Mass Spectrometry-Based Proteomics for Classification and Treatment Optimisation of Triple Negative Breast Cancer
Abstract
:1. Introduction
2. Current Clinical Approach of Triple-Negative Breast Cancer
2.1. Breast Cancer Classification
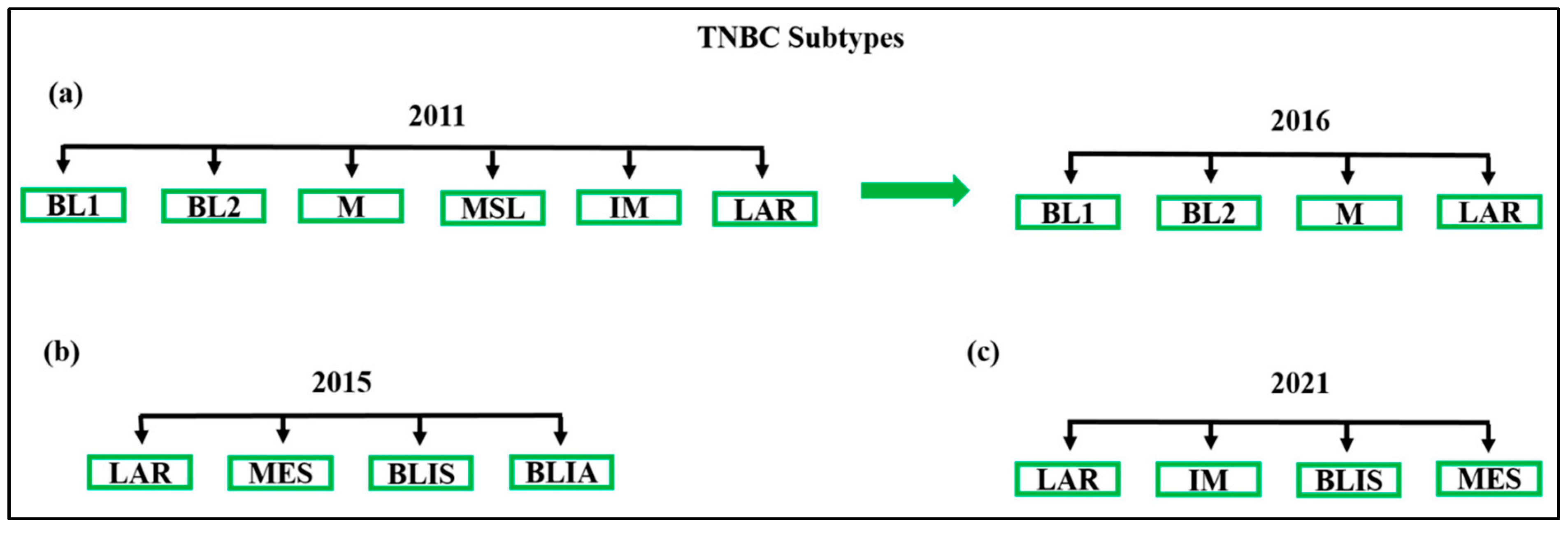
2.2. Subclassification of TNBC: Molecular Insights
2.3. Diagnosis, Staging, and Current Treatment of TNBC
2.4. Need for New Treatment Approaches
2.4.1. Neoadjuvant Chemotherapy versus Adjuvant Chemotherapy for TNBC Patients
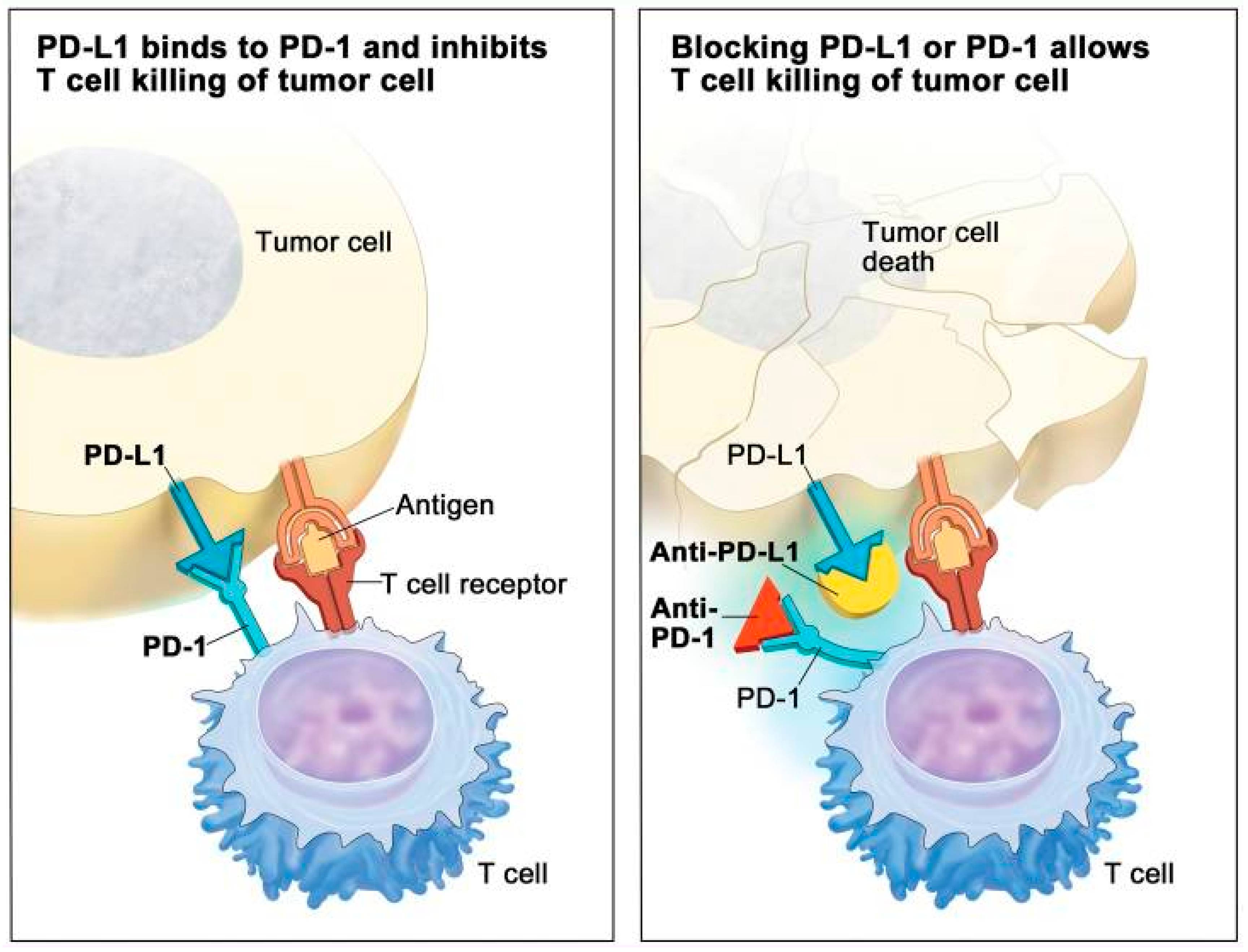
2.4.2. Immunotherapy
2.4.3. Local Therapy: Surgery and Radiation
3. Role of Biomarkers in TNBC
Importance of Predictive Biomarkers
4. Mass Spectrometry-Based Proteomics in TNBC
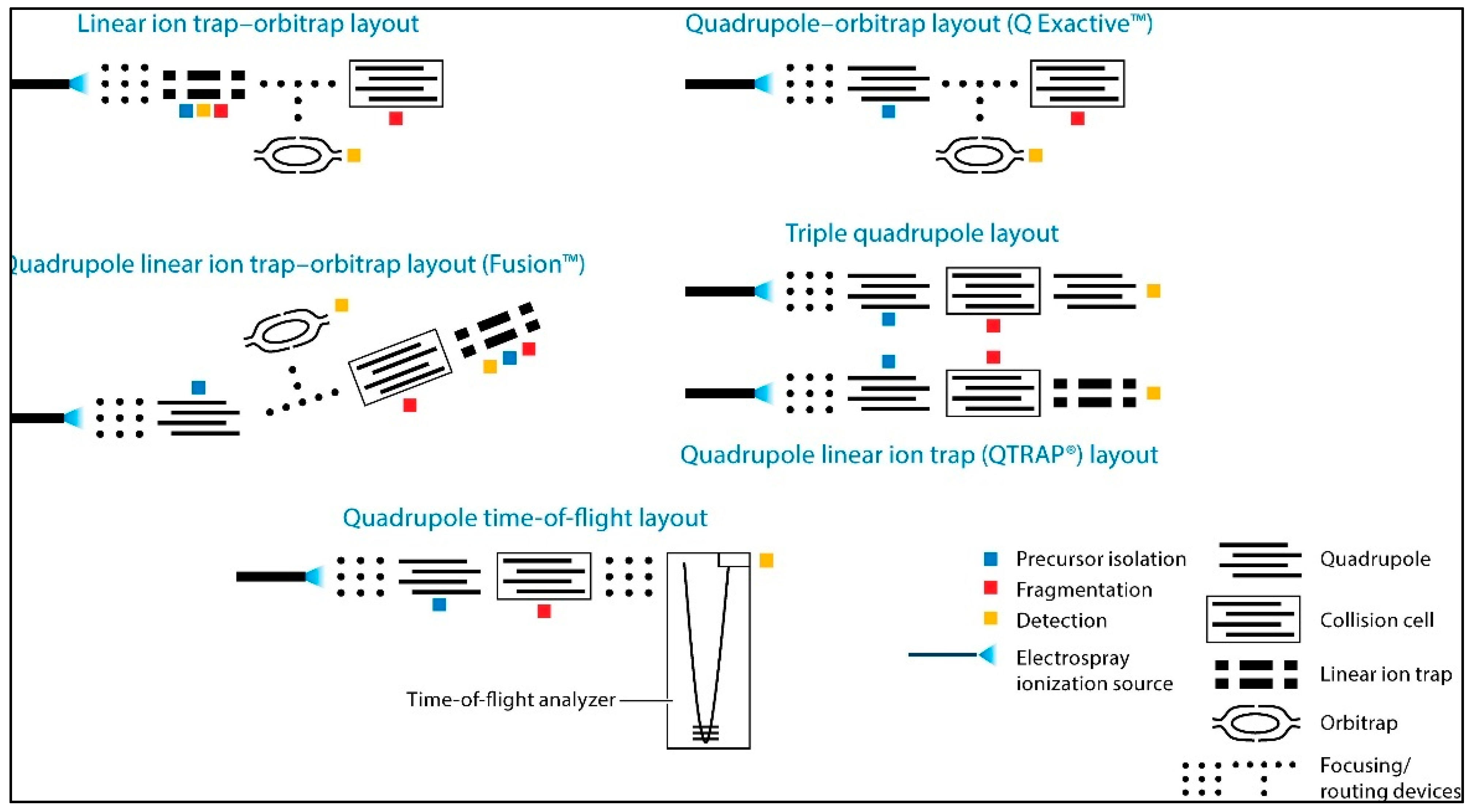
4.1. Technology Overview
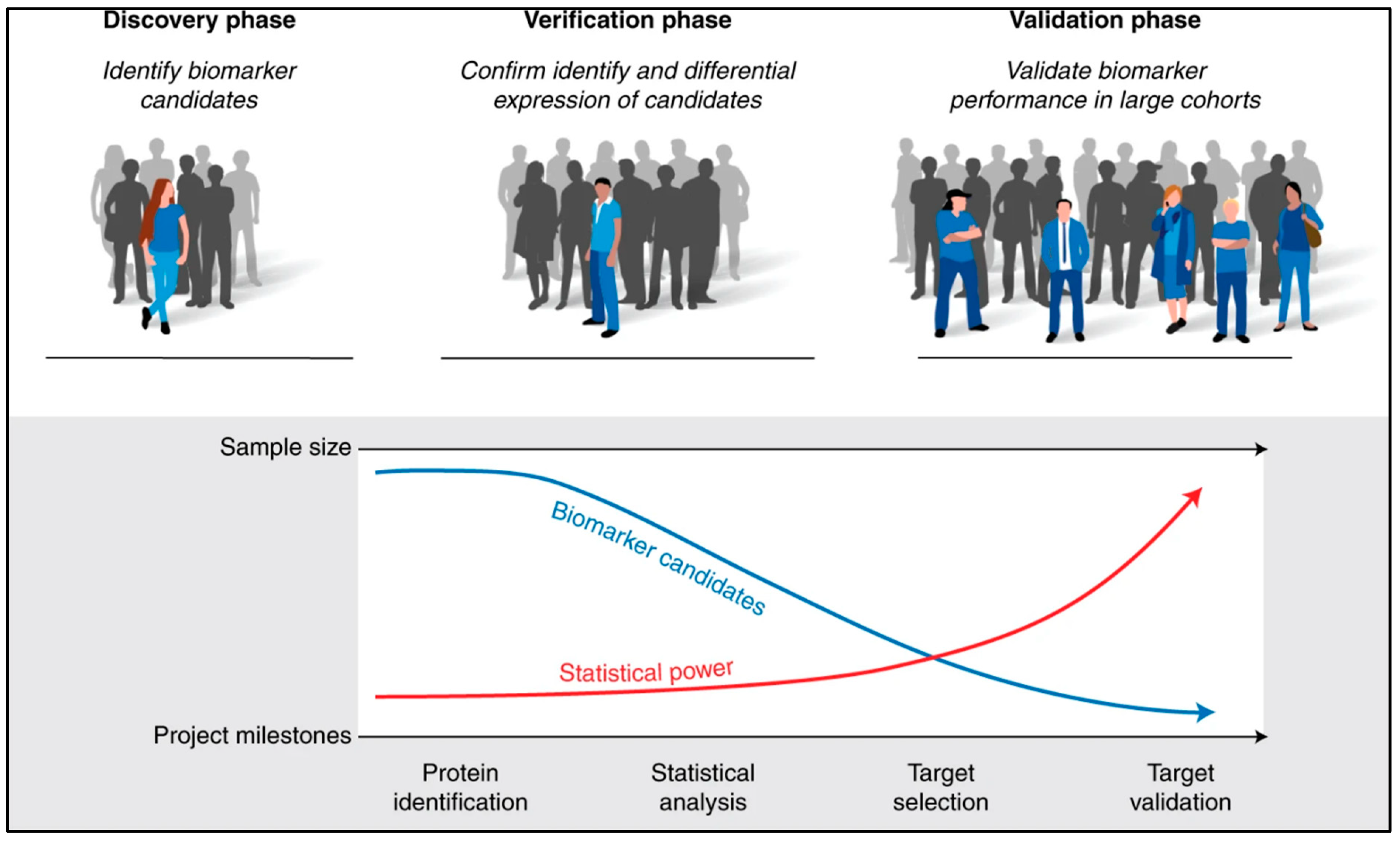
4.2. Identification of Protein Biomarkers in TNBC
4.3. Current Gaps in Biomarker Discovery
5. Clinical Implications
5.1. Advances in Clinical Practices through Biomarker Research and Proteomics
5.2. The Advantages and Disadvantages of MS-Based Proteomics in TNBC
5.2.1. Advantages of MS-Based Proteomics in Clinical Applications
- Comprehensive protein profiling: MS-proteomics allows for the identification and quantification of a wide range of proteins in TNBC tissues. This extensive profiling can uncover novel biomarkers for cancer diagnosis, prediction, and prognosis, which are increasingly sought to enable early cancer detection and to tailor treatment decisions [102].
- Post-translational modifications: Protein PTMs significantly impact protein functions and are vital to nearly all cellular processes. PTMs and their interactions are closely associated with key signalling events that drive cancer development, progression, and