The Long Terminal Repeats of ERV6 Are Activated in Pre-Implantation Embryos of Cynomolgus Monkey
Abstract
:1. Introduction
2. Materials and Methods
2.1. Animal Ethics
2.2. Calculation of the Expression Levels of LTR Elements and Genes
2.3. Oocyte Collection and In Vitro Fertilization
2.4. Lentivirus Production and Purification
2.5. Isolation of Green Fluorescent Protein (GFP) Positive Single Cell and scRNA-Seq
2.6. Single-Cell Cluster Analysis
2.7. TF Binding Site Predictions and Dual Luciferase Expression Assays
2.8. Immunofluorescence
3. Results
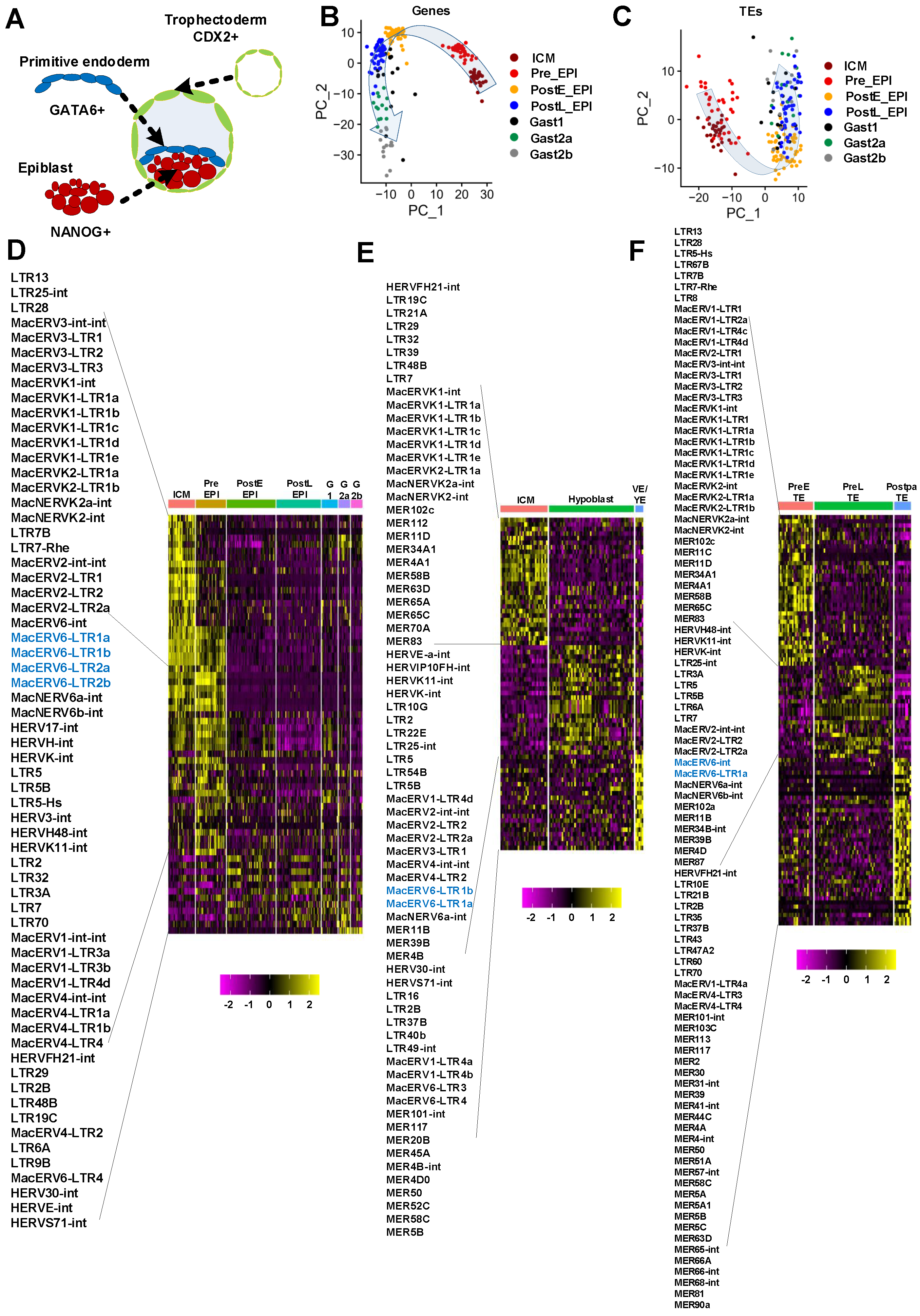
3.1. Temporal Expression of LTR Elements during Monkey Embryo Development
3.2. MacERV6-LTR1a Knockdown in Cynomolgus Monkey Embryos
3.3. Transcriptomic Changes after MacERV6-LTR1a Knockdown
3.4. The Interaction between MacERV6-LTR1a and Pluripotency Factors In Vitro
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| TF | Transcription factor |
| TrB | Trophectoderm |
| TE | Transposable Element |
| ICM | Inner cell mass |
| EPI | Epiblast |
| preEEPI | Pre-implantation early epiblast |
| preETE | Pre-implantation early trophectoderm |
| Pre-EPI | Pre-implantation epiblast |
| postE-EPI | Post-implantation early epiblast |
| postL-EPI | Post-implantation late epiblast |
| PrE | Primitive endoderm |
| ERV | Endogenous retroviruse |
| LTR | Long terminal repeat |
| hESC | Human embryonic stem cell |
| KRAB-ZFP | KRAB-containing zinc finger protein |
| rhFSH | Recombinant human follitropin alpha |
| rhCG | Recombinant human chorionic gonadotropin alpha |
| TALP | Tyrode’s albumin lactate pyruvate |
| BSA | Bovine serum albumin |
| MOR | Morula |
| DEGs | Differentially expressed genes |
| Gast1 | Gastrulating cells 1 |
| Gast2a | Gastrulating cells 2a |
| Gast2b | Gastrulating cells 2b |
References
- Chazaud, C.; Yamanaka, Y. Lineage specification in the mouse preimplantation embryo. Development 2016, 143, 1063–1074. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leung, C.Y.; Zhu, M.; Zernicka-Goetz, M. Polarity in Cell-Fate Acquisition in the Early Mouse Embryo. Curr. Top. Dev. Biol. 2016, 120, 203–234. [Google Scholar] [CrossRef]
- Rossant, J. Genetic Control of Early Cell Lineages in the Mammalian Embryo. Annu. Rev. Genet. 2018, 52, 185–201. [Google Scholar] [CrossRef] [PubMed]
- Wamaitha, S.E.; Niakan, K.K. Human Pre-gastrulation Development. Curr. Top. Dev. Biol. 2018, 128, 295–338. [Google Scholar] [CrossRef]
- Chazaud, C.; Yamanaka, Y.; Pawson, T.; Rossant, J. Early lineage segregation between epiblast and primitive endoderm in mouse blastocysts through the Grb2-MAPK pathway. Dev. Cell. 2006, 10, 615–624. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marikawa, Y.; Alarcon, V.B. Establishment of trophectoderm and inner cell mass lineages in the mouse embryo. Mol. Reprod. Dev. 2009, 76, 1019–1032. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zheng, Y.; Xue, X.; Shao, Y.; Wang, S.; Esfahani, S.N.; Li, Z.; Muncie, J.M.; Lakins, J.N.; Weaver, V.M.; Gumucio, D.L.; et al. Controlled modelling of human epiblast and amnion development using stem cells. Nature 2019, 573, 421–425. [Google Scholar] [CrossRef]
