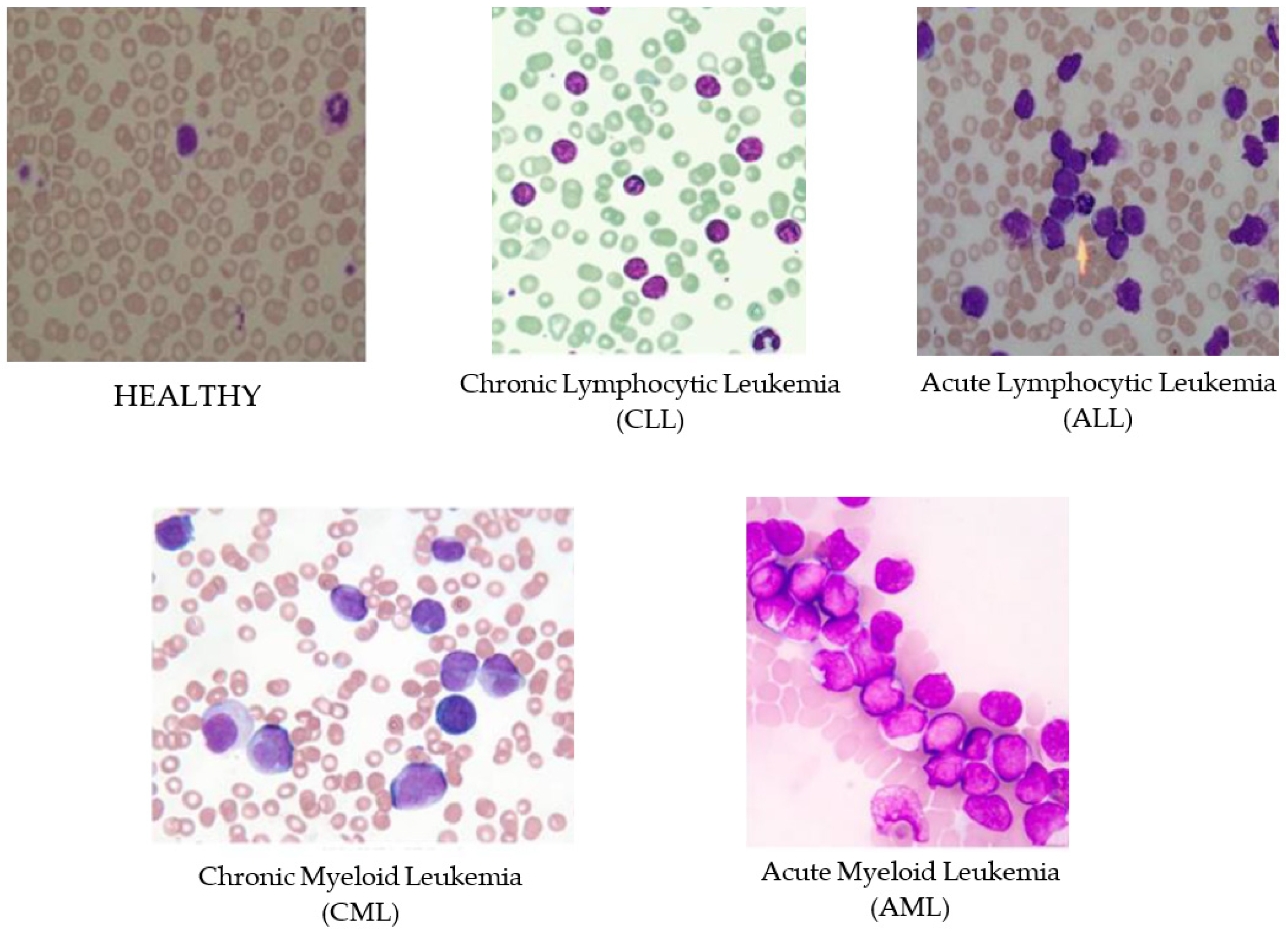
1. Introduction
Leukemia is an aggressive disease related to the white blood cells (WBC) and affects the bone marrow and blood of the human body. This disease can lead to destroying the immune system of the human body. There are two main types of leukemia, acute and chronic leukemia, which is depending on how fast it progresses. In acute leukemia, infected WBC do not perform or act like the normal WBC; while it can act as normal WBC in chronic leukemia. Therefore, chronic leukemia can be severe since it cannot be differentiated from normal WBC. Moreover, there are two subtypes of each leukemia types depending on the size and the shape of the WBC: Lymphoid and myeloid. In general, there are four subtypes of leukemia as shown in
Figure 1, Acute Lymphocytic Leukemia (ALL), Acute Myeloid Leukemia (AML), Chronic Lymphocytic Leukemia (CLL), and Chronic Myeloid Leukemia (CML) [
1,
2]. Identifying the existence of leukemia as well as their specific types is important for hematologists to avoid medical risks and specify the right therapy. Thus, using intelligent ways for diagnosis will facilitate and speed up the discovery of leukemia subtypes using the blood cells images (i.e., blood smears).