metastasis. They play crucial roles in cancer hallmark functions, cancer metabolism, and the regulation of the tumour microenvironment [103]. As a result, by studying PTMs through MS-proteomics, researchers can uncover specific modifications that are associated with TNBC, providing insights into how these changes drive the disease. This knowledge can lead to the identification of novel therapeutic targets and the development of drugs that specifically inhibit or modify these PTM-related pathways. Additionally, PTMs can serve as biomarkers for disease progression and treatment response, further enhancing the ability to tailor treatments to individual patients [66].
- High sensitivity and specificity: MS-proteomics provides high sensitivity and specificity, making it a powerful tool for detecting low-abundance proteins. This capability is especially critical in the context of TNBC, where key proteins involved in signalling pathways, tumour suppression, and drug resistance may be present in very low quantities. The ability to identify these proteins can lead to the discovery of novel biomarkers that could be crucial for early detection and personalised treatment strategies [104]. Moreover, MS-proteomics helps in distinguishing between closely related protein isoforms, which is important for understanding the variations in protein structure and function that can influence TNBC progression and treatment response. Protein isoforms can arise from alternative splicing, PTMs, or genetic mutations, and each isoform may play a distinct role in the disease. By accurately identifying and quantifying these isoforms, researchers can gain deeper insights into the molecular heterogeneity of TNBC, leading to more targeted and effective therapeutic approaches [105].
