In order to complement the work performed during a decade (2008–2018) of surveillance, the objective of this study was to use WGS and SNP analysis to characterize the M. bovis isolates obtained from livestock and wildlife in Catalonia to identify transmission clusters and use this characterization to determine how the isolates from this region fit into the broader global structure of M. bovis.
4.1. Isolates’ Demographic Attributes
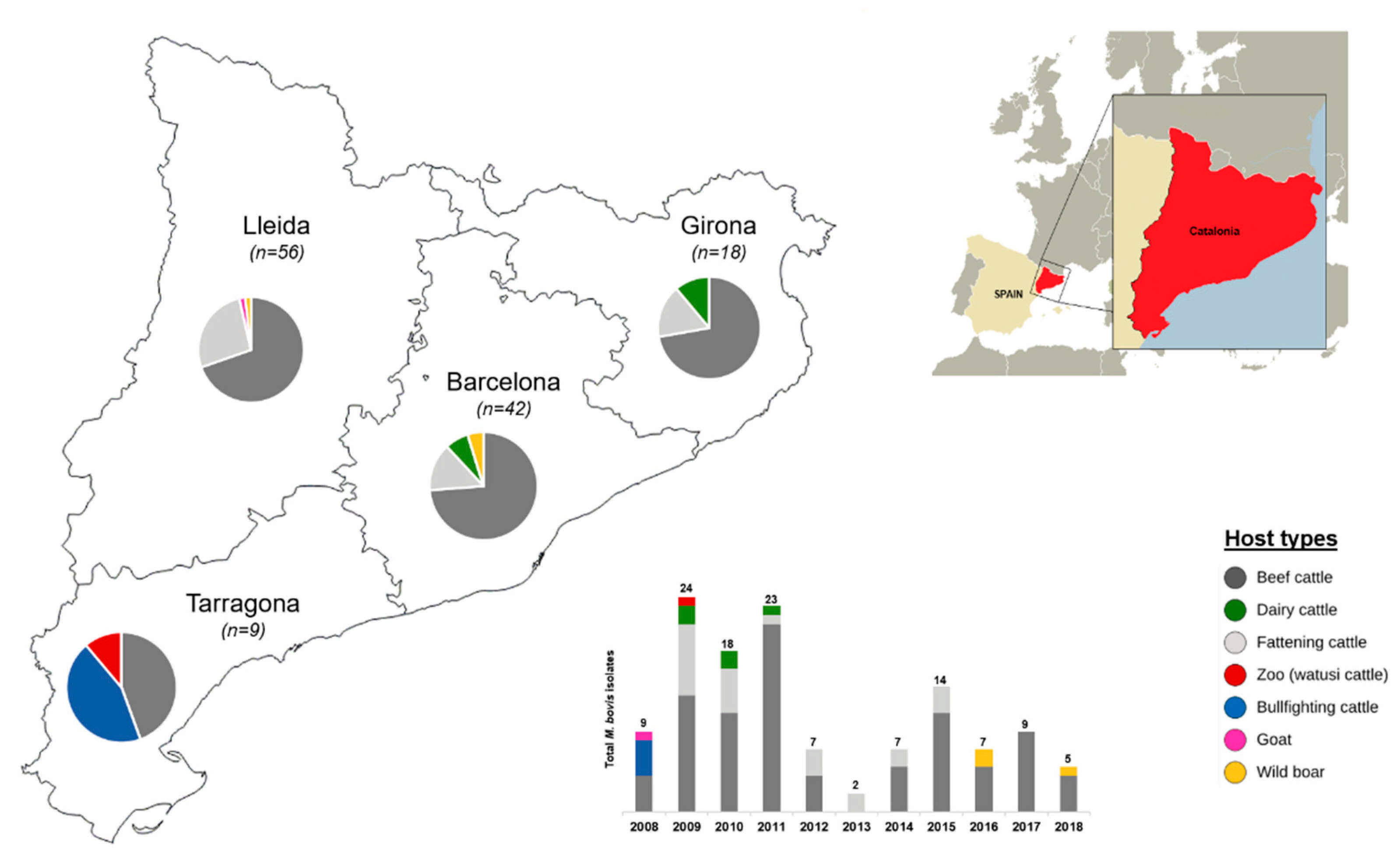
In terms of the host types analyzed in this study, Catalonia’s livestock sector comprises two main cattle production types: dairy and beef. The latter includes “fattening” cattle as it is a stage in the production cycle of beef cattle; additionally, raising bullfighting cattle is traditional in Spain; however, in Catalonia, it is rare (i.e., approximately 12 out of 5000 herds) [
6]. For epidemiological purposes, beef cattle were classified into “fattening” (cattle up to 24 months of age) and “beef” (cattle older than 24 months) and considered as separate host types. From 125 isolates obtained, most were from beef and fattening cattle (88.8%, 111/125). These two host types present a higher TB prevalence compared to dairy cattle, according to the TB surveillance program (2.03% versus 0.71%, respectively) [
6]. This may be directly related to cattle population, for which the Catalan cattle census reports ~85% beef cattle, ~15% dairy, and <0.3% bullfighting cattle. In relation to this, sample submission to the lab for TB diagnostics is higher for beef cattle because more field tests are performed due to the need for animal movements (such as the sale of animals or movement to pasture zones), and in the case for reactor animals that are culled and subsequently sampled for laboratory tests. Dairy cattle were only represented by five isolates (4%), which, as mentioned above, may be in relation to the lower prevalence of TB in this production type [
6,
21,
31]. Bullfighting cattle followed with four isolates (3%), and even though it presents the highest prevalence of the three production types (beef, dairy, and bullfighting) [
6], Catalonia’s bullfighting cattle population is very small in comparison to beef and dairy, as well as the rest of Spain, accounting for approximately 12 out of 1142 bullfighting cattle-raising farms in the country [
6]. As for goats and wild boar (with 1 and 3 isolates, respectively), goats have been found infected mostly with
M. caprae [
30,
47,
48], while the wild boar has been identified as a reservoir for both
M. bovis and
M. caprae [
