TALE and Shape: How to Make a Leaf Different
Abstract
:1. Introduction
| KNOX | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| STM | KNAT1/BP | KNAT2 | KNAT6 | KNAT3 | KNAT4 | KNAT5 | KNAT7 | KNATM | ||
| BLH/BELL | ATH1 | [8] | [8] | [15] | [15] | [16] | [16] | |||
| BEL1 | [16,17,18] | [16,18] | [16,17] | [16] | [17] | [14] | ||||
| BLH1 | [16] | [16] | [16] | [16] | ||||||
| BLH2/SAW1 | [16] | [16] | [16] | [16] | [16] | [16] | [14] | |||
| BLH3 | [8] | [8] | [16] | [16] | [16] | |||||
| BLH4/SAW2 | [16] | [16] | [16] | [16] | [16] | [16] | [14] | |||
| BLH5 | [16] | [16] | [16] | [16] | [16] | |||||
| BLH6 | [16] | [16] | [16] | [16] | [16] | [16] | [16] | |||
| BLH7 | [16,19] | [16] | [16] | [16] | [16] | |||||
| BLH8/PNF | [18] | [16,18] | [16] | |||||||
| BLH9/PNY/BLR | [8] | [8] | [16] | [7,16] | [16] | [16] | [14] | |||
| BLH10 | [16] | |||||||||
| BLH11 | ||||||||||

2. Overview of the Role of TALEs in Shoot Apical Meristem
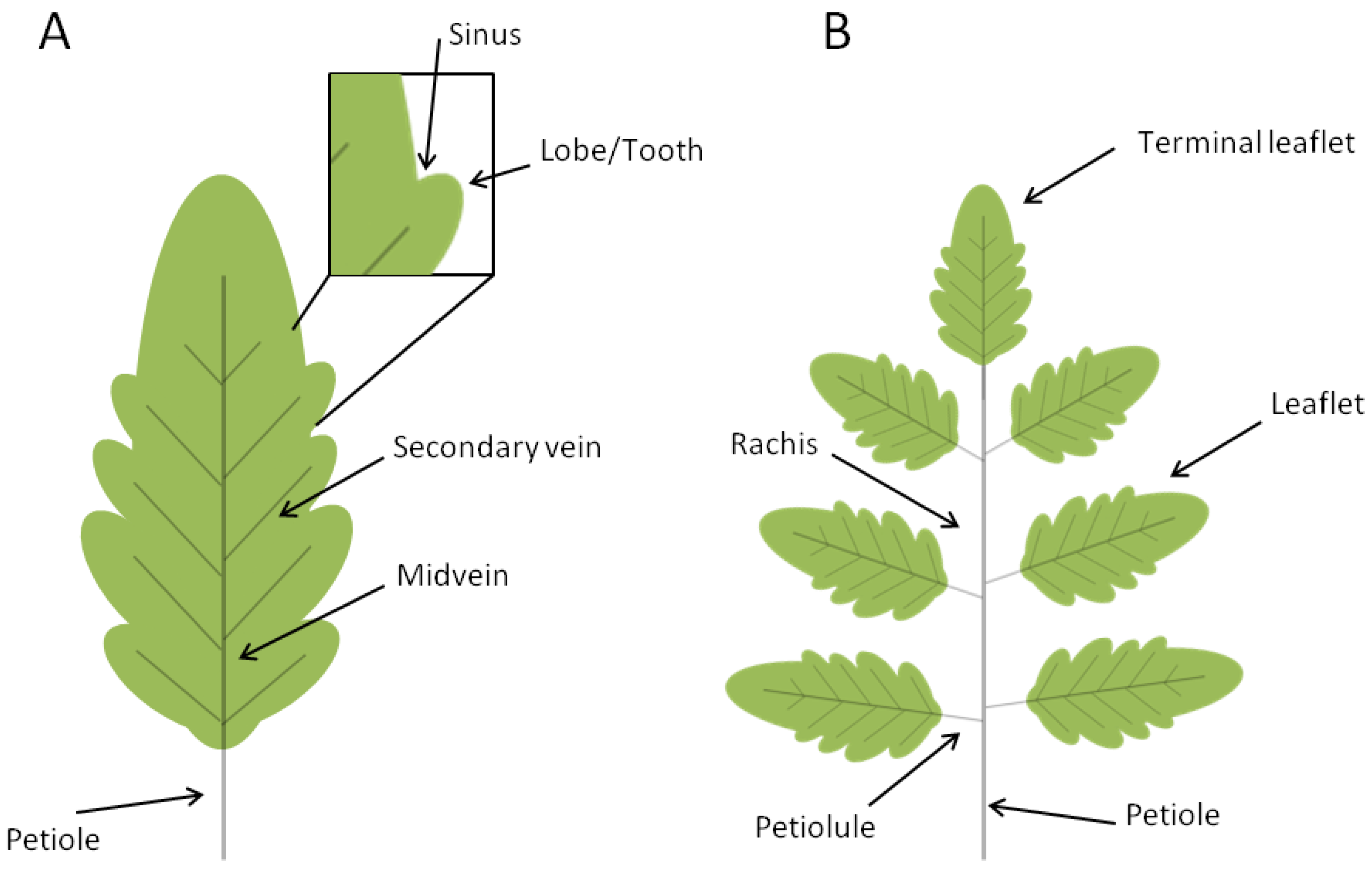
3. TALE Contribution to Leaf Initiation and Morphology in Simple and Compound-Leafed Plant Species
3.1. TALE Genes in Organ Primordia Initiation
3.2. TALE Genes in the Control of Simple Leaf Morphology
3.3. TALE Genes in the Control of Compound Leaf Morphology