- Chang, C.W.; Wakeland, A.K.; Parast, M.M. Trophoblast lineage specification, differentiation and their regulation by oxygen tension. J Endocrinol. 2018, 236, R43–R56. [Google Scholar] [CrossRef] [Green Version]
- Luckett, W.P. Origin and differentiation of the yolk sac and extraembryonic mesoderm in presomite human and rhesus monkey embryos. Am. J. Anat. 1978, 152, 59–97. [Google Scholar] [CrossRef]
- Mole, M.A.; Weberling, A.; Zernicka-Goetz, M. Comparative analysis of human and mouse development: From zygote to pre-gastrulation. Curr. Top. Dev. Biol. 2020, 136, 113–138. [Google Scholar] [CrossRef]
- Boroviak, T.; Stirparo, G.G.; Dietmann, S.; Hernando-Herraez, I.; Mohammed, H.; Reik, W.; Smith, A.; Sasaki, E.; Nichols, J.; Bertone, P. Single cell transcriptome analysis of human, marmoset and mouse embryos reveals common and divergent features of preimplantation development. Development 2018, 145, dev167833. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nichols, J.; Smith, A. Naive and primed pluripotent states. Cell Stem Cell 2009, 4, 487–492. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peng, G.; Suo, S.; Cui, G.; Yu, F.; Wang, R.; Chen, J.; Chen, S.; Liu, Z.; Chen, G.; Qian, Y.; et al. Molecular architecture of lineage allocation and tissue organization in early mouse embryo. Nature 2019, 572, 528–532. [Google Scholar] [CrossRef]
- Frankenberg, S.; Gerbe, F.; Bessonnard, S.; Belville, C.; Pouchin, P.; Bardot, O.; Chazaud, C. Primitive endoderm differentiates via a three-step mechanism involving Nanog and RTK signaling. Dev. Cell. 2011, 21, 1005–1013. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guo, G.; Huss, M.; Tong, G.Q.; Wang, C.; Li Sun, L.; Clarke, N.D.; Robson, P. Resolution of cell fate decisions revealed by single-cell gene expression analysis from zygote to blastocyst. Dev. Cell. 2010, 18, 675–685. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kuijk, E.W.; van Tol, L.T.; Van de Velde, H.; Wubbolts, R.; Welling, M.; Geijsen, N.; Roelen, B.A. The roles of FGF and MAP kinase signaling in the segregation of the epiblast and hypoblast cell lineages in bovine and human embryos. Development 2012, 139, 871–882. [Google Scholar] [CrossRef] [Green Version]
- Roode, M.; Blair, K.; Snell, P.; Elder, K.; Marchant, S.; Smith, A.; Nichols, J. Human hypoblast formation is not dependent on FGF signalling. Dev. Biol. 2012, 361, 358–363. [Google Scholar] [CrossRef] [Green Version]
