IL-13 as Target to Reduce Cholestasis and Dysbiosis in Abcb4 Knockout Mice
Abstract
:1. Introduction
2. Materials and Methods
2.1. Animal Model
2.2. Routine Serum Biochemistry
2.3. Histology
2.4. Immunohistochemistry/Immunofluorescence
2.5. Bile Acid Analysis
2.6. Western Blot Analysis
2.7. Transcriptome Microarray and Gene Set Analysis
2.8. Bacterial Translocation to the Liver
2.9. Quantitative Real-Time PCR
2.10. Hepatic Hydroxyproline Ccontent
2.11. 16 SrDNA Sequencing forMicrobiome Profiling
2.12. Bioinformatics Workflow
2.13. Statistical Analysis
3. Results
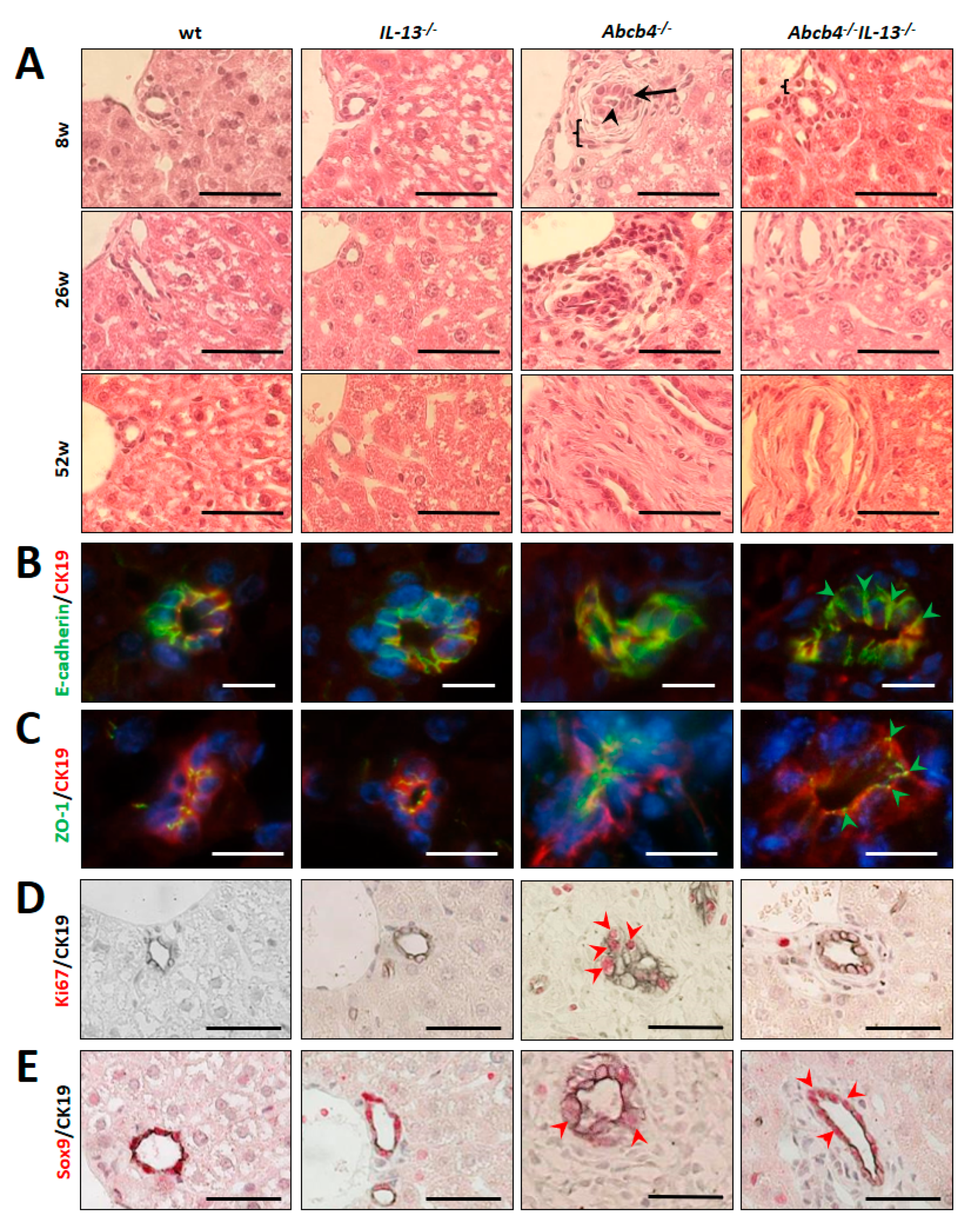
3.1. IL-13 Knockout Restored Intrahepatic Bile Duct Integrity and Biliary Epithelial Cell Barrier Function in Abcb4−/− Mice
3.2. Reduced Serum BA Improved Liver Integrity
3.3. Lack of IL-13 Reduces Hepatic Pathology and Fibrosis
3.4. IL-13 Alters Hepatic Gene Expression
3.5. Lack of IL-13 Alters Bile Acid Excretion Patterns, Improves Ileal Integrity, and Reduces Bacterial Translocation
3.6. IL-13 Depletion is Associated with Profound Compositional Changes in Gut Microbiota
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Rao, R.K.; Samak, G. Bile duct epithelial tight junctions and barrier function. Tissue Barriers 2013, 1, e25718. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sakisaka, S. Alterations in tight junctions differ between primary biliary cirrhosis and primary sclerosing cholangitis. Hepatology 2001, 33, 1460–1468. [Google Scholar] [CrossRef]
- Grosse, B.; Cassio, D.; Yousef, N.; Bernardo, C.; Jacquemin, E.; Gonzales, E. Claudin-1 involved in neonatal ichthyosis sclerosing cholangitis syndrome regulates hepatic paracellular permeability. Hepatology 2012, 55, 1249–1259. [Google Scholar] [CrossRef] [PubMed]
- Fickert, P.; Fuchsbichler, A.; Wagner, M.; Zollner, G.; Kaser, A.; Tilg, H.; Krause, R.; Lammert, F.; Langner, C.; Zatloukal, K.; et al. Regurgitation of bile acids from leaky bile ducts causes sclerosing cholangitis in Mdr2 (Abcb4) knockout mice. Gastroenterology 2004, 127, 261–274. [Google Scholar] [CrossRef] [PubMed]
- Sasatomi, K.; Noguchi, K.; Sakisaka, S.; Sata, M.; Tanikawa, K. Abnormal accumulation of endotoxin in biliary epithelial cells in primary biliary cirrhosis and primary sclerosing cholangitis. J. Hepatol. 1998, 29, 409–416. [Google Scholar] [CrossRef]
- Williamson, K.D.; Chapman, R.W. Primary sclerosing cholangitis. Dig. Dis. 2014, 32, 438–445. [Google Scholar] [CrossRef]
- Konturek, P.C.; Harsch, I.A.; Konturek, K.; Schink, M.; Konturek, T.; Neurath, M.F.; Zopf, Y. Gut-Liver Axis: How Do Gut Bacteria Influence the Liver? Med. Sci. (Basel) 2018, 6, 79. [Google Scholar] [CrossRef] [Green Version]
