (BPS, DO, and control, with n = 6 in each group). The analysis was based on the NGS read counts of 13 selected genes. PCA affirmed that these 13 genes have the capability to distinctly cluster all patients according to their LUTD type (Figure 4A). The scree plot further illustrates that PC1 captured 59.9% of the total variance, while PC2 captured 13.7% of the variance (Figure S4B). Among the 13 genes, MFAP5, PALM, NCALD, TPPP3, and CLEC3B significantly contributed to PC1 (over 10% each), while NRXN3, FAM83A, P2RX2, and ANGPTL7 were the primary contributors to PC2 (Figure S4C).
- Clustering Analysis: The hierarchical clustering algorithm was applied to the Next-Generation Sequencing (NGS) read counts of 13 selected genes. The resulting tree was divided into k clusters. We specifically investigated whether setting k = 3, corresponding to the known groups (BPS, DO, and Control), accurately represented the grouping. The hierarchical cluster dendrogram grouped 18 samples into three distinct clusters: one comprising all BPS samples and one DO sample, another consisting solely of DO samples, and a third including all control samples along with two DO samples (Figure 4B). Furthermore, we utilized both the Elbow method (Figure 4C) and Clustree (Figure 4D) to determine the optimal number of clusters for the given datasets. Both methods consistently identified n = 3 as the optimal number of clusters. We delved deeper into sample clustering using the k-means partitioning clustering method (Figure 4E). Our observations revealed that setting k = 3 effectively separated NGS patients into three groups. Specifically, DO1, DO4, and DO3 clustered together, while DO2, DO5, and DO6 clustered with control samples. Notably, all BPS patients formed a distinct cluster, separate from both control and DO patients.
2.2.3. QPCR Validation of Selected Genes in a Larger Patient Cohort
2.3. Identification of BPS mRNA Signatures through Unsupervised ML Analysis of QPCR Data
2.4. Supervised Machine Learning
2.4.1. Addressing Imbalanced Small Sample Sizes in Multi-Class Classification
2.4.2. ML Model Evaluation

2.4.3. Identification of mRNA Signatures Based on QPCR Data Using Feature Selection Technique
2.4.4. Visualization of Selected mRNA Signatures
3. Discussion
4. Materials and Methods
4.1. Data Depository
4.2. Study Approval
4.3. Patient Selection
4.4. Functional Enrichment Analysis
4.5. QPCR Validation of NGS Studies
4.6. Statistics
4.6.1. Hierarchical Clustering and Heatmaps
4.6.2. Principal Component Analysis (PCA)
4.6.3. D Ellipsoid Chart and Point Identification for Biomarkers
4.7. Deviation Graphs
4.8. Hierarchical Clustering Algorithm
4.9. Prediction of Biological Function of Canonical Pathways
4.10. Running Score and Preranked List of GSEA Result
4.11. Partitioning Clustering Method
4.12. Proposed ML Pipeline
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Irwin, D.E.; Kopp, Z.S.; Agatep, B.; Milsom, I.; Abrams, P. Worldwide prevalence estimates of lower urinary tract symptoms, overactive bladder, urinary incontinence and bladder outlet obstruction. BJU Int. 2011, 108, 1132–1138. [Google Scholar] [CrossRef] [PubMed]
