Transcriptomic Analysis Reveals LncRNAs Associated with Flowering of Angelica sinensis during Vernalization
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. Full-Length Isoform Sequencing and Analysis
2.3. Analysis of Long Noncoding RNAs (lncRNAs)
2.4. Characterization of LncRNAs
2.5. qRT-PCR Validation
2.6. Statistical Analysis
3. Results
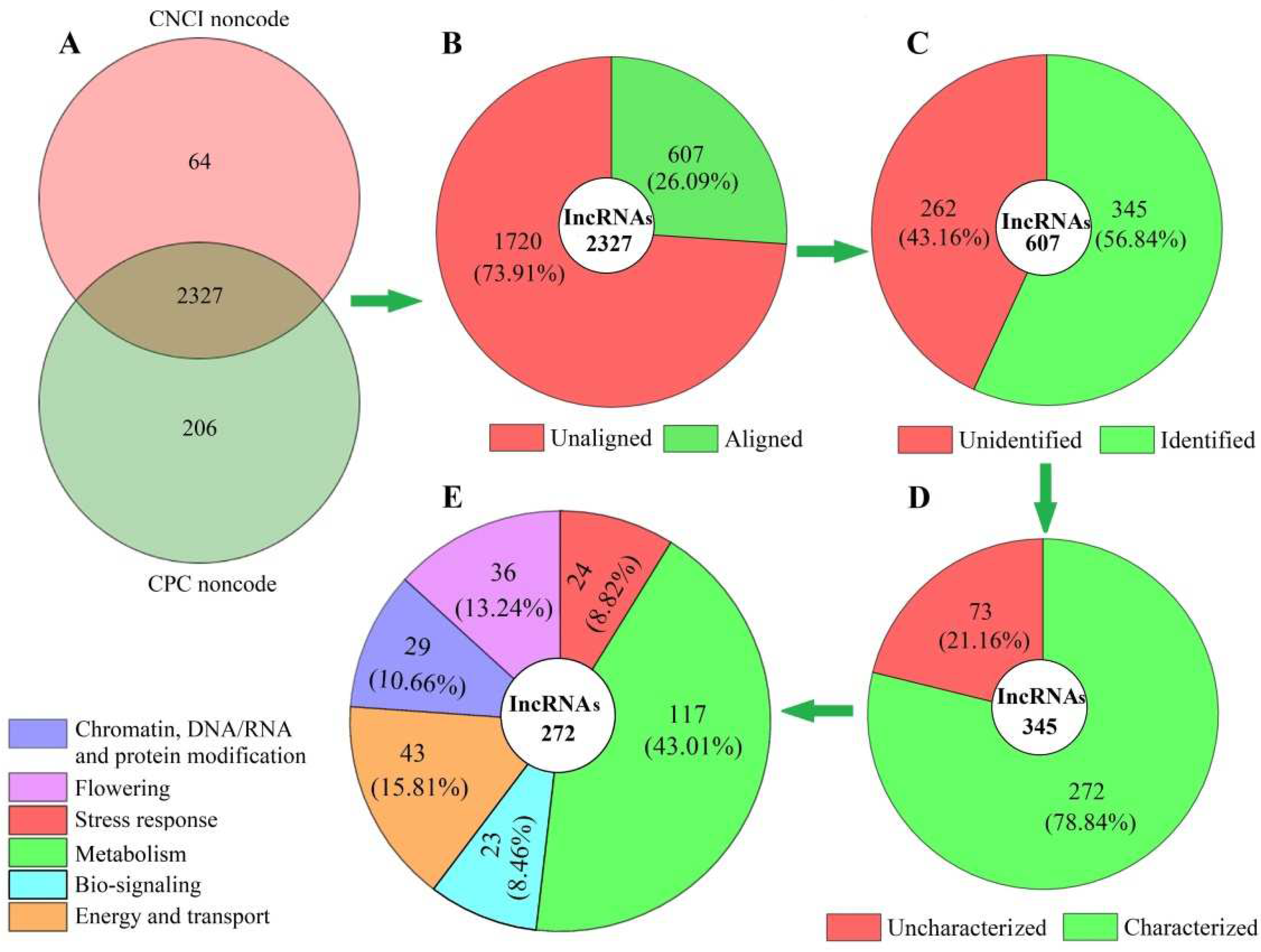
3.1. LncRNAs Analysis
3.2. LncRNAs Linked with Chromatin, DNA/RNA and Protein Modification, as well as Expression Levels of Their Coexpressed mRNAs
3.3. LncRNAs Linked with Flowering and Expression Levels of Their Coexpressed mRNAs
3.4. LncRNAs Linked with Stress Response and Expression Levels of Their Coexpressed mRNAs
3.5. LncRNAs Linked with Metabolism and Expression Levels of Their Coexpressed mRNAs
3.6. LncRNAs Linked with Biosignaling and Expression Levels of Their Coexpressed mRNAs
3.7. LncRNAs Linked with Energy and Transport
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| AP1 | APETALA 1 |
| CCS | Circular consensus sequence |
| CNCI | Coding–Noncoding Index |
| COLDAIR | COLD-ASSISTED INTRONIC NON-CODING RNA |
| COOLAIR | COLD-INDUCED LONG ANTISENSE INTRAGENIC RNAs |
| CPC | Coding-Potential Calculator |
| FLC | FLOWERING LOCUS C |
| FLM | FLOWERING LOCUS M |
| FLNC | Full-length nonchimeric |
| FT | FLOWERING LOCUS T |
| GA | Gibberellin |
| GA2OX1 | Gibberellin 2-β-dioxygenase 1 |
| GO | Gene Ontology |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| KOG | euKaryotic orthologous groups of proteins |
| lncRNAs | Long noncoding RNAs |
| NCBI | National Center for Biotechnology Information |
| ncRNAs | Noncoding RNAs |
| NR | NCBI nonredundant protein |
| PHYA | PHYOCHROME A |
| REL | Relative expression level |
| SOC1 | SUPPRESSOR OF OVEREXPRESSION OF CONSTANS 1 |
| sRNAs | Small RNAs |
| TAIR | The Arabidopsis Information Resource |
References
- Zhang, H.Y.; Bi, W.G.; Yu, Y.; Liao, W.B. Angelica sinensis (Oliv.) Diels in China: Distribution, cultivation, utilization and variation. Genet. Resour. Crop Evol. 2012, 59, 607–613. [Google Scholar] [CrossRef]
- Wang, W.J. Analysis and control of early bolting characteristic of Angelica sinensis. J. Northwest Univ. 1977, 7, 32–39. [Google Scholar]
- Committee for the Pharmacopoeia of PR China. Pharmacopoeia of the People’s Republic of China; Chinese Medical Science and Technology Press: Beijing, China, 2015; p. 133. [Google Scholar]
- Hook, I.L. Danggui to Angelica sinensis root: Are potential benefits to European women lost in translation? A review. J. Ethnopharmacol. 2014, 152, 1–13. [Google Scholar] [CrossRef]
- Upton, R. American Herbal Pharmacopoeia and Therapeutic Compendium: Dang Gui Root-Angelica sinensis (Oliv.); American Herbal Pharmacopoeia: Scotts Valley, CA, USA, 2003; pp. 1–41. [Google Scholar]
- Ma, J.P.; Guo, Z.-B.; Jin, L.; Li, Y.D. Phytochemical progress made in investigations of Angelica sinensis (Oliv.) Diels. Chin. J. Nat. Med. 2015, 13, 241–249. [Google Scholar] [CrossRef]
- Wei, W.L.; Zeng, R.; Gu, C.M.; Qu, Y.; Huang, L.F. Angelica sinensis in China-A review of botanical profile, ethnopharmacology, phytochemistry and chemical analysis. J. Ethnopharmacol. 2016, 190, 116–141. [Google Scholar] [CrossRef]
- Huang, L.Q.; Jin, L. Suitable Technology for Production and Processing of Angelica sinensis; China Pharmaceutical Science and Technology Press: Beijing, China, 2018; pp. 1–14. [Google Scholar]
- Li, M.L.; Cui, X.W.; Jin, L.; Li, M.F.; Wei, J.H. Bolting reduces ferulic acid and flavonoid biosynthesis and induces root lignification in Angelica sinensis. Plant Physiol. Biochem. 2022, 170, 171–179. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.M.; Qiu, D.Y.; Chen, Y. Effect of root diameter on early bolting rate and yield in seedling of Angelica sinensis. Chin. Tradit. Herb. Drugs 2007, 38, 1386–1389. [Google Scholar]
- Wang, W.J. Technology and principle of seedling frozen storage of Angelica sinensis. J. Chin. Med. Mat. 1979, 3, 1–4. [Google Scholar]
- Jia, Z.; Di, S.Q.; Zhao, F.Y.; Li, S.X.; Wang, S.C. Effects of different low overwintering temperatures on Angelica vernalization and premature bolting. Agr. Sci. Tech. 2018, 19, 55–62. [Google Scholar]
- Yao, L. Effect of shading during the nursery of Angelica sinensis on bolting rate and economic characters. Gansu Agr. Sci. Tech. 2005, 10, 54–55. [Google Scholar]
- Yu, G.; Zhou, Y.; Yu, J.J.; Hu, X.Q.; Tang, Y.; Yan, H.; Duan, J.A. Transcriptome and digital gene expression analysis unravels the novel mechanism of early flowering in Angelica sinensis. Sci. Rep. 2019, 9, 10035. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Li, M.L.; Zhu, T.T.; Zhang, X.N.; Li, M.F.; Wei, J.H. Integrated transcriptomics and metabolites at different growth stagesreveals the regulation mechanism of bolting and flowering of Angelica sinensis. Plant Biol. 2021, 23, 574–582. [Google Scholar] [CrossRef] [PubMed]
- Li, M.F.; Li, J.; Wei, J.H.; Paré, P.W. Transcriptional controls for early bolting and flowering in Angelica sinensis. Plants 2021, 10, 1931. [Google Scholar] [CrossRef] [PubMed]
- Zhang, E.H.; Huang, P. Effects of vernalization treatment on physiological character of Angelica sinensis seedlings. J. Gansu Agric. Univ. 1998, 33, 240–243. [Google Scholar]
- Chen, H.G.; Du, T.; Zhu, T.T.; Gao, S.F.; Chai, L.; He, W.W. Study on physiological mechanisms in frozen storage to reduce early bolting of Angelica sinensis. Mod. Tradit. Chin. Med. Mater. Med. World Sci. Tech. 2014, 16, 203–206. [Google Scholar]
- Ponting, C.P.; Oliver, P.L.; Reik, W. Evolution and functions of long noncoding RNAs. Cell 2009, 136, 629–641. [Google Scholar] [CrossRef] [Green Version]
- Axtell, M.J.; Westholm, J.O.; Lai, E.C. Vive la différence: Biogenesis and evolution of microRNAs in plants and animals. Genome Biol. 2011, 12, 221. [Google Scholar] [CrossRef] [Green Version]
