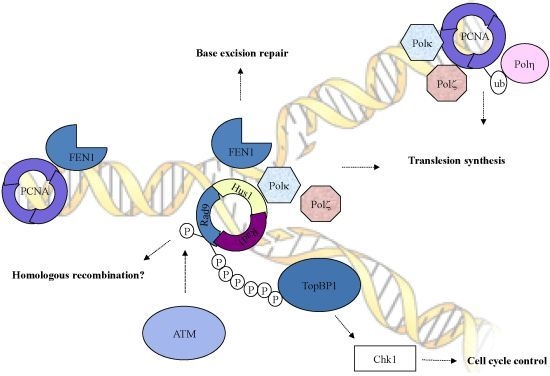
Role of the Checkpoint Clamp in DNA Damage Response
Abstract
:1. Introduction
2. Role of the Checkpoint Clamp in Checkpoint Activation
3. Role of the Checkpoint Clamp in DSB Repair
4. Role of the Checkpoint Clamp in Other Repairs
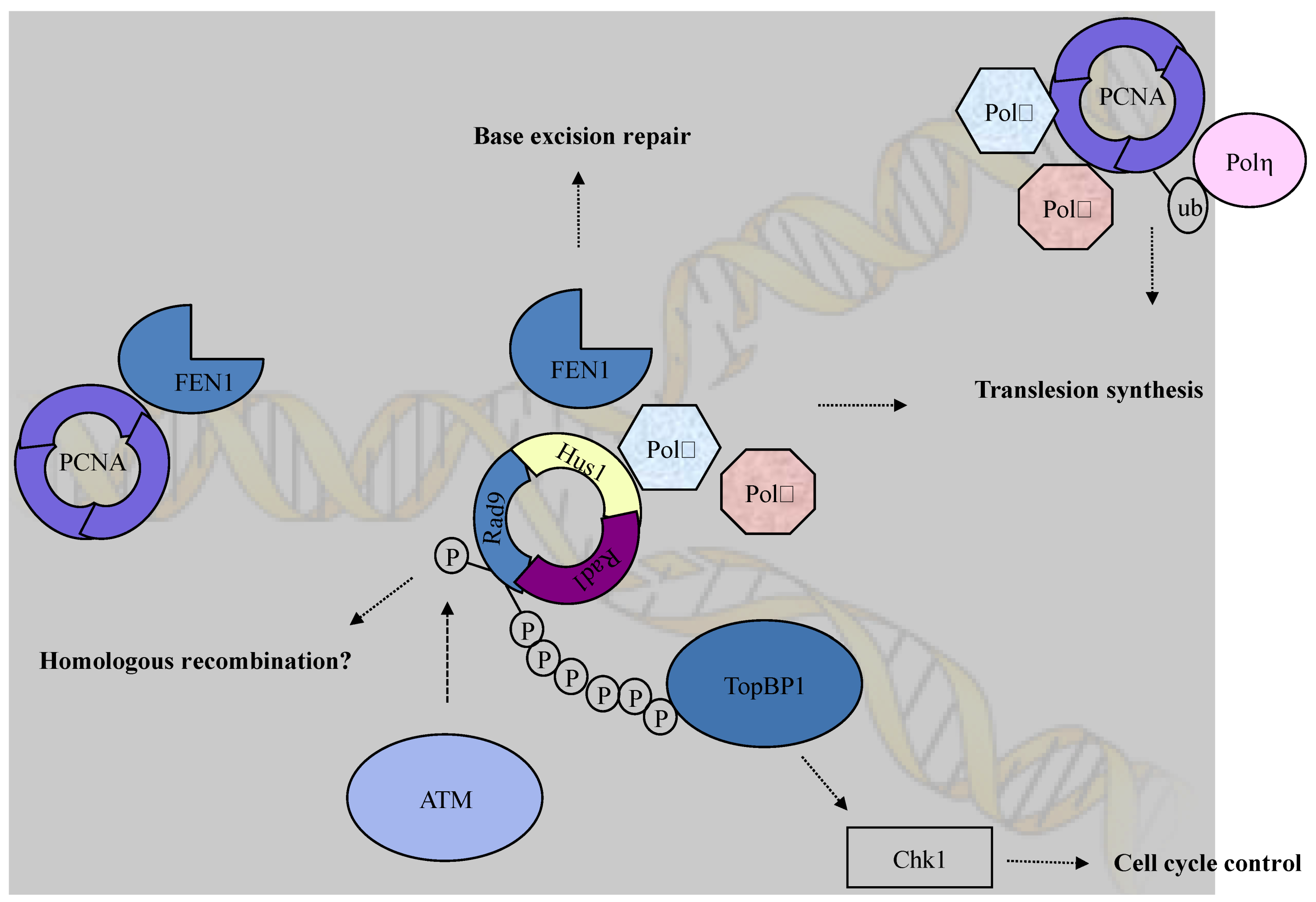
5. Conclusions
References
- Venclovas, C.; Thelen, M.P. Structure-based predictions of rad1, rad9, hus1 and rad17 participation in sliding clamp and clamp-loading complexes. Nucleic Acids Res. 2000, 28, 2481–2493. [Google Scholar] [CrossRef]
- Bermudez, V.P.; Lindsey-Boltz, L.A.; Cesare, A.J.; Maniwa, Y.; Griffith, J.D.; Hurwitz, J.; Sancar, A. Loading of the human 9-1-1 checkpoint complex onto DNA by the checkpoint clamp loader hrad17-replication factor c complex in vitro. Proc. Natl. Acad. Sci. USA 2003, 100, 1633–1638. [Google Scholar]
- Shiomi, Y.; Shinozaki, A.; Nakada, D.; Sugimoto, K.; Usukura, J.; Obuse, C.; Tsurimoto, T. Clamp and clamp loader structures of the human checkpoint protein complexes, rad9-1-1 and rad17-rfc. Genes Cells 2002, 7, 861–868. [Google Scholar] [CrossRef]
- Burtelow, M.A.; Roos-Mattjus, P.M.; Rauen, M.; Babendure, J.R.; Karnitz, L.M. Reconstitution and molecular analysis of the hrad9-hhus1-hrad1 (9-1-1) DNA damage responsive checkpoint complex. J. Bio. Chem. 2001, 276, 25903–25909. [Google Scholar]
- Lindsey-Boltz, L.A.; Bermudez, V.P.; Hurwitz, J.; Sancar, A. Purification and characterization of human DNA damage checkpoint rad complexes. Proc. Natl. Acad. Sci. USA 2001, 98, 11236–11241. [Google Scholar]
- Dore, A.S.; Kilkenny, M.L.; Rzechorzek, N.J.; Pearl, L.H. Crystal structure of the rad9-rad1-hus1 DNA damage checkpoint complex--implications for clamp loading and regulation. Mol. Cell. 2009, 34, 735–745. [Google Scholar] [CrossRef]
- Sohn, S.Y.; Cho, Y. Crystal structure of the human rad9-hus1-rad1 clamp. J. Mol. Biol. 2009, 390, 490–502. [Google Scholar] [CrossRef]
- Xu, M.; Bai, L.; Gong, Y.; Xie, W.; Hang, H.; Jiang, T. Structure and functional implications of the human rad9-hus1-rad1 cell cycle checkpoint complex. J. Bio. Chem. 2009, 284, 20457–20461. [Google Scholar] [CrossRef]
- Lee, S.H.; Hurwitz, J. Mechanism of elongation of primed DNA by DNA polymerase delta, proliferating cell nuclear antigen, and activator 1. Proc. Natl. Acad. Sci. USA 1990, 87, 5672–5676. [Google Scholar] [CrossRef]
- Ng, L.; McConnell, M.; Tan, C.K.; Downey, K.M.; Fisher, P.A. Interaction of DNA polymerase delta, proliferating cell nuclear antigen, and synthetic oligonucleotide template-primers. Analysis by polyacrylamide gel electrophoresis-band mobility shift assay. J. Bio. Chem. 1993, 268, 13571–13576. [Google Scholar]
