1. Introduction
Klebsiella oxytoca is a Gram-negative bacterium that ubiquitously resides in water and soil environments [
1,
2]. The bacterium is also found in the nasopharynx and intestine of healthy individuals in humans [
3].
K. oxytoca was originally isolated from a yogurt specimen in 1886 and named
Bacillus oxytoca. In 1963, this organism was taxonomically classified as a member of the genus
Klebsiella [
4]. Recently, it has been reported that
K. oxytoca is the causative agent of necrotizing enterocolitis [
5], hemorrhagic colitis [
6], sepsis-associated purpura fulminans [
7], and infective endocarditis in humans [
8], arguing that
K. oxytoca is an emerging pathogen [
1].
Outbreak of
K. oxytoca in hospitals involved strains with extended-spectrum beta-lactamases and carbapenemases, which consequently lead to therapeutic problems [
9]. Multidrug resistant (MDR)
K. oxytoca isolates of clinical origins have been reported, particularly in developing nations. For instance, Gunduz et al. isolated
K. oxytoca strains (
n = 33) from different clinical samples collected in the Department of Pediatrics of Turgut Ozal University in Ankara, Turkey from September 2014 to April 2016. High percentages of these
K. oxytoca strains were resistant to ciprofloxacin (CIP) (81%), ceftazidime (CAZ) (72%), amikacin (AK) (71%), gentamicin (GEN) (71%), and imipenem (IPM) (59%) [
10]. Recently, Alemayehu et al. reported that
K. oxytoca strains (
n = 9) isolated from clinical specimens in Sidama, Ethiopia from February 13 to June 7 of 2020 displayed resistance to ampicillin (AMP) (100%), CAZ (100%), CIP (100%), cefuroxime (CRX) (100%), cefotaxime (CTX) (100%), meropenem (MER) (11.1%), GEN (77.8%), cotrimoxazole (COT) (71.4%), nitrofurantoin (NIT) (66.7%), and piperacillin-tazobactam (PIT) (42.9%) [
11]. Nevertheless, the available literature on
K. oxytoca isolates of environmental origins (e.g., aquatic environments and animals) is rare. To our knowledge, few studies have been conducted in this research field. Mann et al. reported on one
K. oxytoca strain, isolated from a wastewater treatment plant in the North West Province of South Africa, showing resistance to oxytetracycline (OT) [
12]. Recently, Håkonsholm et al. collected 476 batches of marine bivalve mollusks along the Norwegian coast from September 2019 to March 2020, including
Mytilus edulis,
Crassostrea gigas,
Pecten maximus,
Modiolus modiolus,
Arctica islandica,
Cerastoderma edule,
Politapes rhomboides,
Mya arenaria, and
Strongylocentrotus droebachiensis. A total of 41
K. oxytoca strains were isolated from
M. edulis and
C. gigas, 39 of which were resistant to AMP [
2]. The presence of
K. oxytoca in aquatic products poses an emergent threat to food safety systems and the public health.
The pollution of aquatic environments by toxic heavy metals has also given rise to one of the most important ecological and organismic problems [
13,
14]. The heavy metals accumulated in consumable aquatic animals may lead to serious hazards to humans through the food chain include cadmium (Cd), chromium (Cr), mercury (Hg), nickel (Ni), and lead (Pb) [
15]. They increase the selection of antibiotic resistance of bacteria to a certain extent, and vice versa [
16]. Heavy metals in the water, in the sediments of rivers, lakes, and oceans, and in fish farming environments worldwide have been reported, particularly in developing countries [
17]. For example, Ni et al. recently reported higher positive sample rates (PSR) of the heavy metals copper (Cu) (100%), Hg (100%), Pb (77.4%), and Cd (34.0%) in aquatic product samples (
n = 108) collected in Shanghai, China in the summer of 2018 and 2019, none of which exceeded individual maximum residue limits (MRLs) [
18]. Notably, Alboghobeish et al. reported one
K. oxytoca strain, ATHA6, isolated from industrial wastewater of Isfahan in Iran, showing resistance to Ni
2+ (24 mM) [
19].