- Integration with other omics data: MS-proteomics data can be integrated with genomics and transcriptomics data to offer a more comprehensive understanding of TNBC biology. This integrative approach helps to elucidate the complex interactions between proteins, genes, and RNA transcripts, providing a deeper understanding of the disease’s molecular mechanisms. For instance, the Human Protein Atlas (HPA) serves as a valuable resource, offering detailed information on the localisation and temporal expression of human proteins across various tissues and cancers. It also provides insights into the availability and quality of antibodies, which can be cross-referenced with genomic and transcriptomic data to link protein behaviour with gene expression patterns [106]. Similarly, the Clinical Proteomic Tumour Analysis Consortium (CPTAC) of the National Cancer Institute (NCI) has made significant contributions by publishing comprehensive multi-omics studies, including MRM (Multiple Reaction Monitoring) assay databases for several cancer types, such as breast [23] and ovarian cancers [107]. The data available through the CPTAC Data Portal include protein sequence databases derived directly from the exome sequences of respective cancer samples, facilitating the integration of proteomic data with genetic information [108].
- Potential for personalised medicine: The advent of personalised medicine offers significant promise for those affected by this challenging disease, as distinct, potentially druggable molecular targets with unique alterations have been identified. Developing treatment strategies for a broad range of TNBC patients requires a thorough understanding of the disease’s underlying mechanisms. Achieving this understanding involves examining and integrating data on TNBC subtypes, focussing on their epigenetic, transcriptomic, proteomic, and phospho-proteomic profiles. For instance, a study has analysed the BRCA1-wild-type MDA-MB-231 TNBC cell line, the BRCA15382insC HCC1937 TNBC cell line, and the MCF10A cell line as a normal breast epithelial control. This multi-omics approach underscores the diversity among different TNBC subtypes and enhances the understanding of the molecular pathways that drive this complex form of BC [102].
- Multiplexing capabilities: MS-based assays can analyse multiple biomarkers simultaneously [109], which is highly efficient and cost-effective compared to other techniques such as enzyme-linked immunosorbent assay (ELISA), which typically measures one biomarker at a time.
- Application to various sample types: Advancements in mass spectrometry technologies, along with enhanced sample preparation techniques, have significantly improved our understanding of the biological complexity across a diverse range of sample types. This includes various organelles, membranes, biofluids (such as blood, cerebrospinal fluid, saliva, and urine), tissues, organs, and microbial communities [110].
5.2.2. Disadvantages of MS-Based Proteomics in Clinical Applications
- Complexity and cost: MS-based proteomics relies on advanced and costly equipment, including high-resolution mass spectrometers. These devices require frequent maintenance and calibration to maintain their precision and dependability. Operating and maintaining such instruments demands specialised training and expertise. Additionally, methods for absolute quantification, such as TMT and iTRAQ, used in TNBC biomarker studies, contribute to further expenses [111].
- Sample preparation challenges: Sample preparation plays a critical role in the proteomic characterisation of clinical samples, and it is essential to establish rigorous standard operating procedures to obtain relevant information about the complex biological processes underlying cancer progression. There is no universal protocol for proteomic sample preparation, as the chosen strategy should be optimised based on factors such as proteomic complexity, available sample quantity, and the study’s goals. Variations in sample handling, storage, and processing can affect the reproducibility and reliability of results [58].
- Data analysis complexity: MS generates vast quantities of data that necessitate the use of advanced analytical techniques and bioinformatics tools for meaningful interpretation. The initial step in data analysis involves the accurate identification and quantification of proteins, a process that relies on advanced algorithms and specialised software such as MaxQuant (version number v2.6.3.0) [112]. This includes the extraction of peptide sequences, matching them against protein databases, and quantifying their abundance. Furthermore, the reliability of results is contingent upon the ac-curacy of the analytical methods used. Limitations in data analysis can significantly impact the interpretation of results, potentially leading to misleading conclusions [113].
- Limited standardisation: Despite significant advancements in technology development, standardisation, and bioinformatics that have enhanced the reliable identification of molecular disease signatures, several major obstacles continue to hinder the effective translation of protein candidates into clinical biomarkers. As a result, only a limited number of biomarkers have received FDA approval in the past two decades [114].
- Validation and translation: The validation of protein biomarkers identified through MS requires extensive testing across different TNBC patient cohorts to ensure that the biomarkers are consistent, reliable, and reflective of the disease state. This process is complicated by the heterogeneity of TNBC, where the variability between tumours can lead to inconsistent biomarker performance [114]. Even when a protein biomarker is validated, translating it into a clinical setting involves overcoming several hurdles. Regulatory approval processes, such as those required by the US Food and Drug Administration (FDA), demand rigorous evidence of clinical utility and cost-effectiveness. Additionally, developing standardised and scalable assays for routine clinical use can be technically challenging and resource-intensive. As a result, despite the identification of promising biomarkers through MS-based proteomics, only a few have successfully made the transition to clinical practice in TNBC. Most FDA-approved tumour markers are blood-based and are used in conjunction with standard imaging techniques to differentiate between malignant and benign conditions. However, many existing cancer screening tests suffer from insufficient sensitivity and/or specificity. As a result, the search for protein biomarkers capable of enabling early cancer diagnosis remains an ongoing effort [114].
5.3. Need for Biomarkers in Predicting the Response to Neoadjuvant Chemotherapy
6. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bianchini, G.; Balko, J.M.; Mayer, I.A.; Sanders, M.E.; Gianni, L. Triple-negative breast cancer: Challenges and opportunities of a heterogeneous disease. Nat. Rev. Clin. Oncol. 2016, 13, 674–690. [Google Scholar] [CrossRef] [PubMed]
- Nascimento, R.G.; Otoni, K.M. Histological and molecular classification of breast cancer: What do we know? Mastology 2020, 30, 1–8. [Google Scholar] [CrossRef]
- Basmadjian, R.B.; Chow, K.; Kim, D.; Kenney, M.; Lukmanji, A.; O’Sullivan, D.E.; Xu, Y.; Quan, M.L.; Cheung, W.Y.; Lupichuk, S.; et al. The Association between Early-Onset Diagnosis and Clinical Outcomes in Triple-Negative Breast Cancer: A Systematic Review and Meta-Analysis. Cancers 2023, 15, 1923. [Google Scholar] [CrossRef] [PubMed]
- Hall, P.E.; Schmid, P. Emerging strategies for TNBC with early clinical data: New chemoimmunotherapy strategies. Breast Cancer Res. Treat. 2022, 193, 21–35. [Google Scholar] [CrossRef] [PubMed]
- Bou Zerdan, M.; Ghorayeb, T.; Saliba, F.; Allam, S.; Bou Zerdan, M.; Yaghi, M.; Bilani, N.; Jaafar, R.; Nahleh, Z. Triple Negative Breast Cancer: Updates on Classification and Treatment in 2021. Cancers 2022, 14, 1253. [Google Scholar] [CrossRef]
- Zhao, S.; Zuo, W.-J.; Shao, Z.-M.; Jiang, Y.-Z. Molecular subtypes and precision treatment of triple-negative breast cancer. Ann. Transl. Med. 2020, 8, 499. [Google Scholar] [CrossRef]
- Pralea, I.E.; Moldovan, R.C.; Țigu, A.B.; Ionescu, C.; Iuga, C.A. Mass spectrometry-based omics for the characterization of triple-negative breast cancer bio-signature. J. Pers. Med. 2020, 10, 277. [Google Scholar] [CrossRef]
- Gong, S.; Wang, Q.; Huang, J.; Huang, R.; Chen, S.; Cheng, X.; Liu, L.; Dai, X.; Zhong, Y.; Fan, C.; et al. LC-MS/MS platform-based serum untargeted screening reveals the diagnostic biomarker panel and molecular mechanism of breast cancer. Methods 2024, 222, 100–111. [Google Scholar] [CrossRef]
- Lawrence, R.T.; Perez, E.M.; Hernández, D.; Miller, C.P.; Haas, K.M.; Irie, H.Y.; Lee, S.I.; Blau, C.A.; Villén, J. The Proteomic Landscape of Triple-Negative Breast Cancer. Cell Rep. 2015, 11, 630–644. [Google Scholar] [CrossRef]
- Perou, C.M.; Sørlie, T.; Eisen, M.B.; Van De Rijn, M.; Jeffrey, S.S.; Rees, C.A.; Pollack, J.R.; Ross, D.T.; Johnsen, H.; Akslen, L.A.; et al. Molecular portraits of human breast tumours. Nature 2000, 406, 747–752. [Google Scholar] [CrossRef]
- Harbeck, N.; Penault-Llorca, F.; Cortes, J.; Gnant, M.; Houssami, N.; Poortmans, P.; Ruddy, K.; Tsang, J.; Cardoso, F. Breast cancer. Nat. Rev. Dis. Prim. 2019, 5, 66. [Google Scholar] [CrossRef]
- National Clinical Effectiveness Committee. Diagnosis, Staging and Treatment of Patients with Gestational Trophoblastic Disease National Clinical Guideline No. 7; National Clinical Effectiveness Committee: Dublin, Ireland, 2015. Available online: https://www.hse.ie/eng/services/list/5/cancer/profinfo/guidelines/breast/treatment-of-patients-with-breast-cancer-radiation-oncology.pdf (accessed on 23 August 2024).
