Molecular Mechanisms Underlying Sugarcane Response to Aluminum Stress by RNA-Seq
Abstract
:1. Introduction
2. Results
2.1. Identification of Sugarcane Genotypes with Contrasting Aluminum Stress Tolerance
2.2. Phytotoxic Effects of Aluminum Ions (Al3+) on the Growth of Sugarcane Roots
2.3. De Novo Assembly of RNA-seq Reads for Generation of Reference Transcriptome
2.4. Differential Expression Analysis
2.5. Functional Annotation of Consensus Sequence (Contigs)
2.6. Differential Gene Expression Analysis of Roots Exposed to Aluminum Stress
2.7. Mapping to Functional Pathways
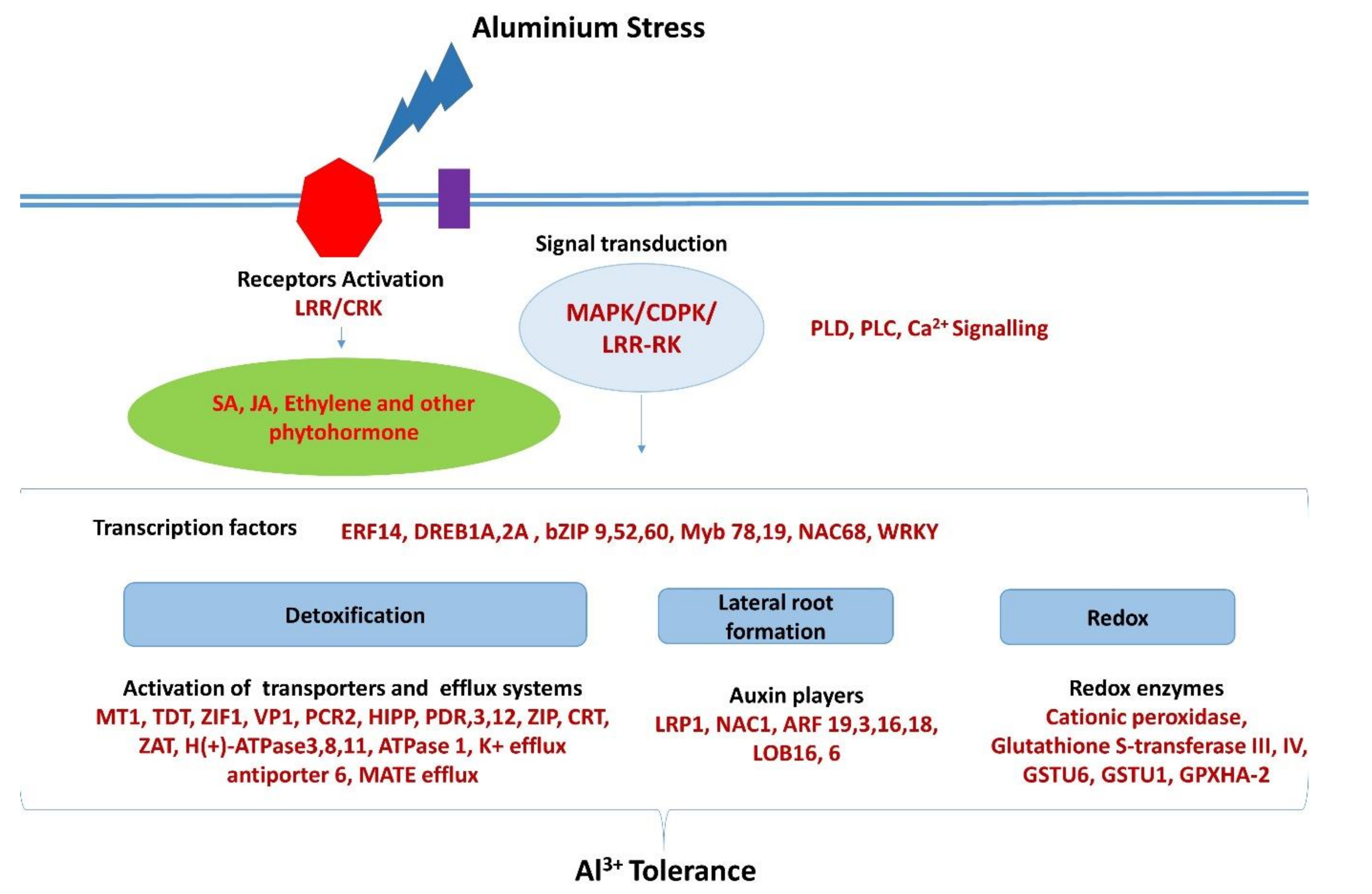
3. Discussion
3.1. Transcriptome Analysis
3.2. Detoxification
3.3. Signaling
3.4. ROS Protection
3.5. Signal Transduction
4. Materials and Methods
4.1. Sample Selection and Preparation of Collection
4.2. Aluminum Abundance in Root
4.3. RNA Extraction, Library Preparation, and Sequencing
4.4. De Novo Transcriptome Assembly and Gene Functional Annotation
4.5. Resequencing for Validation
4.6. Statistical Analysis
4.7. Availability of Data
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| ESTs | expressed sequence tags |
| SUCEST | sugarcane EST |
| SAGE | serial analysis of gene expression |
| TAS | tolerant to aluminum stress |
| SAS | sensitive to aluminum stress |
| X-EDS | energy-dispersive X-ray spectroscopy |
| C | control |
| T | treatment (+Al) |
| DEGs | differentially expressed genes |
| FC | fold-change |
| Reseq | Resequencing |
| NI | not identified |
References
- Rao, I.M.; Miles, J.W.; Beebe, S.E.; Horst, W.J. Root adaptations to soils with low fertility and aluminium toxicity. Ann. Bot. 2016, 118, 593–605. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Von Uexküll, H.R.; Mutert, E. Global extent, development and economic impact of acid soils. Plant Soil 1995, 171, 1–15. [Google Scholar] [CrossRef]
- Vitorello, V.A.; Capaldi, F.R.; Stefanuto, V.A. Recent advances in aluminum toxicity and resistance in higher plants. Braz. J. Plant Physiol. 2005, 17, 129–143. [Google Scholar] [CrossRef]
- Barceló, J.; Poschenrieder, C. Fast root growth responses, root exudates, and internal detoxification as clues to the mechanisms of aluminium toxicity and resistance: A review. Environ. Exp. Bot. 2002, 48, 75–92. [Google Scholar] [CrossRef]
- Klimashevskii, E.L.; Dedov, V.M. Localization of the mechanism of growth inhibiting action of Al3+ in elongating cell walls. Fiziol. Rastenij 1975, 22, 1040–1046. [Google Scholar]
- Matsumoto, H.; Hirasawa, E.; Torikai, H.; Takahashi, E. Localization of absorbed aluminium in pea root and its binding to nucleic acids. Plant Cell Physiol. 1976, 17, 127–137. [Google Scholar] [CrossRef]
- Clarkson, D.T. The effect of aluminium and some other trivalent metal cations on cell division in the root apices of Allium cepa. Ann. Bot. 1965, 29, 309–315. [Google Scholar] [CrossRef]
- Vettore, A.L. Analysis and functional annotation of an expressed sequence tag collection for tropical crop sugarcane. Genome Res. 2003, 13, 2725–2735. [Google Scholar] [CrossRef] [Green Version]
