The Telomere-Telomerase System Is Detrimental to Health at High-Altitude
Abstract
:1. Introduction
2. Materials and Methods
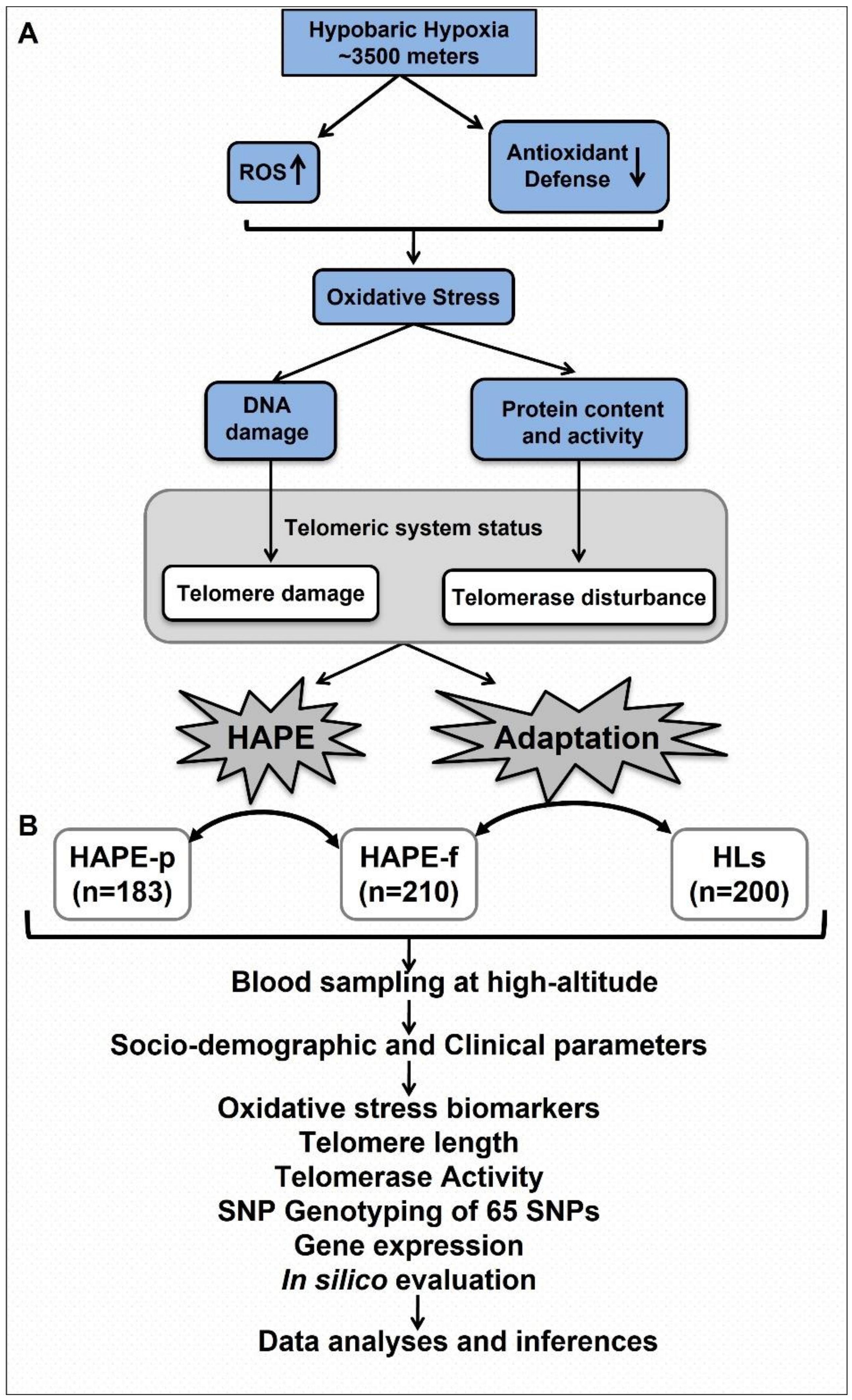
2.1. Study Subjects
2.2. Ethics Approval
2.3. Estimation of Relative Telomere Length
2.4. Estimation of Telomerase Activity and Oxidative Stress Markers
2.5. Genotyping of Single Nucleotide Polymorphisms
2.6. Genetic Analyses
2.7. Gene-Gene Interactions in HAPE and Adaptation
2.8. DNA-Protein Docking
2.9. Protein-Protein Interactions
2.10. Expression Analysis of Telomere Maintaining Genes
2.11. Statistical Analyses
3. Results
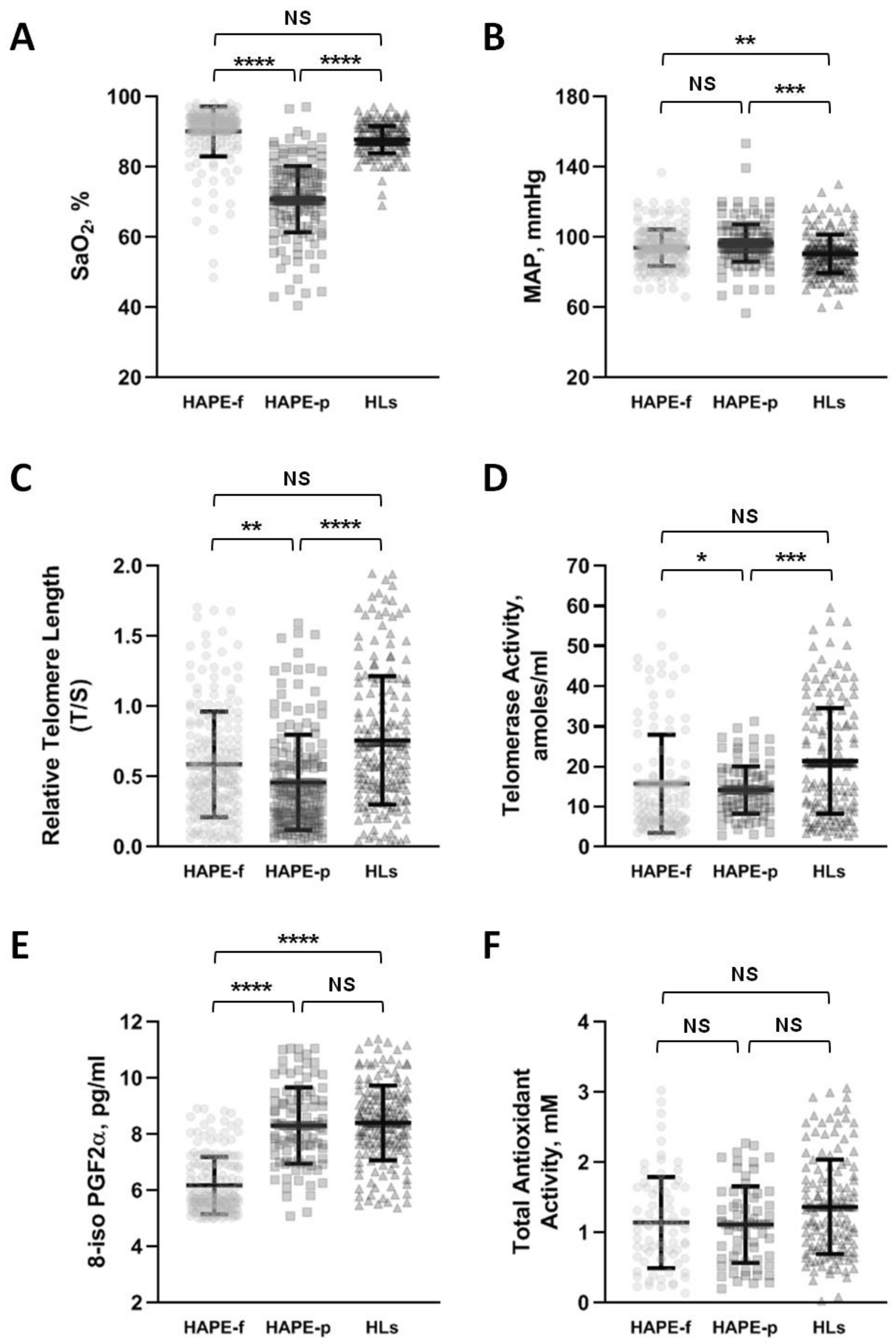
3.1. Low SaO2 Indicates HAPE
3.2. Aberrant Telomere-Telomerase System Inclined with HAPE Susceptibility
3.3. The Oxidative System Shows Stronger Influence
3.4. Genetic Association with HAPE and HA Adaptation
3.5. Haplotypes Distribution Is Specific in HAPE and HA Adaptation
3.6. Gene-Gene Interactions
3.6.1. Best Interaction Models for HAPE
3.6.2. Best Interaction Models for HA Adaptation
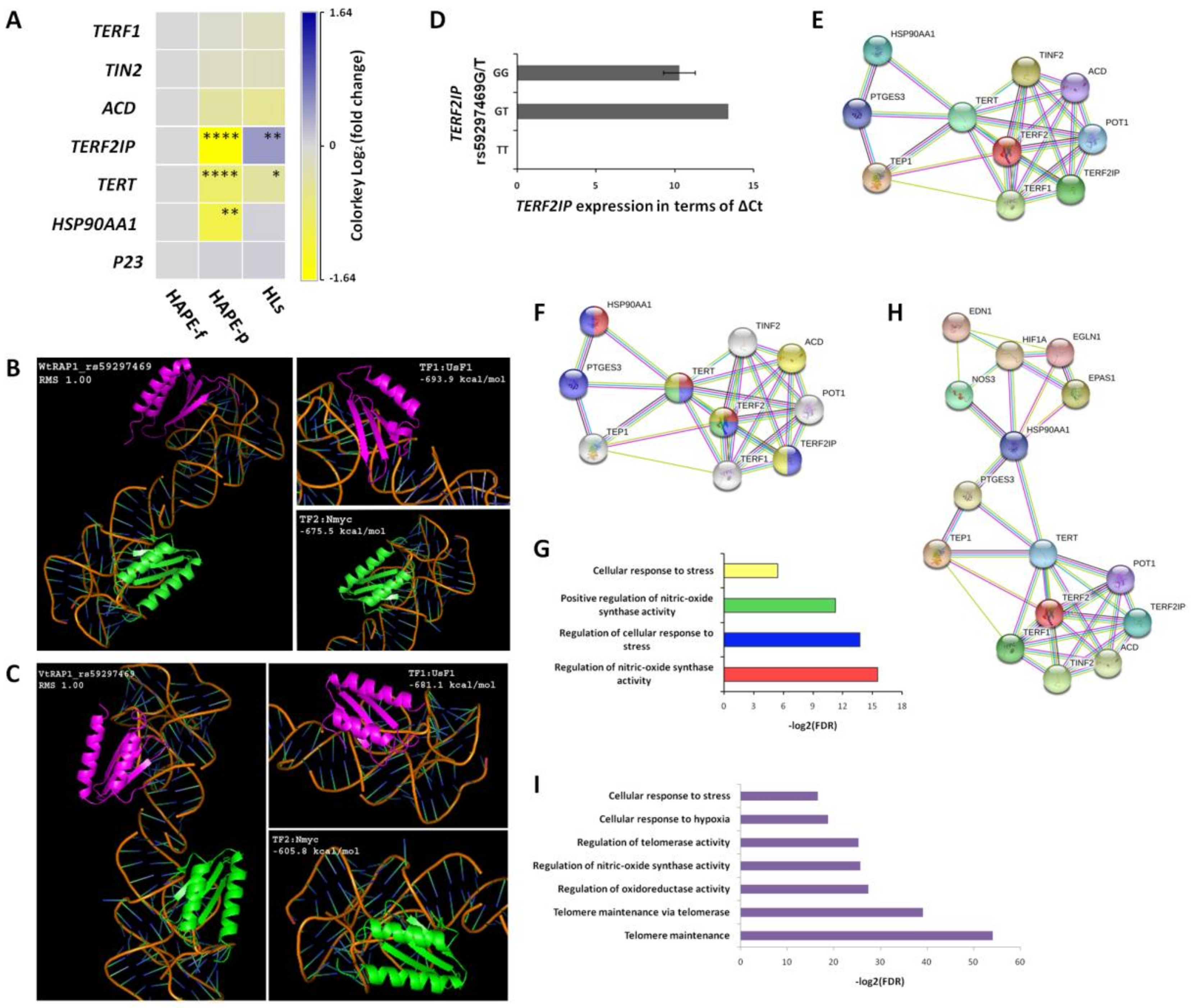
3.7. Expression Profile Differed for Telomere Maintaining Genes
3.8. In Silico Analysis Underlined the Association of Decreased TERF2IP Expression with Risk Alleles in HAPE
3.9. Protein-Protein Interactions
3.10. Correlation between Biomarkers and Clinical Parameters
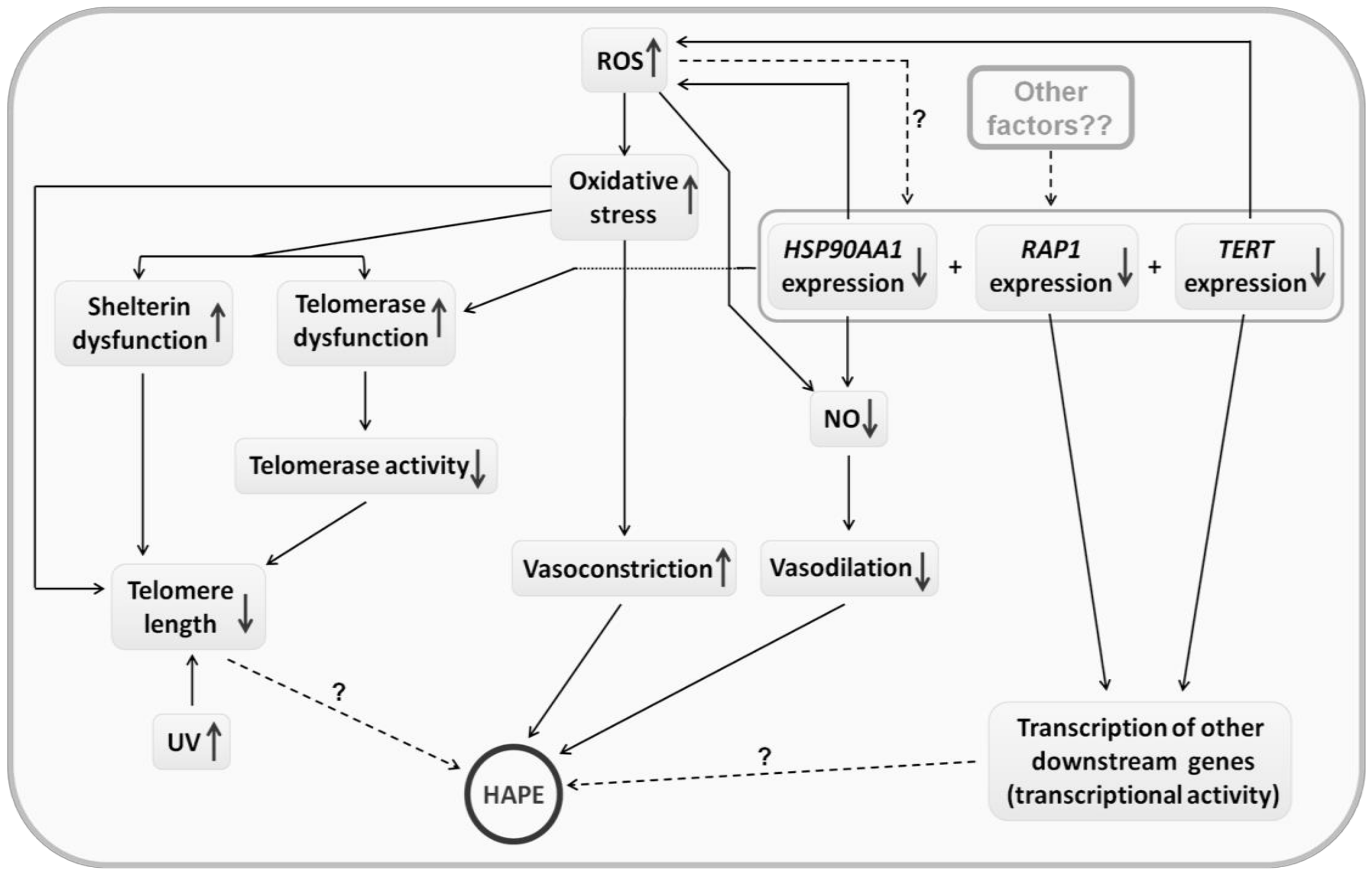
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mishra, A.; Kohli, S.; Dua, S.; Thinlas, T.; Mohammad, G.; Pasha, M.A. Genetic differences and aberrant methylation in the apelin system predict the risk of high-altitude pulmonary edema. Proc. Natl. Acad. Sci. USA 2015, 112, 6134–6139. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beall, C.M. Andean, Tibetan, and Ethiopian patterns of adaptation to high-altitude hypoxia. Integr. Comp. Biol. 2006, 46, 18–24. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bärtsch, P.; Mairbäurl, H.; Swenson, E.R.; Maggiorini, M. High altitude pulmonary oedema. Swiss Med. Wkly. 2003, 133, 377–384. [Google Scholar] [CrossRef] [PubMed]
- Bärtsch, P.; Mairbäurl, H.; Maggiorini, M.; Swenson, E.R. Physiological aspects of high-altitude pulmonary edema. J. Appl. Physiol. 2005, 98, 1101–1110. [Google Scholar] [CrossRef] [Green Version]
- Maggiorini, M. High altitude-induced pulmonary oedema. Cardiovasc. Res. 2006, 72, 41–50. [Google Scholar] [CrossRef] [Green Version]
- Dehnert, C.; Berger, M.M.; Mairbäurl, H.; Bärtsch, P. High altitude pulmonary edema: A pressure-induced leak. Respir. Physiol. Neurobiol. 2007, 158, 266–273. [Google Scholar] [CrossRef]
- Swenson, S.; Maggiorini, M.; Mongovin, S.; Gibbs, S.; Greve, I.; Maierbaurl, H.; Bärtsch, P.J.J. High altitude pulmonary edema is a non-inflammatory high permeability leak of the alveolar-capillary barrier. JAMA 2002, 287, 2226–2235. [Google Scholar]
- Wodopia, R.; Ko, H.S.; Billian, J.; Wiesner, R.; Bärtsch, P.; Mairbäurl, H. Hypoxia decreases proteins involved in epithelial electrolyte transport in A549 cells and rat lung. Am. J. Physiol. Lung Cell Mol. Physiol. 2000, 279, L1110–L1119. [Google Scholar] [CrossRef]
- Irwin, D.C.; McCord, J.M.; Nozik-Grayck, E.; Beckly, G.; Foreman, B.; Sullivan, T.; White, M.; T Crossno, J.; Bailey, D.; Flores, S.C.; et al. A potential role for reactive oxygen species and the HIF-1alpha-VEGF pathway in hypoxia-induced pulmonary vascular leak. Free. Radic. Biol. Med. 2009, 47, 55–61. [Google Scholar] [CrossRef] [Green Version]