China is the largest producer, consumer, and exporter of aquatic products in the world, accounting for about 60% (65,490,200 tons) of the global amount in 2020 (National Bureau of Statistics,
http://www.stats.gov.cn/, accessed on 20 August 2021). In the present study, we conducted a survey on a larger scale to investigate
K. oxytoca contamination in 41 species of commonly consumed aquatic animals, including 21 species of mollusks, 17 species of fish, and 3 species of crustaceans (
Supplementary Material: Table S1). The samples were collected from two large fish markets in July, August, and September of 2018 and 2019 in Shanghai [
18], the largest city for aquatic product transportation and consumption in China. To our knowledge,
K. oxytoca has not ever been detected in these 40 species of aquatic animals. The results in this study provide data to fill gaps for the risk assessment of
K. oxytoca in aquatic products.
2. Results
2.1. Prevalence of K. oxytoca in 41 Species of Consumable Aquatic Animals
A total of 567 red, viscous, and single colonies grown on selective MacConkey Inositol Adonitol Carbenicillin (MIAC) agar plates were randomly picked out, which were recovered from 14 of the 41 species of consumable aquatic animals. Approximately 22.0% (125 of the 567 colonies) were identified as
K. oxytoca by matrix-assisted laser desorption/ionization time of flight mass spectrometry (MALDI-TOF/MS) analysis. Moreover, the 125
K. oxytoca strains were detected positive in the capsular staining but negative in Gram’s staining and the dynamic tests. The results were confirmed by 16S rRNA gene sequencing and analysis. Among the 125
K. oxytoca isolates, approximately 88.0% (
n = 110), 11.2% (
n = 14), and 0.80% (
n = 1) were recovered from the mollusk, fish, and crustacean samples, respectively (
Table S1). Approximately 77.6% (
n = 97) and 22.4% (
n = 28) of the isolates were derived from 11 and 3 species of seawater and freshwater animal samples, respectively. Remarkably, most (96.8%,
n = 121) of the
K. oxytoca isolates were recovered from benthic aquatic animals.
K. oxytoca was present in 14 species of aquatic animals, including 10 species of mollusks:
Anodonta woodiana (
n = 21),
Babylonia areolata (
n = 5),
Cipangopaludina cahayensis (
n = 1),
Haliotis rubra (
n = 10),
Mactra antiquata (
n = 1),
Mytilus edulis (
n = 2),
Neptunea cumingi Crosse (
n = 31),
Scapharca subcrenata (
n = 5),
Sinonovacula constricta (
n = 6),
Tegillarca granosa (
n = 28); 3 species of fish:
Blotchy rock cod,
Carassius auratus (Crucian)
, and
Carassius auratus (Ditrema temmincki Bleeker); and one species of crustacean:
Procambarus clarkii. Additionally, the other 27 species of aquatic animals were absent from
K. oxytoca (
Table S1).
The 125 K. oxytoca isolates were recovered from aquatic animal products originating from four provinces and one city located along the East China Sea, one of the major fishing grounds along China’s coast, which encompasses the Fujian, Jiangsu, Shandong, and Zhejiang provinces, as well as Shanghai City in China. The highest abundance of K. oxytoca isolates were observed in the samples originating from Zhejiang Province (32.8%, 41 of the 125 isolates), followed by Fujian Province (25.6%, 32/125), Shandong Province (20.8%, 26/125), and Shanghai City (19.2, 24/125). Only a few isolates (1.6%, 2/125) originated from Jiangsu Province. Approximately 91.2% of the 125 K. oxytoca were isolated from the samples collected in the Luchao Port Aquatic Market, and 8.8% from the Jiangyang Aquatic Market in Shanghai, China.
2.2. Virulence Associated-Genes in the K. oxytoca Isolates
All of the 125
K. oxytoca isolates were detected negative for virulence-associated genes
mviM,
tisB, and
yqgB by PCR assay. Nevertheless, higher incidence of the genes encoding virulence determinants
brkB (73.6%),
cdcB (66.4%),
pduV (64.8%), and
virk (63.2%) were observed among the isolates. Moreover,
relE (43.2%),
symE (37.6%), and
vagC (23.2%) genes were also detected positive among the isolates. These amplified genes were confirmed by DNA sequencing and analysis, and the obtained sequences were deposited in the GenBank database under the accession numbers MW380322 to MW380328 (
Table S2).
