Microtubule Dynamics Plays a Vital Role in Plant Adaptation and Tolerance to Salt Stress
Abstract
:1. Introduction
2. Results
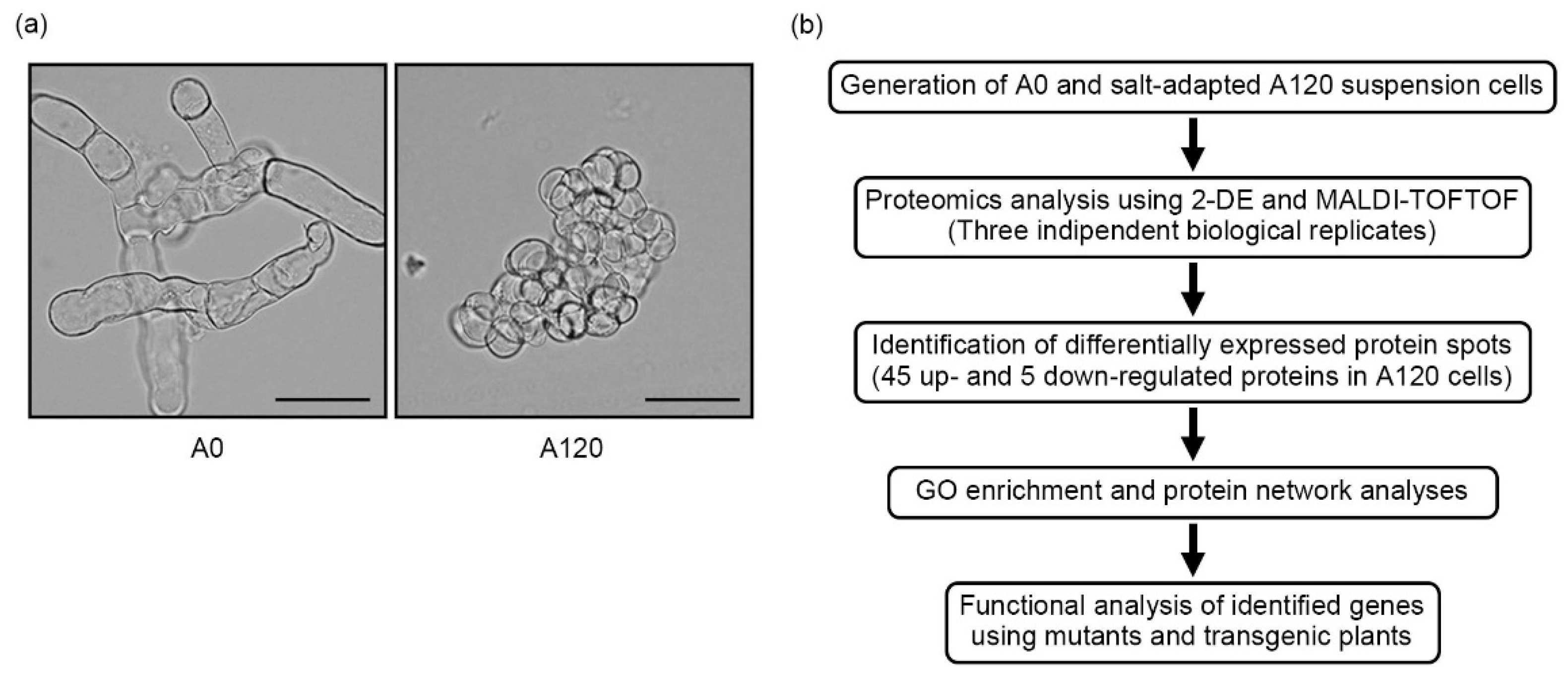
2.1. Morphological Features of Salt-Adapted Callus Suspension-Cultured Cells
2.2. Overview of Proteomic Profiles in Salt-Adapted Cells
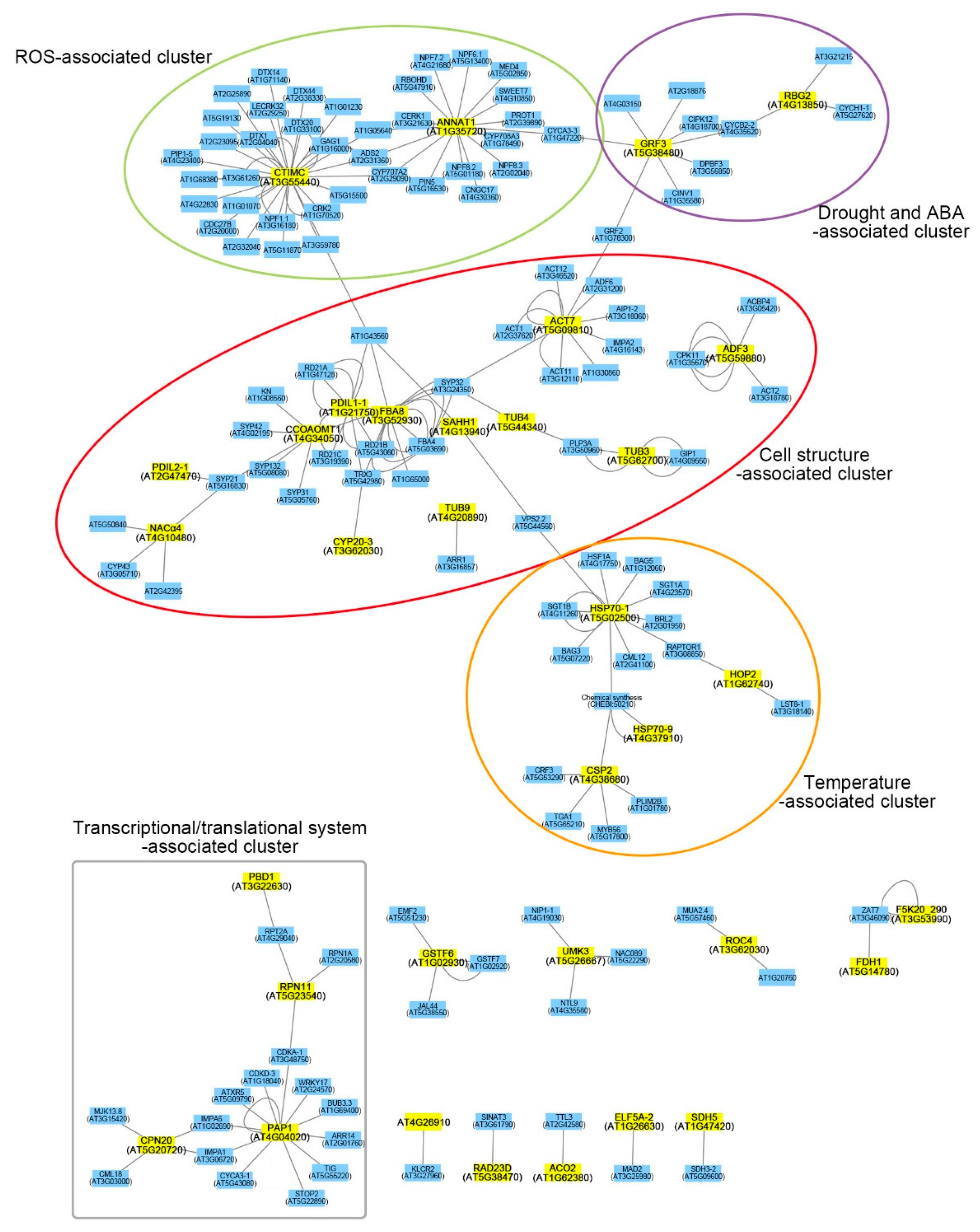
2.3. Functional Network Analysis of Differentially Expressed Proteins
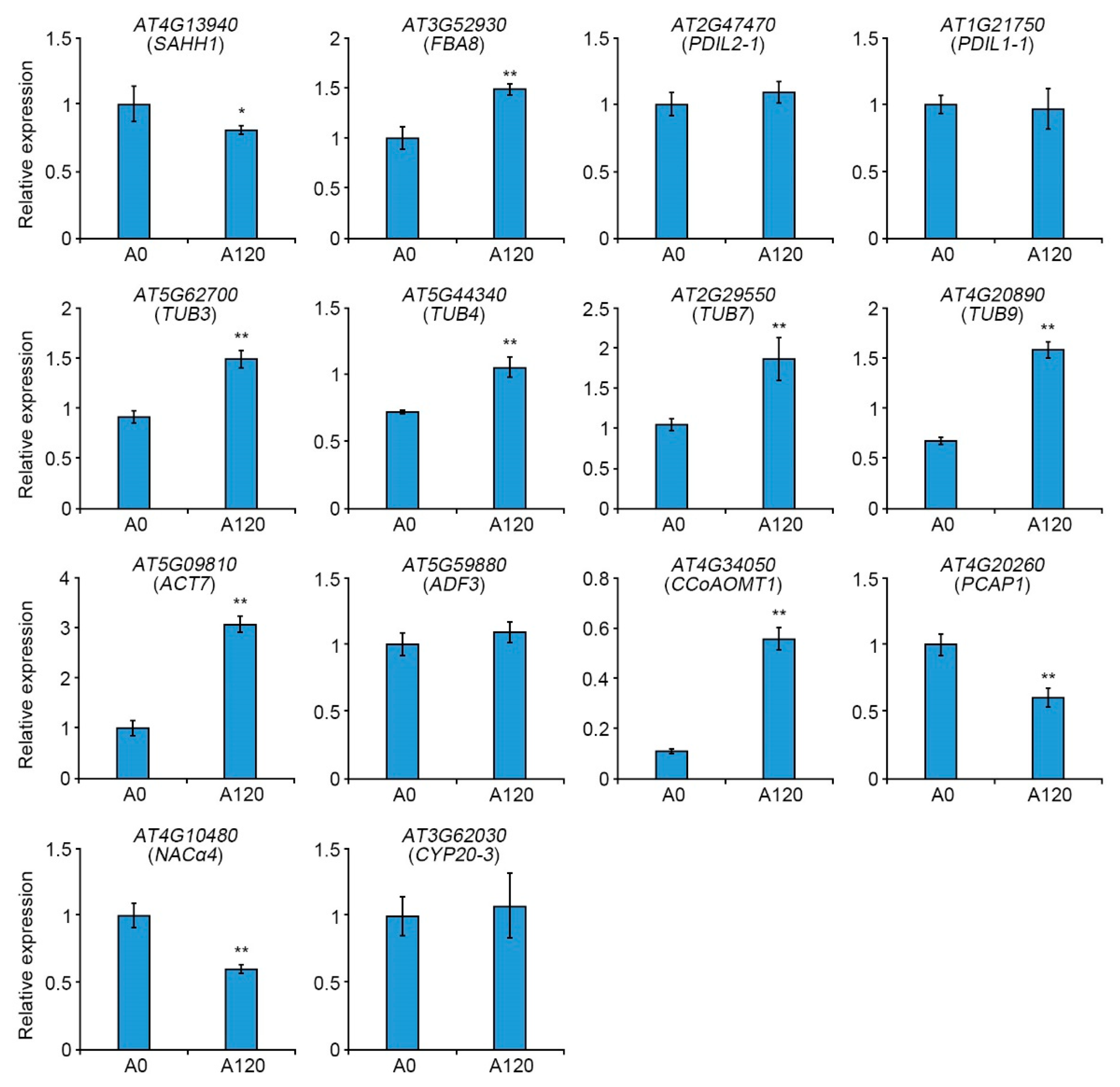
2.4. Expression Patterns of Cytoskeleton-Related Genes in Salt-Adapted Cells
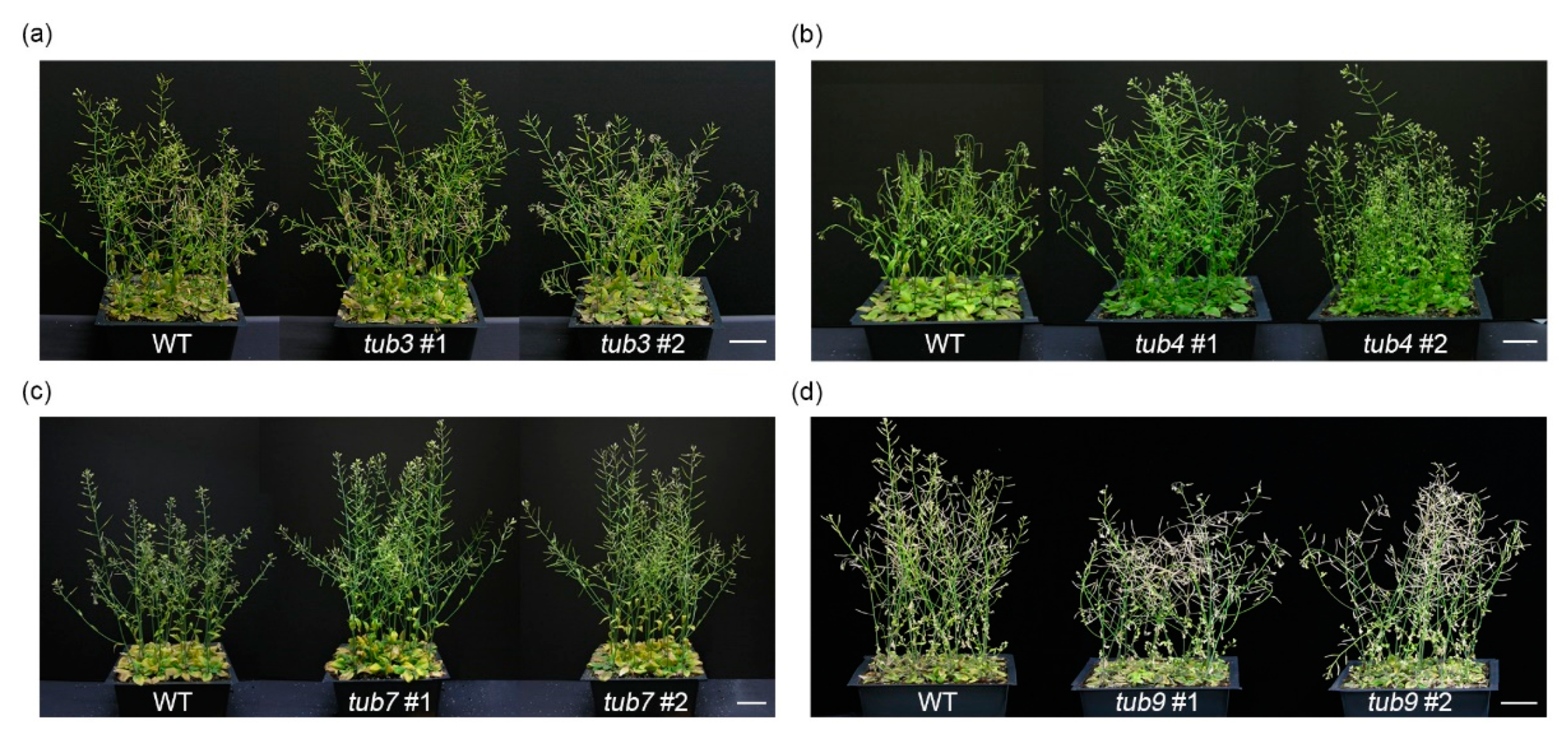
2.5. The Effect of the Loss-of-Function β-Tubulin Genes in Salt Stress Response
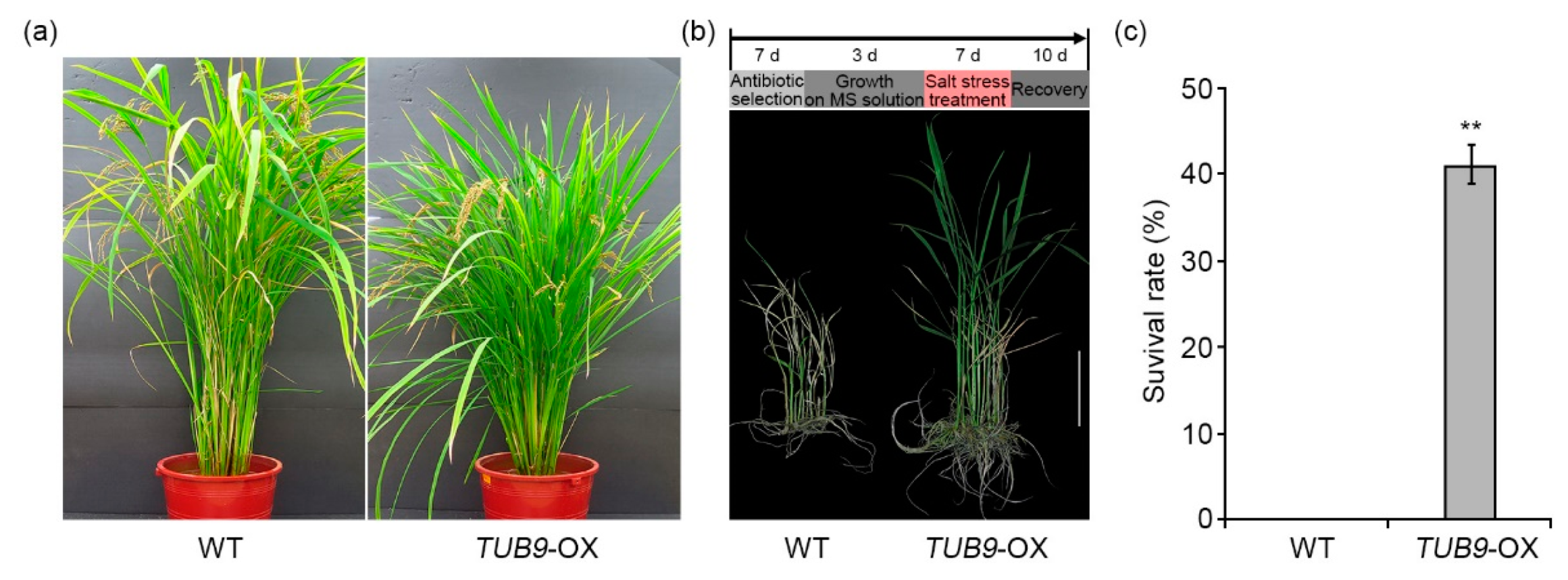
2.6. The Effect of TUB9 Overexpression in Rice during Salt Stress
3. Discussion
3.1. Molecular Functions of Differentially Expressed Proteins in Salt-Adapted Cells
3.1.1. Cell Structure-Associated Cluster
3.1.2. ROS-Associated Cluster
3.1.3. Drought- and ABA-Associated Cluster
3.1.4. Temperature-Associated Cluster
3.1.5. Transcriptional/Translational System-Associated Cluster
3.2. The Role of Microtubules in Plant Adaptation and Tolerance to Salt Stress
4. Materials and Methods
4.1. Growth Conditions of Callus Suspension Cells
4.2. Proteomic Profiling Using Two-Dimensional Gel Electrophoresis
4.3. Bioinformatics Analysis
4.4. Analysis of Quantitative Real Time PCR (qRT-PCR)
4.5. Plant Materials and Growth Conditions
4.6. Generation of Transgenic Rice Plants
4.7. Salt Stress Treatment
4.8. Statistical Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Vinocur, B.; Altman, A. Recent advances in engineering plant tolerance to abiotic stress: Achievements and limitations. Curr. Opin. Biotechnol. 2005, 16, 123–132. [Google Scholar] [CrossRef] [PubMed]
- Tenhaken, R. Cell wall remodeling under abiotic stress. Front. Plant Sci. 2015, 5, 771. [Google Scholar] [CrossRef] [Green Version]
- Zhu, J.-K. Abiotic Stress Signaling and Responses in Plants. Cell 2016, 167, 313–324. [Google Scholar] [CrossRef] [Green Version]
- Lü, P.; Kang, M.; Jiang, X.; Dai, F.; Gao, J.; Zhang, C. RhEXPA4, a rose expansin gene, modulates leaf growth and confers drought and salt tolerance to Arabidopsis. Planta 2013, 237, 1547–1559. [Google Scholar] [CrossRef] [PubMed]
- An, P.; Li, X.; Zheng, Y.; Matsuura, A.; Abe, J.; Eneji, A.E.; Tanimoto, E.; Inanaga, S. Effects of NaCl on Root Growth and Cell Wall Composition of Two Soya bean Cultivars with Contrasting Salt Tolerance. J. Agron. Crop. Sci. 2014, 200, 212–218. [Google Scholar] [CrossRef]
- Zhu, J.; Lee, B.-H.; Dellinger, M.; Cui, X.; Zhang, C.; Wu, S.; Nothnagel, E.A.; Zhu, J.-K. A cellulose synthase-like protein is required for osmotic stress tolerance in Arabidopsis. Plant J. 2010, 63, 128–140. [Google Scholar] [CrossRef] [Green Version]
- Endler, A.; Kesten, C.; Schneider, R.; Zhang, Y.; Ivakov, A.; Froehlich, A.; Funke, N.; Persson, S. A Mechanism for Sustained Cellulose Synthesis during Salt Stress. Cell 2015, 162, 1353–1364. [Google Scholar] [CrossRef] [Green Version]