3.4. ILRC Subgroup of Legumes: An Exception to the TALE Rule?

4. TALEs Work at the Boundaries: How to Establish the Confines between Undifferentiated Cells and Organ Primordia

5. A Matter of Targets: the Secret Agents of TALEs

5.1. Links to Hormones
5.2. Transcription Factors
5.3. Cell Wall Proteins
6. Conclusions and Perspectives
Acknowledgements
References
- Bertolino, E.; Reimund, B.; Wildt-Perinic, D.; Clerc, R.G. A novel homeobox protein which recognizes a TGT core and functionally interferes with a retinoid-responsive motif. J. Biol. Chem. 1995, 270, 31178–31188. [Google Scholar]
- Burglin, T.R. Analysis of TALE superclass homeobox genes (MEIS, PBC, KNOX, Iroquois, TGIF) reveals a novel domain conserved between plants and animals. Nucleic Acids Res. 1997, 25, 4173–4180. [Google Scholar] [CrossRef]
- Burglin, T.R. The PBC domain contains a MEINOX domain: Coevolution of Hox and TALE homeobox genes? Dev. Genes Evol. 1998, 208, 113–116. [Google Scholar] [CrossRef]
- Laurent, A.; Bihan, R.; Omilli, F.; Deschamps, S.; Pellerin, I. PBX proteins: Much more than Hox cofactors. Int. J. Dev. Biol. 2008, 52, 9–20. [Google Scholar] [CrossRef]
- Berthelsen, J.; Kilstrup-Nielsen, C.; Blasi, F.; Mavilio, F.; Zappavigna, V. The subcellular localization of PBX1 and EXD proteins depends on nuclear import and export signals and is modulated by association with PREP1 and HTH. Genes Dev. 1999, 13, 946–953. [Google Scholar] [CrossRef]
- Saleh, M.; Huang, H.; Green, N.C.; Featherstone, M.S. A conformational change in PBX1A is necessary for its nuclear localization. Exp. Cell Res. 2000, 260, 105–115. [Google Scholar]
- Bhatt, A.M.; Etchells, J.P.; Canales, C.; Lagodienko, A.; Dickinson, H. VAAMANA—A BEL1-like homeodomain protein, interacts with KNOX proteins BP and STM and regulates inflorescence stem growth in Arabidopsis. Gene 2004, 328, 103–111. [Google Scholar] [CrossRef]
- Cole, M.; Nolte, C.; Werr, W. Nuclear import of the transcription factor SHOOT MERISTEMLESS depends on heterodimerization with BLH proteins expressed in discrete sub-domains of the shoot apical meristem of Arabidopsis thaliana. Nucleic Acids Res. 2006, 34, 1281–1292. [Google Scholar] [CrossRef]
- Smith, H.M.; Boschke, I.; Hake, S. Selective interaction of plant homeodomain proteins mediates high DNA-binding affinity. Proc. Natl. Acad. Sci. USA 2002, 99, 9579–9584. [Google Scholar] [CrossRef]
- Smaczniak, C.; Immink, R.G.; Muino, J.M.; Blanvillain, R.; Busscher, M.; Busscher-Lange, J.; Dinh, Q.D.; Liu, S.; Westphal, A.H.; Boeren, S.; et al. Characterization of MADS-domain transcription factor complexes in Arabidopsis flower development. Proc. Natl. Acad. Sci. USA 2012, 109, 1560–1565. [Google Scholar] [CrossRef]
- Mukherjee, K.; Brocchieri, L.; Burglin, T.R. A comprehensive classification and evolutionary analysis of plant homeobox genes. Mol. Biol. Evol. 2009, 26, 2775–2794. [Google Scholar] [CrossRef]
- Kerstetter, R.; Vollbrecht, E.; Lowe, B.; Veit, B.; Yamaguchi, J.; Hake, S. Sequence analysis and expression patterns divide the maize knotted1-like homeobox genes into two classes. Plant Cell 1994, 6, 1877–1887. [Google Scholar]
- Reiser, L.; Sanchez-Baracaldo, P.; Hake, S. Knots in the family tree: Evolutionary relationships and functions of knox homeobox genes. Plant Mol. Biol. 2000, 42, 151–166. [Google Scholar] [CrossRef]
- Magnani, E.; Hake, S. KNOX lost the OX: The Arabidopsis KNATM gene defines a novel class of KNOX transcriptional regulators missing the homeodomain. Plant Cell 2008, 20, 875–887. [Google Scholar] [CrossRef]
- Li, Y.; Pi, L.; Huang, H.; Xu, L. ATH1 and KNAT2 proteins act together in regulation of plant inflorescence architecture. J. Exp. Bot. 2012, 63, 1423–1433. [Google Scholar] [CrossRef]