The K. oxytoca isolates recovered from the three types of aquatic products had different virulence-associated gene profiles. Notably, high percentages of the brkB, cdcB, pduV genes were observed in the K. oxytoca isolates recovered from the mollusk (72.7–64.5%) and fish samples (78.6–57.1%). Meanwhile, relatively higher detection frequencies of the relE, and symE genes were found in the mollusks (44.5%, and 40.9%) than in the fish (28.6%, and 7.1%). The vagC and virk genes were only present in the isolates recovered from these two types of aquatic products, but with higher incidence from the mollusks (23.6%, 70.0%) than in the fish (21.4%, 14.3%). Additionally, the isolate (K. o- P. clarkii 8-1-12-7) from the crustacean (P. clarkia) carried the brkB, cdcB, relE, pduV, and symE genes (Figure not shown).
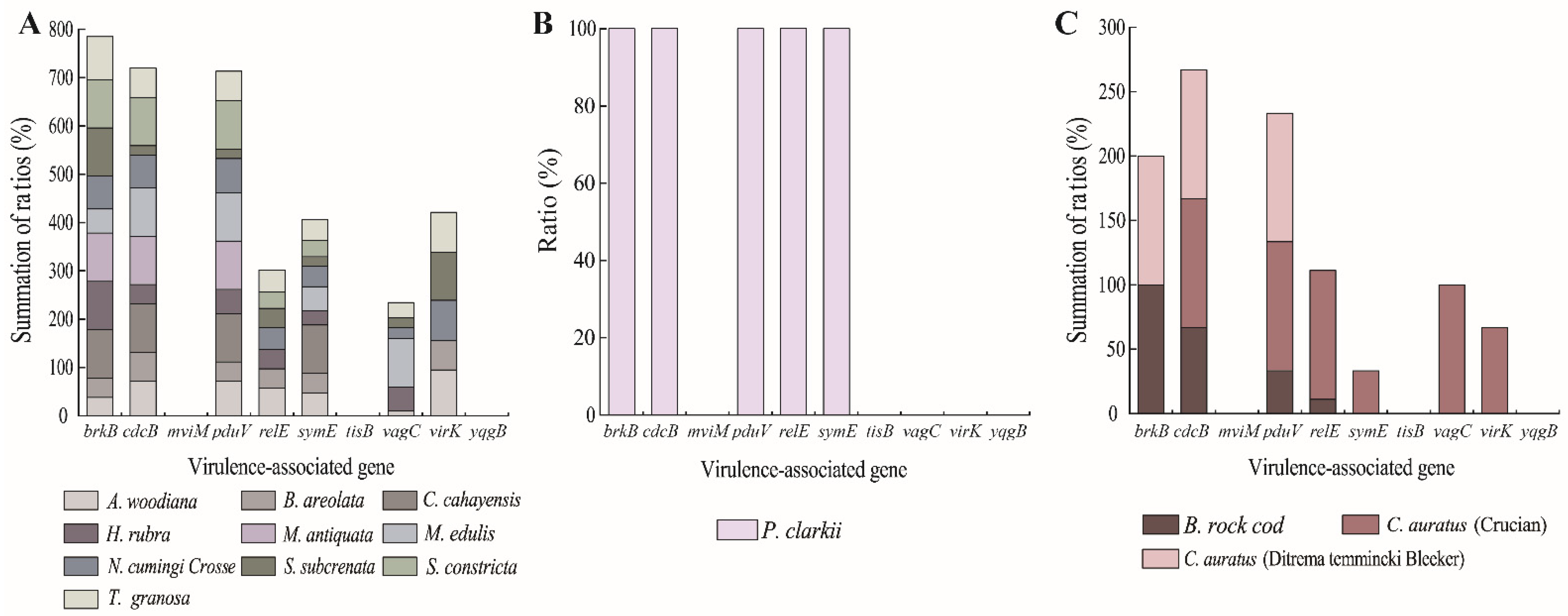
Based on the limited numbers of
K. oxytoca isolates recovered from the 14 species of aquatic animals, different virulence-associated gene profiles were also observed (
Figure 1A–C). For example, six isolates originated from
A. woodiana,
N. cumingi Crosse,
S. subcrenata, and
T. granosa had the maximum number (
n = 7,
brkB+/
cdcB+/
pduV+/
relE+/
symE+/
vagC+/
virk+) of the virulence-associated genes tested. Conversely, 17 isolates from the 14 species of aquatic animals carried the smallest number of these genes (
n = 1,
brkB+ or
virk+).