- Durden, E.; Walker, D.; Gray, S.; Fowler, R.; Juneau, P.; Gooch, K. The economic burden of overactive bladder (OAB) and its effects on the costs associated with other chronic, age-related comorbidities in the United States. Neurourol. Urodyn. 2018, 37, 1641–1649. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.H.; Wu, H.C.; Tseng, C.M.; Ko, T.L.; Weng, T.J.; Chen, Y.F. Health Education and Symptom Flare Management Using a Video-based m-Health System for Caring Women With IC/BPS. Urology 2018, 119, 62–69. [Google Scholar] [CrossRef] [PubMed]
- Di Bello, F.; Scandurra, C.; Muzii, B.; Colla’ Ruvolo, C.; Califano, G.; Mocini, E.; Creta, M.; Napolitano, L.; Morra, S.; Fraia, A.; et al. Are Excessive Daytime Sleepiness and Lower Urinary Tract Symptoms the Triggering Link for Mental Imbalance? An Exploratory Post Hoc Analysis. J. Clin. Med. 2023, 12, 6965. [Google Scholar] [CrossRef] [PubMed]
- Morra, S.; Collà Ruvolo, C.; Napolitano, L.; La Rocca, R.; Celentano, G.; Califano, G.; Creta, M.; Capece, M.; Turco, C.; Cilio, S.; et al. YouTube(TM) as a source of information on bladder pain syndrome: A contemporary analysis. Neurourol. Urodyn. 2022, 41, 237–245. [Google Scholar] [CrossRef] [PubMed]
- Hepner, K.A.; Watkins, K.E.; Elliott, M.N.; Clemens, J.Q.; Hilton, L.G.; Berry, S.H. Suicidal ideation among patients with bladder pain syndrome/interstitial cystitis. Urology 2012, 80, 280–285. [Google Scholar] [CrossRef] [PubMed]
- Moss, M.C.; Rezan, T.; Karaman, U.R.; Gomelsky, A. Treatment of Concomitant OAB and BPH. Curr. Urol. Rep. 2017, 18, 1. [Google Scholar] [CrossRef]
- Oelke, M.; Baard, J.; Wijkstra, H.; de la Rosette, J.J.; Jonas, U.; Hofner, K. Age and bladder outlet obstruction are independently associated with detrusor overactivity in patients with benign prostatic hyperplasia. Eur. Urol. 2008, 54, 419–426. [Google Scholar] [CrossRef]
- Oh, M.M.; Choi, H.; Park, M.G.; Kang, S.H.; Cheon, J.; Bae, J.H.; Moon du, G.; Kim, J.J.; Lee, J.G. Is there a correlation between the presence of idiopathic detrusor overactivity and the degree of bladder outlet obstruction? Urology 2011, 77, 167–170. [Google Scholar] [CrossRef]
- Monastyrskaya, K.; Burkhard, F.C. Urinary Biomarkers for Bladder Outlet Obstruction. Curr. Bladder Dysfunct. Rep. 2017, 12, 129–137. [Google Scholar] [CrossRef]
- Gheinani, A.H.; Kiss, B.; Moltzahn, F.; Keller, I.; Bruggmann, R.; Rehrauer, H.; Fournier, C.A.; Burkhard, F.C.; Monastyrskaya, K. Characterization of miRNA-regulated networks, hubs of signaling, and biomarkers in obstruction-induced bladder dysfunction. JCI Insight 2017, 2, e89560. [Google Scholar] [CrossRef][Green Version]
- Gheinani, A.H.; Akshay, A.; Besic, M.; Kuhn, A.; Keller, I.; Bruggmann, R.; Rehrauer, H.; Adam, R.M.; Burkhard, F.C.; Monastyrskaya, K. Integrated mRNA-miRNA transcriptome analysis of bladder biopsies from patients with bladder pain syndrome identifies signaling alterations contributing to the disease pathogenesis. BMC Urol. 2021, 21, 172. [Google Scholar] [CrossRef]
- Grundy, L.; Caldwell, A.; Brierley, S.M. Mechanisms Underlying Overactive Bladder and Interstitial Cystitis/Painful Bladder Syndrome. Front. Neurosci. 2018, 12, 931. [Google Scholar] [CrossRef]
- Waegeman, W.; Verwaeren, J.; Slabbinck, B.; Baets, B.D. Supervised learning algorithms for multi-class classification problems with partial class memberships. Fuzzy Sets Syst. 2011, 184, 106–125. [Google Scholar] [CrossRef]
- Krawczyk, B. Learning from imbalanced data: Open challenges and future directions. Prog. Artif. Intell. 2016, 5, 221–232. [Google Scholar] [CrossRef]