- Henty-Ridilla, J.L.; Li, J.; Blanchoin, L.; Staiger, C.J. Actin dynamics in the cortical array of plant cells. Curr. Opin. Plant Biol. 2013, 16, 678–687. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ehrhardt, D.W.; Shaw, S. Microtubule Dynamics and Organization in The Plant Cortical Array. Annu. Rev. Plant Biol. 2006, 57, 859–875. [Google Scholar] [CrossRef]
- Dixit, R.; Cyr, R. The Cortical Microtubule Array: From Dynamics to Organization. Plant Cell 2004, 16, 2546–2552. [Google Scholar] [CrossRef] [Green Version]
- Blanchoin, L.; Boujemaa-Paterski, R.; Henty, J.L.; Khurana, P.; Staiger, C.J. Actin dynamics in plant cells: A team effort from multiple proteins orchestrates this very fast-paced game. Curr. Opin. Plant Biol. 2010, 13, 714–723. [Google Scholar] [CrossRef]
- Li, J.; Staiger, C.J. Understanding Cytoskeletal Dynamics During the Plant Immune Response. Annu. Rev. Phytopathol. 2018, 56, 513–533. [Google Scholar] [CrossRef]
- Wang, X.; Mao, T. Understanding the functions and mechanisms of plant cytoskeleton in response to environmental signals. Curr. Opin. Plant Biol. 2019, 52, 86–96. [Google Scholar] [CrossRef]
- Smith, L.G.; Oppenheimer, D.G. Spatial Control of Cell Expansion by The Plant Cytoskeleton. Annu. Rev. Cell Dev. Biol. 2005, 21, 271–295. [Google Scholar] [CrossRef] [Green Version]
- Hashimoto, T.; Kato, T. Cortical control of plant microtubules. Curr. Opin. Plant Biol. 2006, 9, 5–11. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Kurepa, J.; Hashimoto, T.; Smalle, J.A. Salt Stress–Induced Disassembly of Arabidopsis Cortical Microtubule Arrays Involves 26S Proteasome–Dependent Degradation of SPIRAL1. Plant Cell 2011, 23, 3412–3427. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, C.; Li, J.; Yuan, M. Salt Tolerance Requires Cortical Microtubule Reorganization in Arabidopsis. Plant Cell Physiol. 2007, 48, 1534–1547. [Google Scholar] [CrossRef] [Green Version]
- Wang, C.; Zhang, L.-J.; Huang, R.-D. Cytoskeleton and plant salt stress tolerance. Plant Signal. Behav. 2011, 6, 29–31. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Livanos, P.; Galatis, B.; Quader, H.; Apostolakos, P. Disturbance of reactive oxygen species homeostasis induces atypical tubulin polymer formation and affects mitosis in root-tip cells of Triticum turgidum and Arabidopsis thaliana. Cytoskeleton 2012, 69, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Wu, K.; Lin, F.; Qu, Y.; Liu, X.; Zhang, Q. Phosphatidic acid integrates calcium signaling and microtubule dynamics into regulating ABA-induced stomatal closure in Arabidopsis. Planta 2013, 239, 565–575. [Google Scholar] [CrossRef]
- Takatani, S.; Hirayama, T.; Hashimoto, T.; Takahashi, T.; Motose, H. Abscisic acid induces ectopic outgrowth in epidermal cells through cortical microtubule reorganization in Arabidopsis thaliana. Sci. Rep. 2015, 5, 11364. [Google Scholar] [CrossRef] [Green Version]
- Bogoutdinova, L.R.; Lazareva, E.M.; Chaban, I.A.; Kononenko, N.V.; Dilovarova, T.; Khaliluev, M.R.; Kurenina, L.V.; Gulevich, A.A.; Smirnova, E.A.; Baranova, E.N. Salt Stress-Induced Structural Changes Are Mitigated in Transgenic Tomato Plants Over-Expressing Superoxide Dismutase. Biology 2020, 9, 297. [Google Scholar] [CrossRef]