- Reyes, H.; Zapata, R.; Hernández, I.; Gotteland, M.; Sandoval, L.; Jirón, M.I.; Palma, J.; Almuna, R.; Silva, J.J. Is a leaky gut involved in the pathogenesis of intrahepatic cholestasis of pregnancy? Hepatology 2006, 43, 715–722. [Google Scholar] [CrossRef]
- Odenwald, M.A.; Turner, J.R. The intestinal epithelial barrier: A therapeutic target? Nat. Rev. Gastroenterol. Hepatol. 2017, 14, 9–21. [Google Scholar] [CrossRef]
- Tilg, H.; Cani, P.D.; Mayer, E.A. Gut microbiome and liver diseases. Gut 2016, 65, 2035–2044. [Google Scholar] [CrossRef]
- Cabrera-Rubio, R.; Patterson, A.M.; Cotter, P.D.; Beraza, N. Cholestasis induced by bile duct ligation promotes changes in the intestinal microbiome in mice. Sci. Rep. 2019, 9, 12324. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Perez, M.-J.; Briz, O. Bile-acid-induced cell injury and protection. World J. Gastroenterol. 2009, 15, 1677–1689. [Google Scholar] [CrossRef] [PubMed]
- Bessho, K.; Bezerra, J.A. Biliary atresia: Will blocking inflammation tame the disease? Annu. Rev. Med. 2011, 62, 171–185. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bo, X.; Broome, U.; Remberger, M.; Sumitran-Holgersson, S. Tumour necrosis factor alpha impairs function of liver derived T lymphocytes and natural killer cells in patients with primary sclerosing cholangitis. Gut 2001, 49, 131–141. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mano, Y.; Ishii, M.; Okamoto, H.; Igarashi, T.; Kobayashi, K.; Toyota, T. Effect of tumor necrosis factor alpha on intrahepatic bile duct epithelial cell of rat liver. Hepatology 1996, 23, 1602–1607. [Google Scholar] [CrossRef]
- Hanada, S.; Harada, M.; Koga, H.; Kawaguchi, T.; Taniguchi, E.; Kumashiro, R.; Ueno, T.; Ueno, Y.; Ishii, M.; Sakisaka, S.; et al. Tumor necrosis factor-alpha and interferon-gamma directly impair epithelial barrier function in cultured mouse cholangiocytes. Liver Int. 2003, 23, 3–11. [Google Scholar] [CrossRef]
- Müller, T.; Beutler, C.; Picó, A.H.; Otten, M.; Dürr, A.; Al-Abadi, H.; Guckelberger, O.; Zum Meyer Büschenfelde, D.; Jöhrens, K.; Volkmann, M.; et al. Increased T-helper 2 cytokines in bile from patients with IgG4-related cholangitis disrupt the tight junction-associated biliary epithelial cell barrier. Gastroenterology 2013, 144, 1116–1128. [Google Scholar] [CrossRef]
- Zen, Y.; Fujii, T.; Harada, K.; Kawano, M.; Yamada, K.; Takahira, M.; Nakanuma, Y. Th2 and regulatory immune reactions are increased in immunoglobin G4-related sclerosing pancreatitis and cholangitis. Hepatology 2007, 45, 1538–1546. [Google Scholar] [CrossRef]