- Baloglu, E.; Reingruber, T.; Bärtsch, P.; Mairbäurl, H. β2-Adrenergics in hypoxia desensitize receptors but blunt inhibition of reabsorption in rat lungs. Am. J. Respir. Cell. Mol. Biol. 2011, 45, 1059–1068. [Google Scholar] [CrossRef]
- Remillard, C.V.; Yuan, J.X. High altitude pulmonary hypertension: Role of K+ and Ca2+ channels. High Alt. Med. Biol. 2005, 6, 133–146. [Google Scholar] [CrossRef] [PubMed]
- Mishra, A.; Mohammad, G.; Norboo, T.; Newman, J.H.; Pasha, M.A. Lungs at high-altitude: Genomic insights into hypoxic responses. J. Appl. Physiol. 2015, 119, 1–15. [Google Scholar] [CrossRef]
- Ali, Z.; Mishra, A.; Kumar, R.; Alam, P.; Pandey, P.; Ram, R.; Thinlas, T.; Mohammad, G.; Pasha, M.A.Q. Interactions among Vascular-Tone Modulators Contribute to High Altitude Pulmonary Edema and Augmented Vasoreactivity in Highlanders. PLoS ONE 2012, 7, e44049. [Google Scholar] [CrossRef] [PubMed]
- Sinha, S.; Dutta, A.; Singh, S.N.; Ray, U.S. Protein nitration, lipid peroxidation and DNA damage at high altitude in acclimatized lowlanders and native highlanders: Relation with oxygen consumption. Respir. Physiol. Neurobiol. 2010, 171, 115–121. [Google Scholar] [CrossRef] [PubMed]
- Oikawa, S.; Tada-Oikawa, S.; Kawanishi, S. Site-specific DNA damage at the GGG sequence by UVA involves acceleration of telomere shortening. Biochemistry 2001, 40, 4763–4768. [Google Scholar] [CrossRef]
- Denchi, E.L.; de Lange, T. Protection of telomeres through independent control of ATM and ATR by TRF2 and POT1. Nature 2007, 448, 1068–1071. [Google Scholar] [CrossRef]
- Fitzpatrick, A.L.; Kronmal, R.A.; Gardner, J.P.; Psaty, B.M.; Jenny, N.S.; Tracy, R.P.; Walston, J.; Kimura, M.; Aviv, A. Leukocyte telomere length and cardiovascular disease in the cardiovascular health study. Am. J. Epidemiol. 2007, 165, 14–21. [Google Scholar] [CrossRef]
- Zhao, J.; Zhu, Y.; Lin, J.; Matsuguchi, T.; Blackburn, E.; Zhang, Y.; Cole, S.A.; Best, L.G.; Lee, E.T.; Howard, B.V. Short leukocyte telomere length predicts risk of diabetes in american indians: The strong heart family study. Diabetes 2014, 63, 354–362. [Google Scholar] [CrossRef] [Green Version]
- Morgan, R.G.; Ives, S.J.; Walker, A.E.; Cawthon, R.M.; Andtbacka, R.H.; Noyes, D.; Lesniewski, L.A.; Richardson, R.S.; Donato, A.J. Role of arterial telomere dysfunction in hypertension: Relative contributions of telomere shortening and telomere uncapping. J. Hypertens. 2014, 32, 1293–1299. [Google Scholar] [CrossRef] [Green Version]
- Córdoba-Lanús, E.; Cazorla-Rivero, S.; Espinoza-Jiménez, A.; de-Torres, J.P.; Pajares, M.J.; Aguirre-Jaime, A.; Celli, B.; Casanova, C. Telomere shortening and accelerated aging in COPD: Findings from the BODE cohort. Respir. Res. 2017, 18, 59. [Google Scholar] [CrossRef] [Green Version]
- Jin, M.; Lee, E.C.; Ra, S.W.; Fishbane, N.; Tam, S.; Criner, G.J.; Woodruff, P.G.; Lazarus, S.C.; Albert, R.; Connett, J.E.; et al. Relationship of Absolute Telomere Length With Quality of Life, Exacerbations, and Mortality in COPD. Chest 2018, 154, 266–273. [Google Scholar] [CrossRef] [PubMed]
- Wilber, R.L. Application of altitude/hypoxic training by elite athletes. Med. Sci. Sports Exerc. 2007, 39, 1610–1624. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Borghini, A.; Giardini, G.; Tonacci, A.; Mastorci, F.; Mercuri, A.; Mrakic-Sposta, S.; Moretti, S.; Andreassi, M.G.; Pratali, L. Chronic and acute effects of endurance training on telomere length. Mutagenesis 2015, 30, 711–716. [Google Scholar] [CrossRef] [PubMed]
