Characterization of Antagonist Potential of Selected Compost Bacterial Isolates (CBI) against Plant and Human Pathogens
Abstract
:1. Introduction
2. Materials and Methods
3. Results
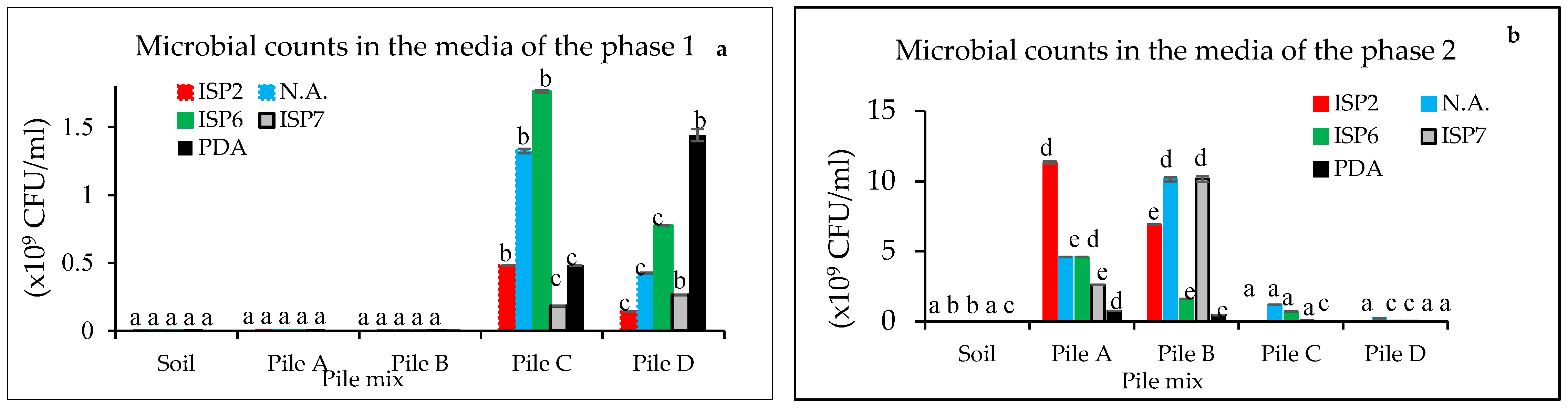
3.1. Microbial Counts at Different Composting Times
3.2. pH and Biomass Variations
3.3. Antagonist Activity—Fungal Pathogens
3.4. Antagonist Activity—Pathogenic Bacteria
3.5. Physiological and Biochemical Testing
3.6. Raw Data Statistics of 16S Sequencing
3.7. Taxonomic Classification and Abundance Ratio (%)
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wilson, D.C.; Rodic, L.; Scheinberg, A.; Velis, C.A.; Alabaster, G. Comparative analysis of solid waste management in 20 cities. Waste. Manag. Res. 2012, 30, 237–254. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- American University of Beirut. Guide to Municipal Waste Management. p. 15. Available online: https://aub.edu.lb/units/natureconservation/gallery/Documents/guide_to_municipal_solid_waste_management.pdf (accessed on 6 June 2017).
