Elucidating Genetic Diversity in Apricot (Prunus armeniaca L.) Cultivated in the North-Western Himalayan Provinces of India Using SSR Markers
Abstract
:1. Introduction
2. Results
2.1. SSR Genotyping and Genetic Diversity Analysis
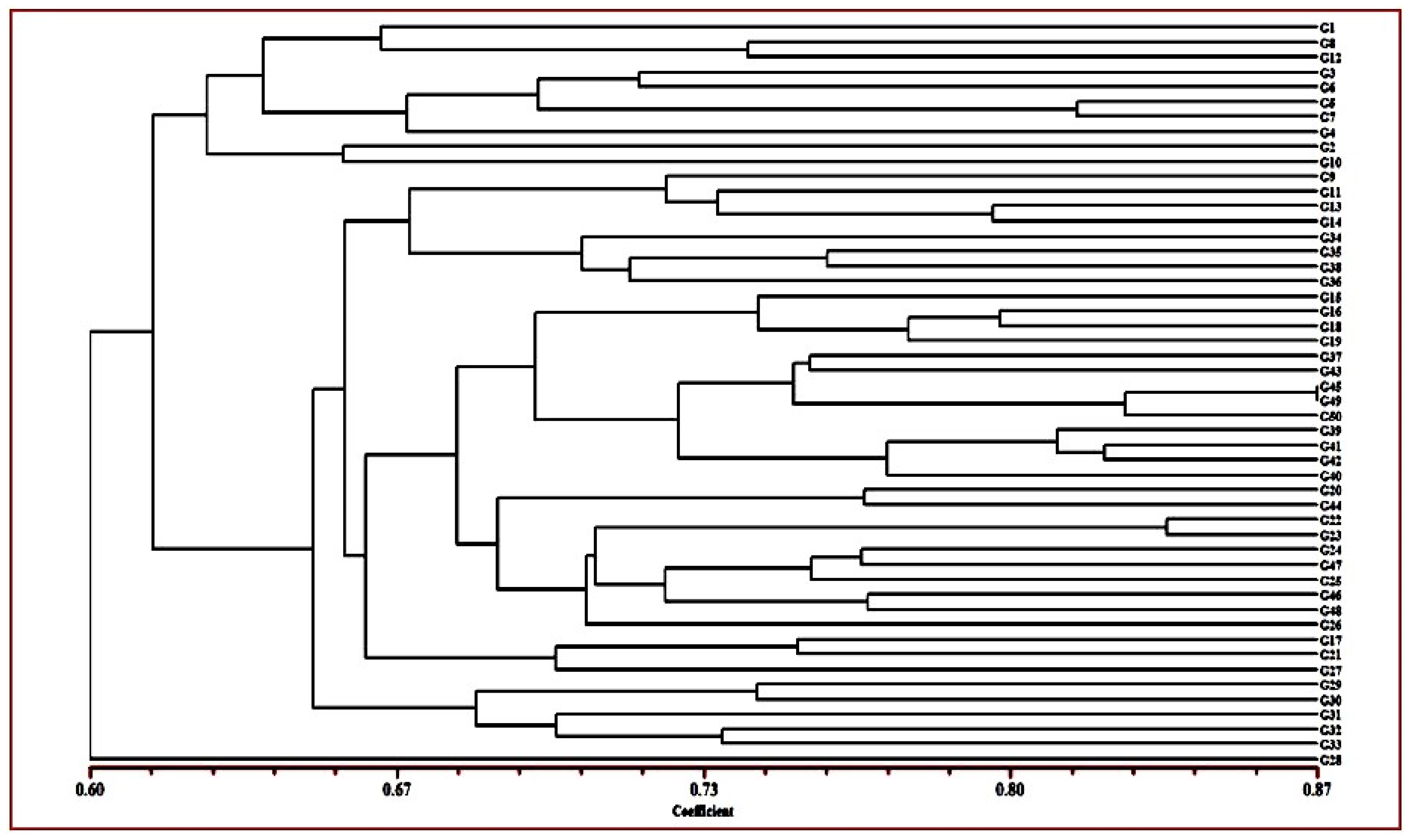
2.2. Cluster and PCoA Analysis
2.3. Population Structure
2.4. AMOVA
3. Discussion
3.1. SSR Genotyping and Genetic Diversity Analysis
3.2. Cluster and PCoA Analysis
3.3. Population Structure
3.4. AMOVA
4. Materials and Methods
4.1. DNA Extraction and Amplification
4.2. Data Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Herrera, S.; Lora, J.; Hormaza, J.; Herrero, M.; Rodrigo, J. Optimizing production in the new generation of Apricot cultivars: Self-incompatibility, S-RNase allele identification, and incompatibility group assignment. Front. Plant Sci. 2018, 27, 527. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bourguiba, H.; Scotti, I.; Sauvage, C.; Zhebentyayeva, T.; Ledbetter, C.; Krska, B.; Remay, A.; Donofrio, C.; Iketani, H.; Christen, D.; et al. Genetic structure of a worldwide germplasm collection of Prunus armeniaca L. reveals three major diffusion routes for varieties coming from the species’ center of origin. Front. Plant Sci. 2020, 25, 638. [Google Scholar] [CrossRef]
- Decroocq, S.; Cornille, A.; Tricon, D.; Babayeva, S.; Chague, A.; Eyquard, P.; Karychev, R.; Dolgikh, S.; Kostritsyna, T.; Liu, S.; et al. New insights into the history of domesticated and wild apricots and its contribution to plum pox virus resistance. Mol. Ecol. 2016, 25, 2712–4729. [Google Scholar] [CrossRef] [PubMed]
- Pedryc, A.; Ruthner, S.; Herman, R.; Krska, B.; Hegedus, A.; Halasz, J. Genetic diversity of apricot revealed by a set of SSR markers from linkage group G1. Sci. Hortic. 2009, 121, 19–26. [Google Scholar] [CrossRef]
- Faostat. 2013. Available online: http://apps.fao.org (accessed on 4 February 2015).