- Denham, J.; O’Brien, B.J.; Prestes, P.R.; Brown, N.J.; Charchar, F.J. Increased expression of telomere-regulating genes in endurance athletes with long leukocyte telomeres. J. Appl. Physiol. (1985) 2016, 120, 148–158. [Google Scholar] [CrossRef] [Green Version]
- Werner, C.M.; Hecksteden, A.; Morsch, A.; Zundler, J.; Wegmann, M.; Kratzsch, J.; Thiery, J.; Hohl, M.; Bittenbring, J.T.; Neumann, F.; et al. Differential effects of endurance, interval, and resistance training on telomerase activity and telomere length in a randomized, controlled study. Eur. Heart J. 2019, 40, 34–46. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ribeiro, V.B.; Pedroso, D.C.C.; Kogure, G.S.; Lopes, I.P.; Santana, B.A.; Dutra de Souza, H.C.; Ferriani, R.A.; Calado, R.T.; Furtado, C.L.M.; Reis, R.M.D. Short-Term Aerobic Exercise Did Not Change Telomere Length While It Reduced Testosterone Levels and Obesity Indexes in PCOS: A Randomized Controlled Clinical Trial Study. Int. J. Environ. Res. Public Health 2021, 18, 11274. [Google Scholar] [CrossRef] [PubMed]
- Schellnegger, M.; Lin, A.C.; Hammer, N.; Kamolz, L.P. Physical Activity on Telomere Length as a Biomarker for Aging: A Systematic Review. Sports Med. Open 2022, 8, 111. [Google Scholar] [CrossRef]
- de Lange, T. Shelterin: The protein complex that shapes and safeguards human telomeres. Genes Dev. 2005, 19, 2100–2110. [Google Scholar] [CrossRef] [Green Version]
- Xin, H.; Liu, D.; Wan, M.; Safari, A.; Kim, H.; Sun, W.; O’Connor, M.S.; Songyang, Z. TPP1 is a homologue of ciliate TEBP-beta and interacts with POT1 to recruit telomerase. Nature 2007, 445, 559–562. [Google Scholar] [CrossRef]
- Sfeir, A.; Kosiyatrakul, S.T.; Hockemeyer, D.; MacRae, S.L.; Karlseder, J.; Schildkraut, C.L.; de Lange, T. Mammalian telomeres resemble fragile sites and require TRF1 for efficient replication. Cell 2009, 138, 90–103. [Google Scholar] [CrossRef] [Green Version]
- Abreu, E.; Aritonovska, E.; Reichenbach, P.; Cristofari, G.; Culp, B.; Terns, R.M.; Lingner, J.; Terns, M.P. TIN2-tethered TPP1 recruits human telomerase to telomeres in vivo. Mol. Cell Biol. 2010, 30, 2971–2982. [Google Scholar] [CrossRef]
- Cesare, A.J.; Reddel, R.R. Alternative lengthening of telomeres: Models, mechanisms and implications. Nat. Rev. Genet. 2010, 11, 319–330. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.-Y.; Zhang, Y.; Zhang, Q.; Li, H.; Luo, Z.; Fang, H.; Kim, S.H.; Qin, L.; Yotnda, P.; Xu, J.; et al. Mitochondrial localization of telomeric protein TIN2 links telomere regulation to metabolic control. Mol. Cell 2012, 47, 839–850. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Santos, J.H.; Meyer, J.N.; Van Houten, B. Mitochondrial localization of telomerase as a determinant for hydrogen peroxide-induced mitochondrial DNA damage and apoptosis. Hum. Mol. Genet. 2006, 15, 1757–1768. [Google Scholar] [CrossRef] [PubMed]
- Haendeler, J.; Dröse, S.; Büchner, N.; Jakob, S.; Altschmied, J.; Goy, C.; Spyridopoulos, I.; Zeiher, A.M.; Brandt, U.; Dimmeler, S. Mitochondrial telomerase reverse transcriptase binds to and protects mitochondrial DNA and function from damage. Arterioscler. Thromb. Vasc. Biol. 2009, 29, 929–935. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Compton, S.A.; Elmore, L.W.; Haydu, K.; Jackson-Cook, C.K.; Holt, S.E. Induction of nitric oxide synthase-dependent telomere shortening after functional inhibition of Hsp90 in human tumor cells. Mol. Cell. Biol. 2006, 26, 1452–1462. [Google Scholar] [CrossRef] [Green Version]