49,
50]. Finally, one isolate was recovered from a Watusi animal from a zoological exhibit; there is only one other report of TB in captive Watusi cattle in the scientific literature from a study performed in Mexico [
51]. In Africa, in its native setting, the Watusi are commonly used for milk, meat and leather, but in Spain, this breed of cattle is mainly seen in zoological settings.
It is important to mention that the isolates used in this study are a subset of a larger collection of isolates maintained at the IRTA-CReSA laboratory and are meant to represent the population genetic diversity of
M. bovis in Catalonia, taking into account two main aspects: first, they are representative of the major spoligotyping-VNTR types and adjacent variants and second, they cover different host types, geographic regions (municipalities) and temporal/epidemiological contexts (years of isolation and farms of origin). Briefly, with respect to year of isolation, the total number of isolates per year (2008–2018) reflects the tendency of TB prevalence, with the most isolates seen for years 2008–2011, in which herd TB prevalence was higher in Catalonia (0.59–0.85%), and then decreasing also as the prevalence decreased. A similar scenario was reported in a study in Portugal [
52]. However, the same cannot be said for the number of isolates per province and TB prevalence in that region.
4.2. Genetic Diversity
A total of 27 spoligotypes are represented by this data set, where SB0121, SB0134, and SB1337 were the three most frequent. Spoligotypes SB0121 and SB0134 are also found in Portugal and France [
52,
53], reflecting the long trade relationships between these countries; however, for SB1337, reports outside of Spain were not found (
www.mbovis.org, accessed on 29 July 2021) and may even be exclusive to Catalonia. Given that this spoligotype is mainly concentrated in the Pyrenees, further studies in Spain and France might provide more insight into this matter. A brief comparison using the profiles reported in the Mbovis.org database showed that 10 of the spoligotypes found here have been reported for Spain, one for Portugal (SB1095), two for France (SB0832 and SB0828), and one for the United Kingdom (SB0142). However, it is possible that not all of the spoligotypes found in these countries are reported within the database, such as SB0140, which is very common in Great Britain [
18], so this may be underestimated. In general, spoligotyping and WGS correlated well, though clearly, WGS resolved phylogenetic relationships at a finer scale.