- Li, J.; Razumilava, N.; Gores, G.J.; Walters, S.; Mizuochi, T.; Mourya, R.; Bessho, K.; Wang, Y.-H.; Glaser, S.S.; Shivakumar, P.; et al. Biliary repair and carcinogenesis are mediated by IL-33-dependent cholangiocyte proliferation. J. Clin. Invest. 2014, 124, 3241–3251. [Google Scholar] [CrossRef] [Green Version]
- Hammerich, L.; Tacke, F. Interleukins in chronic liver disease: Lessons learned from experimental mouse models. Clin. Exp. Gastroenterol. 2014, 7, 297–306. [Google Scholar] [CrossRef] [Green Version]
- Chiaramonte, M.G.; Donaldson, D.D.; Cheever, A.W.; Wynn, T.A. An IL-13 inhibitor blocks the development of hepatic fibrosis during a T-helper type 2-dominated inflammatory response. J. Clin. Invest. 1999, 104, 777–785. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.; Munker, S.; Müllenbach, R.; Weng, H.-L. IL-13 Signaling in liver fibrogenesis. Front. Immunol. 2012, 3, 116. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Long, X.; Chen, Q.; Zhao, J.; Rafaels, N.; Mathias, P.; Liang, H.; Potee, J.; Campbell, M.; Zhang, B.; Gao, L.; et al. An IL-13 promoter polymorphism associated with liver fibrosis in patients with Schistosoma japonicum. PLoS ONE 2015, 10, e0135360. [Google Scholar] [CrossRef] [PubMed]
- Weng, H.-L.; Liu, Y.; Chen, J.-L.; Huang, T.; Xu, L.-J.; Godoy, P.; Hu, J.-H.; Zhou, C.; Stickel, F.; Marx, A.; et al. The etiology of liver damage imparts cytokines transforming growth factor beta1 or interleukin-13 as driving forces in fibrogenesis. Hepatology 2009, 50, 230–243. [Google Scholar] [CrossRef] [PubMed]
- Churin, Y.; Roderfeld, M.; Stiefel, J.; Würger, T.; Schröder, D.; Matono, T.; Mollenkopf, H.-J.; Montalbano, R.; Pompaiah, M.; Reifenberg, K.; et al. Pathological impact of hepatitis B virus surface proteins on the liver is associated with the host genetic background. PLoS ONE 2014, 9, e90608. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McKenzie, G.J.; Emson, C.L.; Bell, S.E.; Anderson, S.; Fallon, P.; Zurawski, G.; Murray, R.; Grencis, R.; McKenzie, A.N.J. Impaired development of Th2 cells in IL-13-deficient mice. Immunity 1998, 9, 423–432. [Google Scholar] [CrossRef] [Green Version]
- Roderfeld, M.; Rath, T.; Pasupuleti, S.; Zimmermann, M.; Neumann, C.; Churin, Y.; Dierkes, C.; Voswinckel, R.; Barth, P.J.; Zahner, D.; et al. Bone marrow transplantation improves hepatic fibrosis in Abcb4−/− mice via Th1 response and matrix metalloproteinase activity. Gut 2012, 61, 907–916. [Google Scholar] [CrossRef]
- Roderfeld, M.; Rath, T.; Voswinckel, R.; Dierkes, C.; Dietrich, H.; Zahner, D.; Graf, J.; Roeb, E. Bone marrow transplantation demonstrates medullar origin of CD34+ fibrocytes and ameliorates hepatic fibrosis in Abcb4−/− mice. Hepatology 2010, 51, 267–276. [Google Scholar] [CrossRef]