- Hackbusch, J.; Richter, K.; Muller, J.; Salamini, F.; Uhrig, J.F. A central role of Arabidopsis thaliana ovate family proteins in networking and subcellular localization of 3-aa loop extension homeodomain proteins. Proc. Natl. Acad. Sci. USA 2005, 102, 4908–4912. [Google Scholar] [CrossRef]
- Bellaoui, M.; Pidkowich, M.S.; Samach, A.; Kushalappa, K.; Kohalmi, S.E.; Modrusan, Z.; Crosby, W.L.; Haughn, G.W. The Arabidopsis BELL1 and KNOX TALE homeodomain proteins interact through a domain conserved between plants and animals. Plant Cell 2001, 13, 2455–2470. [Google Scholar]
- Di Giacomo, E.; Sestili, F.; Iannelli, M.A.; Testone, G.; Mariotti, D.; Frugis, G. Characterization of KNOX genes in Medicago truncatula. Plant Mol. Biol. 2008, 67, 135–150. [Google Scholar] [CrossRef]
- Arabidopsis Interactome Mapping Consortium. Evidence for network evolution in an Arabidopsis interactome map. Science 2011, 333, 601–607. [CrossRef]
- Hake, S.; Smith, H.M.; Holtan, H.; Magnani, E.; Mele, G.; Ramirez, J. The role of knox genes in plant development. Annu. Rev. Cell Dev. Biol. 2004, 20, 125–151. [Google Scholar] [CrossRef]
- Hay, A.; Tsiantis, M. KNOX genes: Versatile regulators of plant development and diversity. Development 2010, 137, 3153–3165. [Google Scholar] [CrossRef]
- Bharathan, G.; Goliber, T.E.; Moore, C.; Kessler, S.; Pham, T.; Sinha, N.R. Homologies in leaf form inferred from KNOXI gene expression during development. Science 2002, 296, 1858–1860. [Google Scholar] [CrossRef]
- Hamant, O.; Pautot, V. Plant development: A TALE story. C. R. Biol. 2010, 333, 371–381. [Google Scholar] [CrossRef]
- Canales, C.; Barkoulas, M.; Galinha, C.; Tsiantis, M. Weeds of change: Cardamine hirsuta as a new model system for studying dissected leaf development. J. Plant Res. 2010, 123, 25–33. [Google Scholar] [CrossRef]
- Blein, T.; Hasson, A.; Laufs, P. Leaf development: What it needs to be complex. Curr. Opin. Plant Biol. 2010, 13, 75–82. [Google Scholar] [CrossRef]
- Efroni, I.; Eshed, Y.; Lifschitz, E. Morphogenesis of simple and compound leaves: A critical review. Plant Cell 2010, 22, 1019–1032. [Google Scholar]
- Uchida, N.; Kimura, S.; Koenig, D.; Sinha, N. Coordination of leaf development via regulation of KNOX1 genes. J. Plant Res. 2010, 123, 7–14. [Google Scholar] [CrossRef]
- Moon, J.; Hake, S. How a leaf gets its shape. Curr. Opin. Plant Biol. 2011, 14, 24–30. [Google Scholar] [CrossRef]
- Burko, Y.; Ori, N. The tomato leaf as a model system for organogenesis. Methods Mol. Biol. 2013, 959, 1–19. [Google Scholar] [CrossRef]
- Hasson, A.; Blein, T.; Laufs, P. Leaving the meristem behind: The genetic and molecular control of leaf patterning and morphogenesis. C. R. Biol. 2010, 333, 350–360. [Google Scholar] [CrossRef]
- Wu, S.; Smith, M.S. Out of step:The function of TALE homeodomain transcription factors that regulate shoot meristem maintenance and meristem identity. Front Biol. 2012, 7, 144–154. [Google Scholar]
- Winter, N.; Kollwig, G.; Zhang, S.; Kragler, F. MPB2C, a microtubule-associated protein, regulates non-cell-autonomy of the homeodomain protein KNOTTED1. Plant Cell 2007, 19, 3001–3018. [Google Scholar] [CrossRef]
- Kim, J.Y.; Yuan, Z.; Cilia, M.; Khalfan-Jagani, Z.; Jackson, D. Intercellular trafficking of a KNOTTED1 green fluorescent protein fusion in the leaf and shoot meristem of Arabidopsis. Proc. Natl. Acad. Sci. USA 2002, 99, 4103–4108. [Google Scholar]
- Kim, J.Y.; Yuan, Z.; Jackson, D. Developmental regulation and significance of KNOX protein trafficking in Arabidopsis. Development 2003, 130, 4351–4362. [Google Scholar] [CrossRef]
- Kim, J.Y.; Rim, Y.; Wang, J.; Jackson, D. A novel cell-to-cell trafficking assay indicates that the KNOX homeodomain is necessary and sufficient for intercellular protein and mRNA trafficking. Genes Dev. 2005, 19, 788–793. [Google Scholar] [CrossRef]
- Xu, X.M.; Wang, J.; Xuan, Z.; Goldshmidt, A.; Borrill, P.G.; Hariharan, N.; Kim, J.Y.; Jackson, D. Chaperonins facilitate KNOTTED1 cell-to-cell trafficking and stem cell function. Science 2011, 333, 1141–1144. [Google Scholar]