- Vettore, A.L.; da Silva, F.R.; Kemper, E.L.; Arruda, P. The libraries that made SUCEST. Genet. Mol. Biol. 2001, 24, 1–7. [Google Scholar] [CrossRef] [Green Version]
- Rodrigues, F.A.; de Laia, M.L.; Zingaretti, S.M. Analysis of gene expression profiles under water stress in tolerant and sensitive sugarcane plants. Plant Sci. 2009, 176, 286–302. [Google Scholar] [CrossRef]
- Rodrigues, F.A.; Graca, J.P.; Laia, M.L.; Nhani-Jr, A.; Galbiati, J.A.; Ferro, M.I.T.; Ferro, J.A.; Zingaretti, S.M. Sugarcane genes differentially expressed during water deficit. Biol. Plant. 2011, 55, 43–53. [Google Scholar] [CrossRef] [Green Version]
- Vantini, J.S.; Dedemo, G.C.; Jovino Gimenez, D.F.; Fonseca, L.F.S.; Tezza, R.I.D.; Mutton, M.A.; Ferro, J.A.; Ferro, M.I.T. Differential gene expression in drought-tolerant sugarcane roots. Genet. Mol. Res. 2015, 14, 7196–7207. [Google Scholar] [CrossRef]
- Cunha, C.P.; Roberto, G.G.; Vicentini, R.; Lembke, C.G.; Souza, G.M.; Ribeiro, R.V.; Machado, E.C.; Lagôa, A.M.M.A.; Menossi, M. Ethylene-induced transcriptional and hormonal responses at the onset of sugarcane ripening. Sci. Rep. 2017, 7, 43364. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Medeiros, C.D.; Ferreira Neto, J.R.C.; Oliveira, M.T.; Rivas, R.; Pandolfi, V.; Kido, É.A.; Baldani, J.I.; Santos, M.G. Photosynthesis, antioxidant activities and transcriptional responses in two sugarcane (Saccharum officinarum L.) cultivars under salt stress. Acta Physiol. Plant. 2014, 36, 447–459. [Google Scholar] [CrossRef]
- Pacheco, C.M.; Pestana-Calsa, M.C.; Gozzo, F.C.; Mansur Custodio Nogueira, R.J.; Menossi, M.; Calsa Junior, T. Differentially delayed root proteome responses to salt stress in sugarcane varieties. J. Proteome Res. 2013, 12, 5681–5695. [Google Scholar] [CrossRef] [PubMed]
- Belesini, A.A.; Carvalho, F.M.S.; Telles, B.R.; de Castro, G.M.; Giachetto, P.F.; Vantini, J.S.; Carlin, S.D.; Cazetta, J.O.; Pinheiro, D.G.; Ferro, M.I.T. De novo transcriptome assembly of sugarcane leaves submitted to prolonged water-deficit stress. Genet. Mol. Res. 2017, 16, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Mattiello, L.; Riaño-Pachón, D.M.; Martins, M.C.M.; da Cruz, L.P.; Bassi, D.; Marchiori, P.E.R.; Ribeiro, R.V.; Labate, M.T.V.; Labate, C.A.; Menossi, M. Physiological and transcriptional analyses of developmental stages along sugarcane leaf. BMC Plant Biol. 2015, 15, 300. [Google Scholar] [CrossRef] [Green Version]
- Li, M.; Liang, Z.; Zeng, Y.; Jing, Y.; Wu, K.; Liang, J.; He, S.; Wang, G.; Mo, Z.; Tan, F.; et al. De novo analysis of transcriptome reveals genes associated with leaf abscission in sugarcane (Saccharum officinarum L.). BMC Genom. 2016, 17, 195. [Google Scholar] [CrossRef] [Green Version]
- Wang, Z.; Li, Y.; Li, C.; Song, X.; Lei, J.; Gao, Y.; Liang, Q. Comparative transcriptome profiling of resistant and susceptible sugarcane genotypes in response to the airborne pathogen Fusarium verticillioides. Mol. Biol. Rep. 2019, 46, 3777–3789. [Google Scholar] [CrossRef]