- Fujita, S.; Pytela, J.; Hotta, T.; Kato, T.; Hamada, T.; Akamatsu, R.; Ishida, Y.; Kutsuna, N.; Hasezawa, S.; Nomura, Y.; et al. An Atypical Tubulin Kinase Mediates Stress-Induced Microtubule Depolymerization in Arabidopsis. Curr. Biol. 2013, 23, 1969–1978. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Q.; Lin, F.; Mao, T.; Nie, J.; Yan, M.; Yuan, M.; Zhang, W. Phosphatidic Acid Regulates Microtubule Organization by Interacting with MAP65-1 in Response to Salt Stress in Arabidopsis. Plant Cell 2012, 24, 4555–4576. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, C.; Zhang, L.; Yuan, M.; Ge, Y.; Liu, Y.; Fan, J.; Ruan, Y.; Cui, Z.; Tong, S.; Zhang, S. The microfilament cytoskeleton plays a vital role in salt and osmotic stress tolerance in Arabidopsis. Plant Biol. 2009, 12, 70–78. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Zhang, W.; Guo, Y. Arabidopsis SOS3 plays an important role in salt tolerance by mediating calcium-dependent microfilament reorganization. Plant Cell Rep. 2013, 32, 139–148. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Pan, Z.; Zhang, Y.; Qu, X.; Zhang, Y.; Yang, Y.; Jiang, X.; Huang, S.; Yuan, M.; Schumaker, K.S.; et al. The Actin-Related Protein2/3 Complex Regulates Mitochondrial-Associated Calcium Signaling during Salt Stress in Arabidopsis. Plant Cell 2013, 25, 4544–4559. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rodríguez-Milla, M.A.; Salinas, J. Prefoldins 3 and 5 Play an Essential Role in Arabidopsis Tolerance to Salt Stress. Mol. Plant 2009, 2, 526–534. [Google Scholar] [CrossRef] [Green Version]
- Barkla, B.J.; Vera-Estrella, R.; Pantoja, O. Progress and challenges for abiotic stress proteomics of crop plants. Proteomics 2013, 13, 1801–1815. [Google Scholar] [CrossRef]
- Silveira, J.A.; Carvalho, F.E. Proteomics, photosynthesis and salt resistance in crops: An integrative view. J. Proteom. 2016, 143, 24–35. [Google Scholar] [CrossRef]
- Lv, X.; Chen, S.; Wang, Y. Advances in Understanding the Physiological and Molecular Responses of Sugar Beet to Salt Stress. Front. Plant Sci. 2019, 10, 1431. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.; Chang, L.; Wang, B.; Wang, D.; Li, P.; Wang, L.; Yi, X.; Huang, Q.; Peng, M.; Guo, A. Comparative Proteomics of Thellungiella halophila Leaves from Plants Subjected to Salinity Reveals the Importance of Chloroplastic Starch and Soluble Sugars in Halophyte Salt Tolerance. Mol. Cell. Proteom. 2013, 12, 2174–2195. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, J.; Meng, Y.; Li, B.; Ma, X.; Lai, Y.; Si, E.; Yang, K.; Xu, X.; Shang, X.; Wang, H.; et al. Physiological and proteomic analyses of salt stress response in the halophyte H alogeton glomeratus. Plant Cell Environ. 2015, 38, 655–669. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cheng, T.; Chen, J.; Zhang, J.; Shi, S.; Zhou, Y.; Lu, L.; Wang, P.; Jiang, Z.; Yang, J.; Zhang, S.; et al. Physiological and proteomic analyses of leaves from the halophyte Tangut Nitraria reveals diverse response pathways critical for high salinity tolerance. Front. Plant Sci. 2015, 6, 30. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shokri-Gharelo, R.; Noparvar, P.M. Molecular response of canola to salt stress: Insights on tolerance mechanisms. Peer J 2018, 6, e4822. [Google Scholar] [CrossRef]