- Carrington, J.C.; Ambros, V. Role of microRNAs in plant and animal development. Science 2003, 301, 336–338. [Google Scholar] [CrossRef] [Green Version]
- Jain, N.; Sinha, N.; Krishna, H.; Singh, P.K.; Gautam, T.; Prasad, P.; Balyan, H.S.; Gupta, P.K. A study of miRNAs and lncRNAs during Lr28-mediated resistance against leaf rust in wheat (Triticum aestivum L.). Physiol. Mol. Plant Pathol. 2020, 112, 101552. [Google Scholar] [CrossRef]
- Sharma, Y.; Sharma, A.; Madhu; Shumayla; Singh, K.; Upadhyay, S.K. Long non-coding RNAs as emerging regulators of pathogen response in plants. Non Coding RNA 2022, 8, 4. [Google Scholar] [CrossRef]
- Qi, L.; Li, X.; Zhang, S.; An, D. Genetic regulation by non-coding RNAs. Sci. China Ser. C Life Sci. 2006, 49, 201–217. [Google Scholar] [CrossRef]
- Mercer, T.R.; Dinger, M.E.; Mattick, J.S. Long non-coding RNAs: Insights into functions. Nat. Rev. Genet. 2009, 10, 155–159. [Google Scholar] [CrossRef] [PubMed]
- Gordon, S.P.; Tseng, E.; Salamov, A.; Zhang, J.; Meng, X.; Zhao, Z.; Kang, D.; Underwood, J.; Grigoriev, I.V.; Figueroa, M.; et al. Widespread polycistronic transcripts in fungi revealed by single-molecule mRNA sequencing. PLoS ONE 2015, 10, e0132628. [Google Scholar] [CrossRef] [Green Version]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.D.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sun, L.; Luo, H.; Bu, D.; Zhao, G.; Yu, K.; Zhang, C.; Liu, Y.; Chen, R.; Zhao, Y. Utilizing sequence intrinsic composition to classify protein-coding and long non-coding transcripts. Nucleic Acids Res. 2013, 41, e166. [Google Scholar] [CrossRef] [PubMed]
- Kong, L.; Zhang, Y.; Ye, Z.Q.; Liu, X.Q.; Zhao, S.Q.; Wei, L.; Gao, G. CPC: Assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res. 2007, 35, W345–W349. [Google Scholar] [CrossRef] [PubMed]
- Poole, R.L. The TAIR database. Methods Mol. Biol. 2007, 406, 179–212. [Google Scholar] [CrossRef] [PubMed]
- Liao, Q.; Xiao, H.; Bu, D.C.; Xie, C.Y.; Miao, R.Y.; Luo, H.T.; Zhao, G.G.; Yu, K.T.; Zhao, H.T.; Skogerbø, G.; et al. ncFANs: A web server for functional annotation of long non-coding RNAs. Nucleic Acids Res. 2011, 39, w118–w124. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mattick, J.S.; Rinn, J.L. Discovery and annotation of long noncoding RNAs. Nat. Struct. Mol. Biol. 2015, 22, 5–7. [Google Scholar] [CrossRef]
- Shumayla; Sharma, S.; Taneja, M.; Tyagi, S.; Singh, K.; Upadhyay, S.K. Survey of high throughput RNA-Seq data reveals potential roles for lncRNAs during development and stress response in bread wheat. Front. Plant Sci. 2017, 8, 1019. [Google Scholar] [CrossRef] [Green Version]
- Xu, R.; Xu, J.; Li, Y.C.; Dai, Y.T.; Zhang, S.P.; Wang, G.; Liu, Z.G.; Dong, L.L.; Chen, S.L. Integrated chemical and transcriptomic analyses unveils synthetic characteristics of different medicinal root parts of Angelica sinensis. Chin. Herb. Med. 2020, 12, 19–28. [Google Scholar] [CrossRef]
- Willems, E.; Leyns, L.; Vandesompele, J. Standardization of real-time PCR gene expression data from independent biological replicates. Anal. Biochem. 2008, 379, 127–129. [Google Scholar] [CrossRef] [PubMed]
- Svec, D.; Tichopad, A.; Novosadova, V.; Pfaffl, M.W.; Kubista, M. How good is a PCR efficiency estimate: Recommendations for precise and robust qPCR efficiency assessments. Biomol. Detect. Quantif. 2015, 3, 9–16. [Google Scholar] [CrossRef] [Green Version]
- Taiz, L.; Zeiger, E. Plant physiology. In The Control of Flowering, 5th ed.; Fosket, D.E., Amasino, R., Eds.; Sinauer Associates, Inc.: Sunderland, MA, USA, 2010; pp. 559–590. [Google Scholar]
- Amasino, R.M. Vernalization and flowering time. Curr. Opin. Biotechnol. 2005, 16, 154–158. [Google Scholar] [CrossRef] [PubMed]
- He, K.; Cao, X.F.; Deng, X. Histone methylation in epigenetic regulation and temperature responses. Curr. Opin. Plant Biol. 2021, 61, 102001. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.K.; Wu, P.; Wang, Q.; Wang, W.L.; Zhang, C.W.; Sun, F.F.; Liu, Z.K.; Li, Y.; Hou, X.L. Comparative transcriptome discovery and elucidation of the mechanism of long noncoding RNAs during vernalization in Brassica rapa. Plant Growth Regul. 2018, 85, 27–39. [Google Scholar] [CrossRef]
- Velanis, C.N.; Goodrich, J. Vernalization and epigenetic inheritance: A game of histones. Curr. Biol. 2017, 27, R1324–R1326. [Google Scholar] [CrossRef]
- Kwak, K.J.; Kim, J.Y.; Kim, Y.O.; Kang, H. Characterization of transgenic Arabidopsis plants overexpressing high mobility group b proteins under high salinity, drought or cold stress. Plant Cell Physiol. 2007, 48, 221–231. [Google Scholar] [CrossRef]
- Theologis, A.; Ecker, J.R.; Palm, C.J.; Federspiel, N.A.; Kaul, S.; White, O.; Alonso, J.; Altafi, H.; Araujo, R.; Bowman, C.L.; et al. Sequence and analysis of chromosome 1 of the plant Arabidopsis thaliana. Nature 2000, 408, 816–820. [Google Scholar] [CrossRef]
- Ohbayashi, I.; Konishi, M.; Ebine, K.; Sugiyama, M. Genetic identification of Arabidopsis RID2 as an essential factor involved in pre-rRNA processing. Plant J. 2011, 67, 49–60. [Google Scholar] [CrossRef]
- Cheng, C.Y.; Krishnakumar, V.; Chan, A.P.; Thibaud-Nissen, F.; Schobel, S.; Town, C.D. Araport11: A complete reannotation of the Arabidopsis thaliana reference genome. Plant J. 2017, 89, 789–804. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dai, X.Z.; Bai, Y.H.; Zhao, L.H.; Dou, X.Y.; Liu, Y.H.; Wang, L.L.; Li, Y.; Li, W.M.; Hui, Y.N.; Huang, X.Y.; et al. H2A.Z represses gene expression by modulating promoter nucleosome structure and enhancer histone modifications in Arabidopsis. Mol. Plant 2017, 10, 1274–1292. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiang, D.; Borg, M.; Lorković, Z.J.; Montgomery, S.A.; Osakabe, A.; Yelagandula, R.; Axelsson, E.; Berger, F. The evolution and functional divergence of the histone H2B family in plants. PLoS Genet. 2020, 16, e1008964. [Google Scholar] [CrossRef] [PubMed]