- Awasthi, M.K.; Zhang, Z.; Wang, Q.; Shen, F.; Li, R.; Li, D.; Ren, X.; Wang, M.; Chen, H.; Zhao, J. New insight with the effects of biochar amendment on bacterial diversity as indicators of biomarkers supports the thermophilic phase during sewage sludge composting. Bioresour. Technol 2017, 238, 589–601. [Google Scholar] [CrossRef] [PubMed]
- El Haggar, S.M. (Ed.) Sustainability of Agricultural and Rural Waste Management. In Sustainable Industrial Design and Waste Management; Academic Press: Cambridge, MA, USA, 2007; pp. 223–260. [Google Scholar]
- Baca, M.T.; Fornasier, F.; de Nobili, M. Mineralization and humification pathways in two composting processes applied to cotton wastes. J. Ferment. Bioeng. 1992, 74, 179–184. [Google Scholar] [CrossRef]
- Ahn, H.K.; Richard, T.L.; Glanville, T.D. Optimum moisture levels for biodegradation of mortality composting envelope materials. Waste Manag. 2008, 28, 1411–1416. [Google Scholar] [CrossRef] [PubMed]
- Sidhu, J.; Gibbs, R.A.; Ho, G.E.; Unkovich, I. The role of indigenous microorganisms in suppression of salmonella regrowth in composted biosolids. Water Res. 2008, 35, 913–920. [Google Scholar] [CrossRef] [PubMed]
- De Gannes, V.; Eudoxie, G.; Hickey, W.J. Prokaryotic successions and diversity in composts as revealed by 454-pyrosequencing. Bioresour. Technol. 2013, 133, 573–580. [Google Scholar] [CrossRef] [PubMed]
- Neher, D.A.; Weicht, T.R.; Bates, S.T.; Leff, J.W.; Fierer, N. Changes in bacterial and fungal communities across compost recipes, preparation methods, and composting times. PLoS ONE 2013, 8, e79512. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Storey, S.; Ni Chualain, D.; Doyle, O.; Clipson, N.; Doyle, E. Comparison of bacterial succession in green waste composts amended with inorganic fertilizer and wastewater treatment plant sludge. Bioresour. Technol. 2015, 179, 71–77. [Google Scholar] [CrossRef] [PubMed]
- Valdrighi, M.M.; Pera, A.; Scatena, S.; Agnolucci, M.; Vallini, G. Effects of humic acids extracted from mined lignite or composted vegetable residues on plant growth and soil microbial populations. Compost Sci. Util. 1995, 3, 30–38. [Google Scholar] [CrossRef]
- Zhang, L.L.; Zhang, H.Q.; Wang, Z.H.; Chen, G.J.; Wang, L.S. Dynamic changes of the dominant functioning microbial community in the compost of a 90-m(3) aerobic solid state fermentor revealed by integrated meta-omics. Bioresour. Technol. 2016, 203, 1–10. [Google Scholar] [CrossRef]
- Nicholson, F.A.; Groves, S.J.; Chambers, B.J. Pathogen survival during livestock manure storage and following land application. Bioresour. Technol. 2005, 96, 135–143. [Google Scholar] [CrossRef] [PubMed]
- You, Y.; Rankin, S.C.; Aceto, H.W.; Benson, C.E.; Toth, J.D.; Dou, Z. Survival of Salmonella enterica serovar Newport in manure and manure amended soils. Appl. Environ. Microbiol. 2006, 72, 5777–5783. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, X.; Li, Q.; Sui, J.; Zhang, J.; Liu, Z.; Du, J.; Xu, R.; Zhou, Y.; Liu, X. Isolation and characterization of antagonistic bacteria Paenibacillus jamilae HS-26 and their effects on plant growth. BioMed Res. Int. 2019, 2019, 3638926. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Suárez-Estrella, F.; Vargas-Garcua, M.C.; López, M.J.; Moreno, J. Survival of Fusarium oxysporum f.sp. melonis on plant waste. Crop Prot. 2004, 23, 127–133. [Google Scholar] [CrossRef]