- Harbeck, N.; Gnant, M. Breast cancer. Lancet 2017, 389, 1134–1150. [Google Scholar] [CrossRef]
- Foulkes, W.D.; Smith, I.E.; Reis-Filho, J.S. Triple-Negative Breast Cancer. N. Engl. J. Med. 2010, 363, 1938–1948. [Google Scholar] [CrossRef] [PubMed]
- Lehmann, B.D.; Bauer, J.A.; Chen, X.; Sanders, M.E.; Chakravarthy, A.B.; Shyr, Y.; Pietenpol, J.A. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J. Clin. Investig. 2011, 121, 2750–2767. [Google Scholar] [CrossRef]
- Ge, J.; Zuo, W.; Chen, Y.; Shao, Z.; Yu, K. The advance of adjuvant treatment for triple-negative breast cancer. Cancer Biol. Med. 2022, 19, 188–201. [Google Scholar] [CrossRef] [PubMed]
- Garufi, G.; Palazzo, A.; Paris, I.; Orlandi, A.; Cassano, A.; Tortora, G.; Scambia, G.; Bria, E.; Carbognin, L. Neoadjuvant therapy for triple-negative breast cancer: Potential predictive biomarkers of activity and efficacy of platinum chemotherapy, PARP- and immune-checkpoint-inhibitors. Expert Opin. Pharmacother. 2020, 21, 687–699. [Google Scholar] [CrossRef] [PubMed]
- Hossain, F.; Majumder, S.; David, J.; Miele, L. Precision Medicine and Triple-Negative Breast Cancer: Current Landscape and Future Directions. Cancers 2021, 13, 3739. [Google Scholar] [CrossRef]
- Weymann, D.; Laskin, J.; Roscoe, R.; Schrader, K.A.; Chia, S.; Yip, S.; Cheung, W.Y.; Gelmon, K.A.; Karsan, A.; Renouf, D.J.; et al. The cost and cost trajectory of whole-genome analysis guiding treatment of patients with advanced cancers. Mol. Genet. Genom. Med. 2017, 5, 251–260. [Google Scholar] [CrossRef]
- Kohler, B.A.; Sherman, R.L.; Howlader, N.; Jemal, A.; Ryerson, A.B.; Henry, K.A.; Boscoe, F.P.; Cronin, K.A.; Lake, A.; Noone, A.M.; et al. Annual Report to the Nation on the Status of Cancer, 1975–2011, Featuring Incidence of Breast Cancer Subtypes by Race/Ethnicity, Poverty, and State. JNCI J. Natl. Cancer Inst. 2015, 107, djv048. [Google Scholar] [CrossRef] [PubMed]
- Heil, J.; Kuerer, H.M.; Pfob, A.; Rauch, G.; Sinn, H.P.; Golatta, M.; Liefers, G.J.; Peeters, M.V. Eliminating the breast cancer surgery paradigm after neoadjuvant systemic therapy: Current evidence and future challenges. Ann. Oncol. 2020, 31, 61–71. [Google Scholar] [CrossRef]
- Sabatier, R.; Finetti, P.; Mamessier, E.; Adelaide, J.; Chaffanet, M.; Ali, H.R.; Viens, P.; Caldas, C.; Birnbaum, D.; Bertucci, F. Prognostic and predictive value of PDL1 expression in breast cancer. Oncotarget 2015, 6, 5449–5464. [Google Scholar] [CrossRef] [PubMed]
- Mertins, P.; Mani, D.R.; Ruggles, K.V.; Gillette, M.A.; Clauser, K.R.; Wang, P.; Wang, X.; Qiao, J.W.; Cao, S.; Petralia, F.; et al. Proteogenomics connects somatic mutations to signalling in breast cancer. Nature 2016, 534, 55–62. [Google Scholar] [CrossRef] [PubMed]
- Losurdo, A.; De Sanctis, R.; Fernandes, B.; Torrisi, R.; Masci, G.; Agostinetto, E.; Gatzemeier, W.; Errico, V.; Testori, A.; Tinterri, C.; et al. Insights for the application of TILs and AR in the treatment of TNBC in routine clinical practice. Nat. Res. 2020, 10, 20100. [Google Scholar] [CrossRef]
- Armaghani, A.J.; Han, H.S. Alpelisib in the treatment of breast cancer: A short review on the emerging clinical data. Breast Cancer Targets Ther. 2020, 12, 251–258. [Google Scholar] [CrossRef]
- Saleh, L.; Wilson, C.; Holen, I. CDK4/6 inhibitors: A potential therapeutic approach for triple negative breast cancer. MedComm 2021, 2, 514–530. [Google Scholar] [CrossRef] [PubMed]
- Eikesdal, H.P.; Yndestad, S.; Elzawahry, A.; Llop-Guevara, A.; Gilje, B.; Blix, E.S.; Espelid, H.; Lundgren, S.; Geisler, J.; Vagstad, G.; et al. Olaparib monotherapy as primary treatment in unselected triple negative breast cancer. Ann. Oncol. 2021, 32, 240–249. [Google Scholar] [CrossRef] [PubMed]
- Chaudhary, L.N. Early stage triple negative breast cancer: Management and future directions. Semin. Oncol. 2020, 47, 201–208. [Google Scholar] [CrossRef]
- Sharma, P. Update on the Treatment of Early-Stage Triple-Negative Breast Cancer. Curr. Treat. Opt. Oncol. 2018, 19, 22. [Google Scholar] [CrossRef]
- Schreiber, A.R.; Kagihara, J.A.; Weiss, J.A.; Nicklawsky, A.; Gao, D.; Borges, V.F.; Kabos, P.; Diamond, J.R. Clinical Outcomes for Patients with Metastatic Breast Cancer Treated with Immunotherapy Agents in Phase I Clinical Trials. Front. Oncol. 2021, 11, 640690. [Google Scholar] [CrossRef]
- Montemurro, F.; Nuzzolese, I.; Ponzone, R. Neoadjuvant or adjuvant chemotherapy in early breast cancer? Expert. Opin. Pharmacother. 2020, 21, 1071–1082. [Google Scholar] [CrossRef]
- Dogan, I.; Aksoy, S.; Cakar, B.; Basaran, G.; Ercelep, O.; Molinas Mandel, N.; Korkmaz, T.; Gokmen, E.; Sener, C.; Aydiner, A.; et al. Demographic and Clinical Features of Patients with Metastatic Breast Cancer: A Retrospective Multicenter Registry Study of the Turkish Oncology Group. Cancers 2023, 15, 1667. [Google Scholar] [CrossRef]
- Burstein, H.J.; Curigliano, G.; Loibl, S.; Dubsky, P.; Gnant, M.; Poortmans, P.; Colleoni, M.; Denkert, C.; Piccart-Gebhart, M.; Regan, M.; et al. Estimating the benefits of therapy for early-stage breast cancer: The St. Gallen International Consensus Guidelines for the primary therapy of early breast cancer 2019. Ann. Oncol. 2019, 30, 1541–1557. [Google Scholar] [CrossRef]
- Landry, I.; Sumbly, V.; Vest, M. Advancements in the Treatment of Triple-Negative Breast Cancer: A Narrative Review of the Literature. Cureus 2022, 14, e21970. [Google Scholar] [CrossRef]
- Vuger, A.T.; Tiscoski, K.; Apolinario, T.; Cardoso, F. Anthracyclines in the treatment of early breast cancer friend or foe? Breast 2022, 65, 67–76. [Google Scholar] [CrossRef]
- Røssevold, A.H.; Andresen, N.K.; Bjerre, C.A.; Gilje, B.; Jakobsen, E.H.; Raj, S.X.; Falk, R.S.; Russnes, H.G.; Jahr, T.; Mathiesen, R.R.; et al. Atezolizumab plus anthracycline-based chemotherapy in metastatic triple-negative breast cancer: The randomized, double-blind phase 2b ALICE trial. Nat. Med. 2022, 28, 2573–2583. [Google Scholar] [CrossRef] [PubMed]
- Ortolan, E.; Appierto, V.; Silvestri, M.; Miceli, R.; Veneroni, S.; Folli, S.; Pruneri, G.; Vingiani, A.; Belfiore, A.; Cappelletti, V.; et al. Blood-based genomics of triple-negative breast cancer progression in patients treated with neoadjuvant chemotherapy. ESMO Open 2021, 6, 100086. [Google Scholar] [CrossRef]
- Howard, J.; Wyse, C.; Argyle, D.; Quinn, C.; Kelly, P.; McCann, A. Exosomes as Biomarkers of Human and Feline Mammary Tumours; A Comparative Medicine Approach to Unravelling the Aggressiveness of TNBC. Biochim. Biophys. Acta—Rev. Cancer 2020, 1874, 188431. [Google Scholar] [CrossRef]