- Cordaux, R.; Batzer, M.A. The impact of retrotransposons on human genome evolution. Nat. Rev. Genet. 2009, 10, 691–703. [Google Scholar] [CrossRef] [Green Version]
- Fort, A.; Hashimoto, K.; Yamada, D.; Salimullah, M.; Keya, C.A.; Saxena, A.; Bonetti, A.; Voineagu, I.; Bertin, N.; Kratz, A.; et al. Deep transcriptome profiling of mammalian stem cells supports a regulatory role for retrotransposons in pluripotency maintenance. Nat. Genet. 2014, 46, 558–566. [Google Scholar] [CrossRef] [PubMed]
- Kapusta, A.; Kronenberg, Z.; Lynch, V.J.; Zhuo, X.; Ramsay, L.; Bourque, G.; Yandell, M.; Feschotte, C. Transposable elements are major contributors to the origin, diversification, and regulation of vertebrate long noncoding RNAs. PLoS Genet. 2013, 9, e1003470. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kelley, D.; Rinn, J. Transposable elements reveal a stem cell-specific class of long noncoding RNAs. Genome Biol. 2012, 13, R107. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lu, X.; Sachs, F.; Ramsay, L.; Jacques, P.E.; Goke, J.; Bourque, G.; Ng, H.H. The retrovirus HERVH is a long noncoding RNA required for human embryonic stem cell identity. Nat. Struct. Mol. Biol. 2014, 21, 423–425. [Google Scholar] [CrossRef]
- Macia, A.; Munoz-Lopez, M.; Cortes, J.L.; Hastings, R.K.; Morell, S.; Lucena-Aguilar, G.; Marchal, J.A.; Badge, R.M.; Garcia-Perez, J.L. Epigenetic control of retrotransposon expression in human embryonic stem cells. Mol. Cell. Biol. 2011, 31, 300–316. [Google Scholar] [CrossRef] [Green Version]
- Santoni, F.A.; Guerra, J.; Luban, J. HERV-H RNA is abundant in human embryonic stem cells and a precise marker for pluripotency. Retrovirology 2012, 9, 111. [Google Scholar] [CrossRef] [Green Version]
- Weiss, R.A. The discovery of endogenous retroviruses. Retrovirology 2006, 3, 67. [Google Scholar] [CrossRef] [Green Version]
- Cohen, C.J.; Lock, W.M.; Mager, D.L. Endogenous retroviral LTRs as promoters for human genes: A critical assessment. Gene 2009, 448, 105–114. [Google Scholar] [CrossRef] [PubMed]
- Jern, P.; Coffin, J.M. Effects of retroviruses on host genome function. Annu. Rev. Genet. 2008, 42, 709–732. [Google Scholar] [CrossRef] [Green Version]
- Maksakova, I.A.; Mager, D.L.; Reiss, D. Keeping active endogenous retroviral-like elements in check: The epigenetic perspective. Cell. Mol. Life Sci. 2008, 65, 3329–3347. [Google Scholar] [CrossRef]
- Macfarlan, T.S.; Gifford, W.D.; Driscoll, S.; Lettieri, K.; Rowe, H.M.; Bonanomi, D.; Firth, A.; Singer, O.; Trono, D.; Pfaff, S.L. Embryonic stem cell potency fluctuates with endogenous retrovirus activity. Nature 2012, 487, 57–63. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thompson, P.J.; Macfarlan, T.S.; Lorincz, M.C. Long Terminal Repeats: From Parasitic Elements to Building Blocks of the Transcriptional Regulatory Repertoire. Mol. Cell. 2016, 62, 766–776. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Wang, L.; Feng, G.; Wang, Y.; Li, Y.; Li, X.; Liu, C.; Jiao, G.; Huang, C.; Shi, J.; et al. Asymmetric Expression of LincGET Biases Cell Fate in Two-Cell Mouse Embryos. Cell 2018, 175, 1887–1901.e18. [Google Scholar] [CrossRef] [Green Version]
