Non-Destructive Soluble Solids Content Determination for ‘Rocha’ Pear Based on VIS-SWNIR Spectroscopy under ‘Real World’ Sorting Facility Conditions
Abstract
1. Introduction
2. Materials and Methods
2.1. Target Fruit
2.2. Experimental Setup and Methodology
2.3. Variability Associated with the Standard Destructive Measurements of SSC
2.4. Models and Pre-Processing Techniques
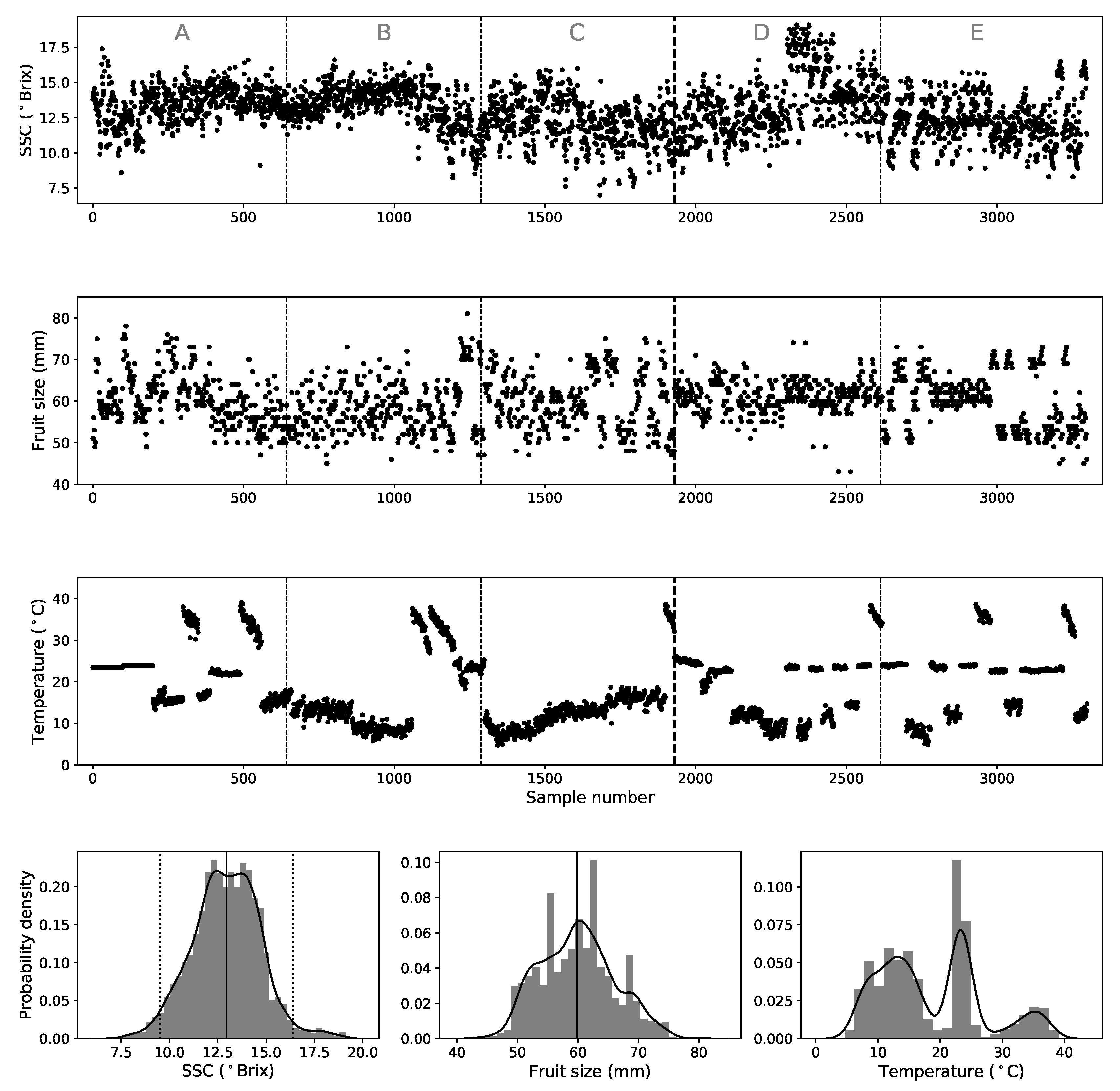
2.5. Validation Strategies and Data Sets
2.6. Performance Metrics and Multivariate Model Optimization
2.6.1. PLS
2.6.2. MLR
2.6.3. SVM
2.6.4. MLP (NN)
3. Results and Discussion
3.1. PLS and MLR Results
3.2. SVM and MLP Results
3.3. Discussion and Remarks
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| SSC | Soluble Solids Content |
| VIS-SWNIR | Visible and Short Wave Near Infrared |
| SNV | Standard Normal Variate |
| MSC | Multiplicative Scatter Correction |
| Chl | Chlorophyll |
| PLS | Partial Least Squares |
| #LV | Number of Latent Variable (PLS model) |
| SVM | Support Vector Machines |
| MLR | Multiple Linear Regressions |
| MLP | Multi-Layer Perceptron |
| NN | Neural Network |
| RGS | Randomized Grid Search |
| GPU | Graphics Processing Unit |
| IV | Internal Validation |
| EV | External Validation |
| RMSEP | Root Mean Squared Error of Prediction |
| RMSEC | Root Mean Squared Error of Calibration |
| RMSECV | Root Mean Squared Error of Calibration using Cross-Validation |
| R | Coefficient of Determination |
| PG | Prediction Gain |
| SDR | Standard Deviation Ratio |
References
- Beddington, J. Food security: Contributions from science to a new and greener revolution. Philos. Trans. R. Soc. B 2010, 365, 61–71. [Google Scholar] [CrossRef] [PubMed]