- Cortazar, P.; Zhang, L.; Untch, M.; Mehta, K.; Costantino, J.P.; Wolmark, N.; Bonnefoi, H.; Cameron, D.; Gianni, L.; Valagussa, P.; et al. Pathological complete response and long-term clinical benefit in breast cancer: The CTNeoBC pooled analysis. Lancet 2014, 384, 164–172. [Google Scholar] [CrossRef] [PubMed]
- Curigliano, G.; Burstein, H.J.; Winer, E.P.; Gnant, M.; Dubsky, P.; Loibl, S.; Colleoni, M.; Regan, M.M.; Piccart-Gebhart, M.; Senn, H.J.; et al. De-escalating and escalating treatments for early-stage breast cancer: The St. Gallen International Expert. Consensus Conference on the Primary Therapy of Early Breast Cancer 2017. Ann. Oncol. 2017, 28, 1700–1712. [Google Scholar] [CrossRef]
- Golshan, M.; Loibl, S.; Wong, S.M.; Huober, J.B.; O’Shaughnessy, J.; Rugo, H.S.; Wolmark, N.; McKee, M.D.; Maag, D.; Sullivan, D.M.; et al. Breast Conservation after Neoadjuvant Chemotherapy for Triple-Negative Breast Cancer: Surgical Results from the BrighTNess Randomized Clinical Trial. JAMA Surg. 2020, 155, e195410. [Google Scholar] [CrossRef] [PubMed]
- Denduluri, N.; Chavez-MacGregor, M.; Telli, M.L.; Eisen, A.; Graff, S.L.; Hassett, M.J.; Holloway, J.N.; Hurria, A.; King, T.A.; Lyman, G.H.; et al. Selection of Optimal Adjuvant Chemotherapy and Targeted Therapy for Early Breast Cancer: ASCO Clinical Practice Guideline Focused Update. J. Clin. Oncol. 2018, 36, 2433–2443. [Google Scholar] [CrossRef] [PubMed]
- Asselain, B.; Barlow, W.; Bartlett, J.; Bergh, J.; Bergsten-Nordström, E.; Bliss, J.; Boccardo, F.; Boddington, C.; Bogaerts, J.; Bonadonna, G.; et al. Long-term outcomes for neoadjuvant versus adjuvant chemotherapy in early breast cancer: Meta-analysis of individual patient data from ten randomised trials. Lancet Oncol. 2018, 19, 27–39. [Google Scholar] [CrossRef] [PubMed]
- Schmid, P.; Cortes, J.; Dent, R.; Pusztai, L.; McArthur, H.; Kümmel, S.; Bergh, J.; Denkert, C.; Park, Y.H.; Hui, R.; et al. Event-free Survival with Pembrolizumab in Early Triple-Negative Breast Cancer. N. Engl. J. Med. 2022, 386, 556–567. [Google Scholar] [CrossRef] [PubMed]
- Spring, L.M.; Tolaney, S.M.; Fell, G.; Bossuyt, V.; Abelman, R.O.; Wu, B.; Maheswaran, S.; Trippa, L.; Comander, A.; Mulvey, T.; et al. Response-guided neoadjuvant sacituzumab govitecan for localized triple-negative breast cancer: Results from the NeoSTAR trial. Ann. Oncol. 2024, 35, 293–301. [Google Scholar] [CrossRef] [PubMed]
- Loibl, S.; Schneeweiss, A.; Huober, J.; Braun, M.; Rey, J.; Blohmer, J.U.; Furlanetto, J.; Zahm, D.M.; Hanusch, C.; Thomalla, J.; et al. Neoadjuvant durvalumab improves survival in early triple-negative breast cancer independent of pathological complete response. Ann. Oncol. 2022, 33, 1149–1158. [Google Scholar] [CrossRef]
- Schmid, P.; Cortes, J.; Pusztai, L.; McArthur, H.; Kümmel, S.; Bergh, J.; Denkert, C.; Park, Y.H.; Hui, R.; Harbeck, N.; et al. Pembrolizumab for Early Triple-Negative Breast Cancer. N. Engl. J. Med. 2020, 382, 810–821. [Google Scholar] [CrossRef]
- Masuda, N.; Lee, S.J.; Ohtani, S.; Im, Y.H.; Lee, E.S.; Yokota, I.; Kuroi, K.; Im, S.A.; Park, B.W.; Kim, S.B.; et al. Adjuvant Capecitabine for Breast Cancer after Preoperative Chemotherapy. N. Engl. J. Med. 2017, 376, 2147–2159. [Google Scholar] [CrossRef]
- Li, J.; Yu, K.; Pang, D.; Wang, C.; Jiang, J.; Yang, S.; Liu, Y.; Fu, P.; Sheng, Y.; Zhang, G.; et al. Adjuvant Capecitabine with Docetaxel and Cyclophosphamide Plus Epirubicin for Triple-Negative Breast Cancer (CBCSG010): An Open-Label, Randomized, Multicenter, Phase III Trial. J. Clin. Oncol. 2020, 38, 1774–1784. [Google Scholar] [CrossRef]
- Lluch, A.; Barrios, C.H.; Torrecillas, L.; Ruiz-Borrego, M.; Bines, J.; Segalla, J.; Guerrero-Zotano, Á.; García-Sáenz, J.A.; Torres, R.; de la Haba, J.; et al. Phase III Trial of Adjuvant Capecitabine after Standard Neo-/Adjuvant Chemotherapy in Patients with Early Triple-Negative Breast Cancer (GEICAM/2003-11_CIBOMA/2004-01). J. Clin. Oncol. 2020, 38, 203–213. [Google Scholar] [CrossRef]
- Luen, S.J.; Salgado, R.; Dieci, M.V.; Vingiani, A.; Curigliano, G.; Gould, R.E.; Castaneda, C.; D’Alfonso, T.; Sanchez, J.; Cheng, E.; et al. Prognostic implications of residual disease tumour-infiltrating lymphocytes and residual cancer burden in triple-negative breast cancer patients after neoadjuvant chemotherapy. Ann. Oncol. 2019, 30, 236–242. [Google Scholar] [CrossRef]
- Wang, S.; Sun, Y.; Zhao, S.; Wei, F.; Yang, G. Breast conserving surgery (BCS) with adjuvant radiation therapy showed improved prognosis compared with mastectomy for early staged triple negative breast cancer patients. Math. Biosci. Eng. 2020, 17, 92–104. [Google Scholar] [CrossRef]
- Gupta, A.; Ohri, N.; Haffty, B.G. Hypofractionated radiation treatment in the management of breast cancer. Expert Rev. Anticancer Ther. 2018, 18, 793–803. [Google Scholar] [CrossRef] [PubMed]
- Tan, M.P.; Silva, E. Addressing the paradox of increasing mastectomy rates in an era of de-escalation of therapy: Communication strategies. Breast 2018, 38, 136–143. [Google Scholar] [CrossRef]
- Connor, C.S.; Kimler, B.F.; Mammen, J.M.; McGinness, M.K.; Wagner, J.L.; Alsop, S.M.; Ward, C.; Fabian, C.J.; Khan, Q.J.; Sharma, P. Impact of neoadjuvant chemotherapy on axillary nodal involvement in patients with clinically node negative triple negative breast cancer. J. Surg. Oncol. 2015, 111, 198–202. [Google Scholar] [CrossRef] [PubMed]
- Montero, A.; Ciérvide, R.; García-Aranda, M.; Rubio, C. Postmastectomy radiation therapy in early breast cancer: Utility or futility? Crit. Rev. Oncol. Hematol. 2020, 147, 102887. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Zhao, J.; Han, X.; Er, P.; Meng, X.; Shi, J.; Sun, H.; Zhu, J.; Zhu, L.; Wu, S.; et al. Is There a Role for Post-Mastectomy Radiotherapy for T1-2N1 Breast Cancers with Node-Positive Pathology after Patients Become Node-Negative Pathology following Neoadjuvant Chemotherapy? Front. Oncol. 2020, 10, 892. [Google Scholar] [CrossRef] [PubMed]
- Macklin, A.; Khan, S.; Kislinger, T. Recent Advances in Mass Spectrometry Based Clinical Proteomics: Applications to Cancer Research. Clin. Proteom. 2020, 17, 1–25. [Google Scholar] [CrossRef]
- da Silva, J.L.; Cardoso Nunes, N.C.; Izetti, P.; de Mesquita, G.G.; de Melo, A.C. Triple negative breast cancer: A thorough review of biomarkers. Crit. Rev. Oncol. Hematol. 2020, 145, 102855. [Google Scholar] [CrossRef]
- Blackley, E.F.; Loi, S. Targeting immune pathways in breast cancer: Review of the prognostic utility of TILs in early stage triple negative breast cancer (TNBC). Breast 2019, 48, 44–48. [Google Scholar] [CrossRef]