- Zhebentyayeva, T.; Ledbetter, C.; Burgos, L.; Llacer, G. Apricot. In Infruit Breeding; Springer: Boston, MA, USA, 2012; pp. 415–458. [Google Scholar] [CrossRef]
- Martín, C.; Herrero, M.; Hormaza, J. Molecular characterization of apricot germplasm from an old stone collection. PLoS ONE 2011, 25, e23979. [Google Scholar] [CrossRef] [Green Version]
- Zhou, Y.; Zhou, H.; Lin-Wang, K.; Vimolmangkang, S.; Espley, V.; Wang, L.; Allan, C.; Han, Y. Transcriptome analysis and transient transformation suggest an ancient duplicated MYB transcription factor as a candidate gene for leaf red coloration in peach. BMC Plant Biol. 2014, 14, 388. [Google Scholar] [CrossRef] [Green Version]
- Shah, R.A.; Bakshi, P.; Sharma, N.; Jasrotia, A.; Itoo, H.; Gupta, R.; Singh, A. Diversity assessment and selection of superior Persian walnut (Juglans regia L.) trees of seedling origin from North-Western Himalayan region. Resour. Environ. Sustain. 2021, 1, 100015. [Google Scholar] [CrossRef]
- Corrado, G.; Forlani, M.; Rao, R.; Basile, B. Diversity and Relationships among Neglected Apricot (Prunus armeniaca L.) Landraces Using Morphological Traits and SSR Markers: Implications for Agro-Biodiversity Conservation. Plants 2021, 10, 1341. [Google Scholar] [CrossRef]
- Tian-Ming, H.; Xue-Sen, C.; Zheng, X.; Jiang-Sheng, G.; Pei-Jun, L.; Wen, L.; Qing, L.; Yan, W. Using SSR markers to determine the population genetic structure of wild apricot (Prunus armeniaca Prunus armeniaca L.) in the Ily Valley of West China. Genet. Resour. Crop Evol. 2007, 54, 563–572. [Google Scholar] [CrossRef]
- Hagen, L.; Khadari, B.; Lambert, P.; Audergon, J.M. Genetic diversity in apricot revealed by AFLP markers: Species and cultivar comparisons. Theor. Appl. Genet. 2002, 105, 298–305. [Google Scholar] [CrossRef] [PubMed]
- Geuna, F.; Toschi, M.; Bassi, D. The use of AFLP markers for cultivar identification in apricot. Plant Breed. 2003, 122, 526–531. [Google Scholar] [CrossRef]
- Kumar, M.; Mishra, G.P.; Singh, R.; Kumar, J.; Naik, P.K.; Singh, S.B. Correspondence of ISSR and RAPD markers for comparative analysis of genetic diversity among different apricot genotypes from cold arid deserts of trans-Himalayas. Physiol. Mol. Biol. Plants 2009, 15, 225–236. [Google Scholar] [CrossRef] [Green Version]
- Krichen, L.; Martins, J.M.; Lambert, P.; Daaloul, A.; Trifi-Farah, N.; Marrakchi, M.; Audergon, J.M. Using AFLP markers for the analysis of the genetic diversity of apricot cultivars in Tunisia. J. Am. Soc. Hortic. 2008, 133, 204–212. [Google Scholar] [CrossRef]
- Fang, J.; Tao, J.; Chao, C.T. Genetic diversity in fruiting-mei, apricot, plum and peach revealed by AFLP analysis. J. Hortic. Sci. Biotechnol. 2006, 81, 898–902. [Google Scholar] [CrossRef]
- Zargar, S.A.; Wani, A.A.; Saggoo, M.I. Analysis of phenotypic diversity of apricot (Prunus armeniaca L.) accessions from Jammu and Kashmir, India. Plant Genet. Resour. 2021, 19, 203–215. [Google Scholar] [CrossRef]
- Sheikh, Z.N.; Sharma, V.; Shah, R.A.; Sharma, N.; Summuna, B.; Al-Misned, F.A.; El-Serehy, H.A.; Mir, J.I. Genetic diversity analysis and population structure in apricot (Prunus armeniaca L.) grown under north-western himalayas using ISSR markers. Saudi J. Biol. Sci. 2021, 28, 5989–5995. [Google Scholar] [CrossRef]
- Bourguiba, H.; Krichen, L.; Audergon, J.M.; Khadari, B.; Trifi-Farah, N. Impact of mapped SSR markers on the genetic diversity of apricot (Prunus armeniaca L.) in Tunisia. Plant Mol. Biol. Rep. 2010, 28, 578–587. [Google Scholar] [CrossRef]
- Shah, R.A.; Baksi, P.; Jasrotia, A.; Bhat, D.J.; Gupta, R.; Bakshi, M. Genetic diversity of walnut (Juglans regia L.) seedlings through SSR markers in north-western Himalayan region of Jammu. Bangladesh J. Bot. 2020, 49, 1003–1012. [Google Scholar] [CrossRef]
- Romero, C.; Pedryc, A.; Munoz, V.; Llacer, G.; Badenes, M.L. Genetic diversity of different apricot geographical groups determined by SSR markers. Genome 2003, 46, 244–252. [Google Scholar] [CrossRef]
- Hormaza, J.I. Molecular characterization and similarity relationships among apricot (Prunus armeniaca L.) genotypes using simple sequence repeats. Theor. Appl. Genet. 2002, 104, 321–328. [Google Scholar] [CrossRef]
- Sanchez-Perez, R.; Martinez-Gomez, P.; Dicenta, F.; Egea, J.; Ruiz, D. Level and transmission of genetic heterozygosity in apricot (Prunus armeniaca L.) explored using simple sequence repeat markers. Genet. Resour. Crop. Evol. 2006, 53, 763–770. [Google Scholar] [CrossRef]
- Zhebentyayeva, T.; Reighard, G.; Gorina, V.; Abbott, A. Simple sequence repeat (SSR) analysis for assessment of genetic variability in apricot germplasm. Theor. Appl. Genet. 2003, 106, 435–444. [Google Scholar] [CrossRef]
- Li, M.; Zheng, P.; Ni, B.; Hu, X.; Miao, X.; Zhao, Z. Genetic diversity analysis of apricot cultivars grown in China based on SSR markers. Eur. J. Hortic. Sci. 2018, 83, 18–27. [Google Scholar] [CrossRef]
- Zhang, Q.P.; Liu, D.C.; Liu, S.; Liu, N.; Wei, X.; Zhang, A.M.; Liu, W.S. Genetic diversity and relationships of common apricot (Prunus armeniaca L.) in China based on simple sequence repeat (SSR) markers. Genet. Resour. Crop. Evol. 2014, 61, 357–368. [Google Scholar] [CrossRef]