- Ou, J.; Ou, Z.; Ackerman, A.W.; Oldham, K.T.; Pritchard, K.A. Inhibition of heat shock protein 90 (hsp90) in proliferating endothelial cells uncouples endothelial nitric oxide synthase activity. Free Radic. Biol. Med. 2003, 34, 269–276. [Google Scholar] [CrossRef]
- Cawthon, R.M. Telomere measurement by quantitative PCR. Nucleic Acids Res. 2002, 30, e47. [Google Scholar] [CrossRef] [Green Version]
- Abecasis, G.R.; Altshuler, D.; Auton, A.; Brooks, L.D.; Durbin, R.M.; Gibbs, R.A.; Hurles, M.E.; McVean, G.A. A map of human genome variation from population-scale sequencing. Nature 2010, 467, 1061–1073. [Google Scholar] [CrossRef] [Green Version]
- Venkatachalam, G.; Surana, U.; Clément, M.V. Replication stress-induced endogenous DNA damage drives cellular senescence induced by a sub-lethal oxidative stress. Nucleic Acids Res. 2017, 45, 10564–10582. [Google Scholar] [CrossRef] [Green Version]
- Tan, J.; Duan, M.; Yadav, T.; Phoon, L.; Wang, X.; Zhang, J.M.; Zou, L.; Lan, L. An R-loop-initiated CSB-RAD52-POLD3 pathway suppresses ROS-induced telomeric DNA breaks. Nucleic Acids Res. 2020, 48, 1285–1300. [Google Scholar] [CrossRef] [PubMed]
- Clouston, S.A.P.; Edelman, N.H.; Aviv, A.; Stewart, C.; Luft, B.J. Shortened leukocyte telomere length is associated with reduced pulmonary function and greater subsequent decline in function in a sample of World Trade Center responders. Sci. Rep. 2019, 9, 8148. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mishra, A.; Ali, Z.; Vibhuti, A.; Kumar, R.; Alam, P.; Ram, R.; Thinlas, T.; Mohammad, G.; Pasha, M.A. CYBA and GSTP1 variants associate with oxidative stress under hypobaric hypoxia as observed in high-altitude pulmonary oedema. Clin. Sci. 2012, 122, 299–309. [Google Scholar] [CrossRef] [PubMed]
- Pandey, P.; Ali, Z.; Mohammad, G.; Pasha, M.A. Elevated blood plasma levels of epinephrine, norepinephrine, tyrosine hydroxylase, TGFβ1, and TNFα associated with high-altitude pulmonary edema in an Indian population. Ther. Clin. Risk. Manag. 2016, 12, 1207–1221. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beall, C.M. Two routes to functional adaptation: Tibetan and Andean high-altitude natives. Proc. Natl. Acad. Sci. USA 2007, 104 (Suppl. 1), 8655–8660. [Google Scholar] [CrossRef] [Green Version]
- Sack, M.N.; Fyhrquist, F.Y.; Saijonmaa, O.J.; Fuster, V.; Kovacic, J.C. Basic Biology of Oxidative Stress and the Cardiovascular System: Part 1 of a 3-Part Series. J. Am. Coll. Cardiol. 2017, 70, 196–211. [Google Scholar] [CrossRef]
- von Zglinicki, T. Oxidative stress shortens telomeres. Trends Biochem. Sci. 2002, 27, 339–344. [Google Scholar] [CrossRef]
- Schiffrin, E.L. Oxidative Stress, Nitric Oxide Synthase, and Superoxide Dismutase. Hypertension 2008, 51, 31–32. [Google Scholar] [CrossRef] [Green Version]
- Ahsan, A.; Charu, R.; Pasha, M.A.; Norboo, T.; Charu, R.; Afrin, F.; Ahsan, A.; Baig, M.A. eNOS allelic variants at the same locus associate with HAPE and adaptation. Thorax 2004, 59, 1000–1002. [Google Scholar] [CrossRef] [Green Version]
- Beall, C.M.; Laskowski, D.; Erzurum, S.C. Nitric oxide in adaptation to altitude. Free Radic. Biol. Med. 2012, 52, 1123–1134. [Google Scholar] [CrossRef] [Green Version]