- Liu, D.; Liu, L.; Duan, K.; Guo, J.; Li, S.; Zhao, Z.; Zhang, X.; Zhou, N.; Zheng, Y. Transcriptional dynamics of transposable elements when converting fibroblast cells of Macaca mulatta to neuroepithelial stem cells. BMC Genom. 2021, 22, 405. [Google Scholar] [CrossRef]
- Jacques, P.E.; Jeyakani, J.; Bourque, G. The majority of primate-specific regulatory sequences are derived from transposable elements. PLoS Genet. 2013, 9, e1003504. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Goke, J.; Lu, X.; Chan, Y.S.; Ng, H.H.; Ly, L.H.; Sachs, F.; Szczerbinska, I. Dynamic transcription of distinct classes of endogenous retroviral elements marks specific populations of early human embryonic cells. Cell Stem Cell 2015, 16, 135–141. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peaston, A.E.; Evsikov, A.V.; Graber, J.H.; de Vries, W.N.; Holbrook, A.E.; Solter, D.; Knowles, B.B. Retrotransposons regulate host genes in mouse oocytes and preimplantation embryos. Dev. Cell. 2004, 7, 597–606. [Google Scholar] [CrossRef]
- Rowe, H.M.; Trono, D. Dynamic control of endogenous retroviruses during development. Virology 2011, 411, 273–287. [Google Scholar] [CrossRef] [Green Version]
- Grow, E.J.; Flynn, R.A.; Chavez, S.L.; Bayless, N.L.; Wossidlo, M.; Wesche, D.J.; Martin, L.; Ware, C.B.; Blish, C.A.; Chang, H.Y.; et al. Intrinsic retroviral reactivation in human preimplantation embryos and pluripotent cells. Nature 2015, 522, 221–225. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, J.; Xie, G.; Singh, M.; Ghanbarian, A.T.; Rasko, T.; Szvetnik, A.; Cai, H.; Besser, D.; Prigione, A.; Fuchs, N.V.; et al. Primate-specific endogenous retrovirus-driven transcription defines naive-like stem cells. Nature 2014, 516, 405–409. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ohnuki, M.; Tanabe, K.; Sutou, K.; Teramoto, I.; Sawamura, Y.; Narita, M.; Nakamura, M.; Tokunaga, Y.; Nakamura, M.; Watanabe, A.; et al. Dynamic regulation of human endogenous retroviruses mediates factor-induced reprogramming and differentiation potential. Proc. Natl. Acad. Sci. USA 2014, 111, 12426–12431. [Google Scholar] [CrossRef] [Green Version]
- Kunarso, G.; Chia, N.Y.; Jeyakani, J.; Hwang, C.; Lu, X.; Chan, Y.S.; Ng, H.H.; Bourque, G. Transposable elements have rewired the core regulatory network of human embryonic stem cells. Nat. Genet. 2010, 42, 631–634. [Google Scholar] [CrossRef] [PubMed]