- Noble, R.; Roberts, S.J. Eradication of plant pathogens and nematodes during composting: A review. Plant Pathol. 2004, 53, 548–568. [Google Scholar] [CrossRef]
- Lemunier, M.; Francou, C.; Rousseaux, S.; Houot, S.; Dantigny, P.; Piveteau, P.; Guzzo, J. Long-Term Survival of Pathogenic and Sanitation Indicator Bacteria in Experimental Biowaste Composts. App. Environ. Microbiol. 2005, 71, 5779–5786. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rynk, R. Monitoring Moisture in Composting Systems. Biocycle 2008, 41, 53–57. [Google Scholar]
- Day, M.; Shaw, K. Biological, chemical and physical processes of composting. In Compost Utilization in Horticultural Cropping Systems; Stoffella, P.J., Kahn, B.A., Eds.; Lewis Publishers: Boca Raton, FL, USA, 2001; pp. 17–50. [Google Scholar]
- Shirling, E.B.; Gottlieb, D. Methods for characterization of Streptomyces species. Int. J. Syst. Bacteriol. 1966, 16, 313–340. [Google Scholar] [CrossRef] [Green Version]
- Badji, B.; Mostefaoui, A.; Sabaou, N.; Mathieu, F.; Lebrihi, A. Identification of a new strain of Actinomadura isolated from Saharan soil and partial characterization of its antifungal compounds. Afr. J. Biotechnol. 2011, 10, 13878–13886. [Google Scholar]
- Joffin, J.N.; Leyral, G. Microbiologie technique, Tome 1- Dictionnaire des Techniques, 4th ed.; Reseau Canope: Chasseneuil-du-Poitou, France, 2006. [Google Scholar]
- Frolund, B.; Griebe, T.; Nielsen, P. Enzymatic activity in the activated sludge floc matrix. Appl. Microbiol. Biotechnol. 1995, 43, 755–761. [Google Scholar] [CrossRef]
- Rizk, Z.; El Rayess, Y.; Ghanem, C.; Mathieu, F.; Taillandier, P.; Nehmé, N. Identification of multiple-derived peptides produces by Saccharomyces cerevisiae Uvaferm BDX inhibiting malolactic fermentation performed by Oenococcus oeni Vitalactic F. FEMS Yeast Res. 2018, 18, 1–10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nehme, N.; Mathieu, F.; Taillandier, P. Impact of the co-culture of Saccharomyces cerevisiae-Oenococcus oeni on malolactic fermentation and partial characterization of yeast derived inhibitory peptidic fraction. Food Microbiol. 2010, 27, 150–157. [Google Scholar] [CrossRef] [PubMed]
- Horvath, G.; Bencsik, T.; Acs, K.; Kocsis, B. Sensitivity of ESBL-Producing Gram-Negative Bacteria to Essential Oils, Plant Extracts, and their isolated compounds. In Antibiotic Resistance; Academic Press: Amsterdam, The Netherlands, 2016; pp. 239–269. [Google Scholar]
- Christenson, J.C.; Korgenski, E.K.; Relich, R.F. Laboratory diagnosis of infection due to bacteria, fungi, parasites and rickettsiae. In Principles and Practice of Pediatric Infectious Diseases, 5th ed.; Elsevier Publishers: Amsterdam, The Netherlands, 2018; pp. 1422–1434. [Google Scholar]