- Sridhar, V.V.; Kapoor, A.; Zhang, K.L.; Zhu, J.J.; Zhou, T.; Hasegawa, P.M.; Bressan, R.A.; Zhu, J.K. Control of DNA methylation and heterochromatic silencing by histone H2B deubiquitination. Nature 2007, 447, 735–738. [Google Scholar] [CrossRef]
- Zhu, J.; Jeong, J.C.; Zhu, Y.; Sokolchik, I.; Miyazaki, S.; Zhu, J.K.; Hasegawa, P.M.; Bohnert, H.J.; Shi, H.; Yun, D.J.; et al. Involvement of Arabidopsis HOS15 in histone deacetylation and cold tolerance. Proc. Natl. Acad. Sci. USA 2008, 105, 4945–4950. [Google Scholar] [CrossRef] [Green Version]
- Lu, F.L.; Cui, X.; Zhang, S.B.; Jenuwein, T.; Cao, X.F. Arabidopsis REF6 is a histone H3 lysine 27 demethylase. Nat. Genet. 2011, 43, 715–719. [Google Scholar] [CrossRef]
- Hou, X.L.; Zhou, J.N.; Liu, C.; Liu, L.; Shen, L.S.; Yu, H. Nuclear factor Y-mediated H3K27me3 demethylation of the SOC1 locus orchestrates flowering responses of Arabidopsis. Nat. Commun. 2014, 5, 4601. [Google Scholar] [CrossRef] [Green Version]
- Zhao, D.Z.; Yu, Q.L.; Chen, M.; Ma, H. The ASK1 gene regulates B function gene expression in cooperation with UFO and LEAFY in Arabidopsis. Development 2001, 128, 2735–2746. [Google Scholar] [CrossRef]
- Liu, F.; Ni, W.; Griffith, M.E.; Huang, Z.; Chang, C.; Peng, W.; Ma, H.; Xie, D. The ASK1 and ASK2 genes are essential for Arabidopsis early development. Plant Cell 2004, 16, 5–20. [Google Scholar] [CrossRef] [Green Version]
- Yoshida, R.; Umezawa, T.; Mizoguchi, T.; Takahashi, S.; Takahashi, F.; Shinozaki, K. The regulatory domain of SRK2E/OST1/SnRK2.6 interacts with ABI1 and integrates abscisic acid (ABA) and osmotic stress signals controlling stomatal closure in Arabidopsis. J. Biol. Chem. 2006, 281, 5310–5318. [Google Scholar] [CrossRef] [Green Version]
- Yan, J.; Wang, P.C.; Wang, B.S.; Hsu, C.C.; Tang, K.; Zhang, H.R.; Hou, Y.J.; Zhao, Y.; Wang, Q.M.; Zhao, C.Z.; et al. The SnRK2 kinases modulate miRNA accumulation in Arabidopsis. PLoS Genet. 2017, 13, e1006753. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sura, W.; Kabza, M.; Karlowski, W.M.; Bieluszewski, T.; Kus-Slowinska, M.; Pawełoszek, Ł.; Sadowski, J.; Ziolkowski, P.A. Dual role of the histone variant H2A.Z in transcriptional regulation of stress-response genes. Plant Cell 2017, 29, 791–807. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hamorsky, K.T.; Kouokam, J.C.; Jurkiewicz, J.M.; Nelson, B.; Moore, L.J.; Husk, A.S.; Kajiura, H.; Fujiyama, K.; Matoba, N. N-Glycosylation of cholera toxin B subunit in Nicotiana benthamiana: Impacts on host stress response, production yield and vaccine potential. Sci. Rep. 2015, 5, 8003. [Google Scholar] [CrossRef] [PubMed]
- Fujii, H.; Verslues, P.E.; Zhu, J.K. Arabidopsis decuple mutant reveals the importance of SnRK2 kinases in osmotic stress re-sponses in vivo. Proc. Natl. Acad. Sci. USA 2011, 108, 1717–1722. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Heo, J.B.; Sung, S. Vernalization-mediated epigenetic silencing by a long intronic noncoding RNA. Science 2011, 331, 76–79. [Google Scholar] [CrossRef] [Green Version]
- Liu, F.; Marquardt, S.; Lister, C.; Swiezewski, S.; Dean, C. Targeted 3′ processing of antisense transcripts triggers Arabidopsis FLC chromatin silencing. Science 2010, 327, 94–97. [Google Scholar] [CrossRef]
- Sun, Q.; Csorba, T.; Skourti-Stathaki, K.; Proudfoot, N.J.; Dean, C. R-loop stabilization represses antisense transcription at the Arabidopsis FLC locus. Science 2013, 340, 619–621. [Google Scholar] [CrossRef] [Green Version]
- Johansson, M.; Staiger, D. SRR1 is essential to repress flowering in non-inductive conditions in Arabidopsis thaliana. J. Exp. Bot. 2014, 65, 5811–5822. [Google Scholar] [CrossRef] [Green Version]
- Endo, M.; Tanigawa, Y.; Murakami, T.; Araki, T.; Nagatani, A. Phytochrome-dependent late-flowering accelerates flowering through physical interactions with phytochrome B and CONSTANS. Proc. Natl. Acad. Sci. USA 2013, 110, 18017–18022. [Google Scholar] [CrossRef] [Green Version]
- Yang, S.W.; Jang, I.C.; Henriques, R.; Chua, N.H. FAR-RED ELONGATED HYPOCOTYL1 and FHY1-LIKE associate with the Arabidopsis transcription factors LAF1 and HFR1 to transmit phytochrome a signals for inhibition of hypocotyl elongation. Plant Cell 2009, 21, 1341–1359. [Google Scholar] [CrossRef] [Green Version]
- Kang, I.H.; Steffen, J.G.; Portereiko, M.F.; Lloyd, A.; Drews, G.N. The AGL62 MADS domain protein regulates cellularization during endosperm development in Arabidopsis. Plant Cell 2008, 20, 635–647. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, R.M.; Wang, Y.; Gruber, M.Y.; Hannoufa, A. miR156/SPL10 modulates lateral root development, branching and leaf morphology in Arabidopsis by silencing AGAMOUS-LIKE 79. Front. Plant Sci. 2017, 8, 2226. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, R.G.; Kroczynska, B.; Miernyk, J.A. The genomic sequence encoding the Arabidopsis thaliana molecular chaperone AtJ3. Plant Physiol. 2000, 122, 291. [Google Scholar] [CrossRef] [Green Version]
- Zhu, Q.H.; Helliwell, C.A. Regulation of flowering time and floral patterning by miR172. J. Exp. Bot. 2011, 62, 487–495. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meng, F.; Liu, H.; Wang, K.; Liu, L.; Wang, S.; Zhao, Y.; Yin, J.; Li, Y. Development-associated microRNAs in grains of wheat (Triticum aestivum L.). BMC Plant Biol. 2013, 13, 140. [Google Scholar] [CrossRef] [Green Version]
- Wu, Q.; Luo, Y.; Wu, X.; Bai, X.; Ye, X.L.; Liu, C.Y.; Wan, Y.; Xiang, D.B.; Li, Q.; Zou, L.; et al. Identification of the specific long-noncoding RNAs involved in night-break mediated flowering retardation in Chenopodium quinoa. BMC Genom. 2021, 22, 284. [Google Scholar] [CrossRef] [PubMed]
