Author Contributions
Conceptualization, A.H.-D. and D.K.; Funding acquisition, D.K.; Investigation, A.Z. and A.H.-D.; Methodology, A.H.-D. and D.K.; Project administration, D.K.; Resources, A.Z., A.H.-D., J.W. and J.N.; Software, A.K.; Supervision, A.K. and D.K.; Validation, D.K.; Visualization, A.Z.; Writing—original draft, A.Z., A.H.-D., J.W., J.N. and D.K.; Writing—review and editing, D.K.
Figure 1.
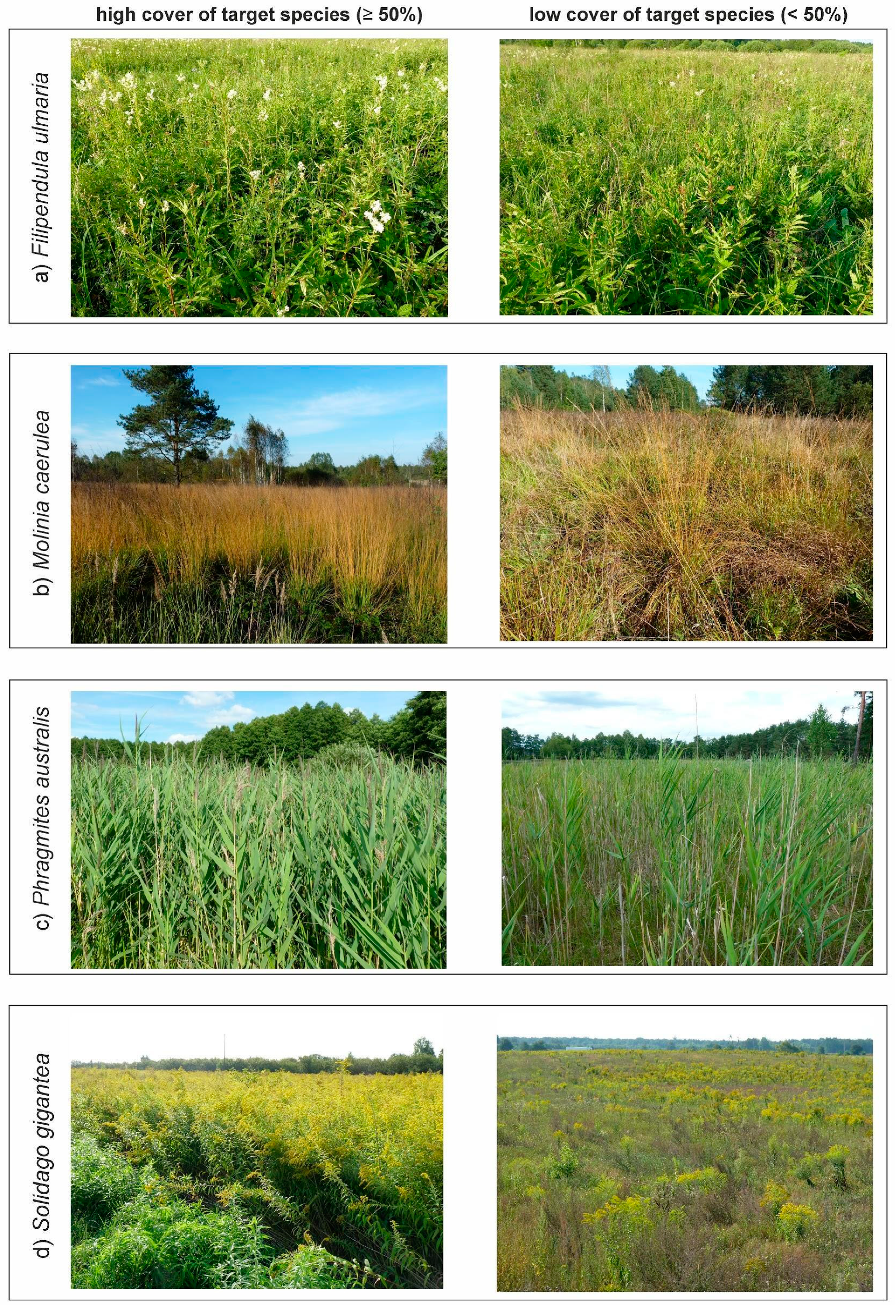
Target species during on-ground sampling, (a) Filipendula ulmaria, (b) Molinia caerulea, (c) Phragmites australis, (d) Solidago gigantea. (Photo. J. Wylazłowska).