- Wei, Y.; Zhao, Y.; Zhou, D.; Qi, D.; Li, K.; Tang, W.; Chen, Y.; Jing, T.; Zang, X.; Xie, J.; et al. A newly isolated Streptomyces sp. YYS-7 with a broad-spectrum antifungal activity improves the banana plant resistance to Fusarium oxysporum f. sp. cubense tropical race 4. Front. Microbiol. 2020, 11, 1712. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Wu, Q.; Piceno, Y.M.; Desantis, T.Z.; Saunders, F.M.; Andersen, G.L.; Liu, W.-T. Diversity of bacterioplankton in contrasting Tibetan lakes revealed by high-density microarray and clone library analysis. FEMS Microbiol. Ecol. 2013, 86, 277–287. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, Y.; Lai, Q.; Shao, Z. Genome analysis-based reclassification of Bacillus weihenstephanensis as a later heterotypic synonym of Bacillus mycoides. Int. J. Syst. Evol. Microbiol. 2018, 68, 106–112. [Google Scholar] [CrossRef] [PubMed]
- Azhar, E.I.; Papadioti, A.; Bibi, F.; Ashshi, A.M.; Raoult, D.; Angelakis, E. ‘Pseudomonas saudimassiliensis’ sp. nov. a new bacterial species isolated from air samples in the urban environment of Makkah, Saudi Arabia. New Microbes New Infect. 2017, 16, 43–44. [Google Scholar] [CrossRef] [PubMed]
- Stanier, R.Y.; Palleroni, N.J.; Doudoroff, M. The Aerobic Pseudomonads a Taxonomic Study. J. GenMicrobiol. 1966, 43, 159–271. [Google Scholar] [CrossRef] [Green Version]
- Salloto, G.R.B.; Cardoso, A.M.; Coutinho, F.H.; Pinot, L.H.; Vieira, R.P.; Chaia, C.; Lima, J.; Albano, R.; Martins, O.; Clementino, M. Pollution impacts on Bacterioplankton diversity in a tropical urban coastal lagoon system. PLoS ONE 2012, 7, e51175. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Crovadore, J.; Soljan, V.; Lefort, F.; Chablais, R. Selected nitrifying micro-organisms from aerobic granular biomass from sewage sludge. Plants and Pathogens Group 2014. NCBI, Project KM251257, 2014. Submitted 5 November 2014.
- Oh, J.; Conlan, S.; Polley, E.C.; Segre, J.A.; Kong, H.H. Shifts in human skin and nares microbiota of healthy children and adults. Genome Med. 2012, 4, 77. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Paek, J.; Shin, J.H.; Shin, Y.; Park, I.-S.; Jin, T.-E.; Kook, J.-K.; Wie, S.-H.; Cho, H.; Park, S.-J.; Chang, Y.-H. Myroïdes injenensis sp. nov., a new member isolated from human urine. Anton. Leeuw. Int. J. G 2015, 107, 201–207. [Google Scholar] [CrossRef]
- Vasudevan, N.; Kanimozhi, R. Combined biological and chemical treatment of anaerobically digested distillery wastewater. J. Adv. Oxxid. Technol. 2010, 13, 221–227. [Google Scholar] [CrossRef]
- Durso, L.M.; Harhay, G.P.; Smith, T.P.L.; Bono, J.L.; DeSantis, T.Z.; Andersen, G.L.; Keen, J.E.; Laegreid, W.W.; Clawson, M.L. Synecology of the primary and secondary feedlot habitats of Escherichia coli O157:H7. USDA, ARS, Meat Animal Research Center, State Spur 18D 2009. NCBI, Project PRJNA 34557. Submitted 3 March 2010.
- Scott, J.J.; Lewin, G.R.; Suen, G.; Tringe, S.G.; Barry, K.W.; Touchon, J.C.; Currie, C.R. Refuse dump 16SrRNA: Leaf-cutter ant refuse dumps are nutrient reservoirs harboring diverse microbial assemblages. NCBI, Project PRJEB7274, 2014 Submitted 2 October 2014.
- Kobayashi, T.; Li, Y.-Y. Bioreactor for desulfurization. National Institute for environmental studies; Onogawa 16–2, Tsukuba, Ibaraki 305-8506 Japan. NCBI, Project AB673200.1, 2011. Submitted 4 October 2011.