- Wang, Q.Q.; Yu, L.; Huang, G.R.; Zhang, L.; Liu, Y.Q.; Wang, T.W.; Lin, H.; Ren, Q.; Liu, P.; Huang, L.; et al. Polymorphisms of angiotensin converting enzyme and nitric oxide synthase 3 genes as risk factors of high-altitude pulmonary edema: A case-control study and meta-analysis. Tohoku J. Exp. Med. 2013, 229, 255–266. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.; Zou, Y.; Gao, Y. Gene polymorphisms and high-altitude pulmonary edema susceptibility: A 2011 update. Respiration 2012, 84, 155–162. [Google Scholar] [CrossRef] [PubMed]
- Teo, H.; Ghosh, S.; Luesch, H.; Ghosh, A.; Wong, E.T.; Malik, N.; Orth, A.; de Jesus, P.; Perry, A.S.; Oliver, J.D.; et al. Telomere-independent Rap1 is an IKK adaptor and regulates NF-kappaB-dependent gene expression. Nat. Cell Biol. 2010, 12, 758–767. [Google Scholar] [CrossRef] [PubMed]
- Martinez, P.; Thanasoula, M.; Carlos, A.R.; Gómez-López, G.; Tejera, A.M.; Schoeftner, S.; Dominguez, O.; Pisano, D.G.; Tarsounas, M.; Blasco, M.A. Mammalian Rap1 controls telomere function and gene expression through binding to telomeric and extratelomeric sites. Nat. Cell Biol. 2010, 12, 768–780. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bachand, F.; Triki, I.; Autexier, C. Human telomerase RNA-protein interactions. Nucleic Acids Res. 2001, 29, 3385–3393. [Google Scholar] [CrossRef] [Green Version]
- Latrick, C.M.; Cech, T.R. POT1-TPP1 enhances telomerase processivity by slowing primer dissociation and aiding translocation. EMBO J. 2010, 29, 924–933. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yuan, X.; Xu, D. Telomerase Reverse Transcriptase (TERT) in Action: Cross-Talking with Epigenetics. Int. J. Mol. Sci. 2019, 20, 3338. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pritchard, K.A., Jr.; Ackerman, A.W.; Gross, E.R.; Stepp, D.W.; Shi, Y.; Fontana, J.T.; Baker, J.E.; Sessa, W.C. Heat shock protein 90 mediates the balance of nitric oxide and superoxide anion from endothelial nitric-oxide synthase. J. Biol. Chem. 2001, 276, 17621–17624. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Keppler, B.R.; Grady, A.T.; Jarstfer, M.B. The Biochemical Role of the Heat Shock Protein 90 Chaperone Complex in Establishing Human Telomerase Activity. J. Biol. Chem. 2006, 281, 19840–19848. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Loayza, D.; De Lange, T. POT1 as a terminal transducer of TRF1 telomere length control. Nature 2003, 423, 1013–1018. [Google Scholar] [CrossRef]
- Deelen, J.; Uh, H.W.; Monajemi, R.; van Heemst, D.; Thijssen, P.E.; Böhringer, S.; van den Akker, E.B.; de Craen, A.J.; Rivadeneira, F.; Uitterlinden, A.G.; et al. Gene set analysis of GWAS data for human longevity highlights the relevance of the insulin/IGF-1 signaling and telomere maintenance pathways. Age 2013, 35, 235–249. [Google Scholar] [CrossRef] [PubMed]
- Collins, M.; Renault, V.; Grobler, L.A.; St Clair Gibson, A.; Lambert, M.I.; Wayne Derman, E.; Butler-Browne, G.S.; Noakes, T.D.; Mouly, V. Athletes with exercise-associated fatigue have abnormally short muscle DNA telomeres. Med. Sci. Sports Exerc. 2003, 35, 1524–1528. [Google Scholar] [CrossRef] [PubMed]
- Rytz, C.L.; Pun, M.; Mawhinney, J.A.; Mounsey, C.A.; Mura, M.; Martin, A.; Pialoux, V.; Hartmann, S.E.; Furian, M.; Rawling, J.M.; et al. Differential effects of high-altitude exposure on markers of oxidative stress, antioxidant capacity, and iron profiles. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2022, 323, R445–R456. [Google Scholar] [CrossRef]
- Garg, S. Computational methods for chromosome-scale haplotype reconstruction. Genome Biol. 2021, 22, 101. [Google Scholar] [CrossRef] [PubMed]