- Hu, X.; Zheng, P.; Ni, B.; Miao, X.; Zhao, Z.; Li, M. Population genetic diversity and structure analysis of wild apricot (Prunus armeniaca L.) revealed by SSR markers in the Tien-Shan mountains of China. Pak. J. Bot. 2018, 50, 609–615. [Google Scholar]
- Bakir, M.; Dumanoglu, H.; Erdogan, V.; Ernim, C.; Macit, T. Characterization of wild apricot (Prunus armeniaca L.) genotypes selected from Cappadocia region (Nevşehir-Turkey) by SSR markers. J. Agric. Sci. 2019, 25, 498–507. [Google Scholar] [CrossRef] [Green Version]
- Yuan, W.Z.; Bai, Y. Analysis of Genetic Diversity in Prunus sibirica L. in Inner Mongolia Using SCoT Molecular Markers. Res. Square 2021, 1–17. [Google Scholar] [CrossRef]
- Wang, L. Genetic Diversity and Population Genetic Structure of Armeniaca Sibirica Clones; Shenyang Agricultural University: Shenyang, China, 2019. [Google Scholar]
- Dirlewanger, E.; Cosson, P.; Tavaud, M.; Aranzana, M.; Poizat, C.; Zanetto, A.; Arus, P.; Laigret, F. Development of microsatellite markers in peach Prunus persica L. Batsch and their use in genetic diversity analysis in peach and sweet cherry (Prunus avium L.). Theor. Appl. Genet. 2002, 105, 127–138. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhang, J.; Sun, H.; Ning, N.; Yang, L. Construction and evaluation of a primary core collection of apricot germplasm in China. Sci. Hortic. 2011, 128, 311–319. [Google Scholar] [CrossRef]
- Aranzana, M.J.; Garcia-Mas, J.; Carbo, J.; Arus, P. Development and variability analysis of microsatellite markers in peach. Plant Breed. 2002, 121, 87–92. [Google Scholar] [CrossRef]
- Ruthner, S.; Pedryc, A.; Kriska, B.; Romero, C. Molecular characterization of apricot (Prunus armenica L.) cultivars using cross species SSR amplification with peach primers. Int. J. Hortic. Sci. 2006, 12, 53–57. [Google Scholar] [CrossRef]
- Dondini, L.; Lain, O.; Geuna, F.; Banfi, R.; Gaiotti, F.; Tartarini, S.; Bassi, D.; Testolin, R. Development of a new SSR-based linkage map in apricot and analysis of synteny with existing Prunus maps. Tree Genet. Genomes 2007, 3, 239–249. [Google Scholar] [CrossRef]
- Wang, Y.J.; Li, X.Y.; Han, J.; Fang, W.M.; Li, X.D.; Wang, S.S.; Fang, J.G. Analysis of genetic relationships and identification of flowering-mei cultivars using EST-SSR markers developed from apricot and fruiting-mei. Sci. Hortic. 2011, 132, 12–17. [Google Scholar] [CrossRef]
- Hagen, L.S.; Chaïb, J.; Fady, B.; Decroocq, V.; Bouchet, J.P.; Lambert, P.; Audergon, J.M. Genomic and cDNA microsatellites from apricot (Prunus armeniaca L.). Mol. Ecol. Notes 2004, 4, 742–745. [Google Scholar] [CrossRef]
- Messina, R.; Lain, O.; Marrazzo, M.T.; Cipriani, G.; Testolin, R. New set of microsatellite loci isolated in apricot. Mol. Ecol. Notes 2004, 3, 432–434. [Google Scholar] [CrossRef]
- Xie, R.; Li, X.; Chai, M.; Song, L.; Jia, H.; Wu, D.; Gao, Z. Evaluation of the genetic diversity of Asian peach accessions using a selected set of SSR markers. Sci. Hortic. 2010, 4, 622–629. [Google Scholar] [CrossRef]
- Bourguiba, H.; Audergon, J.M.; Krichen, L.; Trifi-Farah, N.; Mamouni, A.; Trabelsi, S.; D’Onofrio, C.; Asma, B.M.; Santoni, S.; Khadari, B. Loss of genetic diversity as a signature of apricot domestication and diffusion into the Mediterranean Basin. BMC Plant Biol. 2012, 12, 49. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yilmaz, K.U.; Paydas-Kargi, S.E.; Dogan, Y.; Kafkas, S.A. Genetic diversity analysis based on ISSR, RAPD and SSR among Turkish apricot germplasms in Iran caucasian eco-geographical group. Sci. Hortic. 2012, 138, 138–143. [Google Scholar] [CrossRef]
- Chen, J.; Dong, S.; Zhang, X.; Wu, Y.; Zhang, H.; Sun, Y.; Zhang, J. Genetic diversity of Prunus sibirica L. superior accessions based on the SSR markers developed using restriction-site associated DNA sequencing. Genet. Resour. Crop. Evol. 2021, 68, 615–628. [Google Scholar] [CrossRef]
- Rezaei, M.; Rahmati, M.; Kavand, A.; Hemati, M.; Kazemi, S.R. Screening of some Iranian Commercial Apricot Cultivars by SSR Markers for identification of Synonyms. Int. J. Hortic. Sci. 2021, 35. [Google Scholar] [CrossRef]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software structure: A simulation study. Mol. Ecolo. 2005, 8, 2611–2620. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Groppi, A.; Liu, S.; Cornille, A.; Decroocq, S.; Bui, Q.T.; Tricon, D.; Cruaud, C.; Arribat, S.; Belser, C.; Marande, W.; et al. Population genomics of apricots unravels domestication history and adaptive events. Nat. Commun. 2021, 12, 3956. [Google Scholar] [CrossRef] [PubMed]
- Maghuly, F.; Fernandez, E.B.; Ruthner, S.; Pedryc, A.; Laimer, M. Microsatellite variability in apricots (Prunus armeniaca L.) reflects their geographic origin and breeding history. Tree Genet. Genomes 2005, 4, 151–165. [Google Scholar] [CrossRef]
- Liu, S.; Cornille, A.; Decroocq, S.; Tricon, D.; Chague, A.; Eyquard, J.P.; Liu, W.S.; Giraud, T.; Decroocq, V. The complex evolutionary history of apricots: Species divergence, gene flow and multiple domestication events. Mol. Ecol. 2019, 24, 5299–5314. [Google Scholar] [CrossRef]