- Calixto, C.P.G.; Tzioutziou, N.A.; James, A.B.; Hornyik, C.; Guo, W.B.; Zhang, R.X.; Nimmo, H.G.; Brown, J.W.S. Cold-dependent expression and alternative splicing of Arabidopsis long non-coding RNAs. Front. Plant Sci. 2019, 10, 235. [Google Scholar] [CrossRef] [Green Version]
- Wang, P.; Dai, L.; Ai, J.; Wang, Y.; Ren, F. Identification and functional prediction of cold-related long non-coding RNA (lncRNA) in grapevine. Sci. Rep. 2019, 9, 1–15. [Google Scholar] [CrossRef]
- Bhatia, G.; Singh, A.; Verma, D.; Sharma, S.; Singh, K. Genome-wide investigation of regulatory roles of lncRNAs in response to heat and drought stress in Brassica juncea (Indian mustard). Environ. Exp. Bot. 2020, 171, 103922. [Google Scholar] [CrossRef]
- Wen, X.J.; Ding, Y.; Tan, Z.L.; Wang, J.X.; Zhang, D.Y.; Wang, Y.C. Identification and characterization of cadmium stress-related LncRNAs from Betula platyphylla. Plant Sci. 2020, 299, 110601. [Google Scholar] [CrossRef]
- Xu, J.H.; Fang, M.; Li, Z.H.; Zhang, M.N.; Liu, X.Y.; Peng, Y.Y.; Wan, Y.L.; Chen, J.H. Third-generation sequencing reveals lncRNA-regulated HSP genes in the Populus x Canadensis moench heat stress response. Front. Genet. 2020, 11, 249. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.F.; Xiao, S.; Chye, M.L. Overexpression of the Arabidopsis 10-kilodalton acyl-coenzyme A-binding protein ACBP6 enhances freezing tolerance. Plant Physiol. 2008, 148, 304–315. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, H.; Guo, Y.; Ohta, M.; Xiong, L.; Stevenson, B.; Zhu, J.K. LOS2, a genetic locus required for cold-responsive gene transcription encodes a bifunctional enolase. EMBO J. 2002, 21, 2692–2702. [Google Scholar] [CrossRef] [PubMed]
- Ismond, K.P.; Dolferus, R.; de Pauw, M.; Dennis, E.S.; Good, A.G. Enhanced low oxygen survival in Arabidopsis through increased metabolic flux in the fermentative pathway. Plant Physiol. 2003, 132, 1292–1302. [Google Scholar] [CrossRef] [Green Version]
- Zhang, F.; Maeder, M.L.; Unger-Wallace, E.; Hoshaw, J.P.; Reyon, D.; Christian, M.; Li, X.; Pierick, C.J.; Dobbs, D.; Peterson, T.; et al. High frequency targeted mutagenesis in Arabidopsis thaliana using zinc finger nucleases. Proc. Natl. Acad. Sci. USA 2010, 107, 12028–12033. [Google Scholar] [CrossRef] [Green Version]
- Sasaki, K.; Kim, M.; Imai, R. Arabidopsis cold shock domain protein 2 is a RNA chaperone that is regulated by cold and developmental signals. Biochem. Biophys. Res. Commun. 2007, 364, 633–638. [Google Scholar] [CrossRef]
- Anaraki, Z.E.; Tafreshi, S.A.H.; Shariati, M. Transient silencing of heat shock proteins showed remarkable roles for HSP70 during adaptation to stress in plants. Environ. Exp. Bot. 2018, 155, 142–157. [Google Scholar] [CrossRef]
- Huang, S.; Monaghan, J.; Zhong, X.H.; Lin, L.; Sun, T.J.; Dong, O.X.; Li, X. HSP90s are required for NLR immune receptor accumulation in Arabidopsis. Plant J. 2014, 79, 427–439. [Google Scholar] [CrossRef]
- Lin, W.; Zhou, Q.; Wang, C.Q.; Zhu, L.; Bi, C.; Zhang, S.; Wang, X.; Jin, H. LncRNAs regulate metabolism in cancer. Int. J. Biol. Sci. 2020, 16, 1194–1206. [Google Scholar] [CrossRef]
- Karimi, P.; Bakhtiarizadeh, M.R.; Salehi, A.; Izadnia, H.R. Transcriptome analysis reveals the potential roles of long non-coding RNAs in feed efficiency of chicken. Sci. Rep. 2022, 12, 2558. [Google Scholar] [CrossRef]
- Tello-Flores, V.A.; Beltrán-Anaya, F.O.; Ramírez-Vargas, M.A.; Esteban-Casales, B.E.; Navarro-Tito, N.; Alarcón-Romero, L.d.C.; Luciano-Villa, C.A.; Ramírez, M.; del Moral-Hernández, Ó.; Flores-Alfaro, E. Role of long non-coding RNAs and the molecular mechanisms involved in insulin resistance. Int. J. Mol. Sci. 2021, 22, 7256. [Google Scholar] [CrossRef] [PubMed]
- Slabas, A.R.; Ndimba, B.; Simon, W.J.; Chivasa, S. Proteomic analysis of the Arabidopsis cell wall reveals unexpected proteins with new cellular locations. Biochem. Soc. Trans. 2004, 32, 524–528. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Muñoz-Bertomeu, J.; Cascales-Miñana, B.; Mulet, J.M.; Baroja-Fernández, E.; Pozueta-Romero, J.; Kuhn, J.M.; Segura, J.; Ros, R. Plastidial glyceraldehyde-3-phosphate dehydrogenase deficiency leads to altered root development and affects the sugar and amino acid balance in Arabidopsis. Plant Physiol. 2009, 151, 541–558. [Google Scholar] [CrossRef] [Green Version]
- Zhang, L.; Song, Z.; Li, F.; Li, X.; Ji, H.; Yang, S. The specific MYB binding sites bound by TaMYB in the GAPCp2/3 promoters are involved in the drought stress response in wheat. BMC Plant Biol. 2019, 19, 366. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rius, S.P.; Casati, P.; Iglesias, A.A.; Gomez-Casati, D.F. Characterization of Arabidopsis lines deficient in GAPC-1, a cytosolic NAD-dependent glyceraldehyde-3-phosphate dehydrogenase. Plant Physiol. 2008, 148, 1655–1667. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Egli, B.; Kolling, K.; Kohler, C.; Zeeman, S.C.; Streb, S. Loss of cytosolic phosphoglucomutase compromises gametophyte development in Arabidopsis. Plant Physiol. 2010, 154, 1659–1671. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fulton, D.C.; Stettler, M.; Mettler, T.; Vaughan, C.K.; Li, J.; Francisco, P.; Gil, M.; Reinhold, H.; Eicke, S.; Messerli, G.; et al. Beta-AMYLASE4, a noncatalytic protein required for starch breakdown, acts upstream of three active beta-amylases in Arabidopsis chloroplasts. Plant Cell 2008, 20, 1040–1058. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Oono, Y.; Seki, M.; Satou, M.; Iida, K.; Akiyama, K.; Sakurai, T.; Fujita, M.; Yamaguchi-Shinozaki, K.; Shinozaki, K. Monitoring expression profiles of Arabidopsis genes during cold acclimation and deacclimation using DNA microarrays. Funct. Integr. Genom. 2006, 6, 212–234. [Google Scholar] [CrossRef]