- Zhang, Y.; Wei, M.; Liu, A.; Zhou, R.; Li, D.; Dossa, K.; Wang, L.; Zhang, Y.; Gong, H.; Zhang, X.; et al. Comparative proteomic analysis of two sesame genotypes with contrasting salinity tolerance in response to salt stress. J. Proteom. 2019, 201, 73–83. [Google Scholar] [CrossRef]
- Frukh, A.; Siddiqi, T.O.; Khan, M.I.R.; Ahmad, A. Modulation in growth, biochemical attributes and proteome profile of rice cultivars under salt stress. Plant Physiol. Biochem. 2020, 146, 55–70. [Google Scholar] [CrossRef]
- Parker, R.; Flowers, T.J.; Moore, A.L.; Harpham, N.V.J. An accurate and reproducible method for proteome profiling of the effects of salt stress in the rice leaf lamina. J. Exp. Bot. 2006, 57, 1109–1118. [Google Scholar] [CrossRef]
- Sobhanian, H.; Aghaei, K.; Komatsu, S. Changes in the plant proteome resulting from salt stress: Toward the creation of salt-tolerant crops? J. Proteom. 2011, 74, 1323–1337. [Google Scholar] [CrossRef]
- Chun, H.J.; Baek, D.; Cho, H.M.; Jung, H.S.; Jeong, M.S.; Jung, W.-H.; Choi, C.W.; Lee, S.H.; Jin, B.J.; Park, M.S.; et al. Metabolic Adjustment of Arabidopsis Root Suspension Cells During Adaptation to Salt Stress and Mitotic Stress Memory. Plant Cell Physiol. 2018, 60, 612–625. [Google Scholar] [CrossRef]
- Le Gall, H.; Philippe, F.; Domon, J.-M.; Gillet, F.; Pelloux, J.; Rayon, C. Cell Wall Metabolism in Response to Abiotic Stress. Plants 2015, 4, 112–166. [Google Scholar] [CrossRef]
- Kwon, Y.S.; Kim, S.G.; Chung, W.S.; Bae, H.; Jeong, S.W.; Shin, S.C.; Jeong, M.-J.; Park, S.-C.; Kwak, Y.-S.; Bae, D.-W.; et al. Proteomic analysis of Rhizoctonia solani AG-1 sclerotia maturation. Fungal Biol. 2014, 118, 433–443. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Wang, X.; Qin, T.; Zhang, Y.; Liu, X.; Sun, J.; Zhou, Y.; Zhu, L.; Zhang, Z.; Yuan, M.; et al. MDP25, A Novel Calcium Regulatory Protein, Mediates Hypocotyl Cell Elongation by Destabilizing Cortical Microtubules in Arabidopsis. Plant Cell 2011, 23, 4411–4427. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shoji, T.; Suzuki, K.; Abe, T.; Kaneko, Y.; Shi, H.; Zhu, J.-K.; Rus, A.; Hasegawa, P.M.; Hashimoto, T. Salt Stress Affects Cortical Microtubule Organization and Helical Growth in Arabidopsis. Plant Cell Physiol. 2006, 47, 1158–1168. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Guo, Y. Elucidating the molecular mechanisms mediating plant salt-stress responses. New Phytol. 2018, 217, 523–539. [Google Scholar] [CrossRef] [Green Version]
- Qi, F.; Zhang, F. Cell Cycle Regulation in the Plant Response to Stress. Front. Plant Sci. 2020, 10, 1765. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, H.; Liu, M. The microtubule cytoskeleton acts as a sensor for stress response signaling in plants. Mol. Biol. Rep. 2019, 46, 5603–5608. [Google Scholar] [CrossRef]