The 125
K. oxytoca isolates harbored 41 different virulence gene profiles (
Table S3). Among these, the c
dcB+/
pduV+/
relE+/
symE+/
virk+ profile was the most predominant (11.2%,
n = 14), followed by
BrkB+/Virk+ (10.4%,
n = 13), and
BrkB+ (9.6%,
n = 12). In contrast, one isolate (
K. o-
A. woodiana 8-1-8-11) did not harbor any of the virulence-associated genes tested (
Table S3).
2.3. Antibiotic Resistance of the K. oxytoca Isolates
The susceptibility of the 125
K. oxytoca isolates to nine commonly used antibiotics was determined, and the resulting data are illustrated in
Figure 2. Approximately 28% of the
K. oxytoca isolates were susceptible to all the antibiotics evaluated. Moreover, all the isolates were sensitive to imipenem (IPM) (100%) and meropenem (MEM) (100%), and most isolates were also sensitive to norfloxacin (NOR) (98.4%), GEN (96.8%), chloramphenicol (CHL) (93.6%), tetracycline (TET) (92.0%), CIP (87.2%) and kanamycin (KAN) (56.0%). Conversely, sulphamethoxazole-trimethoprim (SXT) resistance was the most predominant (56.0%) among the
K. oxytoca isolates, followed by CHL (6.4%), TET (5.6%), and KAN (3.2%). Approximately 12.0% and 40.8% of the isolates also exhibited intermediate susceptibility to CIP and KAN, respectively. The resistance trend of the 125
K. oxytoca was SXT > CHL > TET > KAN > CIP = GEN = NOR > MEM = IPM (
Figure 2).
The K. oxytoca isolates recovered from the 3 types of aquatic products had different antibiotic resistance profiles (Figure not shown). Higher rates of resistance to SXT were observed in the isolates recovered from the mollusks (60.9%, 67/110) than those from the fish (21.4%, 3/14) and crustacean (0%, n = 1). The CHL, TET, KAN, and GEN resistance were solely present in the isolates from the mollusks (7.3%, 6.4%, 3.6%, 0.9%), whereas CIP and NOR resistance were only found in the isolates derived from the fish (7.1%, and 7.1%) (Figure not shown).
Additionally, different antibiotic resistance profiles were also observed among the
K. oxytoca isolates recovered from the 14 species of aquatic animals (
Figure 3A,B). Remarkably, resistance to SXT was prevalent among the isolates from 10 of the 14 species, except
C. cahayensis,
C. auratus (Ditrema temmincki Bleeker),
M. antiquate, and
P. clarkii. For example, all isolates (
n = 2) recovered from
M. edulis were resistant to SXT, and more than half of the isolates from
A. woodiana (61.9%, 13/21),
B. areolate (60%, 3/5), and
S. constricta (50%, 3/6) showed resistance to SXT. Resistance to CHL was present in a few isolates originating from
H. rubra (1 of 10 isolates),
M. edulis (2 of 2 isolates)
, N. cumingi Crosse (4 of 31 isolates), and
T. granosa (1 of 28 isolates). One isolate (
K. o-
C. auratus 8-11-1) from
C. auratus (Crucian) showed resistance to CIP and NOR, while GEN resistance was solely observed in one isolate (
K. o-
S. subcrenata 8-2-11) from
S. subcrenata. Additionally, none of the isolates from
C. cahayensis (
n = 1),
C. auratus (Ditrema temmincki Bleeker) (
n = 2),
M. antiquata (
n = 1), and the crustacean
P. clarkii (
n = 1) was resistant to any of the nine antibiotics evaluated in this study.
2.4. MDR Phenotypes of the K. oxytoca Isolates
Approximately 8.0% (n = 10) of the K. oxytoca isolates had MDR phenotypes showing resistance to two or more antimicrobial agents. The isolates originating from the mollusks had the highest occurrence of MDR phenotypes (8.2%, 9/110), followed by the fish (7.1%, 1/14), and the crustacean isolates (0.0%, 0/1). Moreover, the MDR isolates were recovered from 6 of the 14 species of aquatic animals, including M. edulis (n = 2), C. auratus (Crucian) (n =1), S. subcrenata (n = 1), N. cumingi Crosse (n = 4), H. rubra (n = 1), and T. granosa (n = 1). Conversely, no MDR strain was found in the other 8 species of aquatic animals.