- Gupta, G.K.; Collier, A.L.; Lee, D.; Hoefer, R.A.; Zheleva, V.; Siewertsz van Reesema, L.L.; Tang-Tan, A.M.; Guye, M.L.; Chang, D.Z.; Winston, J.S.; et al. Perspectives on Triple-Negative Breast Cancer: Current Treatment Strategies, Unmet Needs, and Potential Targets for Future Therapies. Cancers 2020, 12, 2392. [Google Scholar] [CrossRef]
- Sukumar, J.; Gast, K.; Quiroga, D.; Lustberg, M.; Williams, N. Triple-negative breast cancer: Promising prognostic biomarkers currently in development. Expert Rev. Anticancer Ther. 2021, 21, 135–148. [Google Scholar] [CrossRef] [PubMed]
- Abdel-Razeq, H.; Tamimi, F.; Abujamous, L.; Edaily, S.; Abunasser, M.; Bater, R.; Salama, O. Patterns and Prevalence of BRCA1 and BRCA2 Germline Mutations Among Patients with Triple-Negative Breast Cancer: Regional Perspectives. Cancer Manag. Res. 2021, 13, 4597–4604. [Google Scholar] [CrossRef] [PubMed]
- Edechi, C.; Ikeogu, N.; Uzonna, J.; Myal, Y. Regulation of Immunity in Breast Cancer. Cancers 2019, 11, 1080. [Google Scholar] [CrossRef] [PubMed]
- Borri, F.; Granaglia, A. Pathology of triple negative breast cancer. Semin. Cancer Biol. 2020, 72, 136–145. [Google Scholar] [CrossRef]
- Miah, S.; Banks, C.A.S.; Adams, M.K.; Florens, L.; Lukong, K.E.; Washburn, M.P. Advancement of mass spectrometry-based proteomics technologies to explore triple negative breast cancer. Mol. Biosyst. 2017, 13, 42–55. [Google Scholar] [CrossRef]
- Li, X.; Wetherilt, C.S.; Krishnamurti, U.; Yang, J.; Ma, Y.; Styblo, T.M.; Meisel, J.L.; Peng, L.; Siddiqui, M.T.; Cohen, C.; et al. Stromal PD-L1 Expression Is Associated with Better Disease-Free Survival in Triple-Negative Breast Cancer. Am. J. Clin. Pathol. 2016, 146, 496–502. [Google Scholar] [CrossRef]
- Botti, G.; Collina, F.; Scognamiglio, G.; Rao, F.; Peluso, V.; De Cecio, R.; Piezzo, M.; Landi, G.; De Laurentiis, M.; Cantile, M.; et al. Programmed Death Ligand 1 (PD-L1) Tumour Expression Is Associated with a Better Prognosis and Diabetic Disease in Triple Negative Breast Cancer Patients. Int. J. Mol. Sci. 2017, 18, 459. [Google Scholar] [CrossRef]
- Eno, M.S. PA-CJ Immunotherapy through the Years. J. Adv. Pr. Oncol. 2017, 8, 747–753. [Google Scholar]
- Tramm, T.; Di Caterino, T.; Jylling, A.M.B.; Lelkaitis, G.; Lænkholm, A.V.; Ragó, P.; Tabor, T.P.; Talman, M.L.M.; Vouza, E.; Scientific Committee of Pathology, Danish Breast Cancer Group (DBCG). Standardized assessment of tumour-infiltrating lymphocytes in breast cancer: An evaluation of inter-observer agreement between pathologists. Acta Oncol. 2018, 57, 90–94. [Google Scholar] [CrossRef]
- Gao, G.; Wang, Z.; Qu, X.; Zhang, Z. Prognostic value of tumour-infiltrating lymphocytes in patients with triple-negative breast cancer: A systematic review and meta-analysis. BMC Cancer 2020, 20, 179. [Google Scholar]
- Jang, N.; Kwon, H.J.; Park, M.H.; Kang, S.H.; Bae, Y.K. Prognostic Value of Tumour-Infiltrating Lymphocyte Density Assessed Using a Standardized Method Based on Molecular Subtypes and Adjuvant Chemotherapy in Invasive Breast Cancer. Ann. Surg. Oncol. 2018, 25, 937–946. [Google Scholar] [CrossRef] [PubMed]
- Chrétien, S.; Zerdes, I.; Bergh, J.; Matikas, A.; Foukakis, T. Beyond PD-1/PD-L1 inhibition: What the future holds for breast cancer immunotherapy. Cancers 2019, 11, 628. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Tian, Q.; Wang, B.; Yang, J.; Zhao, S.; Yang, J. The prognostic significance of TILs as a biomarker in triple-negative breast cancer: What is the role of TILs in TME of TNBC? Eur. Rev. Med. Pharmacol. Sci. 2021, 25, 2885–2897. [Google Scholar] [PubMed]
- Denkert, C.; von Minckwitz, G.; Darb-Esfahani, S.; Lederer, B.; Heppner, B.I.; Weber, K.E.; Budczies, J.; Huober, J.; Klauschen, F.; Furlanetto, J.; et al. Tumour-infiltrating lymphocytes and prognosis in different subtypes of breast cancer: A pooled analysis of 3771 patients treated with neoadjuvant therapy. Lancet Oncol. 2018, 19, 40–50. [Google Scholar] [CrossRef]
- Pinard, C.; Debled, M.; Ben Rejeb, H.; Velasco, V.; Tunon de Lara, C.; Hoppe, S.; Richard, E.; Brouste, V.; Bonnefoi, H.; MacGrogan, G. Residual cancer burden index and tumour-infiltrating lymphocyte subtypes in triple-negative breast cancer after neoadjuvant chemotherapy. Breast Cancer Res. Treat. 2020, 179, 11–23. [Google Scholar] [CrossRef]
- Domon, B.; Aebersold, R. Mass Spectrometry and Protein Analysis. Science 2006, 312, 212–217. [Google Scholar] [CrossRef]
- Fu, W.; Sun, H.; Zhao, Y.; Chen, M.; Yang, X.; Liu, Y.; Jin, W. BCAP31 drives TNBC development by modulating ligand-independent EGFR trafficking and spontaneous EGFR phosphorylation. Theranostics 2019, 9, 6468–6484. [Google Scholar] [CrossRef]
- Wu, Y.; Wu, M.; Zheng, X.; Yu, H.; Mao, X.; Jin, Y.; Wang, Y.; Pang, A.; Zhang, J.; Zeng, S.; et al. Discovery of a potent and selective PARP1 degrader promoting cell cycle arrest via intercepting CDC25C-CDK1 axis for treating triple-negative breast cancer. Bioorg. Chem. 2024, 142, 106952. [Google Scholar] [CrossRef]
- Franz, A.; Coscia, F.; Shen, C.; Charaoui, L.; Mann, M.; Sander, C. Molecular response to PARP1 inhibition in ovarian cancer cells as determined by mass spectrometry based proteomics. J. Ovarian Res. 2021, 14, 140. [Google Scholar] [CrossRef]
- Amante, R.J.; Auf der Maur, P.; Richina, V.; Sethi, A.; Iesmantavicius, V.; Bonenfant, D.; Aceto, N.; Bentires-Alj, M. Protein Tyrosine Phosphatase SHP2 Controls Interleukin-8 Expression in Breast Cancer Cells. J. Mammary Gland. Biol. Neoplasia 2022, 27, 145–153. [Google Scholar] [CrossRef]
- Giuliano, S.; Dufies, M.; Ndiaye, P.D.; Viotti, J.; Borchiellini, D.; Parola, J.; Vial, V.; Cormerais, Y.; Ohanna, M.; Imbert, V.; et al. Resistance to lysosomotropic drugs used to treat kidney and breast cancers involves autophagy and inflammation and converges in inducing CXCL5. Theranostics 2019, 9, 1181–1199. [Google Scholar] [CrossRef]
- Qin, G.; Wang, X.; Ye, S.; Li, Y.; Chen, M.; Wang, S.; Qin, T.; Zhang, C.; Li, Y.; Long, Q.; et al. NPM1 upregulates the transcription of PD-L1 and suppresses T cell activity in triple-negative breast cancer. Nat. Commun. 2020, 11, 1669. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Li, S.; Xue, J.; Qi, M.; Liu, X.; Huang, Y.; Hu, J.; Dong, H.; Ling, K. PD-L1 tumour-intrinsic signaling and its therapeutic implication in triple-negative breast cancer. JCI Insight 2021, 6, e131458. [Google Scholar] [CrossRef]