- Kumari, M.; Taylor, G.J.; Deyholos, M.K. Transcriptomic responses to aluminum stress in roots of Arabidopsis thaliana. Mol. Genet. Genom. 2008, 279, 339. [Google Scholar] [CrossRef] [PubMed]
- Maron, L.G.; Kirst, M.; Mao, C.; Milner, M.J.; Menossi, M.; Kochian, L.V. Transcriptional profiling of aluminum toxicity and tolerance responses in maize roots. New Phytol. 2008, 179, 116–128. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mattiello, L.; Kirst, M.; da Silva, F.R.; Jorge, R.A.; Menossi, M. Transcriptional profile of maize roots under acid soil growth. BMC Plant Biol. 2010, 10, 196. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, W.; Xiong, C.; Yan, L.; Zhang, Z.; Ma, L.; Wang, Y.; Liu, Y.; Liu, Z. Transcriptome analyses reveal candidate genes potentially involved in al stress response in alfalfa. Front. Plant Sci. 2017, 8, 26. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, S.C.; Chu, S.J.; Guo, Y.M.; Ji, Y.J.; Hu, D.Q.; Cheng, J.; Lu, G.H.; Yang, R.W.; Tang, C.-Y.; Qi, J.L.; et al. Novel mechanisms for organic acid-mediated aluminium tolerance in roots and leaves of two contrasting soybean genotypes. AoB Plants 2017, 9, 1–12. [Google Scholar] [CrossRef] [Green Version]
- Xu, J.M.; Fan, W.; Jin, J.F.; Lou, H.Q.; Chen, W.W.; Yang, J.L.; Zheng, S.J. Transcriptome analysis of Al-induced genes in buckwheat (Fagopyrum esculentum moench) root apex: New insight into al toxicity and resistance mechanisms in an Al accumulating species. Front. Plant Sci. 2017, 8, 1141. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mantovanini, L.J.; da Silva, R.G.; de Oliveira Leite Silva, J.; dos Santos, T.M.R.; dos Santos, D.M.M.; Zingaretti, S.M. Root system development and proline accumulation in sugarcane leaves under aluminum (Al3+) stress. Aust. J. Crop. Sci. 2019, 13, 208–213. [Google Scholar] [CrossRef]
- Thimm, O.; Bläsing, O.; Gibon, Y.; Nagel, A.; Meyer, S.; Krüger, P.; Selbig, J.; Müller, L.A.; Rhee, S.Y.; Stitt, M. MAPMAN: A user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J. 2004, 37, 914–939. [Google Scholar] [CrossRef]
- D’Hont, A.; Souza, G.M.; Menossi, M.; Vincentz, M.; Van-Sluys, M.A.; Glaszmann, J.C.; Ulian, E. Sugarcane: A major source of sweetness, alcohol, and bio-energy. In Genomics of Tropical Crop Plants; Springer: New York, NY, USA, 2008; pp. 483–513. [Google Scholar]
- Kochian, L.V.; Piñeros, M.A.; Hoekenga, O.A. The Physiology, Genetics and Molecular Biology of Plant Aluminum Resistance and Toxicity. Plant Soil 2005, 274, 175–195. [Google Scholar] [CrossRef]
- Ryan, P.R.; Ditomaso, J.M.; Kochian, L.V. Aluminium toxicity in roots: An investigation of spatial sensitivity and the role of the root cap. J. Exp. Bot. 1993, 44, 437–446. [Google Scholar] [CrossRef]
- Ma, J.F.; Shen, R.; Nagao, S.; Tanimoto, E. Aluminum targets elongating cells by reducing cell wall extensibility in wheat roots. Plant Cell Physiol. 2004, 45, 583–589. [Google Scholar] [CrossRef] [Green Version]