- Sundaram, V.; Cheng, Y.; Ma, Z.; Li, D.; Xing, X.; Edge, P.; Snyder, M.P.; Wang, T. Widespread contribution of transposable elements to the innovation of gene regulatory networks. Genome Res. 2014, 24, 1963–1976. [Google Scholar] [CrossRef] [Green Version]
- Ecco, G.; Cassano, M.; Kauzlaric, A.; Duc, J.; Coluccio, A.; Offner, S.; Imbeault, M.; Rowe, H.M.; Turelli, P.; Trono, D. Transposable Elements and Their KRAB-ZFP Controllers Regulate Gene Expression in Adult Tissues. Dev. Cell 2016, 36, 611–623. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chan, A.W. Progress and prospects for genetic modification of nonhuman primate models in biomedical research. ILAR J. 2013, 54, 211–223. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, H.; Zhai, J.; Wan, H.; Jiang, X.; Wang, X.; Wang, L.; Xiang, Y.; He, X.; Zhao, Z.A.; Zhao, B.; et al. In vitro culture of cynomolgus monkey embryos beyond early gastrulation. Science 2019, 366. [Google Scholar] [CrossRef] [PubMed]
- Niu, Y.; Sun, N.; Li, C.; Lei, Y.; Huang, Z.; Wu, J.; Si, C.; Dai, X.; Liu, C.; Wei, J.; et al. Dissecting primate early post-implantation development using long-term in vitro embryo culture. Science 2019, 366. [Google Scholar] [CrossRef] [PubMed]
- Festuccia, N.; Dubois, A.; Vandormael-Pournin, S.; Gallego Tejeda, E.; Mouren, A.; Bessonnard, S.; Mueller, F.; Proux, C.; Cohen-Tannoudji, M.; Navarro, P. Mitotic binding of Esrrb marks key regulatory regions of the pluripotency network. Nat. Cell Biol. 2016, 18, 1139–1148. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhang, J.; Wang, T.; Esteban, M.A.; Pei, D. Esrrb activates Oct4 transcription and sustains self-renewal and pluripotency in embryonic stem cells. J. Biol. Chem. 2008, 283, 35825–35833. [Google Scholar] [CrossRef] [Green Version]
- Festuccia, N.; Osorno, R.; Halbritter, F.; Karwacki-Neisius, V.; Navarro, P.; Colby, D.; Wong, F.; Yates, A.; Tomlinson, S.R.; Chambers, I. Esrrb is a direct Nanog target gene that can substitute for Nanog function in pluripotent cells. Cell Stem Cell 2012, 11, 477–490. [Google Scholar] [CrossRef] [Green Version]
- Festuccia, N.; Osorno, R.; Wilson, V.; Chambers, I. The role of pluripotency gene regulatory network components in mediating transitions between pluripotent cell states. Curr. Opin. Genet. Dev. 2013, 23, 504–511. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Festuccia, N.; Owens, N.; Navarro, P. Esrrb, an estrogen-related receptor involved in early development, pluripotency, and reprogramming. FEBS Lett. 2018, 592, 852–877. [Google Scholar] [CrossRef] [Green Version]
