1. Introduction
The brain is one of the most complex and delicate systems in nature, and grasping how the brain works is a great challenge for humans to understand nature and themselves. To fully understand the activity of nerve cells and the structure of brain networks, it is often necessary to accurately detect, locate, and quantitatively analyze the number of cells [
1], molecular expression, and neuronal activity in different brain regions of experimental animals. We can do this by analyzing labeled neurons in brain slices. The ARA [
2,
3] contains two types of atlases: the Allen Digital Atlas (ADA) and the Average Template Atlas (ATA). ADA provides detailed localization of each brain region of the mouse, while the ATA is closer to the morphological characteristics of a mouse brain slice. Owing to the differences in the individual size of animals and the distortion, deformation, and tearing of the brain slice in the actual production process, and the modal differences between mouse brain slices and the ARA, it is difficult for the actual mouse brain slice to correspond directly to the ARA. Now, the commonly used mouse brain slice regional localization methods are as follows:
(1) By referring to the ARA, the structure of different brain regions can be manually identified on the brain slice image, and the contour of each brain region can be manually drawn. This method requires not only expert experience but also good painting skills. The workload is huge, and the method is prone to deviation, so only a few samples can be made. (2) Using Photoshop and ImageJ image processing software, the contours of the ARA are extracted and corrected to realize a simple semi-automatic region division of brain slice images. This method also requires expert experience and proficiency in the use of two kinds of software, so it is also impossible to locate brain slice regions rapidly. (3) The multi-modal image registration method is used in computer vision to align the ARA with the actual mouse brain slice. To ensure that the morphology of the labeled neurons in each brain region on the brain slice does not change, the brain slice is taken as the Fixed image (F), and the ARA is taken as the Moving image (M) for registration. Then, the registered ARA outline and brain slice images are fused to complete regional localization. This method is favored by researchers because of its simple and rapid characteristics. For this reason, multi-modal image registration methodology has become an effective method to solve the problem of regional localization of brain slices.
The common idea of multi-modal image registration [
4,
5] is to use joint entropy [
6] or mutual information [
7,
8,
9] as the measure of the local regional similarity of the image and then to perform image deformation for registration. However, the effect is not satisfactory under the conditions of large modal difference, deformation, and noise interference. To better solve the problem of multi-modal image registration, some scholars have proposed a new idea: convert the multi-modal image into a monomodal image through a certain mapping and then achieve accurate multi-modal image registration through the similarity measure of the monomodal registration.
The methods of converting multi-modal images into monomodal images mainly include two types. The first type converts one modal image in the multi-modal image into another modality to obtain two unified modal images. For example, Roche et al. [
10] converted the MR image into a grayscale image similar to Ultrasound (US) based on the Correlation Ratio (CR). Wein et al. [
11] established a reflection model, which converts the CT image simulation into a US image and realizes CT-to-US multi-modal image registration. However, there are still big problems with these kinds of methods. For example, the converted images are only roughly similar, but there may still be large errors after registration, and the model is cumbersome and computationally expensive.
The second type maps the multi-modal image to a common intermediate modality. For example, Wachinger et al. [
12] proposed two registration methods based on image structure representation. The first uses the entropy value to represent the structural features of the image, and the second is based on the structural representation of the manifold learning application. Heinrich et al. [
13] proposed a registration method based on a modal-independent neighborhood descriptor. Yang Feng et al. [
14] proposed combining Weber Local Descriptors (WLD) [
15] with Normalized Mutual Information (NMI) [
16] to realize two-stage image registration. Moreover, Zhu et al. [
17] proposed a structure characterization method based on PCANet, which can extract multilevel features of images and conduct feature fusion and represents the common features of multi-modal images as unified modal images, achieving better performance than similar methods.
Most of the above modal transformation methods based on image representation are tested with T1-weighted, T2-weighted, and PD-weighted MR image data sets as the standard. Although the MR image data set is an image in three different modalities, its key features have a relatively simple correspondence and minimal noise interference. The modal transformation result of the ZMLD method with better performance is shown in
Figure 1a. It can be seen from the figure that the modal transformation method can accurately extract the features of the three MR images and represent them to a unified modal. For the mouse brain slices (upper, left), namely the ATA (upper, middle) and ADA (upper, right) employed in this study, the specific morphological features are shown in the first row of
Figure 1b, and there was a great difference in the corresponding key features between the original brain slices and the two kinds of the ARA. After the grayscale and affine transformation image preprocessing, the modalities between the grayscale brain slice (bottom, left) and ATA (bottom, middle) were closer, and they were selected for registration to use our approach without any processing. The results are shown in the second row and the three columns of
Figure 1b. Regions 1, 2, and 3 in the brain slice images correspond to the ATA images, but the corresponding relationship of color features was very complex. Because of the inconsistency of this corresponding relationship and the difficulty of identifying boundary information, there is no unified standard for algorithm learning. Using the two non-unified modal images for training, the error correspondence was established, as shown in the second line (right) of
Figure 1b. Moreover, the registration results were seriously affected by noise interference in the labeled region 4 in the brain slice. Owing to the differences in non-unified modality, more complex feature correspondence, and noise interference between the brain slice and ATA, conventional direct multi-modal image registration algorithms and existing image registration algorithms based on modal transformation made it difficult to realize the ideal regional localization.