- Kraiem, M.S.; Sánchez-Hernández, F.; Moreno-García, M.N. Selecting the Suitable Resampling Strategy for Imbalanced Data Classification Regarding Dataset Properties. An Approach Based on Association Models. Appl. Sci. 2021, 11, 8546. [Google Scholar] [CrossRef]
- Chawla, N.V. Data Mining for Imbalanced Datasets: An Overview. In Data Mining and Knowledge Discovery Handbook; Maimon, O., Rokach, L., Eds.; Springer: Boston, MA, USA, 2005; pp. 853–867. [Google Scholar] [CrossRef]
- Akshay, A.; Abedi, M.; Shekarchizadeh, N.; Burkhard, F.C.; Katoch, M.; Bigger-Allen, A.; Adam, R.M.; Monastyrskaya, K.; Gheinani, A.H. MLcps: Machine learning cumulative performance score for classification problems. GigaScience 2023, 12, giad108. [Google Scholar] [CrossRef]
- Murray, P.G.; Stevens, A.; De Leonibus, C.; Koledova, E.; Chatelain, P.; Clayton, P.E. Transcriptomics and machine learning predict diagnosis and severity of growth hormone deficiency. JCI Insight 2018, 3, e93247. [Google Scholar] [CrossRef]
- Latkowski, T.; Osowski, S. Computerized system for recognition of autism on the basis of gene expression microarray data. Comput. Biol. Med. 2015, 56, 82–88. [Google Scholar] [CrossRef] [PubMed]
- Greener, J.G.; Kandathil, S.M.; Moffat, L.; Jones, D.T. A guide to machine learning for biologists. Nat. Rev. Mol. Cell Biol. 2022, 23, 40–55. [Google Scholar] [CrossRef] [PubMed]
- Innes, M.; Edelman, A.; Fischer, K.; Rackauckas, C.; Saba, E.; Shah, V.B.; Tebbutt, W. A differentiable programming system to bridge machine learning and scientific computing. arXiv 2019, arXiv:1907.07587. [Google Scholar]
- Akiyama, Y.; Luo, Y.; Hanno, P.M.; Maeda, D.; Homma, Y. Interstitial cystitis/bladder pain syndrome: The evolving landscape, animal models and future perspectives. Int. J. Urol. 2020, 27, 491–503. [Google Scholar] [CrossRef] [PubMed]
- Karamali, M.; Shafabakhsh, R.; Ghanbari, Z.; Eftekhar, T.; Asemi, Z. Molecular pathogenesis of interstitial cystitis/bladder pain syndrome based on gene expression. J. Cell Physiol. 2019, 234, 12301–12308. [Google Scholar] [CrossRef]
- Colaco, M.; Koslov, D.S.; Keys, T.; Evans, R.J.; Badlani, G.H.; Andersson, K.E.; Walker, S.J. Correlation of gene expression with bladder capacity in interstitial cystitis/bladder pain syndrome. J. Urol. 2014, 192, 1123–1129. [Google Scholar] [CrossRef] [PubMed]
- Blalock, E.M.; Korrect, G.S.; Stromberg, A.J.; Erickson, D.R. Gene expression analysis of urine sediment: Evaluation for potential noninvasive markers of interstitial cystitis/bladder pain syndrome. J. Urol. 2012, 187, 725–732. [Google Scholar] [CrossRef] [PubMed]
- Offiah, I.; Didangelos, A.; Dawes, J.; Cartwright, R.; Khullar, V.; Bradbury, E.J.; O’Sullivan, S.; Williams, D.; Chessell, I.P.; Pallas, K.; et al. The Expression of Inflammatory Mediators in Bladder Pain Syndrome. Eur. Urol. 2016, 70, 283–290. [Google Scholar] [CrossRef] [PubMed]
- Izquierdo, L.; Mateu, L.; Lozano, J.J.; Montalbo, R.; Ingelmo-Torres, M.; Gómez, A.; Peri, L.; Mengual, L.; Franco, A.; Alcaraz, A. Urine Gene Expression Profiles in Bladder Pain Syndrome Patients Treated with Triamcinolone. Eur. Urol. Focus 2020, 6, 390–396. [Google Scholar] [CrossRef]
- Akiyama, Y.; Maeda, D.; Katoh, H.; Morikawa, T.; Niimi, A.; Nomiya, A.; Sato, Y.; Kawai, T.; Goto, A.; Fujimura, T.; et al. Molecular Taxonomy of Interstitial Cystitis/Bladder Pain Syndrome Based on Whole Transcriptome Profiling by Next-Generation RNA Sequencing of Bladder Mucosal Biopsies. J. Urol. 2019, 202, 290–300. [Google Scholar] [CrossRef]