- Martinez Mora, C.; Rodríguez, J.; Cenis, J.L.; Ruiz García, L. Genetic variability among local apricots (Prunus armeniaca L.) from the Southeast of Spain. Span. J. Agric. 2009, 7, 855–868. [Google Scholar] [CrossRef] [Green Version]
- Bourguiba, H.; Khadari, B.; Krichen, L.; Trifi-Farah, N.; Mamouni, A.; Trabelsi, S.; Audergon, J.M. Genetic relationships between local North African apricot (Prunus armeniaca L.) germplasm and recently introduced varieties. Sci. Hortic. 2013, 152, 61–69. [Google Scholar] [CrossRef]
- Batnini, M.A.; Krichen, L.; Bourguiba, H.; Trifi-Farah, N.; González, D.R.; Gómez, P.M.; Rubio, M. Comparative analysis of traditional and modern apricot breeding programs: A case of study with Spanish and Tunisian apricot breeding germplasm. Span. J. Agric. 2016, 14, 14. [Google Scholar] [CrossRef] [Green Version]
- Raji, R.; Jannatizadeh, A.; Fattahi, R.; Esfahlani, M.A. Investigation of variability of apricot (Prunus armeniaca L.) using morphological traits and microsatellite markers. Sci. Hortic. 2014, 176, 225–231. [Google Scholar] [CrossRef]
- Sosinski, B.; Gannavarapu, M.; Hager, L.D.; Beck, L.E.; King, G.J.; Ryder, C.D.; Rajapakse, S.; Baird, W.V.; Ballard, R.E.; Abbott, A.G. Characterization of microsatellite markers in peach (Prunus persica L.). Batsch. Theor. Appl. Genet. 2000, 101, 421–428. [Google Scholar] [CrossRef]
- Testolin, R.; Marrazzo, T.; Cipriani, G.; Quarta, R.; Verde, I.; Dettori, M.T.; Pancaldi, M.; Sansavini, S. Microsatellite DNA in peach (Prunus persica L. Batsch) and its use in fingerprinting and testing the genetic origin of cultivars. Genome 2000, 43, 512–520. [Google Scholar] [CrossRef] [PubMed]
- Vavilov, N.I. The origin, variation, immunity and breeding of cultivated plants. LWW. Soil Sci. 1951, 72, 482. [Google Scholar] [CrossRef]
- Vilanova, S.; Soriano, J.M.; Lalli, D.A.; Romero, C.; Abbott, A.G.; Llacer, G.; Badenes, M.L. Development of SSR markers located in the G1 linkage group of apricot (Prunus armeniaca L.) using a bacterial artificial chromosome library. Mol. Ecol. Notes 2006, 3, 789–791. [Google Scholar] [CrossRef]
- Herrera, S.; Rodrigo, J.; Hormaza, J.I.; Lora, J. Identification of self-incompatibility alleles by specific PCR analysis and S-RNase sequencing in apricot. Int. J. Mol. Sci. 2018, 11, 3612. [Google Scholar] [CrossRef] [Green Version]
- Khadivi-Khub, A.; Jafari, H.R.; Zamani, Z. Phenotypic and genotypic variation in Iranian sour and duke cherries. Trees 2013, 27, 1455–1466. [Google Scholar] [CrossRef]
- Gurcan, K.; Ocal, N.; Yılmaz, K.U.; Ullah, S.; Erdogan, A.; Zengin, Y. Evaluation of Turkish apricot germplasm using SSR markers: Genetic diversity assessment and search for Plum pox virus resistance alleles. Sci. Hortic. 2015, 193, 155–164. [Google Scholar] [CrossRef]
- Akpınar, A.E.; Koçal, H.; Ergül, A.; Kazan, K.; Şelli, M.E.; Bakır, M.; Aslantaş, Ş.; Kaymak, S.; Sarıbaş, R. SSR-based molecular analysis of economically important Turkish apricot cultivars. Genet. Mol. Res. 2010, 9, 324–332. [Google Scholar] [CrossRef] [PubMed]
- Dehkordi, M.; Beigzadeh, T.; Sorkheh, K. Novel in silico EST-SSR markers and bioinformatic approaches to detect genetic variation among peach (Prunus persica L.) germplasm. J. For. Res. 2020, 4, 1359–1370. [Google Scholar] [CrossRef]
- Song, Y.; Fan, L.; Chen, H.; Zhang, M.; Ma, Q.; Zhang, S.; Wu, J. Identifying genetic diversity and a preliminary core collection of Pyrus pyrifolia cultivars by a genome-wide set of SSR markers. Sci. Hortic. 2014, 167, 5–16. [Google Scholar] [CrossRef]
- Krichen, H.B.; Jean-Marc, A.; Neila, T.F. Comparative analysis of genetic diversity in Tunisian apricot germplasm using AFLP and SSR markers. Sci. Hortic. 2010, 127, 54–63. [Google Scholar] [CrossRef]
- Zehdi-Azouzi, S.; Cherif, E.; Moussouni, S.; Gros-Balthazard, M.; Abbas, N.S.; Ludena, B.; Castillo, K.; Chabrillange, N.; Bouguedoura, N.; Bennaceur, M.; et al. Genetic structure of the date palm (Phoenix dactylifera) in the old world reveals a strong differentiation between eastern and western populations. Ann. Bot. 2015, 116, 101–112. [Google Scholar] [CrossRef] [Green Version]
- Haouane, H.; Bakkali, A.; Moukhli, A.; Tollon, C.; Santoni, S.; Oukabli, A.; El Modafar, C.; Khadari, B. Genetic structure and core collection of the world olive germplasm bank of marrakech towards the optimised management and use of mediterranean olive genetic resources. Genetica 2011, 139, 1083–1094. [Google Scholar] [CrossRef] [Green Version]
- Pereira-Dias, L.; Vilanova, S.; Fita, A.; Prohens, J.; Rodríguez-Burruezo, A. Genetic diversity, population structure, and relationships in a collection of pepper (Capsicum spp.) landraces from the Spanish centre of diversity revealed by genotyping-by-sequencing (GBS). Hortic. Res. 2019, 6, 54. [Google Scholar] [CrossRef] [Green Version]
- Martinez-Gomez, P.; Arulsekar, S.; Potter, D.; Gradziel, T.M. Relationships among peach, almond, and related species as detected by simple sequence repeat markers. J. Am. Soc. Hortic. 2003, 128, 667–671. [Google Scholar] [CrossRef] [Green Version]