- Silva, S. Aluminium toxicity targets in plants. J. Bot. 2012, 2012, 1–8. [Google Scholar] [CrossRef] [Green Version]
- De Souza, L.T.; Cambraia, J.; Ribeiro, C.; de Oliveira, J.A.; da Silva, L.C. Effects of aluminum on the elongation and external morphology of root tips in two maize genotypes. Bragantia 2015, 75, 19–25. [Google Scholar] [CrossRef]
- Cobbett, C.; Goldsbrough, P. Phytochelatins and Metallotioneins: Roles in heavy metal detoxification and homeostasis. Annu. Rev. Plant Biol. 2002, 53, 159–182. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, J.F. Role of organic acids in detoxification of aluminum in higher plants. Plant Cell Physiol. 2000, 41, 383–390. [Google Scholar] [CrossRef] [Green Version]
- Liu, J.; Li, Y.; Wang, W.; Gai, J.; Li, Y. Genome-wide analysis of MATE transporters and expression patterns of a subgroup of MATE genes in response to aluminum toxicity in soybean. BMC Genom. 2016, 17, 223. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Horst, W.J.; Wang, Y.; Eticha, D. The role of the root apoplast in aluminium-induced inhibition of root elongation and in aluminium resistance of plants: A review. Ann. Bot. 2010, 106, 185–197. [Google Scholar] [CrossRef] [PubMed]
- Hamilton, C.A.; Good, A.G.; Taylor, G.J. Induction of vacuolar ATPase and mitochondrial ATP synthase by aluminum in an aluminum-resistant cultivar of wheat. Plant Physiol. 2001, 125, 2068–2077. [Google Scholar] [CrossRef] [Green Version]
- Francis, D.; Sorrell, D.A. The interface between the cell cycle and plant growth regulators: A mini review. Plant Growth Regul. 2001, 33, 1–12. [Google Scholar] [CrossRef]
- Adamowski, M.; Friml, J. PIN-dependent auxin transport: Action, regulation, and evolution. Plant Cell 2015, 27, 20–32. [Google Scholar] [CrossRef] [Green Version]
- Kazan, K. Auxin and the integration of environmental signals into plant root development. Ann. Bot. 2013, 112, 1655–1665. [Google Scholar] [CrossRef] [Green Version]
- Kollmeier, M.; Felle, H.H.; Horst, W.J. Genotypical differences in aluminum resistance of maize are expressed in the distal part of the transition zone. Is reduced basipetal auxin flow involved in inhibition of root elongation by aluminum? Plant Physiol. 2000, 122, 945–956. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gutierrez, L.; Mongelard, G.; Floková, K.; Pǎcurar, D.I.; Novák, O.; Staswick, P.; Kowalczyk, M.; Pǎcurar, M.; Demailly, H.; Geiss, G.; et al. Auxin controls Arabidopsis adventitious root initiation by regulating jasmonic acid homeostasis. Plant Cell 2012, 24, 2515–2527. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dharmasiri, N. Auxin signaling and regulated protein degradation. Trends Plant Sci. 2004, 9, 302–308. [Google Scholar] [CrossRef]
- Ezaki, B.; Gardner, R.C.; Ezaki, Y.; Matsumoto, H. Expression of aluminum-induced genes in transgenic Arabidopsis plants can ameliorate aluminum stress and/or oxidative stress. Plant Physiol. 2000, 122, 657–665. [Google Scholar] [CrossRef] [Green Version]