- Nurk, S.; Koren, S.; Rhie, A.; Rautiainen, M.; Bzikadze, A.V.; Mikheenko, A.; Vollger, M.R.; Altemose, N.; Uralsky, L.; Gershman, A.; et al. The complete sequence of a human genome. Science 2022, 376, 44–53. [Google Scholar] [CrossRef]
- Jarvis, E.D.; Formenti, G.; Rhie, A.; Guarracino, A.; Yang, C.; Wood, J.; Tracey, A.; Thibaud-Nissen, F.; Vollger, M.R.; Porubsky, D.; et al. Automated assembly of high-quality diploid human reference genomes. bioRxiv 2022. [Google Scholar] [CrossRef] [PubMed]

| A. HAPE Risk/Protection | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Gene/ SNP | Genotype/ Allele | HAPE-f (n = 210) | HAPE-p (n = 183) | HLs (n = 200) | HAPE-p vs. HAPE-f | HLs vs. HAPE-f | HAPE-p vs. HLs | |||
| p Value | OR (95%CI) | p Value | OR (95%CI) | p Value | OR (95%CI) | |||||
| TERF2IP rs59297469 | GG | 73 (34.8%) | 40 (21.9%) | 138 (69.0%) | Ref | - | Ref | - | Ref | - |
| GT | 105 (50.0%) | 92 (50.2%) | 57 (28.5%) | 0.575 | 1.17 (0.68–2.01) | 1.23 × 10−4 | 0.22 (0.10–0.47) | 1.00 × 10−10 | 8.48 (4.43–16.20) | |
| TT | 32 (15.2%) | 51 (27.9%) | 5 (2.5%) | 0.037 | 1.98 (1.04–3.75) | 8.62 × 10−6 | 0.02 (0.004–0.12) | 1.03 × 10−8 | 69.64 (16.29–297.67) | |
| G | 251 (59.8%) | 172 (47.0%) | 333 (83.2%) | Ref | - | Ref | - | Ref | - | |
| T | 169 (40.2%) | 194 (53.0%) | 67 (16.8%) | 0.082 | 1.33 (0.96–1.83) | 2.25 × 10−9 | 0.16 (0.09–0.29) | 3.19 × 10−18 | 6.81 (4.42–10.49) | |
| TERC rs2293607 | AA | 132 (62.8%) | 97 (53.0%) | 32 (16.0%) | Ref | - | Ref | - | Ref | - |
| AG | 65 (31.0%) | 66 (36.1%) | 101 (50.5%) | 0.099 | 1.51 (0.93–2.47) | 1.30 × 10−6 | 9.04 (3.71–22.04) | 3.16 × 10−7 | 0.18 (0.10–0.35) | |
| GG | 13 (6.2%) | 20 (10.9%) | 67 (33.5%) | 0.015 | 2.81 (1.22–6.46) | 8.04 × 10−9 | 26.92 (8.79–82.43) | 2.42 × 10−9 | 0.09 (0.04–0.19) | |
| A | 329 (78.3%) | 260 (71.0%) | 165 (41.2%) | Ref | - | Ref | - | Ref | - | |
| G | 91 (21.7%) | 106 (29.0%) | 235 (58.8%) | 0.005 | 1.70 (1.18–2.45) | 5.12 × 10−12 | 6.04 (3.62–10.06) | 6.81 × 10−12 | 0.27 (0.18–0.39) | |
| TEP1 rs2228036 | GG | 134 (63.8%) | 95 (52.0%) | 127 (63.5%) | Ref | - | Ref | - | Ref | - |
| GT | 69 (32.9%) | 72 (39.3%) | 57 (28.5%) | 0.581 | 1.15 (0.70–1.87) | 0.267 | 0.67 (0.32–1.37) | 0.040 | 1.77 (1.03–3.04) | |
| TT | 7 (3.3%) | 16 (8.7%) | 16 (8.0%) | 0.042 | 2.84 (1.04–7.77) | 0.806 | 1.21 (0.26–5.65) | 0.406 | 1.48 (0.59–3.73) | |
| G | 337 (80.2%) | 262 (71.6%) | 311 (77.8%) | Ref | - | Ref | - | Ref | - | |
| T | 83 (19.8%) | 104 (28.4%) | 89 (22.2%) | 0.087 | 1.38 (0.95–2.01) | 0.447 | 0.80 (0.45–1.42) | 0.053 | 1.50 (0.99–2.27) | |
| ACD rs72556537 | TT | 148 (70.4%) | 151 (82.5%) | 145 (72.5%) | Ref | - | Ref | - | Ref | - |
| TA | 52 (24.8%) | 31 (17.0%) | 48 (24.0%) | 0.073 | 0.59 (0.34–1.05) | 0.138 | 0.53 (0.23–1.23) | 0.183 | 0.65 (0.34–1.23) | |
| AA | 10 (4.8%) | 1 (0.5%) | 7 (3.5%) | 0.038 | 0.11 (0.01–0.88) | 0.346 | 1.89 (0.50–7.08) | 0.008 | 0.05 (0.01–0.45) | |
| T | 348 (82.9%) | 333 (91.0%) | 338 (84.5%) | Ref | - | Ref | - | Ref | - | |
| A | 72 (17.1%) | 33 (9.0%) | 62 (15.5%) | 0.003 | 0.48 (0.29–0.78) | 0.778 | 0.91 (0.49–1.71) | 0.002 | 0.43 (0.25–0.74) | |
| B. HA Adaptation | ||||||||||
| Gene/ SNP | Genotype/ Allele | HAPE-f (n = 210) | HAPE-p (n = 183) | HLs (n = 200) | HAPE-p vs. HAPE-f | HLs vs. HAPE-f | HAPE-p vs. HLs | |||
| p Value | OR (95%CI) | p Value | OR (95%CI) | p Value | OR (95%CI) | |||||
| TERF1 rs2975843 | TT | 76 (36.2%) | 61 (33.3%) | 112 (56.0%) | Ref | - | Ref | - | Ref | - |