- Martello, G.; Sugimoto, T.; Diamanti, E.; Joshi, A.; Hannah, R.; Ohtsuka, S.; Gottgens, B.; Niwa, H.; Smith, A. Esrrb is a pivotal target of the Gsk3/Tcf3 axis regulating embryonic stem cell self-renewal. Cell Stem Cell 2012, 11, 491–504. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nakamura, T.; Okamoto, I.; Sasaki, K.; Yabuta, Y.; Iwatani, C.; Tsuchiya, H.; Seita, Y.; Nakamura, S.; Yamamoto, T.; Saitou, M. A developmental coordinate of pluripotency among mice, monkeys and humans. Nature 2016, 537, 57–62. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Babarinde, I.A.; Sun, L.; Xu, S.; Chen, R.; Shi, J.; Wei, Y.; Li, Y.; Ma, G.; Zhuang, Q.; et al. Identifying transposable element expression dynamics and heterogeneity during development at the single-cell level with a processing pipeline scTE. Nat. Commun. 2021, 12, 1456. [Google Scholar] [CrossRef]
- Niu, Y.; Yu, Y.; Bernat, A.; Yang, S.; He, X.; Guo, X.; Chen, D.; Chen, Y.; Ji, S.; Si, W.; et al. Transgenic rhesus monkeys produced by gene transfer into early-cleavage-stage embryos using a simian immunodeficiency virus-based vector. Proc. Natl. Acad. Sci. USA 2010, 107, 17663–17667. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Picelli, S.; Faridani, O.R.; Bjorklund, A.K.; Winberg, G.; Sagasser, S.; Sandberg, R. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 2014, 9, 171–181. [Google Scholar] [CrossRef] [PubMed]
- Stuart, T.; Butler, A.; Hoffman, P.; Hafemeister, C.; Papalexi, E.; Mauck, W.M., III; Hao, Y.; Stoeckius, M.; Smibert, P.; Satija, R. Comprehensive Integration of Single-Cell Data. Cell 2019, 177, 1888–1902.e21. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [Green Version]
- Zhou, Y.; Zhou, B.; Pache, L.; Chang, M.; Khodabakhshi, A.H.; Tanaseichuk, O.; Benner, C.; Chanda, S.K. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 2019, 10, 1523. [Google Scholar] [CrossRef]
- Bailey, T.L.; Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 1994, 2, 28–36. [Google Scholar]
- Gupta, S.; Stamatoyannopoulos, J.A.; Bailey, T.L.; Noble, W.S. Quantifying similarity between motifs. Genome Biol. 2007, 8, R24. [Google Scholar] [CrossRef] [Green Version]
- Wolf, G.; Greenberg, D.; Macfarlan, T.S. Spotting the enemy within: Targeted silencing of foreign DNA in mammalian genomes by the Kruppel-associated box zinc finger protein family. Mob. DNA 2015, 6, 17. [Google Scholar] [CrossRef] [Green Version]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Morris, S.A.; Teo, R.T.; Li, H.; Robson, P.; Glover, D.M.; Zernicka-Goetz, M. Origin and formation of the first two distinct cell types of the inner cell mass in the mouse embryo. Proc. Natl. Acad. Sci. USA 2010, 107, 6364–6369. [Google Scholar] [CrossRef] [Green Version]
- Yamanaka, Y.; Lanner, F.; Rossant, J. FGF signal-dependent segregation of primitive endoderm and epiblast in the mouse blastocyst. Development 2010, 137, 715–724. [Google Scholar] [CrossRef] [Green Version]
- Feschotte, C.; Gilbert, C. Endogenous viruses: Insights into viral evolution and impact on host biology. Nat. Rev. Genet. 2012, 13, 283–296. [Google Scholar] [CrossRef] [Green Version]