- Tanaka, H.; Nodzylski, T.; Kitakura, S.; Feraru, M.I.; Sasabe, M.; Ishikawa, T.; Kleine-Vehn, J.; Kakimoto, T.; Friml, J. BEX1/ARF1A1C is required for BFA-sensitive recycling of PIN auxin transporters and auxin-mediated development in Ara-bidopsis. Plant Cell Physiol. 2014, 55, 737–749. [Google Scholar] [CrossRef] [Green Version]
- Feng, S.H.; Shen, Y.P.; Sullivan, J.A.; Rubio, V.; Xiong, Y.; Sun, T.P.; Deng, X.W. Arabidopsis CAND1, an unmodified CUL1-interacting protein, is involved in multiple developmental pathways controlled by ubiquitin/proteasome-mediated protein degradation. Plant Cell 2004, 16, 1870–1882. [Google Scholar] [CrossRef] [Green Version]
- Tayengwa, R.; Zhao, J.; Pierce, C.F.; Werner, B.E.; Neff, M.M. Synopsis of the SOFL plant-specific gene family. G3 Genes Genomes Genet. 2018, 8, 1281–1290. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gampala, S.S.; Kim, T.W.; He, J.X.; Tang, W.; Deng, Z.; Bai, M.Y.; Guan, S.; Lalonde, S.; Sun, Y.; Gendron, J.M.; et al. An essential role for 14-3-3 proteins in brassinosteroid signal transduction in Arabidopsis. Dev. Cell 2007, 13, 177–189. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, J.L.; Chen, W.W.; Chen, L.Q.; Qin, C.; Jin, C.W.; Shi, Y.Z.; Zheng, S.J. The 14-3-3 protein general regulatory factor11 (GRF 11) acts downstream of nitric oxide to regulate iron acquisition in Arabidopsis thaliana. New Phytol. 2013, 197, 815–824. [Google Scholar] [CrossRef] [PubMed]
- Ohta, M.; Matsui, K.; Hiratsu, K.; Shinshi, H.; Ohme-Takagi, M. Repression domains of class II ERF transcriptional repressors share an essential motif for active repression. Plant Cell 2001, 13, 1959–1968. [Google Scholar] [CrossRef] [Green Version]
- Jang, Y.H.; Park, H.Y.; Lee, K.C.; Thu, M.P.; Kim, S.K.; Suh, M.C.; Kang, H.; Kim, J.K. A homolog of splicing factor SF1 is essential for development and is involved in the alternative splicing of pre-mRNA in Arabidopsis thaliana. Plant J. 2014, 78, 591–603. [Google Scholar] [CrossRef]
- Shang, S.; Wu, C.; Huang, C.; Tie, W.; Yan, Y.; Ding, Z.; Xia, Z.; Wang, W.; Peng, M.; Tian, L.; et al. Genome-wide analysis of the GRF family reveals their involvement in abiotic stress response in cassava. Genes 2018, 9, 110. [Google Scholar] [CrossRef] [Green Version]
- Fujimoto, S.Y.; Ohta, M.; Usui, A.; Shinshi, H.; Ohme-Takagi, M. Arabidopsis ethylene-responsive element binding factors act as transcriptional activators or repressors of GCC box-mediated gene expression. Plant Cell 2000, 12, 393–404. [Google Scholar] [CrossRef] [Green Version]
- Lee, K.C.; Chung, K.S.; Lee, H.T.; Park, J.H.; Lee, J.H.; Kim, J.K. Role of Arabidopsis splicing factor SF1 in tempera-ture-responsive alternative splicing of FLM pre-mRNA. Front. Plant Sci. 2020, 11, 596354. [Google Scholar] [CrossRef]
- Johnson, X.; Alric, J. Central carbon metabolism and electron transport in Chlamydomonas reinhardtii: Metabolic constraints for carbon partitioning between oil and starch. Eukaryot Cell 2013, 12, 776–793. [Google Scholar] [CrossRef] [Green Version]
- Collakova, E.; Goyer, A.; Naponelli, V.; Krassovskaya, I.; Gregory, J.F.; Hanson, A.D.; Shachar-Hill, Y. Arabidopsis 10-formyl tetrahydrofolate deformylases are essential for photorespiration. Plant Cell 2008, 20, 1818–1832. [Google Scholar] [CrossRef] [Green Version]
- Christ, B.; Schelbert, S.; Aubry, S.; Suessenbacher, I.; Mueller, T.; Kraeutler, B.; Hoertensteiner, S. MES16, a member of the methylesterase protein family, specifically demethylates fluorescent chlorophyll catabolites during chlorophyll breakdown in Arabidopsis. Plant Physiol. 2012, 158, 628–641. [Google Scholar] [CrossRef] [Green Version]
- Baune, M.C.; Lansing, H.; Fischer, K.; Meyer, T.; Charton, L.; Linka, N.; von Schaewen, A. The Arabidopsis plastidial glucose-6-phosphate transporter GPT1 is dually targeted to peroxisomes via the endoplasmic reticulum. Plant Cell 2020, 32, 1703–1726. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sánchez-Fernández, R.; Davies, T.G.E.; Coleman, J.O.; Rea, P.A. The Arabidopsis thaliana ABC protein superfamily, a complete inventory. J. Biol. Chem. 2001, 276, 30231–30244. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Skalitzky, C.A.; Martin, J.R.; Harwood, J.H.; Beirne, J.J.; Adamczyk, B.J.; Heck, G.R.; Cline, K.; Fernandez, D.E. Plastids contain a second sec translocase system with essential functions. Plant Physiol. 2011, 155, 354–369. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kikuchi, S.; Asakura, Y.; Imai, M.; Nakahira, Y.; Kotani, Y.; Hashiguchi, Y.; Nakai, Y.; Takafuji, K.; Bédard, J.; Hirabayashi-Ishioka, Y.; et al. A Ycf2-ftshi heteromeric AAA-ATPase complex is required for chloroplast protein import. Plant Cell 2018, 30, 2677–2703. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reddy, V.S.; Day, I.S.; Thomas, T.; Reddy, A.S.N. KIC, a novel Ca2+ binding protein with one EF-hand motif, interacts with a microtubule motor protein and regulates trichome morphogenesis. Plant Cell 2004, 16, 185–200. [Google Scholar] [CrossRef] [Green Version]
- Bassil, E.; Ohto, M.A.; Esumi, T.; Tajima, H.; Zhu, Z.; Cagnac, O.; Belmonte, M.; Peleg, Z.; Yamaguchi, T.; Blumwald, E. The Arabidopsis intracellular Na+/H+ antiporters NHX5 and NHX6 are endosome associated and necessary for plant growth and development. Plant Cell 2011, 23, 224–239. [Google Scholar] [CrossRef] [Green Version]







| lncRNA ID | Coexpressed mRNAs | mRNA ID | Primer Sequences (5’ to 3’) | Amplicon Size (bp) |
|---|---|---|---|---|
| Isoform0062250 | HMGB2 | NM_001035997.1 | Forward: CAAAGCTGCTGCTAAGGAC | 155 |
| Reverse: GGACTTCCACTTGTCTCCAGC | ||||
| Isoform0061796 | HMGB3 | NM_001035998.1 | Forward: CCTTCCAGTGCCTTCTTCGT | 174 |
| Reverse: CTCAACCTTGCGCTTGTCAG | ||||
| Isoform0001498 | At1g05910 | NM_100472.2 | Forward: AGACCACTCTCTCCGGTTGT | 109 |
| Reverse: TCGTCAACTCCGATGACGTG | ||||