- Baghdasaryan, A.; Fickert, P.; Fuchsbichler, A.; Silbert, D.; Gumhold, J.; Hörl, G.; Langner, C.; Moustafa, T.; Halilbasic, E.; Claudel, T.; et al. Role of hepatic phospholipids in development of liver injury in Mdr2 (Abcb4) knockout mice. Liver Int. 2008, 28, 948–958. [Google Scholar] [CrossRef]
- García-Cañaveras, J.C.; Donato, M.T.; Castell, J.V.; Lahoz, A. Targeted profiling of circulating and hepatic bile acids in human, mouse, and rat using a UPLC-MRM-MS-validated method. J. Lipid Res. 2012, 53, 2231–2241. [Google Scholar] [CrossRef] [Green Version]
- Li, K.; Buchinger, T.J.; Bussy, U.; Fissette, S.D.; Johnson, N.S.; Li, W. Quantification of 15 bile acids in lake charr feces by ultra-high performance liquid chromatography-tandem mass spectrometry. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 2015, 1001, 27–34. [Google Scholar] [CrossRef] [PubMed]
- Reece, S.T.; Loddenkemper, C.; Askew, D.J.; Zedler, U.; Schommer-Leitner, S.; Stein, M.; Mir, F.A.; Dorhoi, A.; Mollenkopf, H.-J.; Silverman, G.A.; et al. Serine protease activity contributes to control of Mycobacterium tuberculosis in hypoxic lung granulomas in mice. J. Clin. Invest. 2010, 120, 3365–3376. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.W.; Sherman, B.T.; Tan, Q.; Collins, J.R.; Alvord, W.G.; Roayaei, J.; Stephens, R.; Baseler, M.W.; Lane, H.C.; Lempicki, R.A. The DAVID Gene Functional Classification Tool: A novel biological module-centric algorithm to functionally analyze large gene lists. Genome Biol. 2007, 8, R183. [Google Scholar] [CrossRef] [Green Version]
- Kanehisa, M.; Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef] [PubMed]
- Irungbam, K.; Churin, Y.; Matono, T.; Weglage, J.; Ocker, M.; Glebe, D.; Hardt, M.; Koeppel, A.; Roderfeld, M.; Roeb, E. Cannabinoid receptor 1 knockout alleviates hepatic steatosis by downregulating perilipin 2. Lab. Invest. 2020, 100, 454–465. [Google Scholar] [CrossRef]
- Jamall, I.S.; Finelli, V.N.; Que Hee, S.S. A simple method to determine nanogram levels of 4-hydroxyproline in biological tissues. Anal. Biochem. 1981, 112, 70–75. [Google Scholar] [CrossRef]
- Kozich, J.J.; Westcott, S.L.; Baxter, N.T.; Highlander, S.K.; Schloss, P.D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 2013, 79, 5112–5120. [Google Scholar] [CrossRef] [Green Version]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [Green Version]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef] [Green Version]
- Demarez, C.; Hubert, C.; Sempoux, C.; Lemaigre, F.P. Expression of Molecular Differentiation Markers Does Not Correlate with Histological Differentiation Grade in Intrahepatic Cholangiocarcinoma. PLoS ONE 2016, 11, e0157140. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Heuman, D.M. Quantitative estimation of the hydrophilic-hydrophobic balance of mixed bile salt solutions. J. Lipid Res. 1989, 30, 719–730. [Google Scholar] [PubMed]