Figure 1.
Target species during on-ground sampling, (a) Filipendula ulmaria, (b) Molinia caerulea, (c) Phragmites australis, (d) Solidago gigantea. (Photo. J. Wylazłowska).
Figure 2.
Location of the study areas. Left side: base map: HySpex image in natural color Red, Green, Blue (RGB) composition with on-ground polygon locations. Right side: surface elevation model. Study areas: No.1 “Dolina Krasnej", No.2 “Ostoja Nadwarciańska”, No.3 “Dolina Dolnego Sanu”.
Figure 2.
Location of the study areas. Left side: base map: HySpex image in natural color Red, Green, Blue (RGB) composition with on-ground polygon locations. Right side: surface elevation model. Study areas: No.1 “Dolina Krasnej", No.2 “Ostoja Nadwarciańska”, No.3 “Dolina Dolnego Sanu”.
Figure 3.
Stage 1 scheme. The number of polygons for the target species and their percentage cover in the polygons and the number of background polygons considered in the training set in each scenario. The group of green rectangles indicates constant and invariable polygons during Stage 1. The group of yellow rectangles indicates scenarios that change during Stage 1, differing in the species percentage cover of in the polygons (FU—Filipendula ulmaria, MC—Molinia caerulea, PA—Phragmites australis, SG—Solidago gigantea).
Figure 3.
Stage 1 scheme. The number of polygons for the target species and their percentage cover in the polygons and the number of background polygons considered in the training set in each scenario. The group of green rectangles indicates constant and invariable polygons during Stage 1. The group of yellow rectangles indicates scenarios that change during Stage 1, differing in the species percentage cover of in the polygons (FU—Filipendula ulmaria, MC—Molinia caerulea, PA—Phragmites australis, SG—Solidago gigantea).
Figure 4.
Stage 2 scheme. Changes in the number of species polygons and the number of background polygons taken into account during the training in particular scenarios. The group of polygons with the green background is constant and invariable during Stage 2. The yellow rectangle indicates the best scenario from Stage 1. The group of blue rectangles indicates scenarios, which change during Stage 2, differing in the number of species training polygons (FU—Filipendula ulmaria, MC—Molinia caerulea, PA—Phragmites australis, SG—Solidago gigantea).
Figure 4.
Stage 2 scheme. Changes in the number of species polygons and the number of background polygons taken into account during the training in particular scenarios. The group of polygons with the green background is constant and invariable during Stage 2. The yellow rectangle indicates the best scenario from Stage 1. The group of blue rectangles indicates scenarios, which change during Stage 2, differing in the number of species training polygons (FU—Filipendula ulmaria, MC—Molinia caerulea, PA—Phragmites australis, SG—Solidago gigantea).