| Isoform0062769 | RID2 | NM_125110.6 | Forward: GCAGGGCTTAGGTCTTCGTT | 100 |
| Reverse: ACGAGGTTCATGCGATGACT | ||||
| Isoform0061049 | At4g26600 | NM_001341820.1 | Forward: TTCCGATTGGTGCAACTCCT | 118 |
| Reverse: GCCATGTCCACAACTCGTTC | ||||
| Isoform0034756 | H2AV | NM_001339683.1 | Forward: CAGTTGGACGAATTCACAGGC | 176 |
| Reverse: CAGATGCCTTGGCGTTATCC | ||||
| Isoform0050517 | At2g28720 | NM_128433.4 | Forward: GCAAGAAGCTTCAAAATTAGC | 107 |
| Reverse: TGCTTAGCAAGTTCACCAGG | ||||
| Isoform0062503 | HTR2 | NM_113651.2 | Forward: CACCGGAGGAGTGAAGAAGC | 189 |
| Reverse: TCTTGAAGAGCTGCGACTGC | ||||
| Isoform0027210 | HOS15 | NM_126132.4 | Forward: TACAGGCGCAGAACCTATGG | 164 |
| Reverse: CTGTTGCATCACCAGACCCT | ||||
| Isoform0062048 | REF6 | NM_148863.4 | Forward: AGGGAACACAGCTTCTGGTG | 124 |
| Reverse: TTCCCCAAGTGAACGGTCTG | ||||
| Isoform0062818 | SKP1A | NM_106245.5 | Forward: GTGCTGCTACCTCCGATGAC | 181 |
| Reverse: GTGCGGATCTCTTCTGGAGT | ||||
| Isoform0061474 | ASK21 | NM_001125404.1 | Forward: CCTGATGACCTTACTGAGGAG | 178 |
| Reverse: CAGGTCATCCACTGAACGCT | ||||
| Isoform0061497 | SRK2G | NM_120946.5 | Forward: ACATCGAGAGAGGTCGCAAG | 110 |
| Reverse: AGGTGTCAGGATCACCTCCTT | ||||
| Isoform0062575 | SRK2H | NM_125760.2 | Forward: TGGTCCTGTGGTGTGACTCT | 164 |
| Reverse: GAGAGAAGGTGTCTGCACTCC | ||||
| Isoform0062220 | SRR1 | NM_125348.4 | Forward: ATCGCATTGTTTGGGAACAGC | 117 |
| Reverse: AGCAAACTCGCTTGTGACTCT | ||||
| Isoform0061284 | PHL | NM_001334526.1 | Forward: CAAAGTCCTCGTTTGTCGGC | 104 |
| Reverse: GCAACTGCTCCATAGTGGGT | ||||
| Isoform0057927 | PHYA | NM_001123784.1 | Forward: GTGCGATATGCTGATGCGTG | 149 |
| Reverse: CCTGCAGGTGGAACTCACTT | ||||
| Isoform0041956 | AGL62 | NM_125437.5 | Forward: CTCCTCACCAACACAACAAC | 197 |
| Reverse: AACGCAAGTTCCTCAACGGG | ||||
| Isoform0045502 | AGL79 | NM_113925.3 | Forward: AATCACCCCATGAGCTTCGC | 107 |
| Reverse: TAGGGTTCCGGCAGCTACTT | ||||
| Isoform0063170 | ATJ3 | NM_114279.4 | Forward: GAATACGCTCACGGAGTTGC | 135 |
| Reverse: GCATCCCACTTGGCTCTCTC | ||||
| Isoform0062470 | ACBP6 | NM_102916.4 | Forward: AATCACCCCATGAGCTTCGC | 107 |
| Reverse: TAGGGTTCCGGCAGCTACTT | ||||
| Isoform0061783 | ENO2 | NM_129209.4 | Forward: CACTGAGTGTGGAACCGAGG | 190 |
| Reverse: GGTCATCACTCCCCAACCTG | ||||
| Isoform0063049 | ADH1 | NM_106362.3 | Forward: TGTGACCGAGTGTGTGAACC | 123 |
| Reverse: TGAATCATGGCCTGAACGCT | ||||
| Isoform0062198 | CSP2 | NM_120029.3 | Forward: GATCTGGAGGTGGATACGGC | 115 |
| Reverse: CAGTCTCTCGCCATGTGACC | ||||
| Isoform0062617 | HSP17.8 | NM_100614.3 | Forward: AACATCGGCGATAACGAACG | 154 |
| Reverse: CTCCACGTGTCTCTCTCCAC | ||||
| Isoform0009507 | HSP70-3 | NM_001202918.1 | Forward: CGACTGCAGGAGACACTCAT | 144 |
| Reverse: TCTCACAGGCGGTTCTCAAC | ||||
| Isoform0034676 | HSP70-10 | NM_120996.4 | Forward: CGTGTCCCCAAGGTTCAGTC | 167 |
| Reverse: CCGAGCGATAGAGGTGTGAC | ||||
| Isoform0042993 | HSP90-3 | NM_124983.4 | Forward: AACAAGGAGGAGTACGCTGC | 193 |
| Reverse: AGACACGACGGACATAGAGC | ||||
| Isoform0061974 | CSY4 | NM_001337082.1 | Forward: GATGCAGAGCTCTACCGACC | 197 |
| Reverse: CCTCTTCCGGGTCAAGCAAT | ||||
| Isoform0062375 | GAPCP2 | NM_101496.3 | Forward: CATTTCTGCACCTTCAGCGG | 198 |
| Reverse: TTCTGAGTAGCTGTGGTCGC | ||||
| Isoform0062268 | At3g52990 | NM_115159.5 | Forward: GACAACTTGCGACCAACTCG | 101 |
| Reverse: AATCCACGAAGGGTCTCAGC | ||||
| Isoform0062585 | PGM2 | NM_001160993.2 | Forward: TGAACTGCGTACCCAAGGAG | 158 |
| Reverse: TCGGTCTGCATCACCATCAG | ||||
| Isoform0062370 | BAM1 | NM_113297.3 | Forward: AACTCTCTCGCTGTTCCTCG | 165 |
| Reverse: GGAGAAGCCCGTCTCACAAT | ||||
| Isoform0062586 | ARF1 | NM_001337250.1 | Forward: GTGACCGTGTTGTTGAAGCC | 148 |
| Reverse: TGAAGCCCAAGCTTGTCAGT | ||||
| Isoform0061377 | CUL1 | NM_001036498.3 | Forward: GTGCCGTGCATTGCTAAGAG | 153 |
| Reverse: TCTTCGGCCTGTTGGACAAG | ||||
| Isoform0057235 | SOFL4 | NM_123240.2 | Forward: AGGTCGTGGATGAGGACTAC | 144 |
| Reverse: GAACCGCTGATAATTTGGCCC | ||||
| Isoform0043114 | SOFL5 | NM_001342234.1 | Forward: TGCGAGTCAGGATGGACTCT | 193 |
| Reverse: TCCTTGGACCAGAAGAAGCAT | ||||
| Isoform0062152 | GRF2 | NM_106479.3 | Forward: AACTCTCCGGAATCTGCGAC | 192 |
| Reverse: GAGCAGATTTGTAAGCGGCG | ||||
| Isoform0062828 | GRF11 | NM_001084180.2 | Forward: GGTGCTAGGAGAGCATCGTG | 198 |
| Reverse: GACGGTGGATTCTCCCGAAG | ||||
| Isoform0061395 | ERF3 | NM_103946.3 | Forward: ATCGTTTAGCGGACCCAGAC | 101 |
| Reverse: CGCAATCGCTGTGACAATCC | ||||
| Isoform0022533 | SF1 | NM_001036978.2 | Forward: GGCTTAGGGTCAACTCCGAC | 164 |
| Reverse: CCAGTCACACGGTCCTTGAT | ||||
| Isoform0044730 | PURU1 | NM_124115.4 | Forward: ACGTCTTCTACTCTCGCAGC | 125 |
| Reverse: AGGCACACGCACAACTGAAT | ||||
| Isoform0047216 | MES16 | NM_117770.5 | Forward: CCATCCCTTCTCCGCATCTT | 195 |
| Reverse: TCATAGGAGCAGGACGCAAC | ||||
| Isoform0063248 | GPT1 | NM_124861.5 | Forward: CGCTGGTTCGTTGATGATGC | 193 |
| Reverse: AAACGCAGGTTCACCACTCT | ||||
| Isoform0062571 | ABCF5 | NM_125882.3 | Forward: TGCTGATAGGCTTGTGGCTT | 103 |
| Reverse: CGGCTCATCAAGTAGCAGCA | ||||
| Isoform0008194 | SECA2 | NM_001198130.1 | Forward: ACTGTGAGGCCCATTGTCTG | 117 |
| Reverse: CTCTGCCACGAAGCTGGTTA | ||||
| Isoform0062373 | VPS26A | NM_124733.4 | Forward: TGTTCCGCTTCCTCCAATCAA | 196 |
| Reverse: TGCTCCAGTTGATTCTCGCC | ||||
| Isoform0062449 | CML19 | NM_119864.5 | Forward: CGAGCTCAACGTTGCTATGAG | 160 |
| Reverse: GTCTATGGAGTCTCGTTCTCCG | ||||
| Isoform0061307 | NHX6 | NM_106609.4 | Forward: GGCATTTGCTCTTGCTCTGC | 112 |
| Reverse: TCCTCCAATCAGCAACACCG |
| lncRNA ID | Coexpressed mRNAs | mRNA ID | Proteins Encoded by Coexpressed mRNAs |
|---|---|---|---|
| Chromatin modification (2) | |||
| Isoform0062250 | HMGB2 | AT1G20693.1 | High mobility group B protein 2 |
| Isoform0061796 | HMGB3 | AT1G20696.1 | High mobility group B protein 3 |
| DNA/RNA modification (3) | |||
| Isoform0001498 | At1g05910 | AT1G05910.1 | ATPase family AAA domain-containing protein At1g05910 |