| TC | 97 (46.2%) | 82 (44.8%) | 81 (40.5%) | 0.729 | 0.91 (0.55–1.52) | 0.046 | 0.46 (0.22–0.99) | 0.033 | 1.82 (1.05–3.16) | |
| CC | 37 (17.6%) | 40 (21.9%) | 7 (3.5%) | 0.806 | 1.08 (0.57–2.06) | 0.007 | 0.15 (0.04–0.60) | 2.98 × 10−5 | 9.07 (3.22–25.55) | |
| T | 249 (59.3%) | 204 (55.7%) | 305 (76.2%) | Ref | - | Ref | - | Ref | - | |
| C | 171 (40.7%) | 162 (44.3%) | 95 (23.8%) | 0.882 | 1.02 (0.74–1.42) | 0.003 | 0.46 (0.28–0.76) | 1.35 × 10−5 | 2.35 (1.60–3.45) | |
| TERF1 rs10099824 | GG | 92 (43.8%) | 81 (44.3%) | 168 (84.0%) | Ref | - | Ref | - | Ref | - |
| GA | 87 (41.4%) | 78 (42.6%) | 28 (14.0%) | 0.762 | 0.93 (0.56–1.53) | 5.52 × 10−6 | 0.06 (0.02–0.20) | 4.46 × 10−9 | 7.71 (3.90–15.27) | |
| AA | 31 (14.8%) | 24 (13.1%) | 4 (2.0%) | 0.256 | 0.67 (0.33–1.34) | 4.49 × 10−5 | 0.02 (0.003–0.13) | 1.77 × 10−4 | 11.08 (3.15–38.96) | |
| G | 271 (64.5%) | 240 (65.6%) | 364 (91.0%) | Ref | - | Ref | - | Ref | - | |
| A | 149 (35.5%) | 126 (34.4%) | 36 (9.0%) | 0.285 | 0.83 (0.59–1.17) | 7.28 × 10−11 | 0.07 (0.03–0.15) | 7.05 × 10−12 | 5.96 (3.58–9.92) | |
| TERF2 rs3785073 | GG | 99 (47.2%) | 76 (41.5%) | 108 (54.0%) | Ref | - | Ref | - | Ref | - |
| GA | 83 (39.5%) | 86 (47.0%) | 80 (40.0%) | 0.864 | 1.04 (0.64–1.71) | 0.033 | 0.44 (0.20–0.94) | 0.001 | 2.64 (1.51–4.62) | |
| AA | 28 (13.3%) | 21 (11.5%) | 12 (6.0%) | 0.899 | 0.96 (0.47–1.94) | 0.018 | 0.16 (0.03–0.73) | 0.070 | 2.70 (0.92–7.90) | |
| G | 281 (66.9%) | 238 (65.0%) | 296 (74.0%) | Ref | - | Ref | - | Ref | - | |
| A | 139 (33.1%) | 128 (35.0%) | 104 (26.0%) | 0.994 | 1.00 (0.71–1.40) | 0.002 | 0.43 (0.24–0.74) | 0.001 | 1.94 (1.30–2.90) | |
| TERF2 rs153058 | TT | 129 (61.4%) | 107 (58.5%) | 92 (46.0%) | Ref | - | Ref | - | Ref | - |
| TC | 73 (34.8%) | 65 (35.5%) | 84 (42.0%) | 0.680 | 1.11 (0.69–1.79) | 0.178 | 1.61 (0.80–3.24) | 0.047 | 0.58 (0.33–0.99) | |
| CC | 8 (3.8%) | 11 (6.0%) | 24 (12.0%) | 0.164 | 2.07 (0.74–5.79) | 0.015 | 4.59 (1.35–15.63) | 0.015 | 0.31 (0.12–0.80) | |
| T | 331 (78.8%) | 279 (76.2%) | 268 (67.0%) | Ref | - | Ref | - | Ref | - | |
| C | 89 (21.2%) | 87 (23.8%) | 132 (33.0%) | 0.253 | 1.25 (0.85–1.82) | 0.014 | 1.89 (1.14–3.15) | 0.004 | 0.55 (0.37–0.83) | |
| POT1 rs7794637 | CC | 90 (42.9%) | 63 (34.4%) | 105 (52.5%) | Ref | - | Ref | - | Ref | - |
| CT | 95 (45.2%) | 88 (48.1%) | 82 (41.0%) | 0.682 | 0.90 (0.54–1.49) | 0.137 | 0.59 (0.29–1.19) | 0.003 | 2.30 (1.32–4.00) | |
| TT | 25 (11.9%) | 32 (17.5%) | 13 (6.5%) | 0.297 | 1.43 (0.73–2.81) | 0.010 | 0.18 (0.05–0.66) | 0.003 | 4.22 (1.62–11.02) | |
| C | 275 (65.5%) | 214 (58.5%) | 292 (73.0%) | Ref | - | Ref | - | Ref | - | |
| T | 145 (34.5%) | 152 (41.5%) | 108 (27.0%) | 0.480 | 1.13 (0.81–1.57) | 0.010 | 0.51 (0.31–0.85) | 2.62 × 10−4 | 2.04 (1.39–2.99) | |
| ACD rs6979 | GG | 67 (31.9%) | 54 (29.5%) | 18 (9.0%) | Ref | - | Ref | - | Ref | - |
| GA | 110 (52.4%) | 101 (55.2%) | 74 (37.0%) | 0.828 | 1.06 (0.64–1.74) | 0.011 | 4.54 (1.41–14.62) | 0.334 | 0.68 (0.31–1.49) | |
| AA | 33 (15.7%) | 28 (15.3%) | 108 (54.0%) | 0.982 | 1.01 (0.50–2.02) | 3.05 × 10−7 | 41.88 (10.03–174.91) | 3.74 × 10−8 | 0.07 (0.03–0.18) | |
| G | 244 (58.1%) | 209 (57.1%) | 110 (27.5%) | Ref | - | Ref | - | Ref | - | |
| A | 176 (41.9%) | 157 (42.9%) | 290 (72.5%) | 0.844 | 1.03 (0.75–1.43) | 3.34 × 10−12 | 7.34 (4.19–12.86) | 5.62 × 10−12 | 0.26 (0.18–0.38) | |
| TEP1 rs2184282 | AA | 138 (65.7%) | 134 (73.2%) | 97 (48.5%) | Ref | - | Ref | - | Ref | - |
