1. Introduction
Salvia miltiorrhiza Bunge (SMB,
danshen in Chinese), belonging to the Labiate family [
1,
2], is an important traditional Chinese herbal plant with a long history as a medicine as well as a health food [
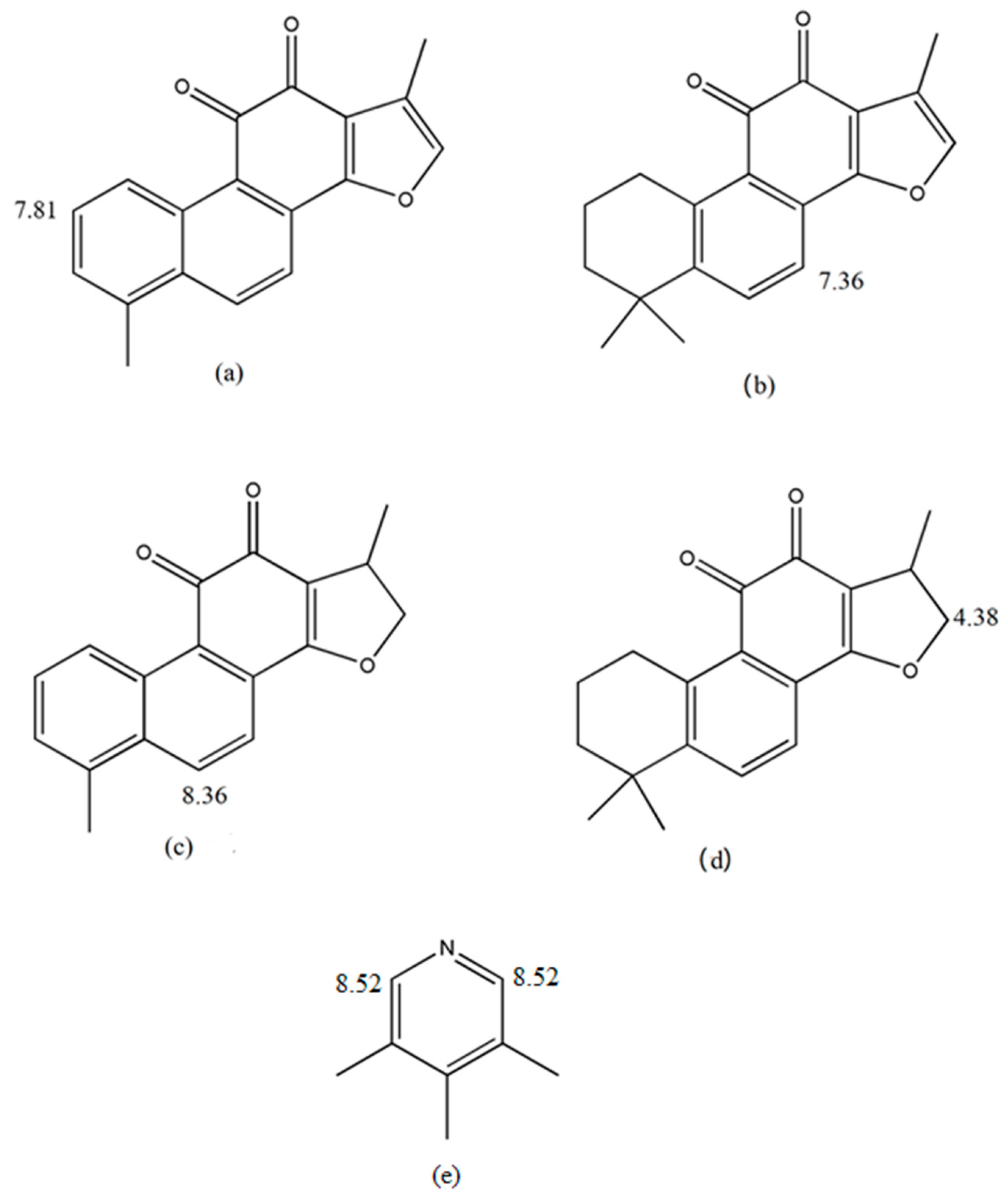
3]. The active ingredients of SMB can be divided into two major groups: lipid-soluble (lipophilic) tanshinones including tanshinone I (Tan I), tanshinone IIA (Tan IIA), tanshinone IIB (Tan IIB), dihydrotanshinone (DTS) and cryptotanshinone (CTS) [
4,
5], and water-soluble (hydrophilic) phenolic acids such as danshensu (DSU), caffeic acid (CA), rosmarinic acid (RA), salvianolic acid A (Sal A) and salvianolic acid B (Sal B) [
6].