- Wang, D.Q.-H.; Tazuma, S.; Cohen, D.E.; Carey, M.C. Feeding natural hydrophilic bile acids inhibits intestinal cholesterol absorption: Studies in the gallstone-susceptible mouse. Am. J. Physiol. Gastrointest. Liver Physiol. 2003, 285, G494–G502. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Billington, D.; Evans, C.E.; Godfrey, P.P.; Coleman, R. Effects of bile salts on the plasma membranes of isolated rat hepatocytes. Biochem. J. 1980, 188, 321–327. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Palmer, R.H. Bile acids, liver injury, and liver disease. Arch. Intern. Med. 1972, 130, 606–617. [Google Scholar] [CrossRef] [PubMed]
- Alaish, S.M.; Smith, A.D.; Timmons, J.; Greenspon, J.; Eyvazzadeh, D.; Murphy, E.; Shea-Donahue, T.; Cirimotich, S.; Mongodin, E.; Zhao, A.; et al. Gut microbiota, tight junction protein expression, intestinal resistance, bacterial translocation and mortality following cholestasis depend on the genetic background of the host. Gut Microbes 2013, 4, 292–305. [Google Scholar] [CrossRef] [Green Version]
- France, M.M.; Turner, J.R. The mucosal barrier at a glance. J. Cell Sci. 2017, 130, 307–314. [Google Scholar] [CrossRef] [Green Version]
- Zihni, C.; Mills, C.; Matter, K.; Balda, M.S. Tight junctions: From simple barriers to multifunctional molecular gates. Nat. Rev. Mol. Cell Biol. 2016, 17, 564–580. [Google Scholar] [CrossRef]
- Németh, Z.; Szász, A.M.; Somorácz, A.; Tátrai, P.; Németh, J.; Gyorffy, H.; Szíjártó, A.; Kupcsulik, P.; Kiss, A.; Schaff, Z. Zonula occludens-1, occludin, and E-cadherin protein expression in biliary tract cancers. Pathol. Oncol. Res. 2009, 15, 533–539. [Google Scholar] [CrossRef]
- Li, J.; Bessho, K.; Shivakumar, P.; Mourya, R.; Mohanty, S.K.; Dos Santos, J.L.; Miura, I.K.; Porta, G.; Bezerra, J.A. Th2 signals induce epithelial injury in mice and are compatible with the biliary atresia phenotype. J. Clin. Invest. 2011, 121, 4244–4256. [Google Scholar] [CrossRef]
- Song, P.; Zhang, Y.; Klaassen, C.D. Dose-response of five bile acids on serum and liver bile Acid concentrations and hepatotoxicty in mice. Toxicol. Sci. 2011, 123, 359–367. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gieseck, R.L.; Ramalingam, T.R.; Hart, K.M.; Vannella, K.M.; Cantu, D.A.; Lu, W.Y.; Ferreira-González, S.; Forbes, S.J.; Vallier, L.; Wynn, T.A. Interleukin-13 Activates Distinct Cellular Pathways Leading to Ductular Reaction, Steatosis, and Fibrosis. Immunity 2016, 45, 145–158. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adachi, T.; Kaminaga, T.; Yasuda, H.; Kamiya, T.; Hara, H. The involvement of endoplasmic reticulum stress in bile acid-induced hepatocellular injury. J. Clin. Biochem. Nutr. 2014, 54, 129–135. [Google Scholar] [CrossRef] [Green Version]