Microscopic blood tests are considered as the main procedure for the diagnosis of leukemia [
2]. Analysis of blood smears is the most common way of discovering leukemia, but not the only one technique. Interventional radiology is an alternative technique for the diagnosis of leukemia. However, radiological techniques, such as percutaneous aspiration, biopsy, and catheter drainage, suffer from inheriting limitations of imaging modality sensitivity and resolution of the radio images [
3]. Moreover, other techniques, such as Molecular Cytogenetics, Long Distance Inverse Polymerase Chain Reaction (LDI-PCR), and Array-based Comparative Genomic Hybridization (aCGH), need extensive work and time to identify leukemia types [
4]. Due to time and cost requirements of these techniques, microscopic blood tests and bone marrow are the most common methods for identification of leukemia subtypes.
A machine learning (ML) algorithm will help to identify the blood cells with leukemia from the HEALTHY cells when a large training set is available. ALL-IDB leukemia image repository [
5] is one of the datasets that is used by a number of medical researchers [
1,
6,
7] as a benchmark. Another leukemia dataset is from the American Society of Hematology (ASH) and it is online at website [
8]. Thanh et al. [
7] used the ASH database for identifying AML leukemia in their research. Google is another source of leukemia image without annotations, where images were collected randomly from websites. Karthikeyan et al. [
9] used the microscopic images collected from Google in their research for identifying leukemia, where authors annotated images by themselves. Successful implementation of machine learning based leukemia diagnosis can be built upon the use of an annotated image dataset.
Identification of leukemia subtypes from the HEALTHY samples is quite a challenging issue. In the literature, many researchers studied only binary classification between one subtype versus HEALTHY samples [
1,
7,
9,
10,
11,
12]. They obtained quite a high performance even more than 96% accuracy. Besides, Shafique et al. [
6] further classified samples with ALL subtypes, according to the size of the cell and the nature of its nucleus. However, dealing with the identification of all subtypes of leukemia is more challenging task than simple binary classification [
13]. To the best of our knowledge, there is no automatic recognition approach dealing with all subtypes of leukemia.
Several ML algorithms help to classify and recognize leukemia disease from microscopic images. For example, Paswan et al. [
10] who used support vector machine (SVM) and k-nearest neighbor (k-NN) to classify AML leukemia subtype, obtained 83% accuracy. Patel et al. [
1] applied SVM for classifying ALL leukemia subtype and achieved 93% accuracy. Karthikeyan et al. [
9] also used SVM and c-mean clustering technique to separate WBC from the background and they reached 90% accuracy. Although using a deep learning (DL) approach seems more effective, its performance highly depends on the quantity and quality of dataset used [
6]. Convolutional Neural Network (CNN) is one of the neural networks architecture that is generally used to deal with image classification and registration problems. Shafique et al. [
6] applied a convolutional neural network (CNN) for identifying ALL leukemia subtypes. Their results recorded 99% for binary classification, between ALL versus HEALTHY samples, and 96% for the further classification of subtypes of ALL only. Thanh et al. [
7] also built a CNN model that consists of five convolutional layers to make a binary classification of ALL leukemia subtypes and obtained 96.6% accuracy. Unfortunately, the performance classification in this kind of neural network needs a large training data to learn how to identify important objects from the whole image. However, developing a large training dataset is very time consuming and is a very labor-intensive task. To avoid this problem, we suggest expanding the limited number of samples by image augmentation. Using an inadequate number of image samples in the training dataset may lead to an overfitting problem [
14]. Hence, most of the researchers in the literature depend on applying some image transformation techniques to increase the number of training set samples synthetically to avoid an overfitting problem. Patel et al. [
1] applied median and Wiener filters to remove noise and blurriness. In the literature, many of the image transformation techniques have been used such as image rotation and mirroring, histogram equalization, image translation, gray scale transformation, blurring images, and image shearing [
6,
9,
10]. Using image augmentation makes it possible to use a DL approach, which requires a large number in the training dataset.