- Jonsson, Z.O.; Hindges, R.; Hubscher, U. Regulation of DNA replication and repair proteins through interaction with the front side of proliferating cell nuclear antigen. EMBO J. 1998, 17, 2412–2425. [Google Scholar] [CrossRef]
- Mozzherin, D.J.; Tan, C.K.; Downey, K.M.; Fisher, P.A. Architecture of the active DNA polymerase delta.Proliferating cell nuclear antigen.Template-primer complex. J. Bio. Chem. 1999, 274, 19862–19867. [Google Scholar]
- Matsumoto, Y. Molecular mechanism of pcna-dependent base excision repair. Prog. Nucleic Acid Res. Mol. Biol. 2001, 68, 129–138. [Google Scholar] [CrossRef]
- Maga, G.; Villani, G.; Ramadan, K.; Shevelev, I.; Tanguy Le Gac, N.; Blanco, L.; Blanca, G.; Spadari, S.; Hubscher, U. Human DNA polymerase lambda functionally and physically interacts with proliferating cell nuclear antigen in normal and translesion DNA synthesis. J. Bio. Chem. 2002, 277, 48434–48440. [Google Scholar]
- Umar, A.; Buermeyer, A.B.; Simon, J.A.; Thomas, D.C.; Clark, A.B.; Liskay, R.M.; Kunkel, T.A. Requirement for pcna in DNA mismatch repair at a step preceding DNA resynthesis. Cell 1996, 87, 65–73. [Google Scholar] [CrossRef]
- Gary, R.; Ludwig, D.L.; Cornelius, H.L.; MacInnes, M.A.; Park, M.S. The DNA repair endonuclease xpg binds to proliferating cell nuclear antigen (pcna) and shares sequence elements with the pcna-binding regions of fen-1 and cyclin-dependent kinase inhibitor p21. J. Bio. Chem. 1997, 272, 24522–24529. [Google Scholar]
- Hosfield, D.J.; Mol, C.D.; Shen, B.; Tainer, J.A. Structure of the DNA repair and replication endonuclease and exonuclease fen-1: Coupling DNA and pcna binding to fen-1 activity. Cell 1998, 95, 135–146. [Google Scholar] [CrossRef]
- Hoege, C.; Pfander, B.; Moldovan, G.L.; Pyrowolakis, G.; Jentsch, S. Rad6-dependent DNA repair is linked to modification of pcna by ubiquitin and sumo. Nature 2002, 419, 135–141. [Google Scholar]
- Essers, J.; Theil, A.F.; Baldeyron, C.; van Cappellen, W.A.; Houtsmuller, A.B.; Kanaar, R.; Vermeulen, W. Nuclear dynamics of pcna in DNA replication and repair. Mol. Cell. Biol. 2005, 25, 9350–9359. [Google Scholar] [CrossRef]
- Lee, J.; Kumagai, A.; Dunphy, W.G. The rad9-hus1-rad1 checkpoint clamp regulates interaction of topbp1 with atr. J. Bio. Chem. 2007, 282, 28036–28044. [Google Scholar] [CrossRef]
- Yan, S.; Michael, W.M. Topbp1 and DNA polymerase-alpha directly recruit the 9-1-1 complex to stalled DNA replication forks. J. Cell. Biol. 2009, 184, 793–804. [Google Scholar] [CrossRef]
- Delacroix, S.; Wagner, J.M.; Kobayashi, M.; Yamamoto, K.; Karnitz, L.M. The rad9-hus1-rad1 (9-1-1) clamp activates checkpoint signaling via topbp1. Genes Dev. 2007, 21, 1472–1477. [Google Scholar] [CrossRef]