- Richards, K.D.; Schott, E.J.; Sharma, Y.K.; Davis, K.R.; Gardner, R.C. Aluminum induces oxidative stress genes in Arabidopsis thaliana. Plant Physiol. 1998, 116, 409–418. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sharma, P.; Dubey, R.S. Involvement of oxidative stress and role of antioxidative defense system in growing rice seedlings exposed to toxic concentrations of aluminum. Plant Cell Rep. 2007, 26, 2027–2038. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Yang, Z.; How, J.; Xu, H.; Chen, L.; Li, K. Overexpression of a peroxidase gene (AtPrx64) of Arabidopsis thaliana in tobacco improves plant’s tolerance to aluminum stress. Plant Mol. Biol. 2017, 95, 157–168. [Google Scholar] [CrossRef] [PubMed]
- Hamel, F.; Breton, C.; Houde, M. Isolation and characterization of wheat aluminum-regulated genes: Possible involvement of aluminum as a pathogenesis response elicitor. Planta 1998, 205, 531–538. [Google Scholar] [CrossRef]
- Hoagland, D.R.; Arnon, D.I. The water-culture method for growing plants without soil. Calif. Agric. Exp. Station. Circ. 1950, 347, 1–32. [Google Scholar]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [Green Version]
- Lohse, M.; Nagel, A.; Herter, T.; May, P.; Schroda, M.; Zrenner, R.; Tohge, T.; Fernie, A.R.; Stitt, M.; Usadel, B. Mercator: A fast and simple web server for genome scale functional annotation of plant sequence data. Plant Cell Environ. 2014, 37, 1250–1258. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Babicki, S.; Arndt, D.; Marcu, A.; Liang, Y.; Grant, J.R.; Maciejewski, A.; Wishart, D.S. Heatmapper: Web-enabled heat mapping for all. Nucleic Acids Res. 2016, 44, 147–153. [Google Scholar] [CrossRef] [PubMed]








| Cultivars | Condition | Aluminum Quantification–by EDS* | |
|---|---|---|---|
| Root Cap | Root Elongation | ||
| TAS | Control | 0.46 ± 0.02 | 0.22 ± 0.02 |
| Treatment | 1.76 ± 0.03 ** | 0.44 ± 0.02 ** | |
| SAS | Control | 0.41 ± 0.02 | 0.05 ± 0.02 |
| Treatment | 1.37 ± 0.03 ** | 0.45 ± 0.02 ** | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rosa-Santos, T.M.; Silva, R.G.d.; Kumar, P.; Kottapalli, P.; Crasto, C.; Kottapalli, K.R.; França, S.C.; Zingaretti, S.M. Molecular Mechanisms Underlying Sugarcane Response to Aluminum Stress by RNA-Seq. Int. J. Mol. Sci. 2020, 21, 7934. https://doi.org/10.3390/ijms21217934
Rosa-Santos TM, Silva RGd, Kumar P, Kottapalli P, Crasto C, Kottapalli KR, França SC, Zingaretti SM. Molecular Mechanisms Underlying Sugarcane Response to Aluminum Stress by RNA-Seq. International Journal of Molecular Sciences. 2020; 21(21):7934. https://doi.org/10.3390/ijms21217934
Chicago/Turabian StyleRosa-Santos, Thiago Mateus, Renan Gonçalves da Silva, Poornasree Kumar, Pratibha Kottapalli, Chiquito Crasto, Kameswara Rao Kottapalli, Suzelei Castro França, and Sonia Marli Zingaretti. 2020. "Molecular Mechanisms Underlying Sugarcane Response to Aluminum Stress by RNA-Seq" International Journal of Molecular Sciences 21, no. 21: 7934. https://doi.org/10.3390/ijms21217934