The SNP-based phylogenetic analysis identified seven clades (I–VII). These corresponded to 123/125 of the isolates and may represent the general population structure and genetic diversity of
M. bovis in Catalonia; however, two additional clades may exist, as two isolates (17-010561-69/SB0130 and 17-010561-61/SB0140) were located on two separate branches that did not cluster with any of the clades identified here. Future studies might benefit from including more isolates with the same spoligotype/VNTR profiles as these to gain more insight into this matter. A correlation of these clades with the main clonal complexes described so far for
M. bovis indicates that Eu1 and Eu2 are well represented in this data set, as well as the “BCG-like” clonal complex. In Spain, one other study used WGS, in which case an outbreak of
M. caprae was investigated [
30]. In Europe, recent studies in France [
26] and Germany [
27] also took advantage of this high-resolution technology to delineate the population structure of
M. bovis in each country, finding 9 and 13 clades, with 87 and 131
M. bovis isolates, respectively. In both, Eu1, Eu2, and the “BCG-like” clonal complexes were well represented, with Af1 and Af2 also present in Germany, plus additional clades that do not fall within the defined complexes. In this study, one clade (associated with spoligotypes SB0134, SB0832, and SB1341) did not correspond to any of the known clonal complexes. The authors of [
54] identified eight new groups in addition to the known complexes, and of these, Clade II from our study correlates with Unknown 7, which was also found in Germany. According to the aforementioned study, this clade has a geographical range within Western and Southern Europe, as well as Northern and Eastern Africa.
4.3. Transmission Clusters
A threshold of 12 SNPs was established to identify strains possibly involved in transmission events, as previous studies have suggested [
27,
28,
29,
46]. This threshold resulted in the identification of 21 putative transmission clusters. Half of these included only a pair of isolates, mostly obtained from one host type (beef cattle) and isolated within a three-year period. However, a few involved transmission between different host types, including beef-dairy and beef-wildlife. In a previous study [
21], seven possible causes of bovine TB breakdowns were assessed in Spain, including: (1) residual infection, (2) introduction of infected cattle, (3) presence of infected goats, (4) contiguous spread, (5) sharing of pastures, (6) interaction with wildlife, and (7) contact with infected humans. In the present study, the explanatory factors attributed to the putative transmission clusters identified through WGS were (1) residual infection, (2) neighborhood (similar municipality), (3) proximity (similar county), (4) movement, (5) shared pasture, (6) livestock-wildlife interaction, and (7) unknown. For the sake of comparison, “neighborhood” and “proximity” are similar to “contiguous spread”, “movement” is homologous to “introduction of infected cattle”, and “residual infection”, “shared pasture” and “livestock-wildlife interaction” are exactly as previously defined (
Table 6). In our study, only two most likely causes were not determined as explanatory factors, “presence of infected goats”, and “contact with infected humans”, which agrees with the results obtained recently by [
19]. After an “Unknown” likely cause, a high proportion of bovine TB breakdowns were attributed to “residual infection” by both [
21] and [
19], while in this study, “contiguous spread” was the most frequent explanatory factor. “Introduction of infected cattle” increased from 5.1% to 13.8%, which is similar to our results of 15%. Finally, the proportion of putative transmission clusters explained by “shared pasture” and “interaction with wildlife”, which were 12% and 15%, respectively in this study, were in agreement with the proportions described by [
21] for the same categories. It is interesting how in spite of the different methodologies used for each of the studies, WGS could lead to comparable results. This highlights the fact that while WGS is a powerful tool for investigating sources of infection, an adequate epidemiological investigation must accompany the findings [
55]. Some key aspects that could partially contribute to the differences in results obtained by [
21], ref. [
19] and this study are, first, that this study analyzed data from Catalonia only, while previous studies analyzed data from different autonomous communities, especially with regard to “wildlife interaction” since the importance of these species in the maintenance of bovine TB is heterogeneous across Spain [
54,
56]; second, in this study, “contiguous spread” encompasses wider criteria, as it also includes farms from the same county, while in the previous studies this only included farms in a 1 km radius.
While these risk factors operate at different scales and may vary across regions, other epidemiological studies have identified a number of risk factors associated with bovine TB herd breakdowns, such as the purchase of cattle, the occurrence of bovine TB in contiguous herds and/or surrounding areas and herd size, and to a lesser degree farm and herd-management practices such as farms having multiple premises, the use of certain housing types, etc. [
57,
58].