- Abraham, R.T. Cell cycle checkpoint signaling through the atm and atr kinases. Genes Dev. 2001, 15, 2177–2196. [Google Scholar] [CrossRef]
- Perry, J.A.; Kornbluth, S. Cdc25 and wee1: Analogous opposites? Cell Div. 2007, 2, 12. [Google Scholar] [CrossRef]
- Bao, S.; Lu, T.; Wang, X.; Zheng, H.; Wang, L.E.; Wei, Q.; Hittelman, W.N.; Li, L. Disruption of the rad9/rad1/hus1 (9-1-1) complex leads to checkpoint signaling and replication defects. Oncogene 2004, 23, 5586–5593. [Google Scholar] [CrossRef]
- Wang, X.; Zou, L.; Lu, T.; Bao, S.; Hurov, K.E.; Hittelman, W.N.; Elledge, S.J.; Li, L. Rad17 phosphorylation is required for claspin recruitment and chk1 activation in response to replication stress. Mol. Cell. 2006, 23, 331–341. [Google Scholar]
- Roos-Mattjus, P.; Hopkins, K.M.; Oestreich, A.J.; Vroman, B.T.; Johnson, K.L.; Naylor, S.; Lieberman, H.B.; Karnitz, L.M. Phosphorylation of human rad9 is required for genotoxin-activated checkpoint signaling. J. Bio. Chem. 2003, 278, 24428–24437. [Google Scholar]
- Puddu, F.; Granata, M.; Di Nola, L.; Balestrini, A.; Piergiovanni, G.; Lazzaro, F.; Giannattasio, M.; Plevani, P.; Muzi-Falconi, M. Phosphorylation of the budding yeast 9-1-1 complex is required for dpb11 function in the full activation of the uv-induced DNA damage checkpoint. Mol. Cell. Biol. 2008, 28, 4782–4793. [Google Scholar] [CrossRef]
- Puddu, F.; Piergiovanni, G.; Plevani, P.; Muzi-Falconi, M. Sensing of replication stress and mec1 activation act through two independent pathways involving the 9-1-1 complex and DNA polymerase epsilon. PLoS Genetics 2011, 7, e1002022. [Google Scholar] [CrossRef]
- Navadgi-Patil, V.M.; Burgers, P.M. The unstructured C-terminal tail of the 9-1-1 clamp subunit ddc1 activates mec1/atr via two distinct mechanisms. Mol. Cell. 2009, 36, 743–753. [Google Scholar] [CrossRef]
- Parker, A.E.; Van de Weyer, I.; Laus, M.C.; Verhasselt, P.; Luyten, W.H. Identification of a human homologue of the schizosaccharomyces pombe rad17+ checkpoint gene. J. Bio. Chem. 1998, 273, 18340–18346. [Google Scholar]
- Ellison, V.; Stillman, B. Biochemical characterization of DNA damage checkpoint complexes: Clamp loader and clamp complexes with specificity for 5' recessed DNA. PLoS Biol. 2003, 1, E33. [Google Scholar]
- Shin, M.H.; Yuan, M.; Zhang, H.; Margolick, J.B.; Kai, M. Atm-dependent phosphorylation of the checkpoint clamp regulates repair pathways and maintains genomic stability. Cell Cycle 2012, 11, 1796–1803. [Google Scholar] [CrossRef]
- Kai, M.; Tsui, E. Role of the checkpoint clamp in double-strand break repair. USA Unpublished Data 2013, Department of Radiation Oncology, Johns Hopkins University: School of Medicine, Baltimore MD 21231, USA .