- Gautam, S.P.; Bundela, P.S.; Pandey, A.K.; Awasthi, M.K.; Sarsaiya, S. Composting of municipal solid waste of Jabalpur city. Glob. J. Environ. Res 2010, 42, 43–46. [Google Scholar]
- Stofella, P.J.; Kahn, B.A. Compost Utilization in Horticultural Cropping Systems; Lewis Publishers: Boca Raton, FL, USA, 2001; Chapter 15; p. 430. [Google Scholar]
- Roletto, E.; Barberis, R.; Consiglio, M.; Jodice, R. Chemical parameters for evaluating compost maturity. BioCycle 1985, 26, 46–47. [Google Scholar]
- Amir, S. Contribution à la Valorisation de boues de Stations D’épuration par Compostage: Devenir des Micropolluants Metalliques et Organiques et Bilan Humique du Compost. Thèse de Doctorat, Institut National Polytechnique de Toulouse, Toulouse, France, 2005. [Google Scholar]
- Gobat, J.M. Le Sol Vivant. Bases de la Pédologie. Biologie des Sols; Collection Gérer l’Environnement N° 14; Presses Polytechniques et Universitaires Romandes: Lausanne, Switzerland, 1998; 519p. [Google Scholar]
- Li, J.; Chen, Y.-T.; Xia, Z.-Y.; Gou, M.; Sun, Z.-Y.; Tang, Y.-Q. Changes in Bacterial Communities during a Pilot-Scale Composting Process of Dairy Manure. J. Environ. Eng. 2020, 146, 1–13. [Google Scholar] [CrossRef]
- Alvarenga, P.; Mourinha, C.; Farto, M.; Santos, T.; Palma, P.; Sengo, J.; Morais, M.-C.; Cunha-Queda, C. Sewage sludge, compost and other representative organic wastes as agricultural soil amendments: Benefits versus limiting factors. Waste Manag. 2015, 40, 44–52. [Google Scholar] [CrossRef] [PubMed]
- Piceno, Y.M.; Pecora-Black, G.; Kramer, S.; Roy, M.; Reid, F.C.; Dubinsky, E.A.; Andersen, G.L. Bacterial community structure transformed after thermophilically composting human waste in Haiti. PLoS ONE 2017, 12, e0177626. [Google Scholar] [CrossRef] [Green Version]
- Duan, M.; Zhang, Y.; Zhou, B.; Qin, Z.; Wu, J.; Wang, Q.J.; Yin, Y. Effects of Bacillus subtilis on carbon components and microbial functional metabolism during cow manure–straw composting. Biores. Technol. 2020, 303, 122868. [Google Scholar] [CrossRef] [PubMed]
- Pan, I.; Dam, B.; Sen, S.K. Composting of common organic wastes using microbial inoculants. Biotech 2012, 2, 127–134. [Google Scholar] [CrossRef] [Green Version]
- Zhao, Y.; Selvaraj, J.; Xing, F.; Zhou, L.; Wang, Y.; Song, H.; Tan, X.; Sun, L.; Sangare, L.; Folly, Y.; et al. Antagonistic action of Bacillus subtilis strain SG6 on Fusarium graminearum. PLoS ONE 2014, 9, e92486. [Google Scholar] [CrossRef] [PubMed]
- Aperce, C.C.; Burkey, T.E.; Kukanich, B.; Crozier-Dodson, B.A.; Dritz, S.S.; Minton, J.E. Interaction of Bacillus species and Salmonella enterica serovar Typhimurium in immune or inflammatory signaling from swine intestinal epithelial cells. J. Anim. Sci 2010, 88, 1649–1656. [Google Scholar] [CrossRef] [Green Version]