Inspired by the fact that human experts automatically ignore the influence of noise when dealing with the regional localization of brain slices, the regional localization of each characteristic part on the brain slices is carried out by referring to the ARA. We proposed a novel idea of building correspondence based on key features. The corresponding key feature maps can be extracted from the brain slice and the ARA at the same time, and they can be converted into unified modal images. Then, the modal transformation problem can be transformed into an image segmentation problem. Since Long [
18] and others first used Fully Convolutional Networks (FCNs) for end-to-end segmentation of natural images, image segmentation has made a breakthrough. SegNet [
19], U-Net [
20], and FC-DenseNet [
21] have been proposed, although subsequent methods have achieved better segmentation results, and these methods are based on encoder–decoder network architecture. Given the satisfactory results achieved by the encoder–decoder architecture in the field of image segmentation, we proposed a new deep learning network architecture JEMI, and through the joint enhancement of multi-modal information, the modal transformation of brain slice and the ARA was effectively realized. Furthermore, the effect of image registration was improved by combining the diffeomorphic registration network based on unsupervised learning. Finally, with reference to the ARA, the automatic regional localization of brain slice images was completed. The brain slice image region localization framework proposed in this study is shown in
Figure 2, but this method can be extended to any non-unified multi-modal image registration task.
4. Experiments and Result Analysis
4.1. Software and Hardware Environment
Software environment: The experiment was built on the PyCharm platform, and the implementation of the Keras deep learning framework was based on Tensorflow. The language version used was Python 3.6.4.
Hardware environment: The experiment was run on the Windows 10 operating system. The CPU was Intel Core-i7 (3.20 GHz), the memory was 64 GB, the graphics card was NVIDIA GeForce GTX 1080Ti, and the video memory was 11 GB.
4.2. Data Set Production and Training
In the experiment, immunohistochemical brain slices of a mouse with confocal microscopy imaging and the second edition of the ARA were used to establish data sets. First, a group of mouse brain slices was selected, and then the ADA and ATA of the corresponding mouse brain slices were found in the ARA data set, and the brain slice images were pre-processed (through denoising, for example) and adjusted to the same size as the ARA. As the image was still too large for the algorithm to run, all three images were compressed to a smaller size (512 × 512) of the same size for fast processing. Then, the grayscale features of ATA, which were more similar to brain slices, were extracted and roughly registered by affine transformation so that the morphological features of the ATA were closer to brain slices. Finally, the brain slice and Brain Atlas were sent to the JEMI network for modal transformation. The key feature segmentation data sets of the brain slice, , and the ATA, , were then generated and sent to the fine registration network for training to obtain the optimal deformation field. A total of 64 groups of images were used for training, 18 groups of images were used for verification, and 18 groups of images were used for testing.
4.3. Experimental Results and Analysis
To verify the effectiveness of the method proposed in this article, some comparative experiments were carried out for each step, and the performance of each method was evaluated from different aspects. In the rough registration stage, the Run Time (RT), Normalized Correlation Coefficient (NCC), and Normalized Mutual Information (NMI) were used to evaluate the performance of different methods. The strategy was calculated according to (
11) and (
12):
where
I and
J represent two images, and
p represents the pixels in the two images.
NCC is one of the most widely used image similarity measures, which can reflect the degree of image correlation. Its value range was between [−1, 1]. The closer the correlation coefficient to 1, the more similar the two images.
where
is the information entropy of the image, and its calculation formula is
;
is the probability of gray value
i, calculated according to the formula
;
represents the total number of pixels with the gray value
i in the image; and
N represents the number of gray levels of the image. Moreover,
is the joint information entropy of the two images, and the calculation formula is
. NMI is currently one of the indicators commonly used in image registration, which can indicate the degree of mutual inclusion of two image information. The larger the normalized mutual information, the better the registration effect between the two images.