- Lisby, M.; Barlow, J.H.; Burgess, R.C.; Rothstein, R. Choreography of the DNA damage response: Spatiotemporal relationships among checkpoint and repair proteins. Cell 2004, 118, 699–713. [Google Scholar] [CrossRef]
- Uziel, T.; Lerenthal, Y.; Moyal, L.; Andegeko, Y.; Mittelman, L.; Shiloh, Y. Requirement of the mrn complex for atm activation by DNA damage. EMBO J. 2003, 22, 5612–5621. [Google Scholar] [CrossRef]
- Carson, C.T.; Schwartz, R.A.; Stracker, T.H.; Lilley, C.E.; Lee, D.V.; Weitzman, M.D. The mre11 complex is required for atm activation and the g2/m checkpoint. EMBO J. 2003, 22, 6610–6620. [Google Scholar] [CrossRef]
- Lee, J.H.; Paull, T.T. Direct activation of the atm protein kinase by the mre11/rad50/nbs1 complex. Science 2004, 304, 93–96. [Google Scholar]
- Lee, J.H.; Paull, T.T. Atm activation by DNA double-strand breaks through the mre11-rad50-nbs1 complex. Science 2005, 308, 551–554. [Google Scholar] [CrossRef]
- Zou, L.; Liu, D.; Elledge, S.J. Replication protein a-mediated recruitment and activation of rad17 complexes. Proc. Natl. Acad. Sci. USA 2003, 100, 13827–13832. [Google Scholar] [CrossRef]
- Zou, L.; Elledge, S.J. Sensing DNA damage through atrip recognition of rpa-ssdna complexes. Science 2003, 300, 1542–1548. [Google Scholar] [CrossRef]
- You, Z.; Shi, L.Z.; Zhu, Q.; Wu, P.; Zhang, Y.W.; Basilio, A.; Tonnu, N.; Verma, I.M.; Berns, M.W.; Hunter, T. Ctip links DNA double-strand break sensing to resection. Mol. Cell. 2009, 36, 954–969. [Google Scholar]
- Yu, X.; Wu, L.C.; Bowcock, A.M.; Aronheim, A.; Baer, R. The C-terminal (brct) domains of brca1 interact in vivo with ctip, a protein implicated in the ctbp pathway of transcriptional repression. J. Bio. Chem. 1998, 273, 25388–25392. [Google Scholar]
- Chen, L.; Nievera, C.J.; Lee, A.Y.; Wu, X. Cell cycle-dependent complex formation of brca1.Ctip.Mrn is important for DNA double-strand break repair. J. Bio. Chem. 2008, 283, 7713–7720. [Google Scholar]
- Yun, M.H.; Hiom, K. Ctip-brca1 modulates the choice of DNA double-strand-break repair pathway throughout the cell cycle. Nature 2009, 459, 460–463. [Google Scholar] [CrossRef]
- Nakamura, K.; Kogame, T.; Oshiumi, H.; Shinohara, A.; Sumitomo, Y.; Agama, K.; Pommier, Y.; Tsutsui, K.M.; Tsutsui, K.; Hartsuiker, E.; Ogi, T.; Takeda, S.; Taniguchi, Y. Collaborative action of brca1 and ctip in elimination of covalent modifications from double-strand breaks to facilitate subsequent break repair. PLoS Genetics 2010, 6, e1000828. [Google Scholar] [CrossRef]
- Mimitou, E.P.; Symington, L.S. DNA end resection: Many nucleases make light work. DNA Repair 2009, 8, 983–995. [Google Scholar] [CrossRef]
- Budd, M.E.; Campbell, J.L. Interplay of mre11 nuclease with dna2 plus sgs1 in rad51-dependent recombinational repair. PLoS One 2009, 4, e4267. [Google Scholar] [CrossRef]