Modern pharmacological studies have shown that tanshinones have many pharmacological activities such as (1) antioxidation: CTS has antioxidant in vivo and in vitro pharmacological activities [
7,
8]; Tan IIA can effectively inhibit the interaction of intracellular lipid peroxidation products with DNA, eliminate the lipid free radicals produced by the lipid peroxidation pathway in the mitochondrial membrane of the myocardium, thus protecting the respiration of mitochondria [
9]. (2) Anti-atherosclerosis: Tan IIA can promote cholesterol efflux, ameliorate lipid accumulation in macrophages, and reduce the development of aortic atherosclerosis [
10]. (3) Antibacterial: Gram-positive bacteria can be significantly inhibited by tanshinones such as CTS and DTS. (4) Antitumor: DTS, Tan I, Tan IIA and CTS in blood can inhibit growth and induce the apoptosis of malignant tumor cells [
11]; Tan I exhibits anti-cancer activity on various human cancers which significantly inhibits osteosarcoma (OS) cancer cell proliferation, migration, invasion and induced cell apoptosis in vitro [
12]; CTS shows significant antitumor effects by inducing apoptosis of tumor cells [
13]; DTS exerts an effective antitumor effect by inhibiting tumor cell proliferation and promoting tumor cell apoptosis [
14]; Tan IIA inhibits cell proliferation and induces cell differentiation by affecting the cell cycle; on the other hand, it increases the expression of bax/bcl-2 protein by the Fas pathway to induce apoptosis. In addition, Tan IIA can also choose to activate members of the Caspase family to exert its anti-cancer effect [
15].
With the increasing research on regulation pathways of tanshinones, it is necessary to develop an accurate and sensitive quantitative method for content determination of the tanshinones. A few analytical techniques including HPLC, UHPLC-Q-Exactive Orbitrap mass spectrometry, LC-MS/MS and NMR have been successfully applied for lipid-soluble ketone determinations [
16,
17,
18,
19]. However, to the best of our knowledge, the methodology of quantitative NMR on the extract of SMB fresh roots has not been verified so far. qNMR method is very efficient for the simultaneous detection and identification of several metabolites in crude extracts or samples [
20,
21,
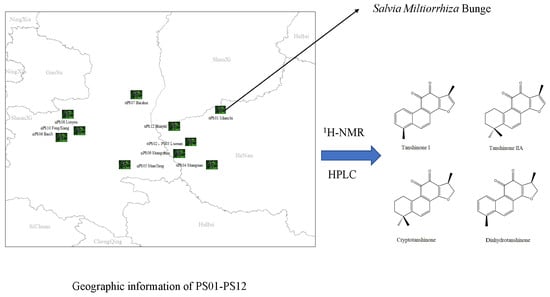
22]. Compared with traditional quantitative methods, qNMR spectroscopy has the following advantages: (1) no calibration standard of the analyte is needed; (2) only an inexpensive internal standard is needed; (3) high selectivity can be achieved under appropriate acquisition conditions; (4) more than one analyte can be determined at one time; (5) reduced measuring time. Thus, establishing a reliable qNMR-based method for measuring lipid-soluble ketone content in SMB is desirable. Furthermore, a quality evaluation model of SMB may be established by combining NMR profiles with chemometrics. Here, we quantified the levels of IS as well as Tan I, Tan IIA, CTS and DTS from fresh SMB root samples using
1H-NMR (
Figure 1). Specifically, 12 batches of SMB were profiled using
1H-NMR. Next, a principal components analysis (PCA) and cluster analysis were conducted to determine the sample correlation. To validate our results, the levels of four ketones were determined using HPLC. We found that the
1H-NMR technique provided a reliable means of quantifying SMB-derived ketones and may be used as a supplementary tool for HPLC-based analyses.
3. Materials and Methods
3.1. Plant Material and Reagents
Fresh SMB roots (PS01–PS12) were obtained from Mianchi (Henan, China), Fengxiang, Baoji, Shangnan, Luonan, Linyou, Baishui, Shanyang, Shangzhou, Tongguan, Huayin (Shaanxi, China) from Shaanxi Tasly Pharmaceutical Co. Ltd. Reference standards Tan I (HPLC ≥ 98%), Tan IIA (HPLC ≥ 98%), CTS (HPLC ≥ 98%) and DTS (HPLC ≥ 98%) were obtained from the Shanghai Yuanye Biotechnology Co. Ltd. (China). Chloroform-D (CDCl3) were purchased from Qingdao Asfirst Science Co. Ltd. (D, 99.8%, Qingdao, China), 3,4,5-trichloropyridine were purchased from Shanghai Aladdin Bio-chemical Technology Co. Ltd. (GC, 98%, Shanghai, China), in which CDCl3 was used as solvent and 3,4,5-trichloropyridine was used as the internal standard (IS). Methanol and acetonitrile of chromatographic grade (TEDIA) were used for extraction and HPLC analysis. Analytical grade phosphoric acid (Damao Chemical Reagent Factory, Tianjin, China) was used for the HPLC analysis. The pure water used in this study was obtained using a Barnstead TII Super Pure Water System (Thermo Fisher Scientific, Boston, MA, USA).