- Vabalas, A.; Gowen, E.; Poliakoff, E.; Casson, A.J. Machine learning algorithm validation with a limited sample size. PLoS ONE 2019, 14, e0224365. [Google Scholar] [CrossRef]
- An, C.; Park, Y.W.; Ahn, S.S.; Han, K.; Kim, H.; Lee, S.K. Radiomics machine learning study with a small sample size: Single random training-test set split may lead to unreliable results. PLoS ONE 2021, 16, e0256152. [Google Scholar] [CrossRef]
- Zhou, W.; Wang, X.; Li, L.; Feng, X.; Yang, Z.; Zhang, W.; Hu, R. Depletion of tubulin polymerization promoting protein family member 3 suppresses HeLa cell proliferation. Mol. Cell Biochem. 2010, 333, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Helmbacher, F. Tissue-specific activities of the Fat1 cadherin cooperate to control neuromuscular morphogenesis. PLoS Biol. 2018, 16, e2004734. [Google Scholar] [CrossRef] [PubMed]
- Hou, R.; Liu, L.; Anees, S.; Hiroyasu, S.; Sibinga, N.E. The Fat1 cadherin integrates vascular smooth muscle cell growth and migration signals. J. Cell Biol. 2006, 173, 417–429. [Google Scholar] [CrossRef] [PubMed]
- Gee, H.Y.; Sadowski, C.E.; Aggarwal, P.K.; Porath, J.D.; Yakulov, T.A.; Schueler, M.; Lovric, S.; Ashraf, S.; Braun, D.A.; Halbritter, J.; et al. FAT1 mutations cause a glomerulotubular nephropathy. Nat. Commun. 2016, 7, 10822. [Google Scholar] [CrossRef] [PubMed]
- Irshad, K.; Srivastava, C.; Malik, N.; Arora, M.; Gupta, Y.; Goswami, S.; Sarkar, C.; Suri, V.; Mahajan, S.; Gupta, D.K.; et al. Upregulation of Atypical Cadherin FAT1 Promotes an Immunosuppressive Tumor Microenvironment via TGF-β. Front. Immunol. 2022, 13, 813888. [Google Scholar] [CrossRef]
- Ivings, L.; Pennington, S.R.; Jenkins, R.; Weiss, J.L.; Burgoyne, R.D. Identification of Ca2+-dependent binding partners for the neuronal calcium sensor protein neurocalcin delta: Interaction with actin, clathrin and tubulin. Biochem. J. 2002, 363, 599–608. [Google Scholar] [CrossRef] [PubMed]
- Riessland, M.; Kaczmarek, A.; Schneider, S.; Swoboda, K.J.; Löhr, H.; Bradler, C.; Grysko, V.; Dimitriadi, M.; Hosseinibarkooie, S.; Torres-Benito, L.; et al. Neurocalcin Delta Suppression Protects against Spinal Muscular Atrophy in Humans and across Species by Restoring Impaired Endocytosis. Am. J. Hum. Genet. 2017, 100, 297–315. [Google Scholar] [CrossRef]
- Upadhyay, A.; Hosseinibarkooie, S.; Schneider, S.; Kaczmarek, A.; Torres-Benito, L.; Mendoza-Ferreira, N.; Overhoff, M.; Rombo, R.; Grysko, V.; Kye, M.J.; et al. Neurocalcin Delta Knockout Impairs Adult Neurogenesis Whereas Half Reduction Is Not Pathological. Front. Mol. Neurosci. 2019, 12, 19. [Google Scholar] [CrossRef]
- Kutzleb, C.; Sanders, G.; Yamamoto, R.; Wang, X.; Lichte, B.; Petrasch-Parwez, E.; Kilimann, M.W. Paralemmin, a prenyl-palmitoyl-anchored phosphoprotein abundant in neurons and implicated in plasma membrane dynamics and cell process formation. J. Cell Biol. 1998, 143, 795–813. [Google Scholar] [CrossRef]
- Suckow, A.T.; Comoletti, D.; Waldrop, M.A.; Mosedale, M.; Egodage, S.; Taylor, P.; Chessler, S.D. Expression of neurexin, neuroligin, and their cytoplasmic binding partners in the pancreatic beta-cells and the involvement of neuroligin in insulin secretion. Endocrinology 2008, 149, 6006–6017. [Google Scholar] [CrossRef]
- Sanchez Freire, V.; Burkhard, F.C.; Kessler, T.M.; Kuhn, A.; Draeger, A.; Monastyrskaya, K. MicroRNAs may mediate the down-regulation of neurokinin-1 receptor in chronic bladder pain syndrome. Am. J. Pathol. 2010, 176, 288–303. [Google Scholar] [CrossRef] [PubMed]