The multiple antimicrobial resistance index (MARI) values of the 125 K. oxytoca isolates ranged from 0.44 to 0.00, which indicated varying degrees of exposure to the nine antibiotics evaluated. The mean MARI values of the isolates originating from the mollusks, fish, and crustacean were 0.21, 0.15, and 0, respectively. Among the 14 species of aquatic animals, the maximum MARI value was found from the isolates recovered from N. cumingi Crosse (0.44), and H. rubra (0.44), followed by C. auratus (Crucian) (0.33), M. edulis (0.33), T. granosa (0.33), and S. subcrenata (0.22), whereas the isolates from the other 8 species had smaller MARI values (≤0.11). Notably, two isolates (K. o- H. rubra 8-2-2-11 and K. o- N. cumingi Crosse 8-6-19), originating from H. rubra and N. cumingi Crosse, respectively, had the largest MARI value of 0.44, showing resistance to four of the nine antibiotics tested.
2.5. Heavy Metal Tolerance of the K. oxytoca Isolates
Tolerance of the 125
K. oxytoca isolates to 8 heavy metals was determined (
Table S4). The maximum observed values of minimal inhibitory concentrations (MICs) among the
K. oxytoca isolates were 3200 μg/mL for Cr
3+, Pb
2+, Mn
2+, and Cu
2+; 1600 μg/mL for Ni
2+, and Zn
2+; 800 μg/mL for Cd
2+; and 50 μg/mL for Hg
2+, when compared with the quality control strain
E. coli K12. Tolerance to Cu
2+ and Pb
2+ were most prevalent among the isolates (84.8%, 80.8%), followed by Cr
3+ (66.4%), Zn
2+ (66.4%), Hg
2+ (49.6%), Mn
2+ (11.2%), Cd
2+ (9.6%), and Ni
2+ (0.8%). The tolerance trend of the 125
K. oxytoca isolates to heavy metals was Cu
2+ > Pb
2+ > Cr
3+ = Zn
2+ > Hg
2+ > Cd
2+ > Mn
2+ > Ni
2+.
K. oxytoca isolates recovered from the three types of aquatic products had different heavy metal tolerance profiles. The majority of the isolates from all three types of aquatic products were tolerant to Cu2+ (100% to 84.5%), Hg2+ (100% to 48.2%), Pb2+ (100% to 79.1%) and Zn2+ (100% to 61.8%). Tolerance to Cr3+ was prevalent among the isolates from the fish (100%) and mollusks (62.7%), whereas lower percentages of Cd2+ tolerance were found in the isolates from the mollusks (10.0%) and fish (7.1%). Tolerance to Mn2+ (12.7%) and Ni2+ (0.9%) were solely observed in the isolate of the mollusks (Figure not shown).
Different heavy metal tolerance profiles were also found among the
K. oxytoca isolates in the 14 species of aquatic animals (
Figure 4A–C). The
K. oxytoca isolates recovered from
S. constricta were tolerant to all eight of the heavy metals tested, followed by
T. granosa, and
H. rubra (7 heavy metals);
S. subcrenata, and
C. auratus (Crucian) (6 heavy metals);
C. auratus (Ditrema temmincki Bleeker),
B. areolata,
B. rock cod, and
N. cumingi Crosse (5 heavy metals). Tolerance to Ni
2+ was only found in one isolate from
S. constricta (16.7%, 1/6).
2.6. Genetic Diversity of the K. oxytoca Isolates
The enterobacterial repetitive intergenic consensus-PCR (ERIC-PCR) was used to analyze the genetic diversity of the 125
K. oxytoca isolates recovered from the 14 species of aquatic animals. The obtained genome fingerprinting profiles comprised various numbers of DNA bands mainly ranging from 100 to 1000 bp (
Figure 5a,b). Based on the fingerprinting profiles, all the isolates were classified into 108 different ERIC-genotypes, 73.2% of which were assigned as singletons (
n = 79). Approximately 36.7% (
n = 29), 30.4% (
n = 24), and 25.3% (
n = 20) of these singletons were derived from the mollusks
N. cumingi Crosse,
T. granosa, and
A. woodiana, respectively. The UPGMA algorithm grouped all the 108 ERIC genotypes into 14 distinct clusters (clusters I to XIV) at a 32.0% similarity cut-off level (
Figure 5). Approximately 14.4% (
n = 18) of the
K. oxytoca isolates were classified into the largest cluster, II, followed by 11.2% (
n = 14), 9.6% (
n = 12), and 8.0% (
n = 10) into clusters VIII, III, and I, respectively. The remaining isolates (56.8%) fell into clusters III–V, VII, and IX–XIV (7.2% to 0.8%). Most isolates had a similarity coefficient of 30.0–85.0%, and a Simpson’s diversity index of 0.8485. These results demonstrate the considerable genetic diversity of the 125