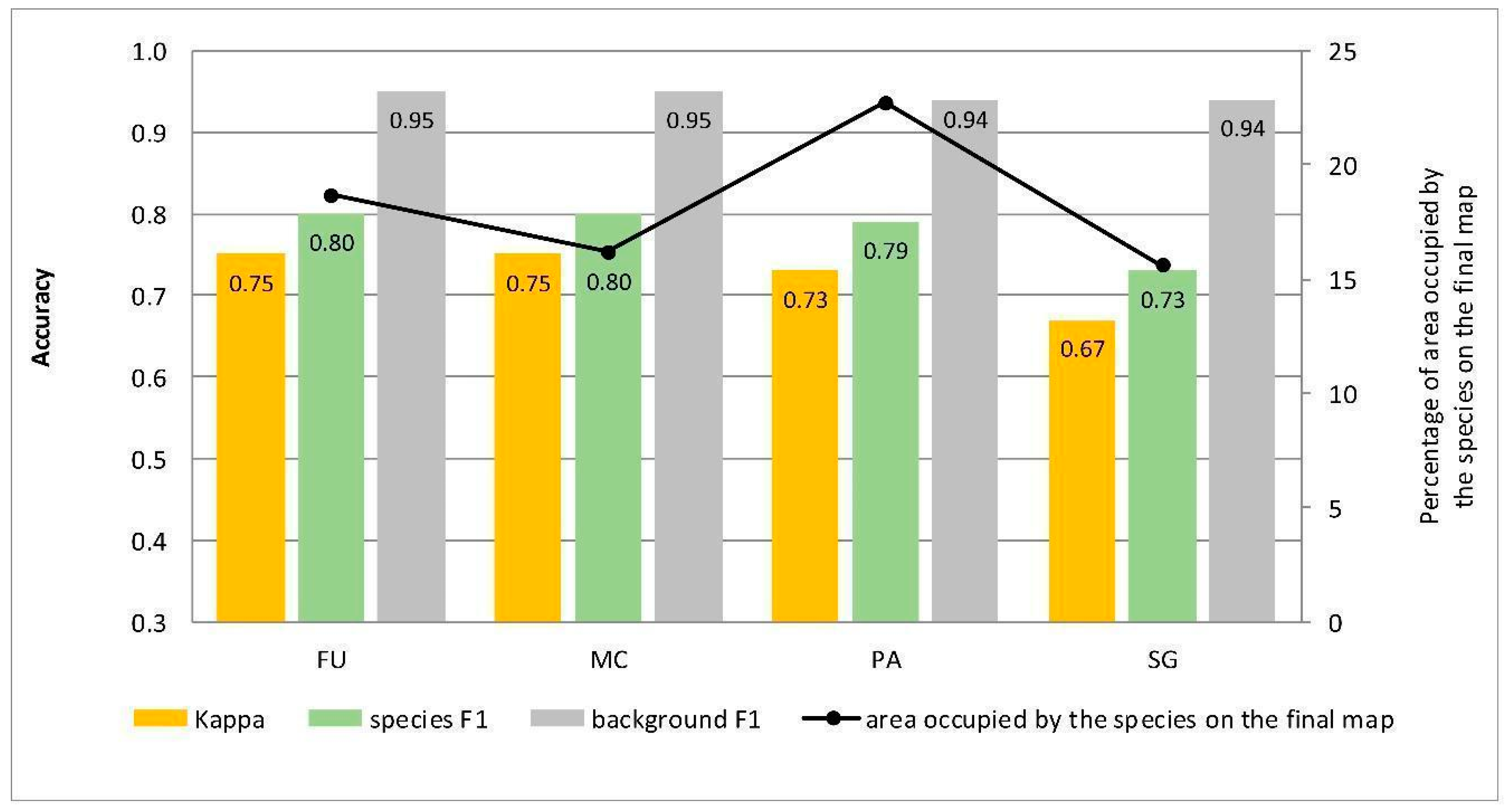
Figure 5.
Classification accuracy described by Kappa and F1 measures (columns) and percentage of area occupied by the species on the final map (the black line) for four target species (FU—Filipendula ulmaria, MC—Molinia caerulea, PA—Phragmites australis, SG—Solidago gigantea) in the classifications performed at Stage 0.
Figure 5.
Classification accuracy described by Kappa and F1 measures (columns) and percentage of area occupied by the species on the final map (the black line) for four target species (FU—Filipendula ulmaria, MC—Molinia caerulea, PA—Phragmites australis, SG—Solidago gigantea) in the classifications performed at Stage 0.
Figure 6.
Classification results of Filipendula ulmaria (FU) in all stages and scenarios. The black line areas show the target species patches identified during the field mapping. The best scenario for Stage 1 is marked by green line. The best scenario for Stage 2 is marked by blue line.
Figure 6.
Classification results of Filipendula ulmaria (FU) in all stages and scenarios. The black line areas show the target species patches identified during the field mapping. The best scenario for Stage 1 is marked by green line. The best scenario for Stage 2 is marked by blue line.
Figure 7.
Classification results of Molinia caerulea (MC) in all stages and scenarios. The black line areas show the target species patches identified during the field mapping. The best scenario for Stage 1 is marked by green line. The best scenario for Stage 2 is marked by blue line.
Figure 7.
Classification results of Molinia caerulea (MC) in all stages and scenarios. The black line areas show the target species patches identified during the field mapping. The best scenario for Stage 1 is marked by green line. The best scenario for Stage 2 is marked by blue line.