- Mimitou, E.P.; Symington, L.S. Sae2, exo1 and sgs1 collaborate in DNA double-strand break processing. Nature 2008, 455, 770–774. [Google Scholar] [CrossRef]
- Zhu, Z.; Chung, W.H.; Shim, E.Y.; Lee, S.E.; Ira, G. Sgs1 helicase and two nucleases dna2 and exo1 resect DNA double-strand break ends. Cell 2008, 134, 981–994. [Google Scholar] [CrossRef]
- Nimonkar, A.V.; Ozsoy, A.Z.; Genschel, J.; Modrich, P.; Kowalczykowski, S.C. Human exonuclease 1 and blm helicase interact to resect DNA and initiate DNA repair. Proc. Natl. Acad. Sci. USA 2008, 105, 16906–16911. [Google Scholar] [CrossRef]
- Gravel, S.; Chapman, J.R.; Magill, C.; Jackson, S.P. DNA helicases sgs1 and blm promote DNA double-strand break resection. Genes Dev. 2008, 22, 2767–2772. [Google Scholar] [CrossRef]
- Yan, H.; McCane, J.; Toczylowski, T.; Chen, C. Analysis of the xenopus werner syndrome protein in DNA double-strand break repair. J. Cell. Biol. 2005, 171, 217–227. [Google Scholar] [CrossRef]
- Toczylowski, T.; Yan, H. Mechanistic analysis of a DNA end processing pathway mediated by the xenopus werner syndrome protein. J. Bio. Chem. 2006, 281, 33198–33205. [Google Scholar] [CrossRef]
- Nimonkar, A.V.; Genschel, J.; Kinoshita, E.; Polaczek, P.; Campbell, J.L.; Wyman, C.; Modrich, P.; Kowalczykowski, S.C. Blm-dna2-rpa-mrn and exo1-blm-rpa-mrn constitute two DNA end resection machineries for human DNA break repair. Genes Dev. 2011, 25, 350–362. [Google Scholar] [CrossRef]
- Warmerdam, D.O.; Freire, R.; Kanaar, R.; Smits, V.A. Cell cycle-dependent processing of DNA lesions controls localization of rad9 to sites of genotoxic stress. Cell Cycle 2009, 8, 1765–1774. [Google Scholar] [CrossRef]
- Shin, M.H.; Yuan, M.; Zhang, H.; Margolick, J.B.; Kai, M. Atm-dependent phosphorylation of the checkpoint clamp regulates repair pathways and maintains genomic stability. Cell Cycle 2012, 11, 1796–1803. [Google Scholar] [CrossRef]
- Zhang, J.; Willers, H.; Feng, Z.; Ghosh, J.C.; Kim, S.; Weaver, D.T.; Chung, J.H.; Powell, S.N.; Xia, F. Chk2 phosphorylation of brca1 regulates DNA double-strand break repair. Mol. Cell. Biol. 2004, 24, 708–718. [Google Scholar]
- Kim, S.S.; Cao, L.; Li, C.; Xu, X.; Huber, L.J.; Chodosh, L.A.; Deng, C.X. Uterus hyperplasia and increased carcinogen-induced tumorigenesis in mice carrying a targeted mutation of the chk2 phosphorylation site in brca1. Mol. Cell. Biol. 2004, 24, 9498–9507. [Google Scholar] [CrossRef]
- Kai, M.; Wang, T.S. Checkpoint activation regulates mutagenic translesion synthesis. Genes Dev. 2003, 17, 64–76. [Google Scholar] [CrossRef]