- Vendramin, G.G.; Anzidei, M.; Madaghiele, A.; Bucci, G. Distribution of genetic diversity in Pinus pinaster Ait. as revealed by chloroplast microsatellites. Theor. Appl. Genet. 1998, 97, 456–463. [Google Scholar] [CrossRef]
- Doyle, J.J.; Doyle, J.L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 1987, 19, 11–15. [Google Scholar]
- Dettori, M.T.; Micali, S.; Giovinazzi, J.; Scalabrin, S.; Verde, I.; Cipriani, G. Mining microsatellites in the peach genome: Development of new long-core SSR markers for genetic analyses in five Prunus species. Springer Plus 2015, 4, 337. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, M.P.; Du, H.Y.; Zhu, G.P.; Fu, D.L.; Tana, W.Y. Genetic diversity analysis of sweet kernel apricot in China based on SSR and ISSR markers. Genet. Mol. Res. 2015, 14, 9722–9729. [Google Scholar] [CrossRef]
- Lambert, P.; Hagen, L.S.; Arus, P.; Audergon, J.M. Genetic linkage maps of two apricot cultivars (Prunus armeniaca L.) compared with the almond Texas×peach Earlygold reference map for Prunus. Theor. Appl. Genet. 2004, 108, 1120–1130. [Google Scholar] [CrossRef]
- Nei, M. Analysis of gene diversity in subdivided populations. Proc. Natl. Acad. Sci. USA 1973, 70, 3321–3323. [Google Scholar] [CrossRef] [Green Version]
- Shannon, C.E. A mathematical theory of communication. ACM SIGMOBILE Mob. Comput. Commun. Rev. 2001, 5, 3–55. [Google Scholar] [CrossRef]
- Yeh, F.C.; Yang, R.C.; Boyle, T.B.; Ye, Z.H.; Mao, J.X. POPGENE, the User-Friendly Shareware for Population Genetic Analysis; Molecular Biology and Biotechnology Centre, University of Alberta: Edmonton, AB, Canada, 1997; Volume 10, pp. 295–301. [Google Scholar]
- Rohlf, F.J. NTSYS-pc Version. 2.02i Numerical Taxonomy and Multivariate Analysis System; Applied Biostatistics Inc.: Setauket, NY, USA, 1997. [Google Scholar]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [CrossRef] [PubMed]
- Earl, D.A. Structure harvester: A website and program for visualizing STRUCTURE output and implementing the evanno method. Conserv. Genet. Resour. 2012, 2, 359–361. [Google Scholar] [CrossRef]
- Jakobsson, M.; Rosenberg, N.A. CLUMPP: Cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 2007, 23, 1801–1806. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rosenberg, N.A. Distruct: A program for the graphical display of population structure. Mol. Ecol. Notes 2004, 1, 137–148. [Google Scholar] [CrossRef]
- Peakall, R.O.; Smouse, P.E. Genalex 6: Genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes 2006, 1, 288–295. [Google Scholar] [CrossRef]




| Marker | No. of Alleles | PIC Value | Obs Hom | Obs Het | Exp Hom | Exp Het | Ave Het | Ne | I | Fst |
|---|---|---|---|---|---|---|---|---|---|---|
| RPPG1-017 | 6 | 0.374 | 0.8333 | 0.1667 | 0.7193 | 0.2807 | 0.2778 | 1.3846 | 0.4506 | 0.05 |
| RPPG1-026 | 6 | 0.350 | 0.5833 | 0.4167 | 0.5088 | 0.4912 | 0.4861 | 1.9459 | 0.6792 | 0.02 |
| RPPG1-032 | 5 | 0.362 | 0.5417 | 0.4583 | 0.5026 | 0.4974 | 0.4922 | 1.9692 | 0.6853 | 0.04 |
| RPPG1-037 | 3 | 0.283 | 0.5208 | 0.4792 | 0.5055 | 0.4945 | 0.4894 | 1.9584 | 0.6825 | 0.07 |
| RPPG1-041 | 2 | 0.196 | 0.7083 | 0.2917 | 0.5509 | 0.4491 | 0.4444 | 1.8 | 0.6365 | 0.02 |
| RPPG2-011 | 1 | 0.104 | 0.4167 | 0.5833 | 0.5825 | 0.4175 | 0.4132 | 1.7041 | 0.6036 | 0 |
| RPPG2-022 | 1 | 0.104 | 0.8542 | 0.1458 | 0.8634 | 0.1366 | 0.1352 | 1.1563 | 0.2611 | 0.08 |
| RPPG3-026 | 2 | 0.186 | 0.2917 | 0.7083 | 0.4982 | 0.5018 | 0.4965 | 1.9862 | 0.6897 | 0.01 |
| RPPG4-059 | 4 | 0.350 | 0.6458 | 0.3542 | 0.5125 | 0.4875 | 0.4824 | 1.9321 | 0.6755 | 0.01 |
| RPPG4-067 | 2 | 0.203 | 0.9375 | 0.0625 | 0.5318 | 0.4682 | 0.4633 | 1.8633 | 0.656 | 0.04 |
| RPPG4-077 | 3 | 0.284 | 0.2917 | 0.7083 | 0.5377 | 0.4623 | 0.4575 | 1.8432 | 0.65 | 0 |
| RPPG4-084 | 2 | 0.194 | 0.7083 | 0.2917 | 0.5263 | 0.4737 | 0.4688 | 1.8824 | 0.6616 | 0.06 |
| RPPG4-091 | 6 | 0.462 | 0.2917 | 0.7083 | 0.4982 | 0.5018 | 0.4965 | 1.9862 | 0.6897 | 0 |
| RPPG5-018 | 4 | 0.326 | 0.4167 | 0.5833 | 0.5825 | 0.4175 | 0.4132 | 1.7041 | 0.6036 | 0 |
| RPPG5-022 | 3 | 0.284 | 0.4792 | 0.5208 | 0.495 | 0.505 | 0.4998 | 1.9991 | 0.6929 | 0.05 |
| RPPG5-023 | 2 | 0.188 | 0.5208 | 0.4792 | 0.5055 | 0.4945 | 0.4894 | 1.9584 | 0.6825 | 0 |
| RPPG5-025 | 1 | 0.105 | 0.7292 | 0.2708 | 0.5055 | 0.4945 | 0.4894 | 1.9584 | 0.6825 | 0.03 |
| RPPG5-030 | 4 | 0.354 | 0.000 | 1.000 | 0.4947 | 0.5053 | 0.500 | 2.000 | 0.6931 | 0.01 |
| RPPG6-009 | 4 | 0.351 | 0.5 | 0.5 | 0.5167 | 0.4833 | 0.4783 | 1.9168 | 0.6713 | 0.02 |
| RPPG6-032 | 4 | 0.354 | 0.5625 | 0.4375 | 0.495 | 0.505 | 0.4998 | 1.9991 | 0.6929 | 0 |
| RPPG6-033 | 2 | 0.190 | 0.7083 | 0.2917 | 0.4947 | 0.5053 | 0.500 | 2.000 | 0.6931 | 0.05 |
| RPPG7-015 | 4 | 0.353 | 0.2708 | 0.7292 | 0.5055 | 0.4945 | 0.4894 | 1.9584 | 0.6825 | 0.02 |
| RPPG7-026 | 4 | 0.354 | 0.5417 | 0.4583 | 0.643 | 0.357 | 0.3533 | 1.5463 | 0.5383 | 0.04 |
| RPPG7-032 | 3 | 0.279 | 0.875 | 0.125 | 0.4956 | 0.5044 | 0.4991 | 1.9965 | 0.6923 | 0 |