- Bazellières, E.; Conte, V.; Elosegui-Artola, A.; Serra-Picamal, X.; Bintanel-Morcillo, M.; Roca-Cusachs, P.; Muñoz, J.J.; Sales-Pardo, M.; Guimerà, R.; Trepat, X. Control of cell-cell forces and collective cell dynamics by the intercellular sdhesome. Nat. Cell Biol. 2015, 17, 409–420. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Halilbasic, E.; Fuchs, C.; Hofer, H.; Paumgartner, G.; Trauner, M. Therapy of Primary Sclerosing Cholangitis--Today and Tomorrow. Dig. Dis. 2015, 33, 149–163. [Google Scholar] [CrossRef]
- Cai, S.-Y.; Mennone, A.; Soroka, C.J.; Boyer, J.L. All-trans-retinoic acid improves cholestasis in α-naphthylisothiocyanate-treated rats and Mdr2−/− mice. J. Pharmacol. Exp. Ther. 2014, 349, 94–98. [Google Scholar] [CrossRef] [Green Version]
- Heller, F.; Florian, P.; Bojarski, C.; Richter, J.; Christ, M.; Hillenbrand, B.; Mankertz, J.; Gitter, A.H.; Bürgel, N.; Fromm, M.; et al. Interleukin-13 is the key effector Th2 cytokine in ulcerative colitis that affects epithelial tight junctions, apoptosis, and cell restitution. Gastroenterology 2005, 129, 550–564. [Google Scholar] [CrossRef]
- Hruz, P.; Zimmermann, C.; Gutmann, H.; Degen, L.; Beuers, U.; Terracciano, L.; Drewe, J.; Beglinger, C. Adaptive regulation of the ileal apical sodium dependent bile acid transporter (ASBT) in patients with obstructive cholestasis. Gut 2006, 55, 395–402. [Google Scholar] [CrossRef]
- Vítek, L. Bile acid malabsorption in inflammatory bowel disease. Inflamm. Bowel Dis. 2015, 21, 476–483. [Google Scholar] [CrossRef]
- Stenman, L.K.; Holma, R.; Forsgård, R.; Gylling, H.; Korpela, R. Higher fecal bile acid hydrophobicity is associated with exacerbation of dextran sodium sulfate colitis in mice. J. Nutr. 2013, 143, 1691–1697. [Google Scholar] [CrossRef] [Green Version]
- Kakiyama, G.; Hylemon, P.B.; Zhou, H.; Pandak, W.M.; Heuman, D.M.; Kang, D.J.; Takei, H.; Nittono, H.; Ridlon, J.M.; Fuchs, M.; et al. Colonic inflammation and secondary bile acids in alcoholic cirrhosis. Am. J. Physiol. Gastrointest. Liver Physiol. 2014, 306, G929–G937. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pavlidis, P.; Powell, N.; Vincent, R.P.; Ehrlich, D.; Bjarnason, I.; Hayee, B. Systematic review: Bile acids and intestinal inflammation-luminal aggressors or regulators of mucosal defence? Aliment. Pharmacol. Ther. 2015, 42, 802–817. [Google Scholar] [CrossRef] [PubMed]