- Kai, M.; Wang, T.S. Checkpoint responses to replication stalling: Inducing tolerance and preventing mutagenesis. Mutation Res. 2003, 532, 59–73. [Google Scholar] [CrossRef]
- Sabbioneda, S.; Minesinger, B.K.; Giannattasio, M.; Plevani, P.; Muzi-Falconi, M.; Jinks-Robertson, S. The 9-1-1 checkpoint clamp physically interacts with polzeta and is partially required for spontaneous polzeta-dependent mutagenesis in saccharomyces cerevisiae. J. Bio. Chem. 2005, 280, 38657–38665. [Google Scholar]
- Smirnova, E.; Toueille, M.; Markkanen, E.; Hubscher, U. The human checkpoint sensor and alternative DNA clamp rad9-rad1-hus1 modulates the activity of DNA ligase i, a component of the long-patch base excision repair machinery. Biochem. J. 2005, 389, 13–17. [Google Scholar] [CrossRef]
- Toueille, M.; El-Andaloussi, N.; Frouin, I.; Freire, R.; Funk, D.; Shevelev, I.; Friedrich-Heineken, E.; Villani, G.; Hottiger, M.O.; Hubscher, U. The human rad9/rad1/hus1 damage sensor clamp interacts with DNA polymerase beta and increases its DNA substrate utilisation efficiency: Implications for DNA repair. Nucleic Acids Res. 2004, 32, 3316–3324. [Google Scholar] [CrossRef]
- Friedrich-Heineken, E.; Toueille, M.; Tannler, B.; Burki, C.; Ferrari, E.; Hottiger, M.O.; Hubscher, U. The two DNA clamps rad9/rad1/hus1 complex and proliferating cell nuclear antigen differentially regulate flap endonuclease 1 activity. J. Mol. Biol. 2005, 353, 980–989. [Google Scholar] [CrossRef]
- Wang, W.; Brandt, P.; Rossi, M.L.; Lindsey-Boltz, L.; Podust, V.; Fanning, E.; Sancar, A.; Bambara, R.A. The human rad9-rad1-hus1 checkpoint complex stimulates flap endonuclease 1. Proc. Natl. Acad. Sci. USA 2004, 101, 16762–16767. [Google Scholar]
- Gary, R.; Kim, K.; Cornelius, H.L.; Park, M.S.; Matsumoto, Y. Proliferating cell nuclear antigen facilitates excision in long-patch base excision repair. J. Bio. Chem. 1999, 274, 4354–4363. [Google Scholar]
- Kedar, P.S.; Kim, S.J.; Robertson, A.; Hou, E.; Prasad, R.; Horton, J.K.; Wilson, S.H. Direct interaction between mammalian DNA polymerase beta and proliferating cell nuclear antigen. J. Bio. Chem. 2002, 277, 31115–31123. [Google Scholar]
- Bunting, K.A.; Roe, S.M.; Pearl, L.H. Structural basis for recruitment of translesion DNA polymerase pol iv/dinb to the beta-clamp. EMBO J. 2003, 22, 5883–5892. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Kai, M. Role of the Checkpoint Clamp in DNA Damage Response. Biomolecules 2013, 3, 75-84. https://doi.org/10.3390/biom3010075
Kai M. Role of the Checkpoint Clamp in DNA Damage Response. Biomolecules. 2013; 3(1):75-84. https://doi.org/10.3390/biom3010075
Chicago/Turabian StyleKai, Mihoko. 2013. "Role of the Checkpoint Clamp in DNA Damage Response" Biomolecules 3, no. 1: 75-84. https://doi.org/10.3390/biom3010075