| AG | 58 (27.6%) | 41 (22.4%) | 69 (34.5%) | 0.108 | 0.64 (0.37–1.10) | 0.015 | 2.43 (1.18–4.97) | 0.002 | 0.39 (0.22–0.70) | |
| GG | 14 (6.7%) | 8 (4.4%) | 34 (17.0%) | 0.550 | 0.74 (0.27–2.01) | 0.004 | 4.60 (1.63–12.95) | 2.38 × 10−5 | 0.11 (0.04–0.31) | |
| A | 334 (79.5%) | 309 (84.4%) | 263 (65.8%) | Ref | - | Ref | - | Ref | - | |
| G | 86 (20.5%) | 57 (15.6%) | 137 (34.2%) | 0.128 | 0.72 (0.47–1.10) | 2.95 × 10−4 | 2.52 (1.53–4.16) | 7.81 × 10−8 | 0.31 (0.20–0.47) | |
| TEP1 rs4246977 | TT | 57 (27.1%) | 45 (24.6%) | 127 (63.5%) | Ref | - | Ref | - | Ref | - |
| TC | 105 (50.0%) | 99 (54.1%) | 60 (30.0%) | 0.493 | 1.21 (0.70–2.11) | 2.88 × 10−4 | 0.22 (0.10–0.50) | 8.99 × 10−7 | 4.20 (2.37–7.44) | |
| CC | 48 (22.9%) | 39 (21.3%) | 13 (6.5%) | 0.525 | 0.81 (0.42–1.56) | 4.58 × 10−5 | 0.09 (0.03–0.28) | 8.50 × 10−6 | 6.76 (2.92–15.69) | |
| T | 219 (52.1%) | 189 (51.6%) | 314 (78.5%) | Ref | - | Ref | - | Ref | - | |
| C | 201 (47.9%) | 177 (48.4%) | 86 (21.5%) | 0.538 | 0.90 (0.66–1.25) | 2.68 × 10−7 | 0.25 (0.15–0.42) | 1.35 × 10−8 | 3.08 (2.09–4.54) | |
| Gene | Haplotype | HAPE-p vs. HAPE-f | HLs vs. HAPE-f | HAPE-p vs. HLs | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| χ2 | OR (95% CI) | p-Value | χ2 | OR (95% CI) | p-Value | χ2 | OR (95% CI) | p-Value | ||
| TERF1 | AAGG | 0.927 | 1.17 (0.85–1.63) | 0.336 | 18.700 | 2.84 (1.77–4.56) | 1.53 × 10−5 | 22.768 | 0.41 (0.29–0.59) | 1.83 × 10−6 |
| GAAA * | 5.190 | 0.58 (0.37–0.93) | 0.023 | 10.145 | 0.22 (0.09–0.56) | 0.001 | 4.581 | 2.18 (1.07–4.44) | 0.032 | |
| TPP1 + TERF2 + RAP1 | AAAAATTCGCA | 6.133 | 4.71 (1.38–16.06) | 0.013 | 7.470 | 0.05 (0.01–0.43) | 0.006 | |||
| AAGTATCCGCA | 1.369 | 0.66 (0.33–1.33) | 0.242 | 6.893 | 2.42 (1.25–4.67) | 0.009 | 12.218 | 0.37 (0.22–0.65) | 4.73 × 10−4 | |
| AAGTATTCGCA | 11.734 | 5.49 (2.07–14.55) | 0.001 | 12.983 | 0.14 (0.05–0.41) | 3.14 × 10−4 | ||||
| AAGTGTCCGCA | 2.512 | 1.77 (0.87–3.57) | 0.113 | 16.480 | 4.44 (2.16–9.12) | 4.92 × 10−5 | 19.034 | 0.29 (0.17–0.51) | 1.28 × 10−5 | |
| TGGTATCCGCA | 0.701 | 0.63 (0.21–1.87) | 0.402 | 5.101 | 3.60 (1.18–10.93) | 0.024 | ||||
| TERT | GATCGCCAG | 1.308 | 0.72 (0.41–1.26) | 0.253 | 22.883 | 3.75 (2.18–6.44) | 1.72 × 10−6 | 38.560 | 0.23 (0.15–0.37) | 5.31 × 10−10 |
| TEP1 + HSP90AA1 | TCCCGACTTA * | 6.003 | 0.23 (0.07–0.74) | 0.014 | 0.043 | 0.89 (0.30–2.63) | 0.835 | |||
| TCCCGGCTCA | 1.063 | 1.74 (0.61–4.97) | 0.303 | 4.621 | 4.40 (1.14–16.98) | 0.032 | 1.215 | 0.58 (0.22–1.52) | 0.270 | |
| TTCCGATCT * | 8.066 | 0.18 (0.05–0.58) | 0.005 | |||||||
| TTCCGGCTCA | 4.482 | 5.58 (1.14–27.44) | 0.034 | 7.513 | 0.25 (0.09–0.68) | 0.006 | ||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pasha, Q.; Rain, M.; Tasnim, S.; Kanipakam, H.; Thinlas, T.; Mohammad, G. The Telomere-Telomerase System Is Detrimental to Health at High-Altitude. Int. J. Environ. Res. Public Health 2023, 20, 1935. https://doi.org/10.3390/ijerph20031935
Pasha Q, Rain M, Tasnim S, Kanipakam H, Thinlas T, Mohammad G. The Telomere-Telomerase System Is Detrimental to Health at High-Altitude. International Journal of Environmental Research and Public Health. 2023; 20(3):1935. https://doi.org/10.3390/ijerph20031935
Chicago/Turabian StylePasha, Qadar, Manjari Rain, Sana Tasnim, Hema Kanipakam, Tashi Thinlas, and Ghulam Mohammad. 2023. "The Telomere-Telomerase System Is Detrimental to Health at High-Altitude" International Journal of Environmental Research and Public Health 20, no. 3: 1935. https://doi.org/10.3390/ijerph20031935