- Vollbrecht, E.; Veit, B.; Sinha, N.; Hake, S. The developmental gene Knotted-1 is a member of a maize homeobox gene family. Nature 1991, 350, 241–243. [Google Scholar] [CrossRef]
- Barton, M.K. Twenty years on: The inner workings of the shoot apical meristem, a developmental dynamo. Dev. Biol. 2010, 341, 95–113. [Google Scholar] [CrossRef]
- Long, J.A.; Moan, E.I.; Medford, J.I.; Barton, M.K. A member of the KNOTTED class of homeodomain proteins encoded by the STM gene of Arabidopsis. Nature 1996, 379, 66–69. [Google Scholar]
- Clark, S.E.; Jacobsen, S.E.; Levin, J.Z.; Meyerowitz, E.M. The CLAVATA and SHOOT MERISTEMLESS loci competitively regulate meristem activity in Arabidopsis. Development 1996, 122, 1567–1575. [Google Scholar]
- Endrizzi, K.; Moussian, B.; Haecker, A.; Levin, J.Z.; Laux, T. The SHOOT MERISTEMLESS gene is required for maintenance of undifferentiated cells in Arabidopsis shoot and floral meristems and acts at a different regulatory level than the meristem genes WUSCHEL and ZWILLE. Plant J. 1996, 10, 967–979. [Google Scholar]
- Kanrar, S.; Onguka, O.; Smith, H.M. Arabidopsis inflorescence architecture requires the activities of KNOX-BELL homeodomain heterodimers. Planta 2006, 224, 1163–1173. [Google Scholar] [CrossRef]
- Scofield, S.; Dewitte, W.; Murray, J.A. The KNOX gene SHOOT MERISTEMLESS is required for the development of reproductive meristematic tissues in Arabidopsis. Plant J. 2007, 50, 767–781. [Google Scholar] [CrossRef]
- Rutjens, B.; Bao, D.; van Eck-Stouten, E.; Brand, M.; Smeekens, S.; Proveniers, M. Shoot apical meristem function in Arabidopsis requires the combined activities of three BEL1-like homeodomain proteins. Plant J. 2009, 58, 641–654. [Google Scholar] [CrossRef]
- Ramirez, J.; Bolduc, N.; Lisch, D.; Hake, S. Distal expression of knotted1 in maize leaves leads to reestablishment of proximal/distal patterning and leaf dissection. Plant Physiol. 2009, 151, 1878–1888. [Google Scholar] [CrossRef]
- Takano, S.; Niihama, M.; Smith, H.M.; Tasaka, M.; Aida, M. gorgon, a novel missense mutation in the SHOOT MERISTEMLESS gene, impairs shoot meristem homeostasis in Arabidopsis. Plant Cell Physiol. 2010, 51, 621–634. [Google Scholar] [CrossRef]
- Long, J.A.; Barton, M.K. The development of apical embryonic pattern in Arabidopsis. Development 1998, 125, 3027–3035. [Google Scholar]
- Aida, M.; Ishida, T.; Tasaka, M. Shoot apical meristem and cotyledon formation during Arabidopsis embryogenesis: Interaction among the CUP-SHAPED COTYLEDON and SHOOT MERISTEMLESS genes. Development 1999, 126, 1563–1570. [Google Scholar]
- Vollbrecht, E.; Reiser, L.; Hake, S. Shoot meristem size is dependent on inbred background and presence of the maize homeobox gene, knotted1. Development 2000, 127, 3161–3172. [Google Scholar]
- Takada, S.; Hibara, K.; Ishida, T.; Tasaka, M. The CUP-SHAPED COTYLEDON1 gene of Arabidopsis regulates shoot apical meristem formation. Development 2001, 128, 1127–1135. [Google Scholar]
- Poethig, R.S. Leaf morphogenesis in flowering plants. Plant Cell 1997, 9, 1077–1087. [Google Scholar] [CrossRef]
- Dengler, N.; Kang, J. Vascular patterning and leaf shape. Curr. Opin. Plant Biol. 2001, 4, 50–56. [Google Scholar]
- Holtan, H.E.; Hake, S. Quantitative trait locus analysis of leaf dissection in tomato using Lycopersicon pennellii segmental introgression lines. Genetics 2003, 165, 1541–1550. [Google Scholar]
- Barkoulas, M.; Galinha, C.; Grigg, S.P.; Tsiantis, M. From genes to shape: Regulatory interactions in leaf development. Curr. Opin. Plant Biol. 2007, 10, 660–666. [Google Scholar] [CrossRef]
- Shani, E.; Burko, Y.; Ben-Yaakov, L.; Berger, Y.; Amsellem, Z.; Goldshmidt, A.; Sharon, E.; Ori, N. Stage-specific regulation of Solanum lycopersicum leaf maturation by class 1 KNOTTED1-LIKE HOMEOBOX proteins. Plant Cell 2009, 21, 3078–3092. [Google Scholar] [CrossRef]