- Liu, D.; Ford, K.L.; Roessner, U.; Natera, S.; Cassin, A.M.; Patterson, J.H.; Bacic, A. Rice suspension cultured cells are evaluated as a model system to study salt responsive networks in plants using a combined proteomic and metabolomic profiling approach. Proteomics 2013, 13, 2046–2062. [Google Scholar] [CrossRef]
- Wang, J.; Yao, L.; Li, B.; Meng, Y.; Ma, X.; Lai, Y.; Si, E.; Ren, P.; Yang, K.; Shang, X.; et al. Comparative Proteomic Analysis of Cultured Suspension Cells of the Halophyte Halogeton glomeratus by iTRAQ Provides Insights into Response Mechanisms to Salt Stress. Front. Plant Sci. 2016, 7, 110. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McDowell, J.M.; Huang, S.; McKinney, E.C.; An, Y.Q.; Meagher, R.B. Structure and evolution of the actin gene family in Ar-abidopsis thaliana. Genetics 1996, 142, 587–602. [Google Scholar] [CrossRef] [PubMed]
- Chang, M.; Huang, S. ArabidopsisACT11 modifies actin turnover to promote pollen germination and maintain the normal rate of tube growth. Plant J. 2015, 83, 515–527. [Google Scholar] [CrossRef]
- Kandasamy, M.K.; McKinney, E.C.; Meagher, R.B. A Single Vegetative Actin Isovariant Overexpressed under the Control of Multiple Regulatory Sequences Is Sufficient for Normal Arabidopsis Development. Plant Cell 2009, 21, 701–718. [Google Scholar] [CrossRef] [Green Version]
- Mondal, H.A.; Louis, J.; Archer, L.; Patel, M.; Nalam, V.J.; Sarowar, S.; Sivapalan, V.; Root, D.D.; Shah, J. Arabidopsis Actin-Depolymerizing Factor3 Is Required for Controlling Aphid Feeding from the Phloem. Plant Physiol. 2018, 176, 879–890. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Salekdeh, G.H.; Siopongco, J.; Wade, L.J.; Ghareyazie, B.; Bennett, J. Proteomic analysis of rice leaves during drought stress and recovery. Proteomics 2002, 2, 1131–1145. [Google Scholar] [CrossRef]
- Yan, S.; Tang, Z.; Su, W.; Sun, W. Proteomic analysis of salt stress-responsive proteins in rice root. Proteomics 2005, 5, 235–244. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.-C.; Huang, W.-L.; Hong, C.-Y.; Lur, H.-S.; Chang, M.-C. Comprehensive analysis of differentially expressed rice actin depolymerizing factor gene family and heterologous overexpression of OsADF3 confers Arabidopsis Thaliana drought tolerance. Rice 2012, 5, 33. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lu, W.; Tang, X.; Huo, Y.; Xu, R.; Qi, S.; Huang, J.; Zheng, C.; Wu, C. Identification and characterization of fructose 1,6-bisphosphate aldolase genes in Arabidopsis reveal a gene family with diverse responses to abiotic stresses. Gene 2012, 503, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.-Q.; Yang, B.; Harris, N.S.; Deyholos, M.K. Comparative proteomic analysis of NaCl stress-responsive proteins in Arabidopsis roots. J. Exp. Bot. 2007, 58, 3591–3607. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chun, H.J.; Baek, D.; Cho, H.M.; Lee, S.H.; Jin, B.J.; Yun, D.-J.; Hong, Y.-S.; Kim, M.C. Lignin biosynthesis genes play critical roles in the adaptation of Arabidopsis plants to high-salt stress. Plant Signal. Behav. 2019, 14, 1625697. [Google Scholar] [CrossRef]