- Vega-Magaña, N.; Delgado-Rizo, V.; García-Benavides, L.; Del Toro-Arreola, S.; Segura-Ortega, J.; Morales, A.S.M.Z.; Zepeda-Nuño, J.S.; Escarra-Senmarti, M.; Gutiérrez-Franco, J.; Haramati, J.; et al. Bacterial translocation is linked to increased intestinal IFN-γ, IL-4, IL-17, and mucin-2 in cholestatic rats. Ann. Hepatol. 2018, 17, 318–329. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Ji, F.; Guo, J.; Shi, D.; Fang, D.; Li, L. Dysbiosis of small intestinal microbiota in liver cirrhosis and its association with etiology. Sci. Rep. 2016, 6, 34055. [Google Scholar] [CrossRef] [PubMed]
- Lam, Y.Y.; Ha, C.W.Y.; Campbell, C.R.; Mitchell, A.J.; Dinudom, A.; Oscarsson, J.; Cook, D.I.; Hunt, N.H.; Caterson, I.D.; Holmes, A.J.; et al. Increased gut permeability and microbiota change associate with mesenteric fat inflammation and metabolic dysfunction in diet-induced obese mice. PLoS ONE 2012, 7, e34233. [Google Scholar] [CrossRef] [Green Version]
- Li, J.; Sung, C.Y.J.; Lee, N.; Ni, Y.; Pihlajamäki, J.; Panagiotou, G.; El-Nezami, H. Probiotics modulated gut microbiota suppresses hepatocellular carcinoma growth in mice. Proc. Natl. Acad. Sci. USA 2016, 113, E1306–E1315. [Google Scholar] [CrossRef] [Green Version]
- Zhang, M.; Wang, X.-Q.; Zhou, Y.-K.; Ma, Y.-L.; Shen, T.-Y.; Chen, H.-Q.; Chu, Z.-X.; Qin, H.-L. Effects of oral Lactobacillus plantarum on hepatocyte tight junction structure and function in rats with obstructive jaundice. Mol. Biol. Rep. 2010, 37, 2989–2999. [Google Scholar] [CrossRef]
- Swann, J.R.; Want, E.J.; Geier, F.M.; Spagou, K.; Wilson, I.D.; Sidaway, J.E.; Nicholson, J.K.; Holmes, E. Systemic gut microbial modulation of bile acid metabolism in host tissue compartments. Proc. Natl. Acad. Sci. USA 2011, 108, 4523–4530. [Google Scholar] [CrossRef] [Green Version]
- Inagaki, T.; Moschetta, A.; Lee, Y.-K.; Peng, L.; Zhao, G.; Downes, M.; Yu, R.T.; Shelton, J.M.; Richardson, J.A.; Repa, J.J.; et al. Regulation of antibacterial defense in the small intestine by the nuclear bile acid receptor. Proc. Natl. Acad. Sci. USA 2006, 103, 3920–3925. [Google Scholar] [CrossRef] [Green Version]
- Wahlström, A.; Sayin, S.I.; Marschall, H.-U.; Bäckhed, F. Intestinal Crosstalk between Bile Acids and Microbiota and Its Impact on Host Metabolism. Cell Metab. 2016, 24, 41–50. [Google Scholar] [CrossRef] [Green Version]
- Tian, Y.; Gui, W.; Koo, I.; Smith, P.B.; Allman, E.L.; Nichols, R.G.; Rimal, B.; Cai, J.; Liu, Q.; Patterson, A.D. The microbiome modulating activity of bile acids. Gut Microbes 2020, 11, 979–996. [Google Scholar] [CrossRef] [PubMed]





| Term | Count | p-Value * | Gene Name |
|---|---|---|---|
| Secondary metabolites biosynthesis, transport, and catabolism | 18 14 | 4.9 × 10−9 1.4× 10−7 | Cyp17a1, Cyp2b13, Cyp2a22, Cyp2c55, Cyp2c67, Cyp2c68, Cyp2g1, Cyp2c37, Cyp26a1, Cyp26b1, Cyp3a11, Cyp3a16, Cyp3a41b, Cyp3a44, Cyp4a10, Cyp4a12b, Cyp4a12a, Cyp4a31, Cyp4f18, Cyp7a1, Cyp7b1 |