In this study, we propose a new approach for leukemia diagnosis from microscopic blood images identifying the four subtypes of leukemia (i.e., ALL, AML, CLL, and CML) by using CNN architecture of deep learning. To the best of our knowledge, this is the first study dealing with all four leukemia subtypes. Furthermore, the deep neural network requires large training datasets, which is not a trivial problem in our case. Thus, we used an image augmentation technique to increase the number of samples. Additionally, we compared our approach with well-known ML algorithms and evaluated their performances with 5-fold cross validation.
The rest of the paper is organized as follows:
Section 2 represents the dataset we used and our method in detail.
Section 3 presents the experimentation setups. Results and discussions are given in
Section 4. Finally,
Section 5 concludes the paper and provides a look at further studies on this topic.
4. Results and Discussion
Table 2 shows the general summary of the results we obtained from each experiment. We obtained the best performance in the binary classification of ALL and HEALTHY classes using augmented samples. The next best score is the multi-classification of the four leukemia subtypes. However, binary classification outperforms the multi-classification problem, since the more classes adhered and involved in a classification process the more complexity will be recognized by the model to differentiate between the classes. We observed that SGD optimizer works much better than ADAM optimizer in terms of both accuracy and loss metrics. Besides, we also observed that longer epoch iterations do not contribute to the performance of the model since
Exp#3a produced a better result than
Exp#3b.
Table 3 and
Table 4 present the results of each fold’s performance for binary and multi-classification, respectively. In binary classification, the best score was achieved in fold 4. In the multi-classification, the best performance was obtained in fold 3 in terms of validation accuracy and loss. We can conclude that the selection of samples for test and training set may fluctuate the performance results significantly. Multiple fold cross validation leads to more reliable results in evaluation, thus that we evaluated our approach using 5-fold cross-validation.
As known, a deep neural network requires large training datasets, since small training set may cause overfitting problem. To overcome this issue, we used data augmentation. The more training samples, the more CNN capability of discovering the patterns which each leukemia type may hold. Although data augmentation plays an important role to increase the sample amounts and to avoid overfitting, they might add some extra noise to the original data. This requires fine-tuning of dropout and batch-normalization parameters; nevertheless, it may not be able to solve problem itself. Besides, it could happen due to very similar structure of blood cells. Performance of multi-classification dropped into 81.74%. This result explains the performance drop and how adding noise with data augmentation degrades the identification of patterns of different leukemia subtypes.
A Comparison with Other Studies and Methods
We performed a general comparison between our model and the other studies in the literature. Additionally, we compared commonly-used machine learning algorithms SVM, NB, DT, and
k-NN (
k = 3) algorithms for the binary and multi-classification as explained in Exp#4. We compared different studies Shafique et al. [
6] and Thanh et al. [
7] in terms of average accuracy, using ALL-DB dataset for the binary classification of leukemia samples.
Table 5 shows a comparison of the binary classification between ALL and HEALTHY samples for two literature studies versus our proposals, in terms of average accuracy. Our model in CNN has the third best score, which is 88.25% in accuracy. Studies by Shafique et al. [
6] and Thanh et al. [
7] outperformed our CNN model in the binary classification problem. However, it should be taken into consideration that they did not use any cross validation in evaluation, it is unknown how test and training sets were selected. When we considered the best individual folds as the final score, the CNN model may be considered to achieve 99.3% accuracy, in fold 3. This result is better than reference [
7] who obtained 96.6% accuracy for the binary classification of leukemia.
The results of several models are given in
Table 6 for the multi-classification problem. Our model obtained the second-best score with 81.74% average accuracy after the study of Shafique et al. [
6] who made a multi-classification of only ALL subtypes, not all subtypes of leukemia. To the best of our knowledge, our study is the first one which explores multi-classification of the four leukemia subtypes. According to
Table 4, we get 98.47% accuracy with fold 4, which ranks our model in the top of the multi-classification results.
Furthermore, our model using SGD optimizer generally outperforms all classical ML classifiers especially NB, which provided the highest score in comparison with DT, SVM, and
k-NN. Moreover, SGD outperforms the ADAM optimizer in our CNN model for the multi-classification. We also noticed that working with 25 epochs performs better than longer epoch periods.
Figure 4 shows a general comparison of all the multi-classification models discussed in this section. Our model with CNN produced 81.74% for multi-classification on average with 5-fold cross validation. The error bar in
Figure 4 indicates the minimum and the maximum values of observations in terms of accuracy. Our model produces competing results when the best fold has been taken into consideration.