In general, the most consistently identified risk factors are biologically plausible and consistent with known transmission routes involving cattle-cattle (or other livestock) and wildlife-cattle pathways [
59,
60]. Clusters I.1, III.1, V.1a, and VI.2 showed within-herd transmission, as the animals from each cluster originated from the same herds. Within-herd transmission is the most problematic due to the challenges that can pose certain herd-management practices that do not prevent direct contact between infected and non-infected animals, in addition to contaminated soil, feed, and water from infected herd-mates [
60]. Congenital transmission is rare, and transmission via milk is easily preventable by pasteurization [
61,
62]. Between-herd transmission can be attributed to all clusters in which the most likely explanatory factors were neighborhood, proximity, movement, and shared pasture. For the first two, this could be due to sharing a common boundary or damaged fences that allow for cattle to mix in an uncontrolled way, or even common access with wildlife reservoir [
60,
63]. With regard to movement, pre-movement testing such as the tuberculin skin test is commonly performed, but the sensitivity of the test may be influenced by the number of animals, as well as the size of the herd of origin, not to mention false-negative results in cases of recent infection where the immune response has not yet developed [
35]. As for shared pastures, this practice predisposes contact with other herds, thereby increasing the risk of disease transmission among herds either by animals that inter-mingle, have access to contaminated pasture and soil, and/or infected wildlife [
64,
65]. Therefore, an important aspect to consider with respect to the potential spread of diseases on communal grazing areas is the level of mixing between animals from different origins. In this regard, [
65] showed that the level of mixing might have an effect on bovine TB spread, which in turn may be influenced by herd size [
66,
67]. More studies are needed to properly estimate the mixing pattern between animals from different herds.
Interestingly, one cluster involved transmission between beef and dairy cattle, which often seems like a rare event since beef and dairy herds are usually raised in separate systems. These isolates were only 2 SNPs apart, which strongly suggests a link between them, and although a direct event of transmission has not been identified, the only link (or explanatory factor) found was “proximity” (i.e., belonged to the same county, <7 km). In a study performed in Northern Ireland, the authors showed borderline significance for spatial proximities of 2 km but none for 5 km [
9]. Besides proximity, another possible explanation for the genetic association between these isolates might be that sometimes male-born calves from dairy farms are sold to beef-raising farms, making it possible for transmission to happen between these systems.
For bullfighting cattle, transmission has been attributed to animals from the same herd and/or wildlife that inhabit the premises of such herds [
68]. Neither spoligotyping nor WGS could match the bullfighting cattle isolates closely to another host type from this data set. Cluster III.1 involved three isolates, separated by only 7 SNPs, though two had identical SNP profiles. While these were more closely related to isolates from beef cattle, there is a more than 32 SNP pairwise distance, so no clear conclusions could be drawn. Previously, SB0295 was identified as the most prevalent in this host type, but the spoligotypes identified here were SB1095, SB1192, and SB1685, which does not match what was previously reported [
69]. Further studies regarding TB in bullfighting cattle may be needed to gain more insight into the epidemiology of the disease in this host type.
The exact origin or direction of infection (i.e., which animal infected which) could not be determined with this data set, but the pairwise SNP distances between isolates (≤12 SNPs) made it possible to follow epidemiological data to identify explanatory factors. WGS provides high-level resolution for strain characterization, which is a powerful tool for building databases against which isolates can be compared to identify closely related strains to understand how disease is introduced into a particular setting or how residual infection persists [
25,
29].
4.4. Catalonia M. bovis in a World Context
Often the comparison of
M. bovis genotypes from different regions of the world has revealed patterns influenced by the historical trade of cattle [
18,
70], as well as the natural migration of domesticated livestock [
71], and the higher resolution of WGS has provided insight into the evolutionary origins of
M. bovis as a whole [
72,
73]. In this study, the comparison of Catalonia
M. bovis isolates against a global data set showed clustering with isolates from the United Kingdom, France, United States, and Mexico, which agrees with previous reports regarding the distribution of
M. bovis lineages throughout the world. For example, we found evidence that showed the introduction of a foreign strain as an isolate from U.K.-imported beef cattle matched an isolate from Northern Ireland within 4 SNP. Similarly, two additional imported beef cattle (U.K. and Ireland) also clustered with these isolates. Aside from historical trade, neighboring countries often maintain close commerce relationships due to the ease of transport. For example, the United States and Mexico have a long-standing trade of cattle, which reflects the high degree of shared
M. bovis genotypes [
74]. For Spain, neighboring countries include Portugal and France, and the results are seen here support this regional distribution of genotypes [
26,
52]. Interestingly, two Catalonia isolates clustered with isolates from South Africa, which may be a function of the Spanish colonization of Africa in the first third of the 20th century [
75], or it could go further back to the routes and scattering of cattle in Africa, which subsequently reached Spain [
76]. Further studies including a larger set of isolates from different regions of Africa may provide more insight into this matter.