| Extracellular matrix | 24 26 | 1.2× 10−9 3.0× 10−13 | Adamtsl2, Fras1, Adamts2, Adamts4, Ctsg, Col1a1, Col1a2, Col3a1, Col5a3, Col6a2, Col6a3, Dmbt1, Dpt, Fbn1, Lgals3, Lpl, Lum, Loxl1, Mgp, Mmp12, Mmp13, Mmp2, Mmp3, Mmp7, Mfap4, Mfge8, Slpi, Serpine1, Thbs1, Timp1, Zg16 |
| Retinol metabolism | 15 11 | 2.4× 10−7 2.4× 10−5 | Ugt2b37, Cyp2b13, Cyp2c55, Cyp2c68, Cyp2c37, Cyp26a1, Cyp26b1, Cyp3a11, Cyp3a16, Cyp3a41b, Cyp3a44, Cyp4a10, Cyp4a12b, Cyp4a12a, Cyp4a31, Dhrs9 |
| PPAR signaling pathway | 13 8 | 3.0× 10−6 1.6× 10−3 | Adipog, Cyp4a10, Cyp4a12b, Cyp4a12a, Cyp4a31, Cyp7a1, Fabp3, Fabp5, Lpl, Plin1, Ppard, Scd1, Scd3, Ucp1 |
| Muscle contraction | 9 9 | 1.2× 10−5 2.4× 10−6 | Cacna1s, Lmod3, Myom2, Mybpc1, Myh1, Myh2, Myh4, Myh7, Myl1, Tmod4 |
| Cell adhesion | 30 23 | 3.5× 10−5 2.1× 10−4 | Cd24a, Frem1, Ajap1, Cdh1, Clstn2, Clstn3, Cgref1, Cx3cl1, Col6a2, Ctgf, Cxadr, Dpt, Emb, Efs, Flrt1, Flrt3, Fbln7, Fblim1, Itga11, Itga8, Itgax, Lamc3, Lamc2, Mfap4, Mfge8, Myh10, Ncam1, Spp1, Siglecf, Sdk1, Svep1, Thbs1, Tinag, Tnfrsf12a, Ttyh1, Vcam1 |
| Steroid hormone biosynthesis | 12 10 | 4.1× 10−5 9.3× 10−5 | Ugt2b37, Cyp17a1, Cyp2b13, Cyp2c55, Cyp2c68, Cyp2c37, Cyp3a11, Cyp3a16, Cyp3a41b, Cyp3a44, Cyp7a1, Cyp7b1, Hsd3b1, Hsd3b5, Sult1e1 |
| Focal adhesion | 18 14 | 3.2× 10−4 4.7× 10−4 | Shc2, Actg1, Cav3, Col1a1, Col1a2, Col3a1, Col5a3, Col6a2, Col6a3, Ccnd1, Itga11, Itga8, Jun, Lamc3, Lamc2, Myl2, Pak6, Pdgfa, Spp1, Thbs1 |
| ECM receptor interaction | 10 10 | 1.0× 10−3 1.0× 10−4 | Col1a1, Col1a2, Col3a1, Col5a3, Col6a2, Col6a3, Itga11, Itga8, Lamc3, Lamc2, Spp1, Thbs1 |
| Chemical carcinogenesis | 10 8 | 1.4× 10−3 3.5× 10−3 | Ugt2b37, Cyp2b13, Cyp2c55, Cyp2c68, Cyp2c37, Cyp3a11, Cyp3a16, Cyp3a41b, Cyp3a44, Gstt3, Sult2a1 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hahn, L.; Helmrich, N.; Herebian, D.; Mayatepek, E.; Drebber, U.; Domann, E.; Olejniczak, S.; Weigel, M.; Hain, T.; Rath, T.; et al. IL-13 as Target to Reduce Cholestasis and Dysbiosis in Abcb4 Knockout Mice. Cells 2020, 9, 1949. https://doi.org/10.3390/cells9091949
Hahn L, Helmrich N, Herebian D, Mayatepek E, Drebber U, Domann E, Olejniczak S, Weigel M, Hain T, Rath T, et al. IL-13 as Target to Reduce Cholestasis and Dysbiosis in Abcb4 Knockout Mice. Cells. 2020; 9(9):1949. https://doi.org/10.3390/cells9091949
Chicago/Turabian StyleHahn, Luisa, Nora Helmrich, Diran Herebian, Ertan Mayatepek, Uta Drebber, Eugen Domann, Stefan Olejniczak, Markus Weigel, Torsten Hain, Timo Rath, and et al. 2020. "IL-13 as Target to Reduce Cholestasis and Dysbiosis in Abcb4 Knockout Mice" Cells 9, no. 9: 1949. https://doi.org/10.3390/cells9091949