- Gleissberg, H.A. Organogenetic capacity of leaves: The significance of marginal blastozones in angiosperms. Plant Syst. Evol. 1996, 199, 31. [Google Scholar]
- Murray, J.A.; Jones, A.; Godin, C.; Traas, J. Systems analysis of shoot apical meristem growth and development: Integrating hormonal and mechanical signaling. Plant Cell 2012, 24, 3907–3919. [Google Scholar]
- Fleming, A.J.; McQueen-Mason, S.; Mandel, T.; Kuhlemeier, C. Induction of leaf primordia by the cell wall protein expansin. Science 1997, 276, 1415–1418. [Google Scholar]
- Reinhardt, D.; Wittwer, F.; Mandel, T.; Kuhlemeier, C. Localized upregulation of a new expansin gene predicts the site of leaf formation in the tomato meristem. Plant Cell 1998, 10, 1427–1437. [Google Scholar]
- Pien, S.; Wyrzykowska, J.; McQueen-Mason, S.; Smart, C.; Fleming, A. Local expression of expansin induces the entire process of leaf development and modifies leaf shape. Proc. Natl. Acad. Sci. USA 2001, 98, 11812–11817. [Google Scholar]
- Peaucelle, A.; Louvet, R.; Johansen, J.N.; Hofte, H.; Laufs, P.; Pelloux, J.; Mouille, G. Arabidopsis phyllotaxis is controlled by the methyl-esterification status of cell-wall pectins. Curr. Biol. 2008, 18, 1943–1948. [Google Scholar]
- Peaucelle, A.; Louvet, R.; Johansen, J.N.; Salsac, F.; Morin, H.; Fournet, F.; Belcram, K.; Gillet, F.; Hofte, H.; Laufs, P.; et al. The transcription factor BELLRINGER modulates phyllotaxis by regulating the expression of a pectin methylesterase in Arabidopsis. Development 2011, 138, 4733–4741. [Google Scholar] [CrossRef]
- Elliott, R.C.; Betzner, A.S.; Huttner, E.; Oakes, M.P.; Tucker, W.Q.; Gerentes, D.; Perez, P.; Smyth, D.R. AINTEGUMENTA, an APETALA2-like gene of Arabidopsis with pleiotropic roles in ovule development and floral organ growth. Plant Cell 1996, 8, 155–168. [Google Scholar]
- Lenhard, M.; Jurgens, G.; Laux, T. The WUSCHEL and SHOOTMERISTEMLESS genes fulfil complementary roles in Arabidopsis shoot meristem regulation. Development 2002, 129, 3195–3206. [Google Scholar]
- Byrne, M.E.; Kidner, C.A.; Martienssen, R.A. Plant stem cells: Divergent pathways and common themes in shoots and roots. Curr. Opin. Genet. Dev. 2003, 13, 551–557. [Google Scholar] [CrossRef]
- Chuck, G.; Lincoln, C.; Hake, S. KNAT1 induces lobed leaves with ectopic meristems when overexpressed in Arabidopsis. Plant Cell 1996, 8, 1277–1289. [Google Scholar]
- Ori, N.; Eshed, Y.; Chuck, G.; Bowman, J.L.; Hake, S. Mechanisms that control knox gene expression in the Arabidopsis shoot. Development 2000, 127, 5523–5532. [Google Scholar]
- Kumar, R.; Kushalappa, K.; Godt, D.; Pidkowich, M.S.; Pastorelli, S.; Hepworth, S.R.; Haughn, G.W. The Arabidopsis BEL1-LIKE HOMEODOMAIN proteins SAW1 and SAW2 act redundantly to regulate KNOX expression spatially in leaf margins. Plant Cell 2007, 19, 2719–2735. [Google Scholar] [CrossRef]
- Piazza, P.; Bailey, C.D.; Cartolano, M.; Krieger, J.; Cao, J.; Ossowski, S.; Schneeberger, K.; He, F.; de Meaux, J.; Hall, N.; et al. Arabidopsis thaliana leaf form evolved via loss of KNOX expression in leaves in association with a selective sweep. Curr. Biol. 2010, 20, 2223–2228. [Google Scholar] [CrossRef]
- Kawamura, E.; Horiguchi, G.; Tsukaya, H. Mechanisms of leaf tooth formation in Arabidopsis. Plant J. 2010, 62, 429–441. [Google Scholar] [CrossRef]
- Phelps-Durr, T.L.; Thomas, J.; Vahab, P.; Timmermans, M.C. Maize rough sheath2 and its Arabidopsis orthologue ASYMMETRIC LEAVES1 interact with HIRA, a predicted histone chaperone, to maintain knox gene silencing and determinacy during organogenesis. Plant Cell 2005, 17, 2886–2898. [Google Scholar] [CrossRef]
- Xu, L.; Shen, W.H. Polycomb silencing of KNOX genes confines shoot stem cell niches in Arabidopsis. Curr. Biol. 2008, 18, 1966–1971. [Google Scholar] [CrossRef]