4.5. Virulence and Antibiotic Resistance Factors
SNPs were investigated to determine if they fell within genes associated with virulence, and out of 181 genes obtained from a previously compiled list [
26], only 68 were found to contain SNPs. After categorizing them into 11 functional classes, genes involved in lipid transport and metabolism had the highest number of SNP-hits. The relationship of lipid and fatty acid metabolism to mycobacteria virulence has been well studied [
77], so the high number of SNPs (52 out of 156) associated with this category may be indicative of how constant remodeling of the cell wall by the mycobacteria has allowed for it to adapt to the various hosts it infects and achieve maximum survival. Within this category, genes pks7 and pks12 were also found at a high degree of mutation by [
26], with 8 and 49 variants, respectively, while in this study, each presents 10 variants, which were the highest number of mutations found in a single gene. The average number of mutations per gene was 2.3, with most genes having between 1 and 3 mutations. Other categories with a high number (above average) of mutations were defense mechanisms (
n = 20), genes and metabolism regulation (
n = 20) and mce families (
n = 20). Within defense mechanisms, oxidative and nitrosative stress genes had the most mutations, which makes sense given that resistance to macrophage-mediated killing by reactive oxygen/nitrogen species is critical to the virulence of mycobacteria [
78]. For genes and metabolism regulation, sigma factors and protein kinases (Pkns) had the most SNP-hits; it has been suggested that crosstalk among Pkns, sigma factors, and two-component systems help the mycobacteria adapt to external stimuli [
77]. Finally, the mce family proteins have been shown to confer mycobacteria the ability to enter into mammalian cells (mce = mammalian cell entry) [
79], so continuous adaptation of these factors through mutations may be key for propagation. Interestingly, 79.5% of all SNPs were nonsynonymous, posing a greater risk of causing modifications in the structure/function of the final product. Previously, a comparable proportion of nonsynonymous SNPs was obtained by [
26] at 69%. While these are only predicted mutations, further investigations on protein structure/chemistry, secreted proteins, cell wall antigens, transmembrane proteins are needed to fully understand the effect of these mutations on phenotype and thus pinpoint specific differences across the main
M. bovis groups.
In this respect, Clade VI presented by far the higher number of variants than the other clades (
Figure 5B), and most of this increase was represented by the secretion system category, represented mainly by the
secA2 and
PPE68 genes. This was the clade with the most isolates, so that might explain the higher number of variants, all of which are nonsynonymous. However, Clade VI also presented the most clusters/subclusters compared to Clades I–V and VII. Studies suggest that the secA2 secretion system, most likely through sodA, inhibits extrinsic and intrinsic apoptosis pathways induced upon mycobacterial macrophages infection and have proposed that it prevents apoptosis and antigen-specific CD8+ cross-presentation, as well as altering the intracellular trafficking in favor of the bacteria [
77].
With respect to antimicrobial resistance,
M. bovis is intrinsically resistant to pyrazinamide (PZA) [
80], so this mutation was expected in all of the genomes. The most common mutation is a CAC > GAC nonsynonymous substitution in the gene
pncA that causes a change of histidine to aspartic acid. In this analysis, we detected a second mutation in one genome, a CTC > CGC nonsynonymous substitution that changes leucine (L) for arginine (R) in the
panD gene [
77,
81]. To our knowledge, this is the only described occurrence of this mutation in a wild-type
M. bovis strain. While it is not rare in
M. tuberculosis, further studies targeting this mutation in
M. bovis might provide more insight into this mechanism of resistance in this species. Similarly, isoniazid (INH) resistance, though less common, has been demonstrated from animal-recovered isolates and is a cause for concern [
82,
83,
84,
85]. INH resistance can be associated with mutations in several genes (
fabG1,
inhA,
iniC,
kasA, and
katG); here, the five genomes with this predicted mutation involved
fabG1. A previous study identified the
katG mutation only among a set of 2074 genomes of
M. bovis lineages in the Americas [
85]. Another study previously performed in Catalonia also identified polyresistant
M. bovis infection in human and sympatric sheep [
86], highlighting the importance of a One Health approach in TB control to prevent the spread of TB between humans and livestock, considering the drug resistance in strains circulating among livestock.