- Goode, G.; Gunda, V.; Chaika, N.V.; Purohit, V.; Yu, F.; Singh, P.K. MUC1 facilitates metabolomic reprogramming in triple-negative breast cancer. PLoS ONE 2017, 12, e0176820. [Google Scholar]
- del Pilar Chantada-Vázquez, M.; López, A.C.; Vence, M.G.; Vázquez-Estévez, S.; Acea-Nebril, B.; Calatayud, D.G.; Jardiel, T.; Bravo, S.B.; Núñez, C. Proteomic investigation on bio-corona of Au, Ag and Fe nanoparticles for the discovery of triple negative breast cancer serum protein biomarkers. J. Proteom. 2020, 212, 103581. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Xue, Q.; Wang, M.; Meng, B.; Jiang, Y.; Zhai, R.; Zhang, Y.; Dai, X.; Fang, X. Evolution of Mass Spectrometry Instruments and Techniques for Blood Proteomics. J. Proteome Res. 2023, 22, 1009–1023. [Google Scholar] [CrossRef] [PubMed]
- Nadler, W.M.; Waidelich, D.; Kerner, A.; Hanke, S.; Berg, R.; Trumpp, A.; Rösli, C. MALDI versus ESI: The Impact of the Ion Source on Peptide Identification. J. Proteome Res. 2017, 16, 1207–1215. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Chu, S.; Tan, S.; Yin, X.; Jiang, Y.; Dai, X.; Gong, X.; Fang, X.; Tian, D. Towards Higher Sensitivity of Mass Spectrometry: A Perspective from the Mass Analyzers. Front. Chem. 2021, 9, 813359. [Google Scholar] [CrossRef]
- Gillet, L.C.; Leitner, A.; Aebersold, R. Mass Spectrometry Applied to Bottom-Up Proteomics: Entering the High-Throughput Era for Hypothesis Testing. Annu. Rev. Anal. Chem. 2016, 9, 449–472. [Google Scholar] [CrossRef]
- De Hoffmann, E.; Stroobant, V. Mass Spectrometry: Principles and Applications, 3rd ed.; John Wiley & Sons: Hoboken, NJ, USA, 2007; pp. 247–253. [Google Scholar]
- Aebersold, R.; Mann, M. Mass-spectrometric exploration of proteome structure and function. Nature 2016, 537, 347–355. [Google Scholar] [CrossRef]
- Kim, M.S.; Pinto, S.M.; Getnet, D.; Nirujogi, R.S.; Manda, S.S.; Chaerkady, R.; Madugundu, A.K.; Kelkar, D.S.; Isserlin, R.; Jain, S.; et al. A draft map of the human proteome. Nature 2014, 509, 575–581. [Google Scholar] [CrossRef] [PubMed]
- Wilhelm, M.; Schlegl, J.; Hahne, H.; Gholami, A.M.; Lieberenz, M.; Savitski, M.M.; Ziegler, E.; Butzmann, L.; Gessulat, S.; Marx, H.; et al. Mass-spectrometry-based draft of the human proteome. Nature 2014, 509, 582–587. [Google Scholar] [CrossRef]
- Nakayasu, E.S.; Gritsenko, M.; Piehowski, P.D.; Gao, Y.; Orton, D.J.; Schepmoes, A.A.; Fillmore, T.L.; Frohnert, B.I.; Rewers, M.; Krischer, J.P.; et al. Tutorial: Best practices and considerations for mass-spectrometry-based protein biomarker discovery and validation. Nat. Protoc. 2021, 16, 3737–3760. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Zheng, N. Accelerating protein biomarker discovery and translation from proteomics research for clinical utility. Bioanalysis 2020, 12, 1469–1481. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2020. CA Cancer J. Clin. 2020, 70, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Yaffe, M.B. Why geneticists stole cancer research even though cancer is primarily a signaling disease. Sci. Signal 2019, 12, eaaw3483. [Google Scholar] [CrossRef] [PubMed]
- Aebersold, R.; Agar, J.N.; Amster, I.J.; Baker, M.S.; Bertozzi, C.R.; Boja, E.S.; Costello, C.E.; Cravatt, B.F.; Fenselau, C.; Garcia, B.A.; et al. How many human proteoforms are there? Nat. Chem. Biol. 2018, 14, 206–214. [Google Scholar] [CrossRef]
- Veyssière, H.; Bidet, Y.; Penault-Llorca, F.; Radosevic-Robin, N.; Durando, X. Circulating proteins as predictive and prognostic biomarkers in breast cancer. Clin. Proteom. 2022, 19, 25. [Google Scholar] [CrossRef]
- Petrera, A.; von Toerne, C.; Behler, J.; Huth, C.; Thorand, B.; Hilgendorff, A.; Hauck, S.M. Multiplatform Approach for Plasma Proteomics: Complementarity of Olink Proximity Extension Assay Technology to Mass Spectrometry-Based Protein Profiling. J. Proteome Res. 2021, 20, 751–762. [Google Scholar] [CrossRef]
- Boys, E.L.; Liu, J.; Robinson, P.J.; Reddel, R.R. Clinical applications of mass spectrometry-based proteomics in cancer: Where are we? Proteomics 2023, 23, 2200238. [Google Scholar] [CrossRef]
- Pan, S.; Chen, R. Pathological implication of protein post-translational modifications in cancer. Mol. Asp. Med. 2022, 86, 10–1097. [Google Scholar] [CrossRef] [PubMed]
- Tyanova, S.; Albrechtsen, R.; Kronqvist, P.; Cox, J.; Mann, M.; Geiger, T. Proteomic maps of breast cancer subtypes. Nat. Commun. 2016, 7, 10259. [Google Scholar] [CrossRef]
- Zhang, Y.; Fonslow, B.R.; Shan, B.; Baek, M.C.; Yates, J.R., III. Protein analysis by shotgun proteomics. Mass. Spectrom. Chem. Proteom. 2013, 113, 2343–2394. [Google Scholar]
- Uhlen, M.; Karlsson, M.J.; Zhong, W.; Tebani, A.; Pou, C.; Mikes, J.; Lakshmikanth, T.; Forsström, B.; Edfors, F.; Odeberg, J.; et al. A genome-wide transcriptomic analysis of protein-coding genes in human blood cells. Science 2019, 366, eaax9198. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Liu, T.; Zhang, Z.; Payne, S.H.; Zhang, B.; McDermott, J.E.; Zhou, J.Y.; Petyuk, V.A.; Chen, L.; Ray, D.; et al. Integrated Proteogenomic Characterization of Human High-Grade Serous Ovarian Cancer. Cell 2016, 166, 755–765. [Google Scholar] [CrossRef]
- Martens, L.; Vizcaíno, J.A. A Golden Age for Working with Public Proteomics Data. Trends Biochem. Sci. 2017, 42, 333–341. [Google Scholar] [CrossRef] [PubMed]
- Hüttenhain, R.; Soste, M.; Selevsek, N.; Röst, H.; Sethi, A.; Carapito, C.; Farrah, T.; Deutsch, E.W.; Kusebauch, U.; Moritz, R.L.; et al. Reproducible Quantification of Cancer-Associated Proteins in Body Fluids Using Targeted Proteomics. Sci. Transl. Med. 2012, 4, 142–194. [Google Scholar] [CrossRef] [PubMed]
- Frampton, I.; Lask, B. Mass spectrometry based proteomics: Existing capabilities and future directions. Eat. Disord. Brain 2012, 41, 207–217. [Google Scholar]
- Zecha, J.; Satpathy, S.; Kanashova, T.; Avanessian, S.C.; Kane, M.H.; Clauser, K.R.; Mertins, P.; Carr, S.A.; Kuster, B. TMT labeling for the masses: A robust and cost-efficient, in-solution labeling approach. Mol. Cell Proteom. 2019, 18, 1468–1478. [Google Scholar] [CrossRef]
- Tyanova, S.; Temu, T.; Cox, J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 2016, 11, 2301–2319. [Google Scholar] [CrossRef]
- Prianichnikov, N.; Koch, H.; Koch, S.; Lubeck, M.; Heilig, R.; Brehmer, S.; Fischer, R.; Cox, J. MaxQuant Software for Ion Mobility Enhanced Shotgun Proteomics. Mol. Cell Proteom. 2020, 19, 1058–1069. [Google Scholar] [CrossRef] [PubMed]
- Birhanu, A.G. Mass spectrometry-based proteomics as an emerging tool in clinical laboratories. Clin. Proteom. 2023, 20, 32. [Google Scholar] [CrossRef]