- Stahle, M.I.; Kuehlich, J.; Staron, L.; von Arnim, A.G.; Golz, J.F. YABBYs and the transcriptional corepressors LEUNIG and LEUNIG_HOMOLOG maintain leaf polarity and meristem activity in Arabidopsis. Plant Cell 2009, 21, 3105–3118. [Google Scholar] [CrossRef]
- Causier, B.; Ashworth, M.; Guo, W.; Davies, B. The TOPLESS interactome: A framework for gene repression in Arabidopsis. Plant Physiol. 2012, 158, 423–438. [Google Scholar]
- Ohta, M.; Matsui, K.; Hiratsu, K.; Shinshi, H.; Ohme-Takagi, M. Repression domains of class II ERF transcriptional repressors share an essential motif for active repression. Plant Cell 2001, 13, 1959–1968. [Google Scholar]
- Kagale, S.; Rozwadowski, K. EAR motif-mediated transcriptional repression in plants: An underlying mechanism for epigenetic regulation of gene expression. Epigenetics 2011, 6, 141–146. [Google Scholar] [CrossRef]
- Hasson, A.; Plessis, A.; Blein, T.; Adroher, B.; Grigg, S.; Tsiantis, M.; Boudaoud, A.; Damerval, C.; Laufs, P. Evolution and diverse roles of the CUP-SHAPED COTYLEDON genes in Arabidopsis leaf development. Plant Cell 2011, 23, 54–68. [Google Scholar] [CrossRef]
- Bilsborough, G.D.; Runions, A.; Barkoulas, M.; Jenkins, H.W.; Hasson, A.; Galinha, C.; Laufs, P.; Hay, A.; Prusinkiewicz, P.; Tsiantis, M. Model for the regulation of Arabidopsis thaliana leaf margin development. Proc. Natl. Acad. Sci. USA 2011, 108, 3424–3429. [Google Scholar] [CrossRef]
- Hareven, D.; Gutfinger, T.; Parnis, A.; Eshed, Y.; Lifschitz, E. The making of a compound leaf: Genetic manipulation of leaf architecture in tomato. Cell 1996, 84, 735–744. [Google Scholar]
- Kimura, S.; Koenig, D.; Kang, J.; Yoong, F.Y.; Sinha, N. Natural variation in leaf morphology results from mutation of a novel KNOX gene. Curr. Biol. 2008, 18, 672–677. [Google Scholar] [CrossRef]
- Hay, A.; Tsiantis, M. The genetic basis for differences in leaf form between Arabidopsis thaliana and its wild relative Cardamine hirsuta. Nat. Genet. 2006, 38, 942–947. [Google Scholar] [CrossRef]
- Champagne, C.E.; Goliber, T.E.; Wojciechowski, M.F.; Mei, R.W.; Townsley, B.T.; Wang, K.; Paz, M.M.; Geeta, R.; Sinha, N.R. Compound leaf development and evolution in the legumes. Plant Cell 2007, 19, 3369–3378. [Google Scholar] [CrossRef]
- Hofer, J.; Turner, L.; Hellens, R.; Ambrose, M.; Matthews, P.; Michael, A.; Ellis, N. UNIFOLIATA regulates leaf and flower morphogenesis in pea. Curr. Biol. 1997, 7, 581–587. [Google Scholar]
- Wang, H.; Chen, J.; Wen, J.; Tadege, M.; Li, G.; Liu, Y.; Mysore, K.S.; Ratet, P.; Chen, R. Control of compound leaf development by FLORICAULA/LEAFY ortholog SINGLE LEAFLET1 in Medicago truncatula. Plant Physiol. 2008, 146, 1759–1772. [Google Scholar] [CrossRef]
- Moyroud, E.; Minguet, E.G.; Ott, F.; Yant, L.; Pose, D.; Monniaux, M.; Blanchet, S.; Bastien, O.; Thevenon, E.; Weigel, D.; et al. Prediction of regulatory interactions from genome sequences using a biophysical model for the Arabidopsis LEAFY transcription factor. Plant Cell 2011, 23, 1293–1306. [Google Scholar] [CrossRef]
- Chen, J.; Yu, J.; Ge, L.; Wang, H.; Berbel, A.; Liu, Y.; Chen, Y.; Li, G.; Tadege, M.; Wen, J.; et al. Control of dissected leaf morphology by a Cys(2)His(2) zinc finger transcription factor in the model legume Medicago truncatula. Proc. Natl. Acad. Sci. USA 2010, 107, 10754–10759. [Google Scholar]
- Peng, J.; Yu, J.; Wang, H.; Guo, Y.; Li, G.; Bai, G.; Chen, R. Regulation of compound leaf development in Medicago truncatula by fused compound leaf1, a class M KNOX gene. Plant Cell 2011, 23, 3929–3943. [Google Scholar] [CrossRef]
- Chen, R. Personal communication, Plant Biology Division, The Samuel Roberts Noble Foundation: Ardmore, OK, USA, 2012.
- Di Giacomo, E. Personal communication, Istituto di Biologia e Biotecnologia Agraria, UOS Roma, Consiglio Nazionale delle Ricerche: Monterotondo Scalo, Roma, Italy, 2011.