- Zambon, I.; Cecchini, M.; Egidi, G.; Saporito, M.G.; Colantoni, A. Revolution 4.0: Industry vs. Agriculture in a Future Development for SMEs. Processes 2019, 7, 36. [Google Scholar] [CrossRef]
- Nicolaï, B.M.; Beullens, K.; Bobelyn, E.; Peirs, A.; Saeys, W.; Theron, K.I.; Lammertyn, J. Nondestructive measurement of fruit and vegetable quality by means of NIR spectroscopy: A review. Postharvest Biol. Technol. 2007, 46, 99–118. [Google Scholar] [CrossRef]
- Ozaki, Y.; McClure, W.F.; Christy, A.A. Near Infrared Spectroscopy in Food Science and Technology; Wiley: Hoboken, NJ, USA, 2007. [Google Scholar]
- Cavaco, A.; Pinto, P.; Antunes, M.D.; Silva, J.M.; Guerra, R. Rocha pear firmness predicted by a Vis/NIR segmented model. Postharvest Biol. Technol. 2009, 51, 311–319. [Google Scholar] [CrossRef]
- Lu, R. Light Scattering Technology for Food Property, Quality and Safety Assessment; Chapman and Hall/CRC: London, UK, 2016. [Google Scholar]
- Bexiga, F.; Rodrigues, D.; Guerra, R.; Brázio, A.; Balegas, T.; Cavaco, A.M.; Antunes, M.D.; de Oliveira, J.V. A TSS classification study of ’Rocha’ pear (Pyrus communis L.) based on non-invasive visible/near infra-red reflectance spectra. Postharvest Biol. Technol. 2017, 132, 23–30. [Google Scholar] [CrossRef]
- Franca, A.S.; Nollet, L.M.L. Spectroscopic Methods in Food Analysis; CRC Press: London, UK, 2018. [Google Scholar]
- Magwaza, L.S.; Opara, U.L. Analytical methods for determination of sugars and sweetness of horticultural products—A review. Sci. Hortic. 2015, 184, 179–192. [Google Scholar] [CrossRef]
- Estatisticas Agricolas 2017; INE I.P. Statistics, P.L.P.: Lisbon, Portugal, 2018.