- Boyle, E.I.; Weng, S.; Gollub, J.; Jin, H.; Botstein, D.; Cherry, J.M.; Sherlock, G. GO::TermFinder--open source software for accessing Gene Ontology information and finding significantly enriched Gene Ontology terms associated with a list of genes. Bioinformatics 2004, 20, 3710–3715. [Google Scholar] [CrossRef] [PubMed]
- Yu, G.; Wang, L.G.; Han, Y.; He, Q.Y. clusterProfiler: An R package for comparing biological themes among gene clusters. Omics 2012, 16, 284–287. [Google Scholar] [CrossRef] [PubMed]
- Cawley, G.C.; Talbot, N.L.C. On Over-fitting in Model Selection and Subsequent Selection Bias in Performance Evaluation. J. Mach. Learn. Res. 2010, 11, 2079–2107. [Google Scholar]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Lemaitre, G.; Nogueira, F.; Aridas, C.K. Imbalanced-learn: A Python Toolbox to Tackle the Curse of Imbalanced Datasets in Machine Learning. J. Mach. Learn. Res. 2017, 18, 1–5. [Google Scholar]
- Rossum, G.V.; Drake, F.L. Python 3 Reference Manual; CreateSpace Publishing Platform; Amazon.com, Inc.: Seattle, WA, USA, 2009. [Google Scholar]
- Gu, Z.; Gu, L.; Eils, R.; Schlesner, M.; Brors, B. “Circlize” implements and enhances circular visualization in R. Bioinformatics 2014, 30, 2811–2812. [Google Scholar] [CrossRef]




| BPS vs. Control | DO vs. Control | BPS vs. DO |
|---|---|---|
| TPPP3 | TPPP3 | TPPP3 |
| FAT1 | CLEC3B | FAT1 |
| SMTN | AIM1 | PALM |
| CLEC3B | P2RX2 | NCALD |
| AIM1 | NRXN2 | NRXN2 |
| NCALD | FAM83A |
| Classification Algorithm | Abbreviation |
|---|---|
| Logistic Regression | LR |
| Linear Discriminant Analysis | LDR |
| Gaussian Naive Bayes | GNB |
| Support Vector Machine | SVM |
| k-nearest neighbors | KNN |
| Decision Tree Classifier | DTC |
| Gaussian Process Classifier | GP |
| Random Forest Classifier | RF |
| Bagging Classifier | BC |
| Extra Trees Classifier | ETC |
| Gradient Boosting Classifier | GBC |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Akshay, A.; Besic, M.; Kuhn, A.; Burkhard, F.C.; Bigger-Allen, A.; Adam, R.M.; Monastyrskaya, K.; Hashemi Gheinani, A. Machine Learning-Based Classification of Transcriptome Signatures of Non-Ulcerative Bladder Pain Syndrome. Int. J. Mol. Sci. 2024, 25, 1568. https://doi.org/10.3390/ijms25031568
Akshay A, Besic M, Kuhn A, Burkhard FC, Bigger-Allen A, Adam RM, Monastyrskaya K, Hashemi Gheinani A. Machine Learning-Based Classification of Transcriptome Signatures of Non-Ulcerative Bladder Pain Syndrome. International Journal of Molecular Sciences. 2024; 25(3):1568. https://doi.org/10.3390/ijms25031568
Chicago/Turabian StyleAkshay, Akshay, Mustafa Besic, Annette Kuhn, Fiona C. Burkhard, Alex Bigger-Allen, Rosalyn M. Adam, Katia Monastyrskaya, and Ali Hashemi Gheinani. 2024. "Machine Learning-Based Classification of Transcriptome Signatures of Non-Ulcerative Bladder Pain Syndrome" International Journal of Molecular Sciences 25, no. 3: 1568. https://doi.org/10.3390/ijms25031568
APA StyleAkshay, A., Besic, M., Kuhn, A., Burkhard, F. C., Bigger-Allen, A., Adam, R. M., Monastyrskaya, K., & Hashemi Gheinani, A. (2024). Machine Learning-Based Classification of Transcriptome Signatures of Non-Ulcerative Bladder Pain Syndrome. International Journal of Molecular Sciences, 25(3), 1568. https://doi.org/10.3390/ijms25031568