In the modal conversion stage, the Pixel Accuracy (PA), the Mean Pixel Accuracy (MPA), the Mean Intersection over Union (MIoU), and the Frequency Weighted Intersection over Union (FWIoU) were used to evaluate the performance of different methods. The evaluation strategy was calculated according to (
13)–(
16):
We supposed there were classes, including a background class, where represents the number of pixels predicted correctly for all classes, represents the number of pixels that belong to class i but are predicted to be class j, and and are called false positives and false negatives, respectively. PA is the ratio of correctly marked pixels to the total pixels. MPA calculates the ratio of correctly classified pixels in each class and then calculates the average of all classes. MIoU is a standard metric for semantic segmentation, which calculates the sum of the intersection of the true value and the predicted value. FWIoU is an improvement of MIoU, and weights were set for each class according to the frequency of appearance to judge the accuracy of the prediction.
In the fine registration stage, image fusion and the aforementioned RT, NCC and NMI were used for performance evaluation.
4.3.1. Rough Registration
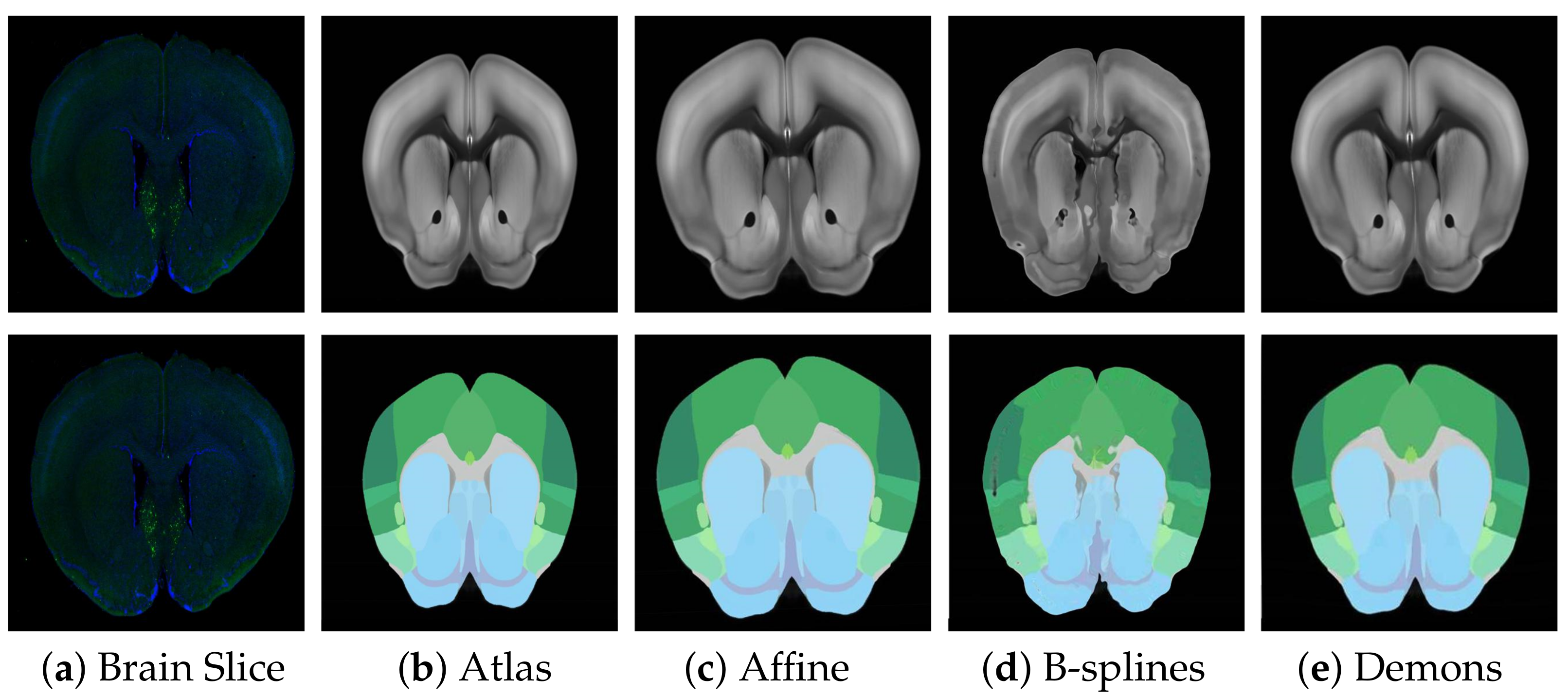
Before fine registration, preliminary rough registration of brain slices and standard brain atlas can effectively reduce the difficulty of registration and improve the effect of fine registration. Therefore, choosing a suitable rough registration method is vital. In this study, affine transform, traditional B-splines [
31], and Demons [
32] were selected for testing, and the experimental results are shown in
Figure 6. The performance comparison results using RT, NCC, and NMI measurement strategies are shown in
Table 1. Obviously, affine transformation was the most suitable rough registration method because of its faster speed and better effect.