- Rossant, J.; Tam, P.P.L. New Insights into Early Human Development: Lessons for Stem Cell Derivation and Differentiation. Cell Stem Cell 2017, 20, 18–28. [Google Scholar] [CrossRef] [Green Version]
- Richter, K.S.; Harris, D.C.; Daneshmand, S.T.; Shapiro, B.S. Quantitative grading of a human blastocyst: Optimal inner cell mass size and shape. Fertil. Steril. 2001, 76, 1157–1167. [Google Scholar] [CrossRef]
- Lindenberg, S.; Hyttel, P.; Sjogren, A.; Greve, T. A comparative study of attachment of human, bovine and mouse blastocysts to uterine epithelial monolayer. Hum. Reprod. 1989, 4, 446–456. [Google Scholar] [CrossRef] [PubMed]
- Xiang, L.; Yin, Y.; Zheng, Y.; Ma, Y.; Li, Y.; Zhao, Z.; Guo, J.; Ai, Z.; Niu, Y.; Duan, K.; et al. A developmental landscape of 3D-cultured human pre-gastrulation embryos. Nature 2020, 577, 537–542. [Google Scholar] [CrossRef] [PubMed]
- Chambers, I.; Tomlinson, S.R. The transcriptional foundation of pluripotency. Development 2009, 136, 2311–2322. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shi, G.; Jin, Y. Role of Oct4 in maintaining and regaining stem cell pluripotency. Stem Cell Res. Ther. 2010, 1, 39. [Google Scholar] [CrossRef] [Green Version]
- Chia, N.Y.; Chan, Y.S.; Feng, B.; Lu, X.; Orlov, Y.L.; Moreau, D.; Kumar, P.; Yang, L.; Jiang, J.; Lau, M.S.; et al. A genome-wide RNAi screen reveals determinants of human embryonic stem cell identity. Nature 2010, 468, 316–320. [Google Scholar] [CrossRef] [PubMed]
- Takashima, Y.; Guo, G.; Loos, R.; Nichols, J.; Ficz, G.; Krueger, F.; Oxley, D.; Santos, F.; Clarke, J.; Mansfield, W.; et al. Resetting transcription factor control circuitry toward ground-state pluripotency in human. Cell 2014, 158, 1254–1269. [Google Scholar] [CrossRef] [Green Version]
- Takahashi, K.; Yamanaka, S. A decade of transcription factor-mediated reprogramming to pluripotency. Nat. Rev. Mol. Cell. Biol. 2016, 17, 183–193. [Google Scholar] [CrossRef] [PubMed]





Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Duan, K.; Si, C.-Y.; Zhao, S.-M.; Ai, Z.-Y.; Niu, B.-H.; Yin, Y.; Xiang, L.-F.; Ding, H.; Zheng, Y. The Long Terminal Repeats of ERV6 Are Activated in Pre-Implantation Embryos of Cynomolgus Monkey. Cells 2021, 10, 2710. https://doi.org/10.3390/cells10102710
Duan K, Si C-Y, Zhao S-M, Ai Z-Y, Niu B-H, Yin Y, Xiang L-F, Ding H, Zheng Y. The Long Terminal Repeats of ERV6 Are Activated in Pre-Implantation Embryos of Cynomolgus Monkey. Cells. 2021; 10(10):2710. https://doi.org/10.3390/cells10102710
Chicago/Turabian StyleDuan, Kui, Chen-Yang Si, Shu-Mei Zhao, Zong-Yong Ai, Bao-Hua Niu, Yu Yin, Li-Feng Xiang, Hao Ding, and Yun Zheng. 2021. "The Long Terminal Repeats of ERV6 Are Activated in Pre-Implantation Embryos of Cynomolgus Monkey" Cells 10, no. 10: 2710. https://doi.org/10.3390/cells10102710