| Isoform0062769 | RID2 | AT5G57280.1 | 18S rRNA (guanine-N(7))-methyltransferase RID2 e |
| Isoform0061049 | At4g26600 | AT4G26600.8 | S-adenosyl-L-methionine-dependent methyltransferases superfamily protein |
| Protein modification (24) | |||
| Isoform0034756 | H2AV | AT3G54560.1 | Histone H2A variant 1 |
| Isoform0050517 | At2g28720 | AT2G28720.1 | Histone H2B.3 |
| Isoform0062503 | HTR2 | AT1G09200.1 | Histone H3.2 |
| Isoform0027210 | HOS15 | AT5G67320.1 | WD40 repeat-containing protein HOS15 |
| Isoform0062048 | REF6 | AT3G48430.1 | Lysine-specific demethylase REF6 |
| Isoform0062818 | SKP1A | AT1G75950.1 | SKP1-like protein 1A |
| Isoform0061474 | ASK21 | AT3G61415.1 | SKP1-like protein 21 |
| Isoform0061497 | SRK2G | AT5G08590.1 | Serine/threonine-protein kinase SRK2G |
| Isoform0062575 | SRK2H | AT5G63650.1 | Serine/threonine-protein kinase SRK2H |
| Isoform0053138 | DET1 | AT4G10180.1 | Light-mediated development protein DET1 |
| Isoform0030135 | BOPAt4g295601 | AT3G57130.2 | Ankyrin repeat family protein/BTB/POZ domain-containing protein |
| Isoform0063715 | At1g45180 | AT1G45180.1 | F27F5.26 |
| Isoform0044146 | At3g50840 | AT3G50840.3 | Phototropic-responsive NPH3 family protein |
| Isoform0034163 | At2g36630 | AT2G36630.1 | Sulfite exporter TauE/SafE family protein 4 |
| Isoform0058941 | At3g47890 | AT3G47890.2 | Ubiquitin carboxyl-terminal hydrolase-related protein |
| Isoform0007659 | UBP7 | A0A1I9LL79 | Ubiquitin carboxyl-terminal hydrolase |
| Isoform0045576 | DER2.1 | AT3G21280.2 | Derlin-2.1 |
| Isoform0062510 | GRP3 | AT2G05520.1 | Glycine-rich protein 3 |
| Isoform0062575 | MDH9.13 | AT5G42440.1 | At5g42440 |
| Isoform0051826 | At3g24715 | AT3G24715.1 | Kinase superfamily with octicosapeptide/Phox/Bem1p domain-containing protein |
| Isoform0061548 | At3g16560 | AT3G16560.4 | Probable protein phosphatase 2C 40 |
| Isoform0049189 | At3g62260 | AT3G62260.1 | Probable protein phosphatase 2C 49 |
| Isoform0060667 | ESMD1 | AT2G01480.1 | Protein ESMERALDA 1 |
| Isoform0062775 | At1g27930 | AT1G27930.1 | Probable methyltransferase At1g27930 |
| lncRNA ID | Coexpressed mRNAs | mRNA ID | Proteins Encoded by Coexpressed mRNAs |
|---|---|---|---|
| Isoform0062220 | SRR1 | AT5G59560.2 | Protein SENSITIVITY TO RED LIGHT REDUCED 1 |
| Isoform0061284 | PHL | AT1G72390.1 | Protein PHYTOCHROME-DEPENDENT LATE-FLOWERING |
| Isoform0057927 | PHYA | AT1G09570.6 | Phytochrome A |
| Isoform0041956 | AGL62 | AT5G60440.1 | Agamous-like MADS-box protein AGL62 |
| Isoform0045502 | AGL79 | AT3G30260.1 | AGAMOUS-like 79 |
| Isoform0063170 | ATJ3 | AT3G44110.1 | Chaperone protein dnaJ 3 |
| Isoform0028325 | BBX29 | AT5G54470.1 | At5g54470 |
| Isoform0054894 | CLE13 | AT1G73965.1 | CLAVATA3/ESR (CLE)-related protein 13 |
| Isoform0061298 | CLE44 | AT4G13195.1 | CLAVATA3/ESR (CLE)-related protein 44 |
| Isoform0056751 | MXC17.10 | AT5G24710.1 | Transducin/WD40 repeatlike superfamily protein |
| Isoform0062524 | At1g06515 | AT1G06515.2 | Transmembrane protein, putative (DUF3317) |
| Isoform0061573 | BHLH30 | AT1G68810.1 | Transcription factor bHLH30 |
| lncRNA ID | Coexpressed mRNAs | mRNA ID | Proteins Encoded by Coexpressed mRNAs |
|---|---|---|---|
| Isoform0062470 | ACBP6 | AT1G31812.1 | Acyl-CoA-binding domain-containing protein 6 |
| Isoform0061783 | ENO2 | AT2G36530.1 | Bifunctional enolase 2/transcriptional activator |
| Isoform0063049 | ADH1 | AT1G77120.1 | Alcohol dehydrogenase class-P |
| Isoform0062198 | CSP2 | AT4G38680.1 | Cold shock protein 2 |
| Isoform0035932 | RH20 | AT1G55150.2 | DEA(D/H)-box RNA helicase family protein |
| Isoform0019851 | RH52 | AT3G58570.1 | DEAD-box ATP-dependent RNA helicase 52 |
| Isoform0062484 | RH53 | AT3G22330.1 | DEAD-box ATP-dependent RNA helicase 53, mitochondrial |
| Isoform0058095 | RAB18 | AT5G66400.1 | Dehydrin Rab18 |
| Isoform0061968 | XERO1 | AT3G50980.1 | Dehydrin Xero 1 |
| Isoform0020919 | MED14 | AT3G04740.1 | Mediator of RNA polymerase II transcription subunit 14 |
| Isoform0062617 | HSP17.8 | AT1G07400.1 | 17.8 kDa class I heat shock protein |
| Isoform0009507 | HSP70-3 | AT3G09440.2 | Heat shock 70 kDa protein 3 |
| Isoform0034676 | HSP70-10 | AT5G09590.1 | Heat shock 70 kDa protein 10, mitochondrial |
| Isoform0042993 | HSP90-3 | AT5G56010.1 | Heat shock protein 90-3 |
| lncRNA ID | Coexpressed mRNAs | mRNA ID | Proteins Encoded by Coexpressed mRNAs |
|---|---|---|---|
| Isoform0061974 | CSY4 | AT2G44350.2 | Citrate synthase 4, mitochondrial |
| Isoform0062251 | FBA3 | AT2G01140.1 | Fructose-bisphosphate aldolase 3, chloroplastic |
| Isoform0062131 | GAPC1 | AT3G04120.1 | Glyceraldehyde-3-phosphate dehydrogenase GAPC1, cytosolic |
| Isoform0062375 | GAPCP2 | AT1G16300.1 | Glyceraldehyde-3-phosphate dehydrogenase GAPCP2, chloroplastic |
| Isoform0062268 | At3g52990 | AT3G52990.1 | Pyruvate kinase |
| Isoform0062585 | PGM2 | AT1G70730.3 | Phosphoglucomutase (alpha-D-glucose-1,6-bisphosphate-dependent) |
| Isoform0063034 | USP | AT5G52560.1 | UDP-sugar pyrophosphorylase |
| Isoform0042435 | GLCNAC1PUT2 | AT2G35020.1 | UDP-N-acetylglucosamine diphosphorylase 2 |
| Isoform0062018 | UXS2 | AT3G62830.1 | UDP-glucuronic acid decarboxylase 2 |
| Isoform0062750 | XYLA | AT5G57655.2 | Xylose isomerase |
| Isoform0060700 | GALS1 | AT2G33570.1 | Galactan beta-1,4-galactosyltransferase GALS1 |
| Isoform0062032 | OFUT31 | AT4G24530.1 | O-fucosyltransferase 31 |
| Isoform0039872 | At5g67460 | AT5G67460.1 | Glucan endo-1,3-beta-D-glucosidase |
| Isoform0045893 | RSW3 | AT5G63840.2 | Glycosyl hydrolases family 31 protein |
| Isoform0036200 | At1g59950 | AT1G59950.1 | Aldo/keto reductase |
| Isoform0042053 | At5g25970 | AT5G25970.2 | Core-2/I-branching beta-1,6-N-acetylglucosaminyl-transferase family protein |
| Isoform0059181 | UGT76E7 | AT5G38040.1 | UDP-glycosyltransferase 76E7 |