- Chun, H.; Lim, L.; Cheong, M.; Baek, D.; Park, M.; Cho, H.; Lee, S.; Jin, B.; No, D.; Cha, Y.; et al. Arabidopsis CCoAOMT1 Plays a Role in Drought Stress Response via ROS- and ABA-Dependent Manners. Plants 2021, 10, 831. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Boavida, L.C.; Ron, M.; McCormick, S. Truncation of a Protein Disulfide Isomerase, PDIL2-1, Delays Embryo Sac Maturation and Disrupts Pollen Tube Guidance in Arabidopsis thaliana. Plant Cell 2009, 20, 3300–3311. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhu, C.; Luo, N.; He, M.; Chen, G.; Zhu, J.; Yin, G.; Li, X.; Hu, Y.; Li, J.; Yan, Y. Molecular Characterization and Expression Profiling of the Protein Disulfide Isomerase Gene Family in Brachypodium distachyon L. PLoS ONE 2014, 9, e94704. [Google Scholar] [CrossRef]
- Dominguez-Solis, J.R.; He, Z.; Lima, A.; Ting, J.; Buchanan, B.B.; Luan, S. A cyclophilin links redox and light signals to cysteine biosynthesis and stress responses in chloroplasts. Proc. Natl. Acad. Sci. USA 2008, 105, 16386–16391. [Google Scholar] [CrossRef] [Green Version]
- Maršálová, L.; Vítámvás, P.; Hynek, R.; Prášil, I.T.; Kosová, K. Proteomic Response of Hordeum vulgare cv. Tadmor and Hordeum marinum to Salinity Stress: Similarities and Differences between a Glycophyte and a Halophyte. Front. Plant Sci. 2016, 7, 1154. [Google Scholar] [CrossRef]
- Yang, Y.; Guo, Y. Unraveling salt stress signaling in plants. J. Integr. Plant Biol. 2018, 60, 796–804. [Google Scholar] [CrossRef] [Green Version]
- Huh, S.M.; Noh, E.K.; Kim, H.G.; Jeon, B.W.; Bae, K.; Hu, H.-C.; Kwak, J.M.; Park, O.K. Arabidopsis Annexins AnnAt1 and AnnAt4 Interact with Each Other and Regulate Drought and Salt Stress Responses. Plant Cell Physiol. 2010, 51, 1499–1514. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, S.; Chen, Q.; Li, X.; Li, Y. MAP65-1 is required for the depolymerization and reorganization of cortical microtubules in the response to salt stress in Arabidopsis. Plant Sci. 2017, 264, 112–121. [Google Scholar] [CrossRef]
- Errabii, T.; Gandonou, C.B.; Essalmani, H.; Abrini, J.; Idaomar, M.; Senhaji, N.S. Effects of NaCl and mannitol induced stress on sugarcane (Saccharum sp.) callus cultures. Acta Physiol. Plant. 2007, 29, 95–102. [Google Scholar] [CrossRef]
- Hiei, Y.; Ohta, S.; Komari, T.; Kumashiro, T. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J. 1994, 6, 271–282. [Google Scholar] [CrossRef] [Green Version]



| Spot No. a | Locus No. | Protein Name | Theo. Mr/pI b | Queries Matched c | Scores d | Expect | Fold (A0 vs. A120) | |
|---|---|---|---|---|---|---|---|---|
| 1, 28 | AT1G56070 | LOS1 | Low expression of osmotically responsive genes 1 | 95.10/ 5.89 | 31 | 566 | 3.70 × 10−50 | 1.554 |
| 2 | AT1G62740 | HOP2 | Stress-inducible protein, putative | 67.63/ 6.24 | 22 | 349 | 1.90 × 10−28 | 2.538 |
| 3 | AT4G13940 | SAHH1 | S-adenosyl-L-homocysteine hydrolase 1 | 53.97/ 5.66 | 27 | 522 | 9.40 × 10−46 | −1.133 |
| 4 | AT1G51710 | UBP6 | Ubiquitin carboxyl-terminal hydrolase 6 | 54.00/ 5.82 | 27 | 467 | 3.00 × 10−40 | 2.473 |
| 5 | AT4G01850 | SAM-2 | S-adenosylmethionine synthase 2 | 43.63/ 5.67 | 28 | 616 | 3.70 × 10−55 | 1.932 |
| 6 | AT2G36880 | MAT3 | Methionine adenosyltransferase 3 | 42.93/ 5.76 | 33 | 699 | 1.90 × 10−63 | 1.249 |