- Smith, H.M.; Ung, N.; Lal, S.; Courtier, J. Specification of reproductive meristems requires the combined function of SHOOT MERISTEMLESS and floral integrators FLOWERING LOCUS T and FD during Arabidopsis inflorescence development. J. Exp. Bot. 2011, 62, 583–593. [Google Scholar] [CrossRef]
- Lal, S.; Pacis, L.B.; Smith, H.M. Regulation of the SQUAMOSA PROMOTER-BINDING PROTEIN-LIKE genes/microRNA156 module by the homeodomain proteins PENNYWISE and POUND-FOOLISH in Arabidopsis. Mol. Plant 2011, 4, 1123–1132. [Google Scholar]
- Waites, R.; Selvadurai, H.R.; Oliver, I.R.; Hudson, A. The PHANTASTICA gene encodes a MYB transcription factor involved in growth and dorsoventrality of lateral organs in Antirrhinum. Cell 1998, 93, 779–789. [Google Scholar] [CrossRef]
- McConnell, J.R.; Barton, M.K. Leaf polarity and meristem formation in Arabidopsis. Development 1998, 125, 2935–2942. [Google Scholar]
- Ha, C.M.; Jun, J.H.; Fletcher, J.C. Control of Arabidopsis leaf morphogenesis through regulation of the YABBY and KNOX families of transcription factors. Genetics 2010, 186, 197–206. [Google Scholar] [CrossRef]
- Jun, J.H.; Ha, C.M.; Fletcher, J.C. BLADE-ON-PETIOLE1 coordinates organ determinacy and axial polarity in arabidopsis by directly activating ASYMMETRIC LEAVES2. Plant Cell 2010, 22, 62–76. [Google Scholar] [CrossRef]
- Reinhardt, D.; Kuhlemeier, C. Plant architecture. EMBO Rep. 2002, 3, 846–851. [Google Scholar] [CrossRef]
- Barkoulas, M.; Hay, A.; Kougioumoutzi, E.; Tsiantis, M. A developmental framework for dissected leaf formation in the Arabidopsis relative Cardamine hirsuta. Nat. Genet. 2008, 40, 1136–1141. [Google Scholar] [CrossRef]
- Ben-Gera, H.; Ori, N. Auxin and LANCEOLATE affect leaf shape in tomato via different developmental processes. Plant Signal. Behav. 2012, 7, 1255–1257. [Google Scholar] [CrossRef]
- Peng, J.; Chen, R. Auxin efflux transporter MtPIN10 regulates compound leaf and flower development in Medicago truncatula. Plant Signal. Behav. 2011, 6, 1537–1544. [Google Scholar] [CrossRef]
- Zhou, C.; Han, L.; Hou, C.; Metelli, A.; Qi, L.; Tadege, M.; Mysore, K.S.; Wang, Z.Y. Developmental analysis of a Medicago truncatula smooth leaf margin1 mutant reveals context-dependent effects on compound leaf development. Plant Cell 2011, 23, 2106–2124. [Google Scholar] [CrossRef]
- Ori, N.; Juarez, M.T.; Jackson, D.; Yamaguchi, J.; Banowetz, G.M.; Hake, S. Leaf senescence is delayed in tobacco plants expressing the maize homeobox gene knotted1 under the control of a senescence-activated promoter. Plant Cell 1999, 11, 1073–1080. [Google Scholar]
- Frugis, G.; Giannino, D.; Mele, G.; Nicolodi, C.; Innocenti, A.M.; Chiappetta, A.; Bitonti, M.B.; Dewitte, W.; van Onckelen, H.; Mariotti, D. Are homeobox knotted-like genes and cytokinins the leaf architects? Plant Physiol. 1999, 119, 371–374. [Google Scholar] [CrossRef]
- Sakamoto, T.; Kamiya, N.; Ueguchi-Tanaka, M.; Iwahori, S.; Matsuoka, M. KNOX homeodomain protein directly suppresses the expression of a gibberellin biosynthetic gene in the tobacco shoot apical meristem. Genes Dev. 2001, 15, 581–590. [Google Scholar] [CrossRef]
- Yanai, O.; Shani, E.; Dolezal, K.; Tarkowski, P.; Sablowski, R.; Sandberg, G.; Samach, A.; Ori, N. Arabidopsis KNOXI proteins activate cytokinin biosynthesis. Curr. Biol. 2005, 15, 1566–1571. [Google Scholar] [CrossRef]
- Jasinski, S.; Piazza, P.; Craft, J.; Hay, A.; Woolley, L.; Rieu, I.; Phillips, A.; Hedden, P.; Tsiantis, M. KNOX action in Arabidopsis is mediated by coordinate regulation of cytokinin and gibberellin activities. Curr. Biol. 2005, 15, 1560–1565. [Google Scholar] [CrossRef]
- Bolduc, N.; Hake, S. The maize transcription factor KNOTTED1 directly regulates the gibberellin catabolism gene ga2ox1. Plant Cell 2009, 21, 1647–1658. [Google Scholar] [CrossRef]
- Hewelt, A.; Prinsen, E.; Thomas, M.; van Onckelen, H.; Meins, F., Jr. Ectopic expression of maize knotted1 results in the cytokinin-autotrophic growth of cultured tobacco tissues. Planta 2000, 210, 884–889. [Google Scholar] [CrossRef]
- Frugis, G.; Giannino, D.; Mele, G.; Nicolodi, C.; Chiappetta, A.; Bitonti, M.B.; Innocenti, A.M.; Dewitte, W.; van Onckelen, H.; Mariotti, D. Overexpression of KNAT1 in lettuce shifts leaf determinate growth to a shoot-like indeterminate growth associated with an accumulation of isopentenyl-type cytokinins. Plant Physiol. 2001, 126, 1370–1380. [Google Scholar] [CrossRef]