Figure 8.
Classification results of Solidago gigantea (SG) in all stages and scenarios. The black line areas show the target species patches identified during the field mapping. The best scenario for Stage 1 is marked by green line. The best scenario for Stage 2 is marked by blue line.
Figure 8.
Classification results of Solidago gigantea (SG) in all stages and scenarios. The black line areas show the target species patches identified during the field mapping. The best scenario for Stage 1 is marked by green line. The best scenario for Stage 2 is marked by blue line.
Figure 9.
Classification results of Phragmites australis (PA) in all stages and scenarios. The black line areas show the target species patches identified during the field mapping. The best scenario for Stage 1 is marked by green line. The best scenario for Stage 2 is marked by blue line.
Figure 9.
Classification results of Phragmites australis (PA) in all stages and scenarios. The black line areas show the target species patches identified during the field mapping. The best scenario for Stage 1 is marked by green line. The best scenario for Stage 2 is marked by blue line.
Figure 10.
Classification accuracy described by Kappa and F1 measures (columns), percentage of area occupied by the target species on the final map (the black line) for four target species (FU—Filipendula ulmaria, MC—Molinia caerulea, PA—Phragmites australis, SG—Solidago gigantea) in five classification scenarios performed at Stage 1.
Figure 10.
Classification accuracy described by Kappa and F1 measures (columns), percentage of area occupied by the target species on the final map (the black line) for four target species (FU—Filipendula ulmaria, MC—Molinia caerulea, PA—Phragmites australis, SG—Solidago gigantea) in five classification scenarios performed at Stage 1.
Figure 11.
Percentage of correctly classified pixels (columns) and percentage of false-positive pixels (the orange line) in the set of validation polygons for four target species (FU—Filipendula ulmaria, MC—Molinia caerulea, PA—Phragmites australis, SG—Solidago gigantea) in four scenarios of Stage 1.
Figure 11.
Percentage of correctly classified pixels (columns) and percentage of false-positive pixels (the orange line) in the set of validation polygons for four target species (FU—Filipendula ulmaria, MC—Molinia caerulea, PA—Phragmites australis, SG—Solidago gigantea) in four scenarios of Stage 1.
Figure 12.
Percentage of correctly classified pixels (columns) and percentage of false-positive pixels (the orange line) in the set of validation polygons for four target species (FU—Filipendula ulmaria, MC—Molinia caerulea, PA—Phragmites australis, SG—Solidago gigantea) in three scenarios of Stage 2.
Figure 12.
Percentage of correctly classified pixels (columns) and percentage of false-positive pixels (the orange line) in the set of validation polygons for four target species (FU—Filipendula ulmaria, MC—Molinia caerulea, PA—Phragmites australis, SG—Solidago gigantea) in three scenarios of Stage 2.
Figure 13.
Classification accuracy described by Kappa and F1 measures (columns), percentage of area occupied by the target species on the final map (the black line) for four target species (FU—Filipendula ulmaria, MC—Molinia caerulea, PA—Phragmites australis, SG—Solidago gigantea) in three classification scenarios performed at Stage 2.
Figure 13.
Classification accuracy described by Kappa and F1 measures (columns), percentage of area occupied by the target species on the final map (the black line) for four target species (FU—Filipendula ulmaria, MC—Molinia caerulea, PA—Phragmites australis, SG—Solidago gigantea) in three classification scenarios performed at Stage 2.
Table 1.
Characteristics of the field and remote sensing data for each study area. Orientation of flight: North-South (NS), West-East (WE).
Table 1.
Characteristics of the field and remote sensing data for each study area. Orientation of flight: North-South (NS), West-East (WE).