| RPPG8-007 | 1 | 0.107 | 0.875 | 0.125 | 0.5026 | 0.4974 | 0.4922 | 1.9692 | 0.6853 | 0.03 |
| RPPG8-028 | 3 | 0.260 | 0.1875 | 0.8125 | 0.5125 | 0.4875 | 0.4824 | 1.9321 | 0.6755 | 0.01 |
| Aprigms18 | 5 | 0.412 | 0.3542 | 0.6458 | 0.495 | 0.505 | 0.4998 | 1.9991 | 0.6929 | 0 |
| UDP98-405 | 6 | 0.447 | 0.5833 | 0.4167 | 0.5825 | 0.4175 | 0.4132 | 1.7041 | 0.6036 | 0 |
| UDP98-406 | 6 | 0.462 | 0.375 | 0.625 | 0.5026 | 0.4974 | 0.4922 | 1.9692 | 0.6853 | 0.02 |
| UDP98-409 | 4 | 0.337 | 0.125 | 0.875 | 0.5026 | 0.4974 | 0.4922 | 1.9692 | 0.6853 | 0.05 |
| UDP98-411 | 6 | 0.464 | 0.0833 | 0.9167 | 0.4982 | 0.5018 | 0.4965 | 1.9862 | 0.6897 | 0.05 |
| Pchgms4 | 6 | 0.463 | 0.8333 | 0.1667 | 0.7789 | 0.2211 | 0.2188 | 1.2800 | 0.3768 | 0.01 |
| Pchgms5 | 5 | 0.414 | 1.000 | 0.000 | 0.8114 | 0.1886 | 0.1866 | 1.2295 | 0.3341 | 0.06 |
| Bppct007 | 6 | 0.464 | 0.5833 | 0.4167 | 0.5509 | 0.4491 | 0.4444 | 1.800 | 0.6365 | 0.01 |
| Bppct025 | 2 | 0.193 | 0.375 | 0.625 | 0.5658 | 0.4342 | 0.4297 | 1.7534 | 0.6211 | 0 |
| Bppct030 | 6 | 0.462 | 0.3542 | 0.6458 | 0.5441 | 0.4559 | 0.4512 | 1.8221 | 0.6435 | 0.05 |
| PacA10 | 6 | 0.462 | 0.000 | 1.000 | 0.4947 | 0.5053 | 0.5000 | 2.000 | 0.6931 | 0 |
| PacA18 | 6 | 0.464 | 0.2083 | 0.7917 | 0.4956 | 0.5044 | 0.4991 | 1.9965 | 0.6923 | 0.04 |
| PacA33 | 6 | 0.463 | 0.2292 | 0.7708 | 0.5213 | 0.4787 | 0.4737 | 1.9002 | 0.6667 | 0.01 |
| PacA22 | 6 | 0.460 | 0.125 | 0.875 | 0.4947 | 0.5053 | 0.500 | 2.000 | 0.6931 | 0 |
| PacA26 | 6 | 0.462 | 0.375 | 0.625 | 0.4947 | 0.5053 | 0.500 | 2.000 | 0.6931 | 0.02 |
| PacA35 | 4 | 0.35 | 0.500 | 0.500 | 0.6211 | 0.3789 | 0.375 | 1.600 | 0.5623 | 0.01 |
| PacC3 | 2 | 0.20 | 0.4167 | 0.5833 | 0.5088 | 0.4912 | 0.4861 | 1.9459 | 0.6792 | 0.02 |
| PacC25 | 3 | 0.27 | 0.4792 | 0.5208 | 0.5739 | 0.4261 | 0.4217 | 1.7291 | 0.6126 | 0.02 |
| PacA58 | 2 | 0.190 | 0.3125 | 0.6875 | 0.495 | 0.505 | 0.4998 | 1.9991 | 0.6929 | 0 |
| PdavW3 | 4 | 0.352 | 0.3542 | 0.6458 | 0.5441 | 0.4559 | 0.4512 | 1.8221 | 0.6435 | 0.02 |
| Mean | 3.89 | 0.320 | 0.4774 | 0.5226 | 0.5470 | 0.4530 | 0.4483 | 1.8221 | 0.6371 | 0.0228 |
| Code | Genotype | K1 | K2 | K3 | K4 | Sub-Population |
|---|---|---|---|---|---|---|
| 1 | G1 | 0.049 | 0.045 | 0.861 | 0.045 | 3 |
| 2 | G2 | 0.029 | 0.025 | 0.923 | 0.024 | 3 |
| 3 | G3 | 0.028 | 0.025 | 0.92 | 0.027 | 3 |
| 4 | G4 | 0.022 | 0.023 | 0.935 | 0.021 | 3 |
| 5 | G5 | 0.021 | 0.02 | 0.937 | 0.022 | 3 |
| 6 | G6 | 0.024 | 0.025 | 0.928 | 0.024 | 3 |
| 7 | G7 | 0.024 | 0.024 | 0.928 | 0.024 | 3 |
| 8 | G8 | 0.032 | 0.028 | 0.915 | 0.026 | 3 |
| 9 | G9 | 0.035 | 0.027 | 0.911 | 0.027 | 3 |
| 10 | G10 | 0.029 | 0.026 | 0.918 | 0.027 | 3 |
| 11 | G11 | 0.033 | 0.03 | 0.903 | 0.034 | 3 |
| 12 | G12 | 0.084 | 0.057 | 0.807 | 0.051 | 3 |
| 13 | G13 | 0.075 | 0.047 | 0.833 | 0.046 | 3 |
| 14 | G14 | 0.101 | 0.077 | 0.751 | 0.071 | Admixture of 1,2,3,4 |
| 15 | G15 | 0.101 | 0.069 | 0.76 | 0.07 | Admixture of 1,2,3,4 |
| 16 | G16 | 0.116 | 0.081 | 0.719 | 0.084 | Admixture of 1,2,3,4 |
| 17 | G17 | 0.094 | 0.062 | 0.783 | 0.061 | Admixture of 1,2,3,4 |
| 18 | G18 | 0.036 | 0.034 | 0.897 | 0.033 | 3 |
| 19 | G19 | 0.178 | 0.146 | 0.53 | 0.146 | Admixture of 1,2,3,4 |
| 20 | G20 | 0.16 | 0.123 | 0.595 | 0.122 | Admixture of 1,2,3,4 |
| 21 | G21 | 0.168 | 0.156 | 0.516 | 0.16 | Admixture of 1,2,3,4 |
| 22 | G22 | 0.15 | 0.136 | 0.579 | 0.134 | Admixture of 1,2,3,4 |
| 23 | G23 | 0.248 | 0.239 | 0.287 | 0.227 | Admixture of 1,2,3,4 |
| 24 | G24 | 0.3 | 0.328 | 0.047 | 0.325 | Admixture of 1,2,3,4 |
| 25 | G25 | 0.296 | 0.317 | 0.078 | 0.309 | Admixture of 1,2,3,4 |
| 26 | G26 | 0.312 | 0.326 | 0.057 | 0.306 | Admixture of 1,2,3,4 |
| 27 | G27 | 0.323 | 0.318 | 0.024 | 0.335 | Admixture of 1,2,3,4 |
| 28 | G28 | 0.329 | 0.311 | 0.028 | 0.333 | Admixture of 1,2,3,4 |
| 29 | G29 | 0.329 | 0.313 | 0.027 | 0.331 | Admixture of 1,2,3,4 |
| 30 | G30 | 0.298 | 0.339 | 0.03 | 0.333 | Admixture of 1,2,3,4 |
| 31 | G31 | 0.318 | 0.321 | 0.024 | 0.337 | Admixture of 1,2,3,4 |
| 32 | G32 | 0.324 | 0.319 | 0.02 | 0.337 | Admixture of 1,2,3,4 |
| 33 | G33 | 0.315 | 0.332 | 0.03 | 0.322 | Admixture of 1,2,3,4 |
| 34 | G34 | 0.307 | 0.327 | 0.028 | 0.338 | Admixture of 1,2,3,4 |
| 35 | G35 | 0.315 | 0.335 | 0.022 | 0.328 | Admixture of 1,2,3,4 |
| 36 | G36 | 0.311 | 0.321 | 0.025 | 0.342 | Admixture of 1,2,3,4 |
| 37 | G37 | 0.314 | 0.317 | 0.024 | 0.345 | Admixture of 1,2,3,4 |
| 38 | G38 | 0.308 | 0.337 | 0.025 | 0.329 | Admixture of 1,2,3,4 |
| 39 | G39 | 0.323 | 0.32 | 0.032 | 0.325 | Admixture of 1,2,3,4 |
| 40 | G40 | 0.308 | 0.337 | 0.033 | 0.322 | Admixture of 1,2,3,4 |
| 41 | G41 | 0.316 | 0.33 | 0.038 | 0.316 | Admixture of 1,2,3,4 |
| 42 | G42 | 0.319 | 0.323 | 0.029 | 0.329 | Admixture of 1,2,3,4 |
| 43 | G43 | 0.318 | 0.328 | 0.025 | 0.329 | Admixture of 1,2,3,4 |
| 44 | G44 | 0.33 | 0.32 | 0.029 | 0.322 | Admixture of 1,2,3,4 |
| 45 | G45 | 0.299 | 0.316 | 0.029 | 0.356 | Admixture of 1,2,3,4 |
| 46 | G46 | 0.312 | 0.327 | 0.033 | 0.328 | Admixture of 1,2,3,4 |
| 47 | G47 | 0.314 | 0.326 | 0.026 | 0.334 | Admixture of 1,2,3,4 |