- Nicolaï, B.M.; Verlinden, B.E.; Desmet, M.; Saevels, S.; Saeys, W.; Theron, K.; Cubeddu, R.; Pifferi, A.; Torricelli, A. Time-resolved and continuous wave NIR reflectance spectroscopy to predict soluble solids content and firmness of pear. Postharvest Biol. Technol. 2008, 47, 68–74. [Google Scholar] [CrossRef]
- Liu, Y.; Chen, X.; Ouyang, A. Nondestructive determination of pear internal quality indices by visible and near-infrared spectrometry. LWT Food Sci. Technol. 2008, 41, 1720–1725. [Google Scholar] [CrossRef]
- Adebayo, S.E.; Hashim, N.; Hass, R.; Reich, O.; Regen, C.; Münzberg, M.; Abdan, K.; Hanafi, M.; Zude-Sasse, M. Using absorption and reduced scattering coefficients for non-destructive analyses of fruit flesh firmness and soluble solids content in pear (Pyrus communis ‘Conference’)—An update when using diffusion theory. Postharvest Biol. Technol. 2017, 130, 56–63. [Google Scholar] [CrossRef]
- Wang, J.; Wang, J.; Chen, Z.; Han, D. Development of multi-cultivar models for predicting the soluble solid content and firmness of European pear (Pyrus communis L.) using portable VIS-NIR spectroscopy. Postharvest Biol. Technol. 2017, 129, 143–151. [Google Scholar] [CrossRef]
- Li, J.; Wang, Q.; Xu, L.; Tian, X.; Xia, Y.; Fan, S. Comparison and Optimization of Models for Determination of Sugar Content in Pear by Portable Vis-NIR Spectroscopy Coupled with Wavelength Selection Algorithm. Food Anal. Methods 2018, 12, 12–22. [Google Scholar] [CrossRef]
- Lu, M.; Li, C.R.; Li, L.; Wu, Y.; Yang, Y. Rapid Detecting Soluble Solid Content of Pears Based on Near-Infrared Spectroscopy. In Proceedings of the 2018 2nd IEEE Advanced Information Management, Communicates, Electronic and Automation Control Conference (IMCEC), Xi’an, China, 25–27 May 2018; pp. 819–823. [Google Scholar] [CrossRef]
- Wulfert, F.; Kok, W.T.; Smilde, A.K. Influence of Temperature on Vibrational Spectra and Consequences for the Predictive Ability of Multivariate Models. Anal. Chem. 1998, 70, 1761–1767. [Google Scholar] [CrossRef] [PubMed]
- Wulfert, F.; Kok, W.T.; de Noord, O.E.; Smilde, A.K. Linear techniques to correct for temperature-induced spectral variation in multivariate calibration. Chemom. Intell. Lab. 2000, 51, 189–200. [Google Scholar] [CrossRef]
- Chen, Z.P.; Morris, J.; Martin, E. Modelling Temperature-Induced Spectral Variations in Chemical Process Monitoring. IFAC Proc. Vol. 2004, 37, 553–558. [Google Scholar] [CrossRef]
- Hageman, J.; Westerhuis, J.; Smilde, A. Temperature Robust Multivariate Calibration: An Overview of Methods for Dealing with Temperature Influences on near Infrared Spectra. J. Near Infrared Spec. 2005, 13, 53–62. [Google Scholar] [CrossRef]
- Xie, L.; Ying, Y.; Sun, T.; Xu, H. Influence of temperature on visible and near-infrared spectra and the predictive ability of multivariate models. In Sensing for Agriculture and Food Quality and Safety II; Kim, M.S., Tu, S.I., Chao, K., Eds.; International Society for Optics and Photonics, SPIE: Bellingham, WA, USA, 2010; Volume 7676, pp. 9–17. [Google Scholar] [CrossRef]
- Kemps, B.J.; Saeys, W.; Mertens, K.; Darius, P.; Baerdemaeker, J.G.D.; Ketelaere, B.D. The Importance of Choosing the Right Validation Strategy in Inverse Modelling. J. Near Infrared Spec. 2010, 18, 231–237. [Google Scholar] [CrossRef]
- Xu, Y.; Goodacre, R. On Splitting Training and Validation Set: A Comparative Study of Cross-Validation, Bootstrap and Systematic Sampling for Estimating the Generalization Performance of Supervised Learning. J. Anal. Test. 2018, 2, 249–262. [Google Scholar] [CrossRef]