| 7 | AT1G77120 | ADH1 | Alcohol dehydrogenase class-P | 41.84/ 5.83 | 27 | 798 | 2.40 × 10−73 | −1.614 |
| 8 | AT4G02930 | GTP binding Elongation factor Tu family protein | 49.61/ 6.25 | 33 | 1100 | 1.50 × 10−103 | 1.129 | |
| 9 | AT3G51800 | CPR | Metallopeptidase M24 family protein | 43.28/ 6.36 | 17 | 280 | 1.50 × 10−21 | 4.172 |
| 10 | AT5G14780 | FDH1 | Formate dehydrogenase, chloroplastic/mitochondrial | 42.67/ 7.12 | 25 | 560 | 1.50 × 10−49 | 2.048 |
| 11 | AT4G26910 | Dihydrolipoamide succinyltransferase | 50.03/ 9.21 | 13 | 185 | 4.70 × 10−12 | 2.026 | |
| 12, 50 | AT3G04120 | GAPC1 | Glyceraldehyde-3-phosphate dehydrogenase 1, cytosolic | 37.01/ 6.62 | 30 | 1040 | 1.50 × 10−97 | 4.510 |
| 13 | AT5G43330 | MDH2 | Lactate/malate dehydrogenase family protein | 35.98/ 7.00 | 24 | 675 | 4.70 * 10−61 | 1.407 |
| 14 | AT5G23540 | RPN11 | 26S proteasome non-ATPase regulatory subunit 14 homolog | 34.39/ 6.31 | 18 | 241 | 1.20 × 10−17 | 5.121 |
| 15 | AT3G52930 | FBA8 | Fructose-bisphosphate aldolase 8, cytosolic | 38.86/ 6.05 | 31 | 1090 | 1.50 × 10−102 | 1.031 |
| 16 | AT5G65020 | ANNAT2 | Annexin D2, calcium binding proteins | 36.36/ 5.76 | 27 | 339 | 1.9 × 10−27 | 1.426 |
| 17 | AT2G47470 | PDIL2-1 | Disulfide isomerase-like (PDIL) protein | 39.81/ 5.80 | 20 | 576 | 3.70 × 10−51 | 1.308 |
| 18 | AT5G02500 | HSP70-1 | Arabidopsis thaliana heat shock cognate protein 70-1 | 57.54/ 5.01 | 34 | 844 | 5.90 × 10−78 | −4.195 |
| 19 | AT1G21750 | PDIL1-1 | Disulfide isomerase-like (PDIL) protein | 55.85/ 4.81 | 23 | 457 | 3.00 × 10−39 | 1.466 |
| 20, 21 | AT5G62700 | TUB3 | Tubulin beta chain 3 | 51.27/ 4.73 | 38 | 778 | 2.40 × 10−71 | 3.032 |
| 22 | AT2G29550 | TUB7 | Tubulin beta-7 chain | 51.34/ 4.74 | 36 | 674 | 5.90 × 10−61 | 3.857 |
| 23 | AT5G38470 | RAD23D | Rad23 UV excision repair protein family | 40.10/ 4.58 | 17 | 382 | 9.40 * 10−32 | 2.453 |
| 24 | AT4G20890 | TUB9 | Tubulin beta-9 chain | 50.31/ 4.69 | 38 | 715 | 4.70 × 10−65 | 1.649 |
| 25 | AT5G44340 | TUB4 | Tubulin beta chain 4 | 50.36/ 4.76 | 31 | 490 | 1.50 ×× 10−42 | 2.158 |
| 26 | AT4G37910 | HSP70-9 | Heat shock 70 kDa protein 9, mitochondrial | 73.32/ 5.51 | 29 | 625 | 4.70 × 10−56 | 2.615 |
| 27 | AT5G09810 | ACT7 | Actin 7 | 41.94/ 5.31 | 32 | 1100 | 1.50 × 10−103 | 1.110 |
| 29 | AT1G35720 | ANNAT1 | Annexin D1, calcium binding proteins | 36.30/ 5.21 | 27 | 754 | 5.90 × 10−69 | 2.469 |
| 30 | AT1G79230 | STR1 | Thiosulfate/3-mercaptopyruvate sulfurtransferase 1, mitochondrial | 42.15/ 5.95 | 21 | 529 | 1.90 × 10−46 | 1.487 |
| 31 | AT3G53970 | Probable proteasome inhibitor | 32.15/ 4.94 | 15 | 329 | 1.90 × 10−26 | 3.353 | |
| 32 | AT1G62380 | ACO2 | 1-aminocyclopropane-1-carboxylate oxidase 2 | 36.39/ 4.98 | 21 | 722 | 9.40 × 10−66 | −2.900 |
| 33, 34 | AT4G20260 | PCAP1 | Plasma membrane associated cation-binding protein 1 | 18.98/ 9.88 | 9 | 168 | 2.40 × 10−10 | −1.309 |
| 37 | AT5G38480 | GRF3 | 14-3-3-like protein GF14 psi | 32.00/ 4.91 | 8 | 149 | 1.90 × 10−08 | 1.052 |
| 38 | AT4G04020 | PAP1 | Probable plastid-lipid-associated protein 1, chloroplastic | 34.99/ 5.45 | 23 | 786 | 3.70 × 10−72 | 5.711 |