| Isoform0037698 | At1g26850 | AT1G26850.2 | Probable methyltransferase PMT2 |
| Isoform0062370 | BAM1 | AT3G23920.1 | Beta-amylase 1, chloroplastic |
| lncRNA ID | Coexpressed mRNAs | mRNA ID | Proteins Encoded by Coexpressed mRNAs |
|---|---|---|---|
| Isoform0062586 | ARF1 | AT2G47170.1 | ADP-ribosylation factor 1 |
| Isoform0061377 | CUL1 | AT4G02570.1 | Cullin-1 |
| Isoform0015752 | T4L20.330 | AT4G34750.1 | SAUR-like auxin-responsive protein family |
| Isoform0057235 | SOFL4 | AT5G38790.1 | Protein SOB FIVE-LIKE 4 |
| Isoform0043114 | SOFL5 | AT4G33800.2 | Protein SOB FIVE-LIKE 5 |
| Isoform0062152 | GRF2 | AT1G78300.1 | 14-3-3-like protein GF14 omega |
| Isoform0062828 | GRF11 | AT1G34760.1 | 14-3-3-like protein GF14 omicron |
| Isoform0061395 | ERF3 | AT1G50640.1 | Ethylene-responsive transcription factor 3 |
| Isoform0007735 | CIPK20 | AT5G45820.1 | CBL-interacting serine/threonine-protein kinase 20 |
| Isoform0022533 | SF1 | AT5G51300.2 | Splicing factorlike protein 1 |
| Isoform0046311 | AGD9 | AT5G46750.1 | Probable ADP-ribosylation factor GTPase-activating protein AGD9 |
| Isoform0057859 | TIFY4B | AT4G14720.1 | Protein TIFY 4B |
| Isoform0062437 | At2g34810 | AT2G34810.1 | Berberine bridge enzyme-like 16 |
| lncRNA ID | Coexpressed mRNAs | mRNA ID | Proteins Encoded by Coexpressed mRNAs |
|---|---|---|---|
| Energy (5) | |||
| Isoform0044730 | PURU1 | AT5G47435.2 | Formyltetrahydrofolate deformylase 1, mitochondrial |
| Isoform0031698 | ndhB1 | ATCG01250.1 | NAD(P)H-quinone oxidoreductase subunit 2 A, chloroplastic |
| Isoform0029144 | NDB1 | AT4G28220.2 | NADH:ubiquinone reductase (nonelectrogenic) |
| Isoform0046995 | WNK9 | AT5G28080.3 | Nonspecific serine/threonine protein kinase |
| Isoform0047216 | MES16 | AT4G16690.1 | pFDCC methylesterase MES16 |
| Transport (38) | |||
| Isoform0047603 | At5g11230 | AT5G11230.1 | Probable sugar phosphate/phosphate translocator At5g11230 |
| Isoform0063248 | GPT1 | AT5G54800.1 | Glucose-6-phosphate/phosphate translocator 1, chloroplastic |
| Isoform0053163 | SMXL5 | AT5G57130.1 | Protein SMAX1-LIKE 5 |
| Isoform0062571 | ABCF5 | AT5G64840.1 | ABC transporter F family member 5 |
| Isoform0062182 | SFH8 | AT2G21520.2 | Phosphatidylinositol/phosphatidylcholine transfer protein SFH8 |
| Isoform0022607 | BASS6 | AT4G22840.1 | Probable sodium/metabolite cotransporter BASS6, chloroplastic |
| Isoform0008194 | SECA2 | AT1G21650.3 | Protein translocase subunit SECA2, chloroplastic |
| Isoform0014041 | At4g14160 | AT4G14160.1 | Protein transport protein SEC23 |
| Isoform0062660 | ycf2-A | ATCG01280.1 | Protein Ycf2 |
| Isoform0036773 | At4g22990 | AT4G22990.1 | SPX domain-containing membrane protein At4g22990 |
| Isoform0032676 | At3g49350 | AT3G49350.1 | At3g49350 |
| Isoform0062443 | TMT2 | AT4G35300.5 | Tonoplast monosaccharide transporter2 |
| Isoform0003274 | VPS24-1 | AT5G22950.1 | Vacuolar protein sorting-associated protein 24 homolog 1 |
| Isoform0062373 | VPS26A | AT5G53530.1 | Vacuolar protein sorting-associated protein 26A |
| Isoform0028575 | VPS52 | AT1G71270.1 | Vacuolar protein sorting-associated protein 52 A |
| Isoform0038654 | VPS60-2 | AT5G04850.1 | Vacuolar protein sorting-associated protein 60.2 |
| Isoform0062996 | At5g19500 | AT5G19500.1 | At5g19500 |
| Isoform0062800 | PIP1-5 | AT4G23400.1 | Probable aquaporin PIP1-5 |
| Isoform0061992 | SULTR2;2 | AT1G77990.1 | Sulfate transporter 2.2 |
| Isoform0062606 | MT2A | AT3G09390.2 | Metallothionein-like protein 2A |
| Isoform0057136 | At3g52300 | AT3G52300.1 | ATP synthase subunit d, mitochondrial |
| Isoform0061450 | ABCB23 | AT4G28630.1 | ABC transporter B family member 23, mitochondrial |
| Isoform0062449 | CML19 | AT4G37010.1 | Calcium-binding protein CML19 |
| Isoform0061763 | JJJ1 | AT1G74250.1 | DNAJ protein JJJ1 homolog |
| Isoform0061955 | PAP1 | AT1G13750.1 | Probable inactive purple acid phosphatase 1 |
| Isoform0062074 | HIPP04 | AT1G29000.2 | Heavy metal-associated isoprenylated plant protein 4 |
| Isoform0062972 | HIPP26 | AT4G38580.1 | Heavy metal-associated isoprenylated plant protein 26 |
| Isoform0014712 | KINUA | AT1G12430.1 | Kinesin-like protein KIN-UA |
| Isoform0063051 | IQD30 | AT1G18840.2 | Protein IQ-DOMAIN 30 |
| Isoform0016777 | At3g18430 | AT3G18430.1 | Calcineurin b subunit (Protein phosphatase 2b regulatory subunit)-like protein |
| Isoform0063921 | CBL10 | AT4G33000.6 | Calcineurin B-like protein 10 |
| Isoform0061307 | NHX6 | AT1G79610.1 | Sodium/hydrogen exchanger 6 |
| Isoform0024682 | CNGC15 | AT2G28260.2 | Cyclic nucleotide-gated channel 15 |
| Isoform0034638 | 2-Oct | AT1G79360.1 | Organic cation/carnitine transporter 2 |
| Isoform0033810 | CNGC13 | AT4G01010.2 | Putative cyclic nucleotide-gated ion channel 13 |
| Isoform0051075 | MTG13.4 | AT5G16680.1 | RING/FYVE/PHD zinc finger superfamily protein |
| Isoform0051563 | SUF4 | AT1G30970.3 | Zinc finger (C2H2 type) family protein |
| Isoform0037483 | AHA4 | AT3G47950.2 | ATPase 4, plasma membranetype |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, X.; Luo, M.; Li, M.; Wei, J. Transcriptomic Analysis Reveals LncRNAs Associated with Flowering of Angelica sinensis during Vernalization. Curr. Issues Mol. Biol. 2022, 44, 1867-1888. https://doi.org/10.3390/cimb44050128
Liu X, Luo M, Li M, Wei J. Transcriptomic Analysis Reveals LncRNAs Associated with Flowering of Angelica sinensis during Vernalization. Current Issues in Molecular Biology. 2022; 44(5):1867-1888. https://doi.org/10.3390/cimb44050128
Chicago/Turabian StyleLiu, Xiaoxia, Mimi Luo, Mengfei Li, and Jianhe Wei. 2022. "Transcriptomic Analysis Reveals LncRNAs Associated with Flowering of Angelica sinensis during Vernalization" Current Issues in Molecular Biology 44, no. 5: 1867-1888. https://doi.org/10.3390/cimb44050128