K. oxytoca isolates recovered from the 14 species of aquatic animals.
Approximately 23.2% (n = 29) of the 125 K. oxytoca isolates shared 12 ERIC-genotypes. For example, four isolates with the ERIC-genotype K. o- 00044 were derived from S. constricta, suggesting near-present relatives or clonal relatedness. Additionally, there were four ERIC-genotypes (K. o- 00011, K. o- 00071, K. o- 00083, and K. o- 00108) containing the isolates derived from different species of aquatic animals. For example, the two isolates K. o-H. rubra 8-2-2-16, and K. o- S. constricta 8-2-3-1, which share the same ERIC-genotype K. o- 00071, were recovered from H. rubra and S. constricta, respectively.
2.7. Comparison of the MDR and Heavy Metal Tolerance
To get insights into co-selection between antibiotics and heavy metals in
K. oxytoca, we further analyzed the 10 isolates with MDR phenotypes by phylogenetic analysis (
Figure 6). The resulting data revealed the genetic diversity of the MDR
K. oxytoca isolates with a Simpson’s index of 0.8556. Moreover, the MDR isolates belonging to 8 ERIC-genotypes were classified into 5 distinct clusters (clusters A–E) (
Figure 6).
Most of the MDR K. oxytoca isolates (90.0%, n = 9) were tolerant to two or more heavy metals. Specifically, one K. oxytoca isolate (K. o- N. cumingi crosse 8-1-6-19) displayed resistance to 4 antibiotics (CHL/SXT/KAN/TET), and was also tolerant to 3 heavy metals (Pb2+/Hg2+/Zn2+). Another K. oxytoca isolate (K. o- H. rubra 8-2-2-11) was resistant to 4 antibiotics (CHL/SXT/KAN/TET) and 6 heavy metals (Cr3+/Cu2+/Hg2/Mn2+/Pb2+/Zn2+). These results provided direct evidence for the cross-resistance between the MDR and heavy metals in the K. oxytoca isolates originating from consumable aquatic animals.
3. Discussion
K. oxytoca is the second most common
Klebsiella species after
K. pneumoniae [
5]. Nevertheless, systematic research on
K. oxytoca is still in its infancy, and current literature on
K. oxytoca in aquatic products is rare [
1,
20]. Recently, Håkonsholm et al. reported 40
K. oxytoca strains isolated from
M. edulis and one from
C. gigas [
2]. In this study, we surveyed
K. oxytoca contamination in 41 species of commonly consumed aquatic animal products sold in Shanghai in July, August, and September of 2018 and 2019.
K. oxytoca was for the first time isolated from 14 species of aquatic animals, including 10 species of mollusks:
A. woodiana,
B. areolata,
C. cahayensis,
H. rubra,
M. antiquata,
M. edulis,
N. cumingi Crosse,
S. subcrenata,
S. constricta and
T. granosa; 3 species of fish:
B. rock cod,
C. auratus (Crucian), and
C. auratus (Ditrema temmincki Bleeker); and one species of crustacean:
P. clarkii. Our data also provided the first experimental evidence for high detection frequencies of
K. oxytoca in benthic aquatic animals, such as
N. cumingi crosse,
T. granosa, and
A. woodiana. These results suggested a potential health risk of the bacterial transmission to communities through the aquatic animals.
The cytotoxin-producing
K. oxytoca has recently been identified as a new candidate etiologic agent in the pathogenesis of necrotizing enterocolitis [
5]. In fact, information in genes associated with the virulence of
K. oxytoca still remains to be discovered. Previous studies have indicated that the genes encoding adhesins, siderophores, and invasins of the genus
Klebsiella were associated with deleterious traits [
21]. In this study, virulence-related genes (
brkB,
cdcB,
pduV,
relE,
symE,
vagC, and
virK) in the 125
K. oxytoca isolates recovered from 14 species of aquatic animals were detected simultaneously. High incidence of the
brkB (73.6%),
cdcB (66.4%),
pduV (64.8%), and
virk (63.2%) genes was observed. The
brkB gene encodes the YihY/virulence factor BrkB family protein, while
cdcB,
pduV, and
virK encode a cytotoxin, a type I toxin-antitoxin system hok family toxin, and the virulence factor VirK, respectively [
22,
23]. The RelE toxin in
Escherichia coli was a global inhibitor of translation [
24]. SymE was toxic in the inhibition of protein synthesis and RNA degradation [
25]. The virulence factor VagC was involved in the bacteriocin secretion system and the type II toxin-antitoxin system in Turkish
Salmonella serovar Infantis isolates [
26]. In this study, the virulence-associated genes
mviM, tisB, and
yqgB were absent from the
K. oxytoca isolates. As a virulence gene,
mviM was associated with the exercise, biofilm formation and antimicrobial resistance of
Cronobacter sakazakii [
27]. TisB is a component of the TisB/IstR-1 toxin-antitoxin system in
E.