4.3.2. Modal Transformation
To better solve the problem of non-unified multi-modal image registration, using the modal transformation method to convert a multi-modal image into a monomodal image is an effective method. In this study, a Joint Enhancement of Multimodal Information image segmentation method was proposed. It segments the corresponding key features in the image to complete the modal transformation. To verify the performance of the proposed method, the U-Net network suitable for small samples was selected for comparison, and the results are shown in
Figure 7. The commonly used evaluation indicators of PA, MPA, MIOU and FWIOU for image segmentation were used. The results were evaluated, and the performance analysis is shown in
Figure 8.
The experimental results indicate that the U-Net network can accurately segment the ATA images with obvious features and less noise. As for some brain slice images with no obvious features and a lot of noise, they could not recognize the key features or achieve satisfactory segmentation results. The JEMI network proposed in this study trains two multi-modal images together, and, by fusing the input multi-modal information, it can both promote and enhance the learning of key features and also restrict the influence of marker neurons to achieve a better effect of modal transformation ultimately.
4.3.3. Fine Registration
To verify the performance of the modal transformation method proposed in this study and the performance of the diffeomorphic registration method, three comparative experiments were carried out.
- (1)
Modal transformation effects
To verify the impact of modal transformation on registration results, the data set was registered once in the absence of modal transformation and then again with the same registration method with other parameters unchanged. The experimental registration results are shown in
Figure 9.
The experimental results indicate that modal transformation plays an important role in the registration results. Without modal transformation, the corresponding features between the brain slices and Brain Atlas were both of the mutual reference and non-reference types. Some were easy to identify, and some were difficult to distinguish from the surrounding environment, and there might be noises such as labeled neurons and staining on the brain slices. The registration results are shown in
Figure 9c. The registration algorithm could not accurately identify the complex corresponding relationship between brain slices and the Brain Atlas, pull and deform the internal features, or retain the topological structure of the original image. The image noise also interfered with the results. After the modal transformation, the registration algorithm could effectively deform the Brain Atlas Image features and align with the brain slice, overcoming the complex feature correspondence and image noise, and achieving a better registration effect.
- (2)
Different modal transformation methods
To further verify the superiority of the JEMI method, we compared the PCA modal transformation method with the JEMI modal transformation method. The PCA modal transformation method re-characterizes the extracted feature information by extracting multi-level features of the image and performing feature fusion, which can convert images of different modalities into similar modalities, reducing the difference between modalities. Two different modal transformation images were sent into the registration network for registration, and the results are shown in
Figure 10.
The experimental results indicate that different modal transformation methods had a great influence on the registration results. Although the PCA modal transformation method represents the images of different modalities as new images with a similar modality, which reduced the effects of modal differences and labeled neuron noise, it also blurred many features, making it difficult to accurately identify feature location and causing a local blockage. It was better than the registration result without modal transformation, but it was still far from achieving the ideal effect. The JEMI method can completely transform the images of different modalities into a unified one, the feature correspondence is simple, and there is no noise interference. Thus, it could effectively identify the corresponding features and carry out the deformation with diffeomorphism. Without changing the topological structure of the image, the fine registration of brain slices and the standard Brain Atlas was realized, and the regional localization could be completed accurately.
- (3)
Different registration methods
To verify the superiority of the registration algorithm in this study, the traditional B-splines registration method, Demons registration method, and Reg-Net Registration method using Deep Learning were compared in the experiment. The fusion results of the brain slice and Brain Atlas after registration of each method are shown in
Figure 11.
According to the fusion effect of images, the results of the regional localization method in this study were noticeably better than others. The external edge and internal feature contour fit well with the brain slice images, and the original topological structure was left unchanged. The regional localization effect of some brain slices is shown in
Figure 12.
To quantify the performance of different methods, RT, NCC, and NMI were used to compare the brain slice and the deformed ATA. Using 18 groups of images, we carry out calculation of the above three indicators and compare them with average values, as shown in
Table 2.
The experimental results indicate that the performance of the proposed method is better than that of existing methods. Compared with the PCA + REG-NET method, the RT of the registration results of this method was reduced by 66.66%, the NCC and NMI were increased by 3.20% and 0.16%, respectively, and excellent performance was achieved in each evaluation index. The experimental results indicate that this method can carry out unsupervised diffeomorphic registration for brain slice images with non-unified modality and can complete the localization of mouse brain slices accurately and quickly.