| 48 | G48 | 0.305 | 0.34 | 0.035 | 0.32 | Admixture of 1,2,3,4 |
| 49 | G49 | 0.314 | 0.328 | 0.031 | 0.327 | Admixture of 1,2,3,4 |
| 50 | G50 | 0.296 | 0.327 | 0.028 | 0.349 | Admixture of 1,2,3,4 |
| S. No | Sub-Population | Exp Het | Fst |
|---|---|---|---|
| 01 | 1 | 1.77 | 0.005 |
| 02 | 2 | 1.81 | 0.143 |
| 03 | 3 | 1.77 | 0.006 |
| 04 | 4 | 1.77 | 0.006 |
| Average | 1.78 | 0.04 | |
| Source | df | SS | MS | Est. Var. | % |
|---|---|---|---|---|---|
| Among populations | 1 | 32.385 | 32.385 | 0.557 | 5% |
| Within populations | 98 | 973.665 | 9.935 | 9.935 | 95% |
| Total | 99 | 1006.050 | 10.492 | 100% |
| S.NO | Genotype Name | Code | Location | District | Latitude | Longitude | Origin |
|---|---|---|---|---|---|---|---|
| 1 | ‘Harcot’ | G1 | CITH | Budgam | 33.9749° N | 74.7895° E | Exotic |
| 2 | ‘Hartlay’ | G2 | CITH | Budgam | 33.9741° N | 74.7889° E | Exotic |
| 3 | ‘Irani’ | G3 | CITH | Budgam | 33.9739° N | 74.7884°E | Exotic |
| 4 | ‘Communis-Holi’ | G4 | CITH | Budgam | 33.9725° N | 74.7882° E | Exotic |
| 5 | ‘Tilton’ | G5 | CITH | Budgam | 33.9719° N | 74.7875° E | Exotic |
| 6 | ‘Rival’ | G6 | CITH | Budgam | 33.9716° N | 74.7872° E | Exotic |
| 7 | ‘Tokpopanimu’ | G7 | CITH | Budgam | 33.9711° N | 74.7863° E | Exotic |
| 8 | ‘Fair medister’ | G8 | CITH | Budgam | 33.9706° N | 74.7858° E | Exotic |
| 9 | ‘Viva Gold’ | G9 | CITH | Budgam | 33.9701° N | 74.7850° E | Exotic |
| 10 | ‘Cummins’ | G10 | CITH | Budgam | 33.9721° N | 74.7877° E | Exotic |
| 11 | ‘Turkey’ | G11 | CITH | Budgam | 33.9714° N | 74.7867° E | Exotic |
| 12 | ‘New-Castle’ | G12 | CITH | Budgam | 33.9731° N | 74.7881°E | Exotic |
| 13 | ‘Chinese Apricot’ | G13 | CITH | Budgam | 33.9752° N | 74.7497° E | Exotic |
| 14 | Unknown | G14 | Hardas | Ladakh | 34.6061° N | 76.0981° E | Indigenous |
| 15 | Unknown | G15 | Ushkara | Baramulla | 34.2504° N | 74.3788° E | Indigenous |
| 16 | Unknown | G16 | Chardari | Baramulla | 34.1852° N | 74.3634° E | Indigenous |
| 17 | Unknown | G17 | Kantibag | Baramulla | 34.2406° N | 74.3674° E | Indigenous |
| 18 | Unknown | G18 | Uri | Baramulla | 34.0831° N | 74.0543° E | Indigenous |
| 19 | Unknown | G19 | Rangwar | Baramulla | 34.2343° N | 74.3676° E | Indigenous |
| 20 | Unknown | G20 | Beerwah | Budgam | 34.0128° N | 74.5956° E | Indigenous |
| 21 | Unknown | G21 | Katiyawali | Baramulla | 34.1754° N | 74.3531° E | Indigenous |
| 22 | unknown | G22 | Gatha Baderwah | Doda | 32.9973° N | 75.7007° E | Indigenous |
| 23 | unknown | G23 | Khanpora | Baramulla | 34.2086° N | 74.3275° E | Indigenous |
| 24 | unknown | G24 | Brazllo | Kulgam | 33.6467° N | 75.0589° E | Indigenous |
| 25 | unknown | G25 | Shiva | Baramulla | 34.3521° N | 74.4748° E | Indigenous |
| 26 | unknown | G26 | Dogar | Baramulla | 33.1829° N | 74.3619° E | Indigenous |
| 27 | unknown | G27 | Narapora | Shopian | 34.7611° N | 74.8019° E | Indigenous |
| 28 | unknown | G28 | Buniyar | Baramulla | 34.1009° N | 74.2004° E | Indigenous |
| 29 | unknown | G29 | Gozahama | Ganderbal | 34.1934° N | 74.6755° E | Indigenous |
| 30 | unknown | G30 | Kokarnag | Anantnag | 33.6801° N | 75.3895° E | Indigenous |
| 31 | unknown | G31 | Malpora | Baramulla | 34.3528° N | 74.4732° E | Indigenous |
| 32 | unknown | G32 | Dangerpora | Pulwama | 33.8756° N | 74.9793° E | Indigenous |
| 33 | unknown | G33 | Sopore | Baramulla | 34.2604° N | 74.4681° E | Indigenous |
| 34 | unknown | G34 | Duroo | Baramulla | 34.3516° N | 74.4633° E | Indigenous |
| 35 | unknown | G35 | Pazelpora | Baramulla | 34.3587° N | 74.4831° E | Indigenous |
| 36 | unknown | G36 | Kanispora | Baramulla | 34.2184° N | 74.3998° E | Indigenous |
| 37 | unknown | G37 | Darpora | Baramulla | 34.3570° N | 74.4323° E | Indigenous |
| 38 | unknown | G38 | Goripora | Baramulla | 34.3465° N | 74.4212° E | Indigenous |
| 39 | unknown | G39 | Mundji | Baramulla | 34.3607° N | 74.4738° E | Indigenous |
| 40 | unknown | G40 | Handwara | Kupwara | 34.4043° N | 74.2831° E | Indigenous |
| 41 | unknown | G41 | Brath Kalan | Baramulla | 34.3446° N | 74.4065° E | Indigenous |
| 42 | unknown | G42 | Wadura | Baramulla | 34.3528° N | 74.4018° E | Indigenous |
| 43 | unknown | G43 | Badwenchak | Qazigund | 33.5927° N | 75.1658° E | Indigenous |
| 44 | unknown | G44 | Sheeri | Baramulla | 34.1107° N | 74.1837° E | Indigenous |
| 45 | unknown | G45 | Krawah | Banihal | 33.2518° N | 75.1048° E | Indigenous |
| 46 | unknown | G46 | Chadoora | Budgam | 33.9453° N | 75.7967° E | Indigenous |
| 47 | unknown | G47 | Kralpora | Budgam | 34.4997° N | 74.1177° E | Indigenous |
| 48 | unknown | G48 | Bhangra | Doda | 32.9831° N | 75.7116° E | Indigenous |
| 49 | unknown | G49 | KapraBaderwah | Doda | 32.9833° N | 75.7112° E | Indigenous |
| 50 | unknown | G50 | Rawalpora | Srinagar | 34.0042° N | 74.4676° E | Indigenous |
| SSR Marker | Primer Sequence 5′→3′ | Reference | Size Range (bp) |
|---|---|---|---|
| RPPG1-017 | F:GCTCATCAAAACTCTCAACCA R:CCCTTTCTTCAATCCCATC | Dettori et al., 2015 | 90–220 |
| RPPG1-026 | F:CTTCTGGCACTCTTCCATTT R:GTTCCCAAGTTTTCCTCTCA | Dettori et al., 2015 | 90–220 |