| Area | No. 1 | No. 1 | No. 2 | No. 3 |
|---|
| Target species | Molinia caerulea (MC) | Filipendula ulmaria (FU) | Phragmites australis (PA) | Solidago gigantea (SG) |
| Flight dates and start of field sampling | 27 September 2017 | 07 July 2017 | 16 July 2017 | 09 September 2017 |
| End of field sampling | 29 September 2017 | 13 July 2017 | 18 July 2017 | 19 September 2017 |
| Dominant development phase of target species | fruiting | flowering | flowering | flowering |
| Phenological and structural traits of plants during acquisition | Plants form a dense tussock, floral stems become pinkish–orange as the leaves turn yellow | Plants form a bushy clump, bearing sprays of creamy–white flowers | Plants form dense stands that include flowering bushy panicles and standing dead stems from previous year’s growth | Plants form a bushy mound of deep-green leaves, bearing large clusters of golden-yellow flowers |
| Number of established target species reference polygons | 110 | 110 | 110 | 110 |
| Number of established background polygons | 200 | 200 | 200 | 200 |
| Data collection area [km2] | 40.59 | 40.59 | 10.37 | 35.45 |
| Number of flight lines | 26 | 25 | 11 | 20 |
| Orientation of flight | NS | WE | WE | NS |
Table 2.
Flight parameters and hyperspectral data.
Table 2.
Flight parameters and hyperspectral data.
| Sensor Type | Data Parameters | Flight Lines Overlap | Swath Width |
|---|
| HySpex VNIR-1800 0.4–0.9 µm | GSD 0.49 [m] | 35 [%] | 440 m |
| HySpex SWIR-384 0.9–2.5 µm | GSD 1.07 [m] | 30 [%] | 410 m |
Table 3.
The number of polygons established for each target species.
Table 3.
The number of polygons established for each target species.
| Number of the Target Species Reference Polygons and Background Reference Polygons |
|---|
| Percentage Cover of the Target Species |
|---|
| 20–40% | 50–70% | 80–100% |
| 30 | 30 | 50 |
Table 4.
Division of the classification results from each stage and scenario to the area of the target species and their background and to the percentage of True Positive, False Positive, True Negative, and False Negative in the control area selected for each target species (FU—Filipendula ulmaria, MC—Molinia caerulea, PA—Phragmites australis, SG—Solidago gigantea).
Table 4.
Division of the classification results from each stage and scenario to the area of the target species and their background and to the percentage of True Positive, False Positive, True Negative, and False Negative in the control area selected for each target species (FU—Filipendula ulmaria, MC—Molinia caerulea, PA—Phragmites australis, SG—Solidago gigantea).
| Target Species | Stage and Scenario | Control Area of Target Species [m2] | Control Area of Back-Ground [m2] | True Positive [m2] | False Positive [m2] | True Negative [m2] | False Negative [m2] | True Positive [%] | False Positive [%] | True Negative [%] | False Negative [%] |
|---|
| MC | Stage 0 | 14189 | 85667 | 9784 | 1946 | 83739 | 4387 | 69 | 14 | 98 | 31 |
| Stage 1 SC0 | 10840 | 3449 | 82236 | 3331 | 76 | 24 | 96 | 23 |
| Stage 1 SC1 | 9812 | 1774 | 83911 | 4359 | 69 | 13 | 98 | 31 |
| Stage 1 SC2 | 7578 | 878 | 84807 | 6593 | 53 | 6 | 99 | 46 |
| Stage 1 SC3 | 6953 | 694 | 84991 | 7218 | 49 | 5 | 99 | 51 |
| Stage 1 SC4 | 5102 | 460 | 85225 | 9069 | 36 | 3 | 99 | 64 |