- Pathma, J.; Sakthivel, N. Microbial diversity of vermicompost bacteria that exhibit useful agricultural traits and waste management potential. Springerplus 2012, 1, 1–26. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dzionek, A.; Wojcieszyńska, D.; Guzik, U. Natural carriers in bioremediation: A review. Elect. J. Biotechnol. 2016, 23, 28–36. [Google Scholar] [CrossRef] [Green Version]
- Feng, N.-X.; Feng, Y.-X.; Liang, Q.-F.; Chen, X.; Xiang, L.; Zhao, H.-M.; Liu, B.-L.; Cao, G.; Li, Y.-W.; Li, H.; et al. Complete biodegradation of di-n-butyl phtalate (DBP) by a novel Pseudomonas sp. YJB6. Sci. Total Environ. 2021, 76, 143208. [Google Scholar] [CrossRef] [PubMed]
- Pathma, J.; Rahul, G.R.; Kennedy, R.; Subashri, R.; Sakthivel, N. Secondary metabolite production by bacterial antagonists. J. Biol. Control 2011, 25, 165–181. [Google Scholar]
- Zhao, H.-M.; Du, H.; Feng, N.-X.; Mo, C.h.-M. Biodegradation of di-n-butylphthalate and phthalic acid by a novel Providencia sp. 2D and its stimulation in a compost-amended soil. Biol. Fertil. Soils 2016, 52, 65–76. [Google Scholar] [CrossRef]
- Ayilara, M.S.; Olanrewaju, O.S.; Babalola, O.O.; Odeyemi, O. Waste Management through Composting: Challenges and Potentials. Sustainability 2020, 12, 4456. [Google Scholar] [CrossRef]
- Deng, Y.; Mao, Y.; Li, B.; Yang, C.; Zhang, T. Aerobic degradation of sulfadiazine by Arthrobacter spp.: Kinetics, pathways, and genomic characterization. Environ. Sci. Technol. 2016, 50, 9566–9575. [Google Scholar] [CrossRef] [PubMed]
- Sylvia, D.M.; Hartel, P.G.; Fuhrmann, J.J.; Zuberer, D.A. Principles and Applications of Soil Microbiology, 2nd ed.; Sylvia, D.M., Ed.; Pearson Prentice Hall: Upper Saddle River, NJ, USA, 2005. [Google Scholar]
- Zhao, X.; Wang, L.; Fang, M.F.; Yang, J. Characterisation of an efficient atrazine-degrading bacterium, Arthrobacter sp. ZXY-2: An attempt to lay the foundation for potential bioaugmentation applications. Biotechnol. Biofuels 2018, 11, 113. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lewis, S.E.; Brown, A.V. Comparative leaf decomposition rates including a non-native species in an urban Ozark stream. J. Arkansas Acad. Sci. 2010, 64, 92–96. [Google Scholar]
- Felestrino, E.B.; Assis, R.; Lemes, C.; Cordeiro, I.; Fonseca, N.; Villa, M.; Vieira, I.; Kamino, L.; Carmo, F.; Moreira, L. Alcaligenes faecalis associated with Mimosa calodendron rizhosphere assist plant survival in arsenic rich soils. J. Soil Sci. Plant Nutr. 2017, 17, 1102–1115. [Google Scholar] [CrossRef] [Green Version]
- Li, Z.; Yang, Y.; Xia, Y.; Wu, T.; Zhu, J.; Wang, Z.; Yang, J. The succession pattern of bacterial diversity in compost using pig manure mixed with wood chips analyzed by 16S rRNA gene analysis. BioRxiv 2019. [CrossRef] [Green Version]