- Bhawal, R.; Oberg, A.L.; Zhang, S.; Kohli, M. Challenges and opportunities in clinical applications of blood-based proteomics in cancer. Cancers 2020, 12, 2428. [Google Scholar] [CrossRef] [PubMed]
- Meyer, J.G.; Schilling, B. Clinical applications of quantitative proteomics using targeted and untargeted data-independent acquisition techniques. Expert Rev. Proteom. 2017, 14, 419–429. [Google Scholar] [CrossRef] [PubMed]
- Li, X.J.; Hayward, C.; Fong, P.Y.; Dominguez, M.; Hunsucker, S.W.; Lee, L.W.; McLean, M.; Law, S.; Butler, H.; Schirm, M.; et al. A Blood-Based Proteomic Classifier for the Molecular Characterization of Pulmonary Nodules. Sci. Transl. Med. 2013, 5, 207ra142. [Google Scholar] [CrossRef]
| Gene | Function | Frequency in TNBC | Clinical Implications |
|---|---|---|---|
| TP53 mutation | Tumour suppressor gene | 41% of tumours | The inactivation of tumour suppressor genes, resulting in the advancement of tumour growth. |
| PIK3CA mutation | Catalytic subunit of PI3K | 30% | Activates the PI3K/AKT/mTOR pathway, promoting cell survival and growth. |
| MYC overexpression | Oncogene | 20% | Stimulates cell division and supports tumour development. |
| PTEN inactivation | Tumour suppressor gene | 16% | Activates the PI3K/AKT pathway due to the loss of regulatory inhibition. |
| BRCA1/BRCA2 Germline mutation | DNA repair, tumour suppressor genes | 72% | Strong correlation with hereditary TNBC. |
| FGFR1 overexpression | Receptor tyrosine kinase | 11% | Overexpression of FGFR1 contributes to tumour aggressiveness. |
| Molecular Target | Targeted Pathway/Process | Associated Therapy | Clinical Status | Remarks |
|---|---|---|---|---|
| PD-L1 | Immune checkpoint inhibition | Pembrolizumab, Atezolizumab | FDA-approved for specific TNBC cases | Enhances immune response against tumours [22] |
| PARP1/2 | DNA damage repair | Olaparib, Talazoparib | FDA-approved for BRCA-mutated TNBC | Exploits synthetic lethality in BRCA-deficient tumours [15] |
| EGFR | Growth factor signalling | Cetuximab (in trials) | Under investigation | Overexpressed in some TNBC subtypes [23] |
| Androgen Receptor (AR) | Hormone receptor signalling | Enzalutamide (in trials) | Under investigation | Targeted in AR-positive TNBC [24] |
| PI3K/AKT/mTOR | Cell growth and survival | Alpelisib (in trials) | Under investigation | Pathway frequently activated in TNBC [25] |
| CDK4/6 | Cell cycle regulation | Palbociclib, Ribociclib (in trials) | Under investigation | Inhibition can block cell proliferation in TNBC [26] |
| BRCA1/2 | DNA repair | Olaparib | FDA-approved for BRCA-mutated TNBC | Germline mutations can drive tumour development [27] |
| Trials | Design | Population | Intervention | Outcomes |
|---|---|---|---|---|
| KEYNOTE-522 | Phase III, Randomised, Double-Blind, Placebo-Controlled | Early-stage, high-risk TNBC patients | Pembrolizumab + NAC vs. Placebo | Improved pCR and EFS, supporting pembrolizumab with NAC for high-risk TNBC. |
| NeoSTAR | Phase II, Single-Arm | Stage II/III TNBC patients | Neoadjuvant Nivolumab + Chemotherapy | Trial is ongoing. Promising pCR rates and immune activation, suggesting neoadjuvant nivolumab’s potential in TNBC, though further studies needed to confirm efficacy |
| GeparNuevo | Phase II, Randomised, Double-Blind, Placebo-Controlled | Early-stage TNBC patients | Neoadjuvant Durvalumab + Chemotherapy vs. Placebo | Durvalumab improved pCR rates, particularly when administered before chemotherapy, highlighting potential timing considerations for ICI in TNBC treatment. |
| CREATE-X | Phase III, Randomised, Open-Label | HER2-negative breast cancer patients with residual disease | Adjuvant Capecitabine vs. Observation | Significantly improved DFS and OS in patients with residual disease after NAC, demonstrating the benefit of adjuvant capecitabine, especially in the TNBC subpopulation. |
| ADAPT-TN | Phase II, Randomised | Early-stage TNBC | Neoadjuvant Pembrolizumab + Nab-Paclitaxel + Epirubicin vs. Chemotherapy Alone | Trial is ongoing. Evaluates the impact of adding pembrolizumab to NAC on pCR rates, with positive results suggesting enhanced efficacy in TNBC. |
| SASCIA | Phase III, Randomised, Open-Label | Patients with early-stage TNBC are at high risk of recurrence | Sacituzumab Govitecan vs. Treatment of Physician’s Choice | Trial is still ongoing. Aims to assess the efficacy of Sacituzumab Govitecan in reducing recurrence rates and improving survival in high-risk TNBC patients following standard neoadjuvant therapy. |
| Protein Biomarker/Pathway | Role in TNBC | Clinical Relevance | MS-Based Studies: References |
|---|---|---|---|
| EGFR | Growth factor signalling | Potential target for targeted therapy | Studies have confirmed that EGFR expression is elevated in TNBC, suggesting its potential as a therapeutic target [23,78]. |
| PARP1/2 | DNA damage repair | Targeted by PARP inhibitor | Studies have validated the critical role of PARP1/2 in BRCA-mutated TNBC, thereby guiding treatment strategies [79,80]. |
| CXCL8 (IL-8) | Chemokine signalling | Linked to poor prognosis and metastasis | Studies have linked elevated CXCL8 levels to poorer clinical outcomes in TNBC patients [81,82]. |
| PD-L1 | Immune checkpoint regulation | Targeted by immune checkpoint inhibitors | Studies have revealed the upregulation of PD-L1, supporting the use of immune checkpoint blockade in TNBC [83,84]. |
| MUC1 | Cell surface glycoprotein | Potential target for immunotherapy | Identified as an overexpressed marker in TNBC, it is considered a potential target for novel therapies [66,85]. |
| S100A7 | Calcium-binding protein | Associated with invasion and metastasis | Identified as a key contributor to TNBC progression, particularly in its more invasive forms [86]. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Metwali, E.; Pennington, S. Mass Spectrometry-Based Proteomics for Classification and Treatment Optimisation of Triple Negative Breast Cancer. J. Pers. Med. 2024, 14, 944. https://doi.org/10.3390/jpm14090944
Metwali E, Pennington S. Mass Spectrometry-Based Proteomics for Classification and Treatment Optimisation of Triple Negative Breast Cancer. Journal of Personalized Medicine. 2024; 14(9):944. https://doi.org/10.3390/jpm14090944
Chicago/Turabian StyleMetwali, Essraa, and Stephen Pennington. 2024. "Mass Spectrometry-Based Proteomics for Classification and Treatment Optimisation of Triple Negative Breast Cancer" Journal of Personalized Medicine 14, no. 9: 944. https://doi.org/10.3390/jpm14090944