coli [
28]. The adaptive factor YqgB in
Bacillus thuringiensis facilitated the bacterial colonization of the host [
29]. In this study, the
K. oxytoca isolates recovered from the 3 types and 14 species of aquatic animals harbored different virulence-associated gene profiles. Remarkably, the isolates originating from
A. woodiana,
N. cumingi crosse,
S. subcrenata, and
T. granosa harbored the maximum number (
n = 7) of the virulence-associated genes tested, which suggests a health risk from the potentially virulent
K. oxytoca in these consumable aquatic animals.
The propagation and spread of resistant pathogenic bacteria pose serious threats to the public heath for humans and animals [
30,
31]. It is estimated that antibiotic resistance may lead to 10 million deaths per year by 2050 [
31]. Misuse of antimicrobial agents is the main cause of antibiotic resistance in pathogenic bacteria, particularly in developing nations [
32,
33,
34]. In this study, our results indicate that SXT resistance was the most predominant (56.0%) among the 125
K. oxytoca isolates of aquatic animal origins. SXT-resistant
K. oxytoca isolates in humans has also been reported. For example, Maharjan et al. recently reported that 9.4% of
K. oxytoca strains (
n = 48) isolated from stool specimens of healthy adult volunteers (
n = 510) in Kathmandu in Nepal showed the highest resistance toward SXT (45.8%) [
35]. In this study, incidences of intermediate susceptibility to KAN (40.8%), and CIP (12.0%) were observed, which suggests a potential resistance trend of
K. oxytoca in aquaculture environments.
MARI is commonly used to determine the health risk associated with antibiotic resistance [
17]. In this study, the mean MARI values for
K. oxytoca isolates derived from the mollusks, fish, and crustacean samples were 0.21, 0.15 and 0, respectively. Among the 14 species of aquatic animals, the maximum MARI value was derived from the isolates in
H. rubra (0.44), and
N. cumingi Crosse (0.44). These results suggested likely extended exposure of the mollusks, particularly
H. rubra, and
N. cumingi Crosse, to the antibiotic drugs evaluated in this study. Recently, Ni et al. reported that 10 antibiotics (AMP, CHL, CIP, ENR, OFX, OT, pefloxacin (PEF), sulfadiazine (SDZ), sulfisomidine (SIM), and sulfapyridine (SP)) were detected positive in the 41 species of aquatic animal samples (
n = 108) with an overall detection frequency of 61.3%. The residual CHL (47.2%), AMP (31.1%), and SP (1.9%) exceeded their MRLs [