- Lorenzin, G.; Piccinelli, G.; Carlassara, L.; Scolari, F.; Caccuri, F.; Caruso, A.; De Francesco, M.A. Myroides odoratimimus urinary tract infection in an immunocompromised patient: An emerging multidrug-resistant micro-organism. Antimicrob. Resist. Infect. Control 2018, 7, 96. [Google Scholar] [CrossRef] [PubMed]




| Phase 1 | ||||||||||
| Soil | Pile A | Pile B | Pile C | Pile D | ISP 2 | NA | ISP 6 | ISP7 | PDA | |
| Soil | 1.000 | |||||||||
| Pile A | −0.250 | 1.000 | ||||||||
| Pile B | −0.250 | −0.250 | 1.000 | |||||||
| Pile C | −0.250 | −0.250 | −0.250 | 1.000 | ||||||
| Pile D | −0.250 | −0.250 | −0.250 | −0.250 | 1.000 | |||||
| ISP 2 | −0.332 | −0.336 | −0.332 | 0.956 | 0.044 | 1.000 | ||||
| NA | −0.339 | −0.341 | −0.339 | 0.948 | 0.072 | 1.000 | 1.000 | |||
| ISP 6 | −0.365 | −0.365 | −0.364 | 0.902 | 0.191 | 0.989 | 0.993 | 1.000 | ||
| ISP7 | −0.396 | −0.398 | −0.397 | 0.413 | 0.779 | 0.661 | 0.681 | 0.765 | 1.000 | |
| PDA | −0.343 | −0.343 | −0.342 | 0.087 | 0.943 | 0.374 | 0.400 | 0.507 | 0.943 | 1.000 |
| Phase 2 | ||||||||||
| Soil | Pile A | Pile B | Pile C | Pile D | ISP 2 | NA | ISP 6 | ISP7 | PDA | |
| Soil | 1.000 | |||||||||
| Pile A | −0.250 | 1.000 | ||||||||
| Pile B | −0.250 | −0.250 | 1.000 | |||||||
| Pile C | −0.250 | −0.250 | −0.250 | 1.000 | ||||||
| Pile D | −0.250 | −0.250 | −0.250 | −0.250 | 1.000 | |||||
| ISP 2 | −0.390 | 0.821 | 0.347 | −0.390 | −0.389 | 1.000 | ||||
| NA | −0.421 | 0.178 | 0.903 | −0.268 | −0.393 | 0.704 | 1.000 | |||
| ISP 6 | −0.406 | 0.940 | 0.062 | −0.200 | −0.397 | 0.947 | 0.482 | 1.000 | ||
| ISP7 | −0.328 | 0.004 | 0.967 | −0.320 | −0.324 | 0.574 | 0.980 | 0.312 | 1.000 | |
| PDA | −0.393 | 0.828 | 0.335 | −0.390 | −0.381 | 1.000 | 0.695 | 0.951 | 0.564 | 1.000 |
| Phase 3 | ||||||||||
| Soil | Pile A | Pile B | Pile C | Pile D | ISP 2 | NA | ISP 6 | ISP7 | PDA | |
| Soil | 1.000 | |||||||||
| Pile A | −0.250 | 1.000 | ||||||||
| Pile B | −0.250 | −0.250 | 1.000 | |||||||
| Pile C | −0.250 | −0.250 | −0.250 | 1.000 | ||||||
| Pile D | −0.250 | −0.250 | −0.250 | −0.250 | 1.000 | |||||
| ISP 2 | −0.291 | −0.168 | −0.265 | 0.996 | −0.272 | 1.000 | ||||
| NA | −0.427 | −0.310 | 0.877 | 0.240 | −0.380 | 0.229 | 1.000 | |||
| ISP 6 | −0.301 | −0.211 | −0.259 | 0.998 | −0.226 | 0.998 | 0.234 | 1.000 | ||
| ISP7 | −0.475 | −0.225 | 0.796 | 0.362 | −0.458 | 0.360 | 0.985 | 0.359 | 1.000 | |
| PDA | −0.298 | −0.293 | 0.979 | 0.998 | −0.298 | −0.115 | 0.940 | −0.108 | 0.876 | 1.000 |
| Pathogenic Bacteria Inhibition Halo (cm) | ||||||
|---|---|---|---|---|---|---|
| Salmonella Typhimurium | Listeria monocytogenes | Escherichia coli | ||||
| 5–10 K | ≥ 10 K | 5–10 K | ≥ 10 K | 5–10 K | ≥ 10 K | |
| CBI1 | 1 | 0.8 | 0 | 0 | 0.8 | 0.8 |
| CBI2 | 0 | 0 | 0 | 0 | 0 | 1 |
| CBI3 | 0 | 0 | 0 | 0 | 1 | 1 |
| CBI4 | 1 | 1 | 0 | 0 | 0 | 0 |