| Stage 2 SC1_20 | 6123 | 402 | 85283 | 8048 | 43 | 3 | 100 | 57 |
| Stage 2 SC1_30 | 9812 | 1774 | 83911 | 4359 | 69 | 13 | 98 | 31 |
| Stage 2 SC1_40 | 9343 | 1737 | 83948 | 4828 | 66 | 12 | 98 | 34 |
| FU | Stage 0 | 10797 | 89203 | 8134 | 5910 | 83149 | 2663 | 75 | 55 | 93 | 25 |
| Stage 1 SC0 | 6486 | 4406 | 84653 | 4311 | 60 | 41 | 95 | 40 |
| Stage 1 SC1 | 7102 | 5006 | 84053 | 3695 | 66 | 46 | 94 | 34 |
| Stage 1 SC2 | 6679 | 4590 | 84469 | 4118 | 62 | 43 | 95 | 38 |
| Stage 1 SC3 | 6287 | 3454 | 85605 | 4510 | 58 | 32 | 96 | 42 |
| Stage 1 SC4 | 6457 | 3126 | 85933 | 4340 | 60 | 29 | 96 | 40 |
| Stage 2 SC4_20 | 4684 | 1482 | 87577 | 6113 | 43 | 14 | 98 | 57 |
| Stage 2 SC4_30 | 6457 | 3126 | 85933 | 4340 | 60 | 29 | 96 | 40 |
| Stage 2 SC4_40 | 6453 | 3057 | 86002 | 4344 | 60 | 28 | 96 | 40 |
| SG | Stage 0 | 11477 | 88523 | 7076 | 3398 | 84981 | 4401 | 62 | 30 | 96 | 38 |
| Stage 1 SC0 | 4208 | 10431 | 77948 | 7269 | 37 | 91 | 88 | 63 |
| Stage 1 SC1 | 1488 | 4128 | 84251 | 9989 | 13 | 36 | 95 | 87 |
| Stage 1 SC2 | 1916 | 581 | 87798 | 9561 | 17 | 5 | 99 | 83 |
| Stage 1 SC3 | 4061 | 665 | 87714 | 7416 | 35 | 6 | 99 | 65 |
| Stage 1 SC4 | 2575 | 568 | 87811 | 8902 | 22 | 5 | 99 | 78 |
| Stage 2 SC3_20 | 1636 | 211 | 88168 | 9841 | 14 | 2 | 100 | 86 |
| Stage 2 SC3_30 | 4061 | 665 | 87714 | 7416 | 35 | 6 | 99 | 65 |
| Stage 2 SC3_40 | 4770 | 638 | 87741 | 6707 | 42 | 6 | 99 | 58 |
| PA | Stage 0 | 33106 | 66894 | 24417 | 3534 | 63216 | 8689 | 74 | 11 | 95 | 26 |
| Stage 1 SC0 | 18279 | 3409 | 63341 | 14827 | 55 | 10 | 95 | 45 |
| Stage 1 SC1 | 15426 | 1700 | 65050 | 17680 | 47 | 5 | 97 | 53 |
| Stage 1 SC2 | 10979 | 657 | 66093 | 22127 | 33 | 2 | 99 | 67 |
| Stage 1 SC3 | 8604 | 356 | 66394 | 24502 | 26 | 1 | 99 | 74 |
| Stage 1 SC4 | 10450 | 473 | 66277 | 22656 | 32 | 1 | 99 | 68 |
| Stage 2 SC1_20 | 3904 | 315 | 66435 | 29202 | 12 | 1 | 99 | 88 |
| Stage 2 SC1_30 | 15426 | 1700 | 65050 | 17680 | 47 | 5 | 97 | 53 |
| Stage 2 SC1_40 | 16393 | 1532 | 65218 | 16713 | 50 | 5 | 97 | 50 |
Table 5.
A comparison of the three measures with the best independently chosen scenario of for each measure for Stage 1.
Table 5.
A comparison of the three measures with the best independently chosen scenario of for each measure for Stage 1.
| Target Species | RS Accuracy Measures (Kappa, F1) | Correctly Classified Species Pixels (%) | Compatibility with Field Mapping (Control Area) | Chosen Scenario |
|---|
| Molinia caerulea (MC) | SC1 | SC1 | SC1 | SC1 |
| Filipendula ulmaria (FU) | SC4 | SC4 | SC4 | SC4 |
| Solidago gigantea (SG) | SC3 | SC3 | SC3 | SC3 |
| Phragmites australis (PA) | SC1 | SC1 | SC1 | SC1 |
Table 6.
A comparison of the three measures with the best independently chosen scenario of for each measure for Stage 2.
Table 6.
A comparison of the three measures with the best independently chosen scenario of for each measure for Stage 2.
| Target Species | RS Accuracy Measures (Kappa, F1) | Correctly Classified Species Pixels (%) | Compatibility with Field Mapping | Chosen Scenario |
|---|
| Molinia caerulea (MC) | SC1_30 | SC1_40 | SC1_30 | SC1_30 |
| Filipendula ulmaria (FU) | SC4_40 | SC4_40 | SC4_40 | SC4_40 |
| Solidago gigantea (SG) | SC3_40 | SC3_40 | SC3_40 | SC3_40 |
| Phragmites australis (PA) | SC1_30 | SC1_40 | SC1_40 | SC1_40 |