3.2. Instruments and Parameters
Fresh SMB roots samples were milled using a Q-250B1mill (Shanghai, China). A KH5200DE ultrasonic bath (Nanjing, China) was used for extraction.
The samples for qNMR analysis were measured on a 600 MHz Avance III HD spectrometer with a TXI probe at 298K (Bruker Corporation, Faellanden, Switzerland). 1H-NMR experimental parameters were shown as follows: zg30 pulse sequence with 32 scans of 32 K data points in a spectral width of 12019.2 (20 ppm), acquisition time 5.0 s, relaxation delay 25 s. All of the data processing was performed by using MestReNova11.0 and SPSS21.0: first, we use MestReNova’s superposition function to superimpose the internal standard, solvent, sample, and 4 standards, and select the appropriate signal peak. Then, the sample is integrated according to the selected signal peak, and the content of the four components in the sample is calculated by combining with the standard curve. After the above process is completed, SPSS is used for principal component analysis and cluster analysis.
3.3. Preparation of Sample Solution
Approximately 2.0 g of the SMB root powder was added into 20 mL 70% methanol aqueous solution and settled for 12 h at room temperature, then extracted for 45 min in an ultrasonic bath. The supernatant was cooled to room temperature and filtered. Next, 1500 μL of filtered liquor was transferred into a clean Eppendorf tube and nitrogen at 35 °C. The 600 μL of CDCl3 (containing 0.03% v/v TMS) was then added to dissolve the sample for NMR analysis. The NMR sample tube was assembled for analysis at 298 K and sealed prior to the 1H-NMR measurement. For HPLC analysis, the filter liquor was directly injected into the HPLC system. All of the extractions and subsequent NMR measurements were performed in triplicate.
3.4. Internal Standards
The extracted sample PS02 with 600 μL CDCl3, containing 0.13 mg 3,4,5-trichloropyridine was analyzed as IS.
3.5. Quantitative NMR Analysis
The most important fundamental relation of qNMR is the signal response (integrated signal area)
Ix in a spectrum that is directly proportional to the number of nuclei
Nx generating the corresponding resonance line [
27,
28]:
Ks is an unknown spectrometer constant, which is a constant for all resonance lines in the same
1H single-pulse NMR spectrum. Accordingly, the determination of the relative area ratios
Ix/
Iy is the most efficient way to obtain quantitative results by using Equation (5) when
Ks cancels for the ratio:
For the purity determination of a substance, an internal standard with known purity is needed. Based on Equation (5), the component purity can be calculated from the NMR intensity via Equations (6) and (7):
Wx and Px represent the mass and purity of the analyte. Mx and MStd are the molar masses of the analyte and the standard (3,4,5-trichloropyridine: 182.44 g/mol). m is the weighed mass of the investigated sample. mStd and PStd are the weighed mass and the purity (99.5%) of the standard. NStd and IStd correspond to the number of protons for the standard (in this experiment is 2) and the integrated signal area of a typical NMR line (which was 2 in this experiment). Nx and Ix correspond to the number of protons for the analyte 1H.
3.6. HPLC Analysis
The qNMR results were verified using HPLC. The HPLC experiments were performed using an Agilent 1260 HPLC system (Agilent, Palo Alto, CA, USA) equipped with a Diode-array detector and a Sunfire C18 column (5 μm, 250 mm × 4.6 mm i.d., Waters, Milford, MA, USA). The mobile phase consisted of 0.02% phosphoric acid aqueous solution (A) and acetonitrile (B), programmed as follows: 0–10 min, 5–20% B; 10–15 min, 20–25% B; 20–25 min, 25–20% B; 25–28 min, 20–30% B; 28–40 min, 30% B; 40–45 min, 30–45% B; 45–58 min, 45–58% B; 58–67 min, 58–50% B; 67–70 min, 50–60% B; 70–80 min, 60–65% B; 80–85 min, 65–95% B; 85–95 min, 95–95% B; 85–96 min, 95–5% B. The detection wavelength was set at 270 nm and flow rate was 1 mL/min. The content of DTS, Tan IIA, Tan I and CTS in each sample was determined the external standard method and the calibration curve of the corresponding standards.
3.7. Chemometrics Methods
Principal component analysis (PCA) and hierarchical cluster analysis were used to analyze the NMR data.
4. Conclusions
A selective and accurate qNMR method was established for quantitatively determining and validating four lipid-soluble ketones in SMB extracts. The results of the analysis of linearity, precision, stability, accuracy, LOD and LOQ demonstrated that 1H-NMR can be used to accurately determine the content of ketones. We found that the results generated using the qNMR method were consistent with those obtained using HPLC analysis. Whilst HPLC took more than 60 min to complete, qNMR took only 19 min for each sample. Due to the good linearity between IS and the references, qNMR can be performed without any standard references. Although future studies will be required to maximize its application potential, NMR profiles can be combined with chemometric approaches to give insights into the relationship between the quality, properties and geographic origins of plant-based materials. In summary, qNMR presents a rapid and effective method of analyzing various plant extracts and their by-products.