| CBI5 | 1 | 1 | 0 | 0 | 1 | 1 |
| CBI6 | 3 | 3 | 0 | 0 | 0 | 0 |
| CBI7 | 0 | 0 | 0 | 0 | 0 | 0 |
| CBI8 | 0 | 0 | 1 | 1 | 0 | 0 |
| CBI9 | 1 | 1.5 | 0 | 0 | 1 | 1 |
| CBI10 | 0 | 0 | 0 | 0 | 1 | 1 |
| CBI11 | 0 | 0 | 0 | 0 | 0 | 0 |
| Next Related Type Strain | ||||||
|---|---|---|---|---|---|---|
| Sample Name | Sample Accession Number | Number of Bases | Accession Code 1 | Accession Code 2 | Accession Code 3 | Highest 16S rRNA Sequence Similarity |
| CBI1 | SRX9104367 | 467 | EU257444 (GenBank) | FJ526332 (GenBank) | FJ772081 (GenBank) | Bacillus subtilis (70.83%) |
| CBI2 | SRX9104368 | 466 | EU257444 (GenBank) | FJ526332 (GenBank) | FJ772081 (GenBank) | Bacillus subtilis (66.68%) |
| CBI3 | SRX9104370 | 440 | HM127153 (GenBank) | HM127159 (GenBank) | HM127158 (GenBank) | Providencia sp. (41.13%) |
| CBI4 | SRX9104371 | 440 | AB205631 (GenBank) | JF692663 (GenBank) | KM251257 (GenBank) | Alcaligens sp. (62.40%) |
| CBI5 | SRX9104372 | 440 | EF586016 (GenBank) | LK021121 (GenBank) | CCSF01000001 (GenBank) | Pseudomonas sp. 20_BN (78.41%) |
| CBI6 | SRX9104373 | 451 | FJ786044 (GenBank) | HQ683838 (GenBank) | CP007626 (NCBI) | Bacillus pseudomycoides (97.73%) |
| CBI7 | SRX9104374 | 466 | EU257444 (GenBank) | FJ526332 (GenBank) | FJ772081 (GenBank) | Bacillus subtilis (95.09%) |
| CBI8 | SRX9104375 | 451 | JF218251 (GenBank) | Jf218331 (GenBank) | JF218225 (GenBank) | Arthrobacter sp. (95.66%) |
| CBI9 | SRX9104376 | 440 | HQ671078 (GenBank) | JX966100 (GenBank) | FJ649673 (GenBank) | Myroides sp. (99.51%) |
| CBI10 | SRX9104377 | 440 | FJ672787 (GenBank) | AB673200 (GenBank) | LN565711 (GenBank) | Pseudomonas sp. (54.01%) |
| CBI11 | SRX9104369 | 440 | HM127153 (GenBank) | HM127159 (GenBank) | HM127158 (GenBank) | Providencia sp. (48.98%) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tannouri, A.; Rizk, Z.; Al Daccache, M.; Ghanem, C.; Azzi, V.; Haddad, R.; Maroun, R.G.; Hobaika, Z.; Badra, R.; Salameh, D. Characterization of Antagonist Potential of Selected Compost Bacterial Isolates (CBI) against Plant and Human Pathogens. Agronomy 2022, 12, 2977. https://doi.org/10.3390/agronomy12122977
Tannouri A, Rizk Z, Al Daccache M, Ghanem C, Azzi V, Haddad R, Maroun RG, Hobaika Z, Badra R, Salameh D. Characterization of Antagonist Potential of Selected Compost Bacterial Isolates (CBI) against Plant and Human Pathogens. Agronomy. 2022; 12(12):2977. https://doi.org/10.3390/agronomy12122977
Chicago/Turabian StyleTannouri, Abdo, Ziad Rizk, Marina Al Daccache, Chantal Ghanem, Valérie Azzi, Rami Haddad, Richard G. Maroun, Zeina Hobaika, Rebecca Badra, and Dominique Salameh. 2022. "Characterization of Antagonist Potential of Selected Compost Bacterial Isolates (CBI) against Plant and Human Pathogens" Agronomy 12, no. 12: 2977. https://doi.org/10.3390/agronomy12122977