- Sakamoto, T.; Sakakibara, H.; Kojima, M.; Yamamoto, Y.; Nagasaki, H.; Inukai, Y.; Sato, Y.; Matsuoka, M. Ectopic expression of KNOTTED1-like homeobox protein induces expression of cytokinin biosynthesis genes in rice. Plant Physiol. 2006, 142, 54–62. [Google Scholar] [CrossRef]
- Shani, E.; Ben-Gera, H.; Shleizer-Burko, S.; Burko, Y.; Weiss, D.; Ori, N. Cytokinin regulates compound leaf development in tomato. Plant Cell 2010, 22, 3206–3217. [Google Scholar] [CrossRef]
- Hay, A.; Kaur, H.; Phillips, A.; Hedden, P.; Hake, S.; Tsiantis, M. The gibberellin pathway mediates KNOTTED1-type homeobox function in plants with different body plans. Curr. Biol. 2002, 12, 1557–1565. [Google Scholar] [CrossRef]
- Chen, H.; Banerjee, A.K.; Hannapel, D.J. The tandem complex of BEL and KNOX partners is required for transcriptional repression of ga20ox1. Plant J. 2004, 38, 276–284. [Google Scholar] [CrossRef]
- Kessler, S.; Townsley, B.; Sinha, N. L1 division and differentiation patterns influence shoot apical meristem maintenance. Plant Physiol. 2006, 141, 1349–1362. [Google Scholar] [CrossRef]
- Jasinski, S.; Tattersall, A.; Piazza, P.; Hay, A.; Martinez-Garcia, J.F.; Schmitz, G.; Theres, K.; McCormick, S.; Tsiantis, M. PROCERA encodes a DELLA protein that mediates control of dissected leaf form in tomato. Plant J. 2008, 56, 603–612. [Google Scholar] [CrossRef]
- Bolduc, N.; Yilmaz, A.; Mejia-Guerra, M.K.; Morohashi, K.; O’Connor, D.; Grotewold, E.; Hake, S. Unraveling the KNOTTED1 regulatory network in maize meristems. Genes Dev. 2012, 26, 1685–1690. [Google Scholar] [CrossRef]
- Hay, A.; Barkoulas, M.; Tsiantis, M. PINning down the connections: Transcription factors and hormones in leaf morphogenesis. Curr. Opin. Plant Biol. 2006, 9, 443. [Google Scholar] [CrossRef]
- Heisler, M.G.; Ohno, C.; Das, P.; Sieber, P.; Reddy, G.V.; Long, J.A.; Meyerowitz, E.M. Patterns of auxin transport and gene expression during primordium development revealed by live imaging of the Arabidopsis inflorescence meristem. Curr. Biol. 2005, 15, 1899–1911. [Google Scholar] [CrossRef]
- Gallavotti, A.; Yang, Y.; Schmidt, R.J.; Jackson, D. The Relationship between auxin transport and maize branching. Plant Physiol. 2008, 147, 1913–1923. [Google Scholar] [CrossRef]
- Vernoux, T.; Brunoud, G.; Farcot, E.; Morin, V.; van den Daele, H.; Legrand, J.; Oliva, M.; Das, P.; Larrieu, A.; Wells, D.; et al. The auxin signalling network translates dynamic input into robust patterning at the shoot apex. Mol. Syst. Biol. 2011, 7, 508. [Google Scholar]
- Chapman, E.J.; Estelle, M. Mechanism of auxin-regulated gene expression in plants. Annu. Rev. Genet. 2009, 43, 265–285. [Google Scholar] [CrossRef]
- Di Giacomo, E.; Serino, G.; Frugis, G. Emerging role of the ubiquitin proteasome system in the control of shoot apical meristem function. J. Integr. Plant Biol. 2013, 55, 7–20. [Google Scholar] [CrossRef]
- Yadav, R.K.; Girke, T.; Pasala, S.; Xie, M.; Reddy, G.V. Gene expression map of the Arabidopsis shoot apical meristem stem cell niche. Proc. Natl. Acad. Sci. USA 2009, 106, 4941–4946. [Google Scholar]
- Spinelli, S.V.; Martin, A.P.; Viola, I.L.; Gonzalez, D.H.; Palatnik, J.F. A mechanistic link between STM and CUC1 during Arabidopsis development. Plant Physiol. 2011, 156, 1894–1904. [Google Scholar]
- Tsuda, K.; Ito, Y.; Sato, Y.; Kurata, N. Positive autoregulation of a KNOX gene is essential for shoot apical meristem maintenance in rice. Plant Cell 2011, 23, 4368–4381. [Google Scholar] [CrossRef]
- Mele, G.; Ori, N.; Sato, Y.; Hake, S. The knotted1-like homeobox gene BREVIPEDICELLUS regulates cell differentiation by modulating metabolic pathways. Genes Dev. 2003, 17, 2088–2093. [Google Scholar] [CrossRef]
- Groover, A.; Jones, A.M. Tracheary element differentiation uses a novel mechanism coordinating programmed cell death and secondary cell wall synthesis. Plant Physiol. 1999, 119, 375–384. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Di Giacomo, E.; Iannelli, M.A.; Frugis, G. TALE and Shape: How to Make a Leaf Different. Plants 2013, 2, 317-342. https://doi.org/10.3390/plants2020317
Di Giacomo E, Iannelli MA, Frugis G. TALE and Shape: How to Make a Leaf Different. Plants. 2013; 2(2):317-342. https://doi.org/10.3390/plants2020317
Chicago/Turabian StyleDi Giacomo, Elisabetta, Maria Adelaide Iannelli, and Giovanna Frugis. 2013. "TALE and Shape: How to Make a Leaf Different" Plants 2, no. 2: 317-342. https://doi.org/10.3390/plants2020317