| 40 | AT4G10480 | NACα4 | Nascent polypeptide-associated complex (NAC), alpha subunit family protein | 23.10/ 4.25 | 8 | 202 | 9.40 × 10−14 | 2.031 |
| 41, 42 | AT4G02450 | P23-1 | HSP20-like chaperones superfamily protein | 25.38/ 4.46 | 10 | 248 | 2.40 × 10−18 | 3.315 |
| 43 | AT4G34050 | CCoAOMT1 | S-adenosyl-L-methionine-dependent methyltransferases superfamily protein | 29.25/ 5.13 | 20 | 659 | 1.90 × 10−59 | 7.222 |
| 45 | AT1G47420 | SDH5 | Succinate dehydrogenase subunit 5, mitochondrial | 28.15/ 6.19 | 15 | 410 | 1.50 × 10−34 | 5.433 |
| 46, 48 | AT3G55440 | CTIMC | Triosephosphate isomerase, cytosolic | 27.38/ 5.39 | 26 | 904 | 5.90 × 10−84 | 1.155 |
| 47 | AT5G20720 | CPN20 | 20 kDa chaperonin, chloroplastic | 26.79/ 8.86 | 8 | 87 | 0.033 | 1.638 |
| 49 | AT5G26667 | UMK3 | P-loop containing nucleoside triphosphate hydrolases superfamily protein | 22.58/ 5.79 | 13 | 298 | 2.40 × 10−23 | 2.088 |
| 51 | AT1G02930 | GSTF6 | Glutathione S-transferase F6 | 23.47/ 5.80 | 18 | 467 | 3.00 * 10−40 | 1.795 |
| 52 | AT3G22630 | PBD1 | Proteasome subunit beta type-2-A | 22.64/ 5.95 | 15 | 138 | 2.40 * 10−07 | 1.952 |
| 53 | AT4G38680 | CSP2 | Glycine-rich protein 2, Arabidopsis thaliana cold shock protein 2 | 19.49/ 5.62 | 5 | 97 | 0.0031 | 4.117 |
| 54 | AT3G62030 | CYP20-3 /ROC4 | Peptidyl-prolyl cis-trans isomerase CYP20-3, chloroplastic | 26.73/ 8.63 | 16 | 418 | 2.40 × 10−35 | 1.402 |
| 55 | AT1G26630 | ELF5A-2 | Eukaryotic translation initiation factor 5A-1 (eIF-5A 1) protein | 17.36/ 5.55 | 7 | 232 | 9.40 × 10−17 | 2.141 |
| 56 | AT5G59880 | ADF3 | Actin-depolymerizing factor 3 | 16.03/ 5.93 | 10 | 545 | 4.70 × 10−48 | 1.290 |
| 57 | AT3G53990 | F5K20_290 | Adenine nucleotide alpha hydrolases-like superfamily protein | 17.90/ 5.66 | 21 | 575 | 4.70 × 10−51 | 1.502 |
| 58 | AT3G23490 | CYN | Cyanate hydratase | 18.64/ 5.49 | 15 | 574 | 5.90 × 10−51 | 1.316 |
| 59 | AT4G13850 | RBG2 | Glycine-rich RNA-binding protein 2, mitochondrial | 14.74/ 6.73 | 7 | 268 | 2.40 × 10−20 | 13.161 |
| 60 | AT5G18060 | SAUR23 | SAUR-like auxin-responsive protein family | 72.78/ 5.87 | 21 | 69 | 1.9 | 2.814 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chun, H.J.; Baek, D.; Jin, B.J.; Cho, H.M.; Park, M.S.; Lee, S.H.; Lim, L.H.; Cha, Y.J.; Bae, D.-W.; Kim, S.T.; et al. Microtubule Dynamics Plays a Vital Role in Plant Adaptation and Tolerance to Salt Stress. Int. J. Mol. Sci. 2021, 22, 5957. https://doi.org/10.3390/ijms22115957
Chun HJ, Baek D, Jin BJ, Cho HM, Park MS, Lee SH, Lim LH, Cha YJ, Bae D-W, Kim ST, et al. Microtubule Dynamics Plays a Vital Role in Plant Adaptation and Tolerance to Salt Stress. International Journal of Molecular Sciences. 2021; 22(11):5957. https://doi.org/10.3390/ijms22115957
Chicago/Turabian StyleChun, Hyun Jin, Dongwon Baek, Byung Jun Jin, Hyun Min Cho, Mi Suk Park, Su Hyeon Lee, Lack Hyeon Lim, Ye Jin Cha, Dong-Won Bae, Sun Tae Kim, and et al. 2021. "Microtubule Dynamics Plays a Vital Role in Plant Adaptation and Tolerance to Salt Stress" International Journal of Molecular Sciences 22, no. 11: 5957. https://doi.org/10.3390/ijms22115957