18], which provided direct evidence for the antibiotic-resistant phenotypes of
K. oxytoca isolates observed in this study.
A high level of the bioaccumulation of toxic heavy metals (e.g., Cd
2+, Cr
3+, Hg
2+, Ni
2+, and Pb
2+) through the food chain is a grave threat to human health, due to their non-degradable nature [
36]. Numerous studies have reported heavy metal residues in various aquatic environments and species of food animals (shellfish, fish, crustaceans, and crabs) sampled worldwide, particularly in developing nations [
13,
14,
15,
18,
37]. In the recent surveys by our research group [
17,
38,
39], the heavy metal-tolerant waterborne pathogen
Vibrio cholerae has been discovered in many species of aquatic animals. In this study, the results indicated that the
K. oxytoca isolates originated from the 3 types and 14 species of aquatic animals had different heavy metal tolerance profiles. Tolerance to Cu
2+ and Pb
2+ was the most prevalent among the isolates (84.8% and 80.8%). Cu is an essential element for various biological functions in many organisms, and it is also a cofactor for hemocyanin in aquatic arthropods and mollusks. However, excessive levels of Cu can be toxic or disrupting metabolic processes [
40]. In this study, our data also revealed a higher detection frequency of Cr
3+ tolerance in the
K. oxytoca isolates from the fish samples than that from the mollusks. Moreover, the isolates recovered from
S. constricta were tolerant to all 8 heavy metals, followed by
H. rubra, and
T. granosa (7 heavy metals);
C. auratus (Crucian), and
S. subcrenata (6 heavy metals);
B. areolata,
B. rock cod, C. auratus (Ditrema temmincki Bleeker), and
N. cumingi Crosse (5 heavy metals). Tolerance to Ni
2+ was only detected in the isolates from
S. constricta. These data suggest serious heavy metal pollution likely occurred in the aquaculture environments, consistent with previous reports [
18,
38,
39]. Recently, Ni et al. reported that the heavy metals Cu, Hg, Pb, and Cd were observed in the 41 species of aquatic animals with PSRs of 100%, 100%, 77.4%, and 34.0%, respectively, none of which exceeded their MRLs [
18], which provided direct evidence for the high incidence of
K. oxytoca tolerance to heavy metals in this study. Additional attention should be paid to the potential health risk of heavy metal pollution in consumable aquatic animals sold in Shanghai, China.
In this study, ten
K. oxytoca isolates had MDR phenotypes, nine of which also exhibited tolerance to two or more than two heavy metals. For example, the
K. oxytoca isolate
K. o- H. rubra 8-2-2-11, showing resistance to the four antibiotics CHL, SXT, KAN, and TET, also tolerated the 6 heavy metals Cr
3+, Cu
2+, Hg
2+, Mn
2+, Pb
2+, and Zn
2+. These results suggested that heavy metal pollution likely co-selected for antibiotic resistance in
K. oxytoca, and vice versa. In our previous research, antibiotic resistance correlating positively with heavy metal resistance was also observed in the
V. cholerae isolates recovered from the 41 species of aquatic animals [
18]. Cross-resistance due to co-selection can be inferred as the most likely mechanism of the rising antibiotic-resistant pathogens [
41].
5. Conclusions
In the present study, we surveyed K. oxytoca contamination in 41 species of consumable aquatic animals sold in July, August, and September of 2018 and 2019 in Shanghai, China, including 21 species of mollusks, 17 species of fish, and 3 species of crustaceans. Among these, 40 species of the bacterium had not been detected previously.. K. oxytoca was for the first time isolated from 14 species, including 10 species of mollusks: A. woodiana, B. areolata, C. cahayensis, H. rubra, M. antiquata, M. edulis, N. cumingi Crosse, S. subcrenata, S. constricta, and T. granosa; 3 species of fish: B. rock cod, C. auratus (Crucian), and C. auratus (Ditrema temmincki Bleeker); and one species of crustacean: P. clarkii. The other 27 species of aquatic animals were without K. oxytoca. Remarkably, K. oxytoca was found in high abundance in benthic animals, such as N. cumingi crosse, T. granosa, and A. woodiana.
None of K. oxytoca isolates (n = 125) harbored the toxin genes mviM, tisB, and yqgB. However, a high occurrence of virulence-associated genes was observed, including brkB (73.6%), cdcB (66.4%), pduV (64.8%), and virk (63.2%). Resistance to SXT (56.0%) was the most predominant among the isolates, followed by CHL (6.4%), TET (5.6%), and KAN (3.2%). Incidence of intermediate susceptibility to KAN (40.8%), and CIP (12.0%) were also observed. Approximately 8.0% of the isolates displayed MDR phenotypes. Meanwhile, high percentages of the isolates were tolerant to the heavy metals Cu2+ (84.8%), Pb2+ (80.8%), Cr3+ (66.4%), Zn2+ (66.4%), and Hg2+ (49.6%). Different virulence and resistance profiles were observed among the K. oxytoca isolates in 3 types and 14 species of aquatic animals. The ERIC-PCR-based genome fingerprinting of the 125 K. oxytoca isolates revealed 108 ERIC genotypes with 79 singletons, which demonstrated the considerable genetic diversity of the isolates.
Overall, the results of this study provided first experimental evidence for the presence of potentially virulent K. oxytoca in 14 species of commonly consumed aquatic animals, which poses a potential threat to food safety systems and the public health. In future research, the molecular mechanisms underlying the cross-resistance between antibiotics and heavy metals should be further examined in the emerging pathogen K. oxytoca.