| RPPG1-032 | F:ATGGCAGAGAGCACAACAA R:TTGAGAGGTAACAGCGAGAA | Dettori et al., 2015 | 90–250 |
| RPPG1-037 | F:GTCTCTGATCCAAGCCAACT R:ACGCTGCCATTGTTTCTATT | Dettori et al., 2015 | 100–250 |
| RPPG1-041 | F:TGTTGTAATGGATGGTGTCTTC R:CTTGGTCTTGGTTTCATTCA | Dettori et al., 2015 | 120–220 |
| RPPG2-011 | F:TTTACAGGTGCCTCAACAAA R:GTACAGCCGATGGAGAGAAA | Dettori et al., 2015 | 180 |
| RPPG2-022 | F:CTGCTGCGTCTGATGATG R:ACAGGACAGGACCACTTTCT | Dettori et al., 2015 | 200 |
| RPPG3-026 | F:AGAACGCTATTCCCCTGTAA R:TCATCCTCTCCAAATGTCAA | Dettori et al., 2015 | 90–200 |
| RPPG4-059 | F:GACGGCTGTTTATTTGCATT R:TGCATTTGTGATCTCGTTTC | Dettoriet al., 2015 | 100–180 |
| RPPG4-067 | F:AGAAGGGAGGGTGAGAGAAG R:CACGAAGGAAGAAACGAAGT | Dettori et al., 2015 | 100–210 |
| RPPG4-077 | F:CCTCGTCTTCAGTCTTTTCTG R:CTGTCCCTTCTGTGTTCCTAA | Dettori et al., 2015 | 90–150 |
| RPPG4-084 | F:TCCTCAAAAGTTACCCCAAG R:CTTGCTGTGGAAGAAGAACC | Dettori et al., 2015 | 120–200 |
| RPPG4-091 | F:GGAGGGTAGAGAACAGAGCA R:CGGAAGATGTGATTGTGAGA | Dettori et al., 2015 | 90–220 |
| RPPG5-018 | F:GCATGAAATTGACCCATACA R:TAATTGCTTTGGGGAGGAC | Dettori et al., 2015 | 90–200 |
| RPPG5-022 | F:CTTGTGAACTGGCATCTGTC R:AGTTGTATGGGCATGTTGTG | Dettori et al., 2015 | 90–180 |
| RPPG5-023 | F:TTGTTTGCACTAGGCTTTGA R:TTCTTCTTGCATGTCCTTGA | Dettori et al., 2015 | 90–150 |
| RPPG5-025 | F:GTGTCTCCTCCTCAAAGCAA R:TACGGCAACCAAGAACATC | Dettori et al., 2015 | 120 |
| RPPG5-030 | F:AAGGCAAGGAATTGGGTAGT R:TGGTTTGTCGTAAGAGTCCA | Dettori et al., 2015 | 90–280 |
| RPPG6-009 | F:GGGCTTGGCTGATAAAATAA R:TGGTAAAATAGAAGAGCGAGAAG | Dettori et al., 2015 | 100–120 |
| RPPG6-032 | F:TCCTATGGCAAAAACAAAATC R:TGAAGAGATGGAGTGGAAGAG | Dettori et al., 2015 | 90–150 |
| RPPG6-033 | F:CATTATCAAACCACGACCAA R:AAAGCTCAACAGCGACTTCT | Dettori et al., 2015 | 100–200 |
| RPPG7-015 | F:TCTTGGTGGTGGTGAAGTAA R:GAGAGATGGAGGAGGCTGA | Dettori et al., 2015 | 90–180 |
| RPPG7-026 | F:TTTGGTGAGTGGGCTCTATT R:CTATCGTTCGCTGGTCTTCT | Dettori et al., 2015 | 90–180 |
| RPPG7-032 | F:AAGGGAGGAGGATTGTGAA R:TGGTAGACGGGTAGATGTTG | Dettori et al., 2015 | 90–180 |
| RPPG8-007 | F:ACCACCACCTCTTCCAATC R:ACCTCAAAGTGTCCCAGAAA | Dettori et al., 2015 | 150 |
| RPPG8-028 | F:AAGGAGCCGACATCAGAAC R:TGACCAGAAGCCAAATACATC | Dettori et al., 2015 | 120–180 |
| Aprigms18 | F:TCTGAGTTCAGTGGGTAGCA R:ACAGAATGTGCGTTGCTTTA | Liu et al., 2015 | 90–200 |
| UDP98-405 | F:ACGTCATGAACTGACACCCA R:GAGTCTTTGCTCTGCCATCC | Liu et al., 2015 | 90–120 |
| UDP98-406 | F:TCGGAAACTGGTAGTATGAACAGA R:ATGGGTCGTATGCACAGTCA | Liu et al., 2015 | 90–120 |
| UDP98-409 | F:GCTGATGGGTTTTATGGTTTTC R:CGGACTCTTATCCTCTATCAACA | Liu et al., 2015 | 90–150 |
| UDP98-411 | F:AAGCCATCCACTCAGCACTC R:CCAAAAACCAAAACCAAAGG | Liu et al., 2015 | 90–180 |
| Pchgms4 | F:ATCTTCACAACCCTAATGTC R:GTTGAGGCAAAAGACTTCAAT | Liu et al., 2015 | 90–280 |
| Pchgms5 | F:CGCCCATGACAAACTTA R:GTCAAGAGGTACACCAG | Liu et al., 2015 | 150–280 |
| Bppct007 | F:TCATTGCTCGTCATCAGC R:CAGATTTCTGAAGTTAGCGGTA | Liu et al., 2015 | 150–420 |
| Bppct025 | F:TCCTGCGTAGAAGAAGGTAGC R:CGACATAAAGTCCAAATGGC | Liu et al., 2015 | 90–150 |
| Bppct030 | F:AATTGTACTTGCCAATGCTATGA R:CTGCCTTCTGCTCACACC | Liu et al., 2015 | 90–180 |
| PacA10 | F:TGAGCATAATTGGGGCAG R:GCCAGAGAAGCCATTTCAGT | Lambert et al., 2004 | 120–250 |
| PacA18 | F:TCCAAACCTACCGTTTCTCAT R:CAACAGCACAAACAGAACCAC | Lambert et al., 2004 | 180–250 |
| PacA33 | F:TCAGTCTCATCCTGCATACG R:CATGTGGCTCAAGGATCAAA | Lambert et al., 2004 | 90–250 |
| PacA22 | F:AACCAGTTGCCTCTAGATTTTG R:AGCTGAAAGTCAATTCAGAGTAGTT | Lambert et al., 2004 | 100–180 |
| PacA26 | F:CCAATCATGAAAATCATAAAAGCAA R:TGGGATGTCCTATTGTTTTCA | Lambert et al., 2004 | 100–200 |
| PacA35 | F:ATTGCGATTTCGGTCTGTT R:CCATCCCAAATTGCTTACTT | Lambert et al., 2004 | 120–180 |
| PacC3 | F:TGACTTGATCAGACTCGACA R:TTGCATTTGCATTTACAATAGA | Lambert et al., 2004 | 90–200 |
| PacC25 | F:GTGTTTTGACAAGAAATGAATTG R:TCCATTCGCAGTAAAATTAAAC | Lambert et al., 2004 | 100–200 |
| PacA58 | F:GACATTGCGATTTCGGTC R:TCCATCCCAAATTGCTTACT | Lambert et al., 2004 | 100–180 |
| PdavW3 | F:GAGGGCTGGATCATGACG R:AACCCAGTGGCACAATCGTA | Lambert et al., 2004 | 90–200 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sheikh, Z.N.; Sharma, V.; Shah, R.A.; Raina, S.; Aljabri, M.; Mir, J.I.; AlKenani, N.; Hakeem, K.R. Elucidating Genetic Diversity in Apricot (Prunus armeniaca L.) Cultivated in the North-Western Himalayan Provinces of India Using SSR Markers. Plants 2021, 10, 2668. https://doi.org/10.3390/plants10122668
Sheikh ZN, Sharma V, Shah RA, Raina S, Aljabri M, Mir JI, AlKenani N, Hakeem KR. Elucidating Genetic Diversity in Apricot (Prunus armeniaca L.) Cultivated in the North-Western Himalayan Provinces of India Using SSR Markers. Plants. 2021; 10(12):2668. https://doi.org/10.3390/plants10122668
Chicago/Turabian StyleSheikh, Zahid Nabi, Vikas Sharma, Rafiq Ahmad Shah, Shilpa Raina, Maha Aljabri, Javid Iqbal Mir, Naser AlKenani, and Khalid Rehman Hakeem. 2021. "Elucidating Genetic Diversity in Apricot (Prunus armeniaca L.) Cultivated in the North-Western Himalayan Provinces of India Using SSR Markers" Plants 10, no. 12: 2668. https://doi.org/10.3390/plants10122668