- Tian, X.; Wang, Q.; Li, J.; Peng, F.; Huang, W. Non-destructive prediction of soluble solids content of pear based on fruit surface feature classification and multivariate regression analysis. Infrared Phys. Tech. 2018, 92, 336–344. [Google Scholar] [CrossRef]
- Balabin, R.M.; Lomakina, E.I. Support vector machine regression (SVR/LS-SVM)—An alternative to neural networks (ANN) for analytical chemistry? Comparison of nonlinear methods on near infrared (NIR) spectroscopy data. Analyst 2011, 136, 1703. [Google Scholar] [CrossRef]
- Li, J.; Zhao, C.; Huang, W.; Zhang, C.; Peng, Y. A combination algorithm for variable selection to determine soluble solid content and firmness of pears. Anal. Methods 2014, 6, 2170–2180. [Google Scholar] [CrossRef]
- Balabin, R.M.; Lomakina, E.I.; Safieva, R.Z. Neural network (ANN) approach to biodiesel analysis: Analysis of biodiesel density, kinematic viscosity, methanol and water contents using near infrared (NIR) spectroscopy. Fuel 2011, 90, 2007–2015. [Google Scholar] [CrossRef]
- Goldshleger, N.; Chudnovsky, A.; Ben-Dor, E. Using Reflectance Spectroscopy and Artificial Neural Network to Assess Water Infiltration Rate into the Soil Profile. Appl. Environ. Soil Sci. 2012, 2012, 439567. [Google Scholar] [CrossRef]
- Tekin, Y.; Tümsavas, Z.; Mouazen, A.M. Comparing the artificial neural network with parcial least squares for prediction of soil organic carbon and pH at different moisture content levels using visible and near-infrared spectroscopy. Revista Brasileira de Ciência do Solo 2014, 38, 1794–1804. [Google Scholar] [CrossRef]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Rinnan, Å.; van den Berg, F.; Engelsen, S.B. Review of the most common pre-processing techniques for near-infrared spectra. TRAC Trend. Anal. Chem. 2009, 28, 1201–1222. [Google Scholar] [CrossRef]
- Savitzky, A.; Golay, M.J.E. Smoothing and Differentiation of Data by Simplified Least Squares Procedures. Anal. Chem. 1964, 36, 1627–1639. [Google Scholar] [CrossRef]
- Barnes, R.J.; Dhanoa, M.S.; Lister, S.J. Standard Normal Variate Transformation and De-Trending of Near-Infrared Diffuse Reflectance Spectra. Appl. Spectrosc. 1989, 43, 772–777. [Google Scholar] [CrossRef]
- Golic, M.; Walsh, K.; Lawson, P. Short-Wavelength Near-Infrared Spectra of Sucrose, Glucose, and Fructose with Respect to Sugar Concentration and Temperature. Appl. Spectrosc. 2003, 57, 139–145. [Google Scholar] [CrossRef]
- Cavaco, A.M.; Pires, R.; Antunes, M.D.; Panagopoulos, T.; Brázio, A.; Afonso, A.M.; Silva, L.; Lucas, M.R.; Cadeiras, B.; Cruz, S.P.; et al. Validation of short wave near infrared calibration models for the quality and ripening of Newhall orange on tree across years and orchards. Postharvest Biol. Technol. 2018, 141, 86–97. [Google Scholar] [CrossRef]
- Geladi, P.; Kowalski, B.R. Partial least-squares regression: A tutorial. Anal. Chim. Acta 1986, 185, 1–17. [Google Scholar] [CrossRef]
- Mehmood, T.; Liland, K.H.; Snipen, L.; Sæbø, S. A review of variable selection methods in Partial Least Squares Regression. Chemom. Intell. Lab. 2012, 118, 62–69. [Google Scholar] [CrossRef]
- Pellicia, D. A Variable Selection Method for PLS in Python; Instruments & Data Tools Pty Ltd.: Rowville, Australia, 2018. [Google Scholar]
- Drucker, H.; Burges, C.J.C.; Kaufman, L.; Smola, A.; Vapnik, V. Support Vector Regression Machines. In Proceedings of the 9th International Conference on Neural Information Processing Systems, NIPS’96, Denver, CO, USA, 3–5 December 1996; MIT Press: Cambridge, MA, USA, 1996; pp. 155–161. [Google Scholar]
- Smola, A.J.; Schölkopf, B. A tutorial on support vector regression. Stat. Comput. 2004, 14, 199–222. [Google Scholar] [CrossRef]
- Tsirikoglou, P.; Abraham, S.; Contino, F.; Lacor, C.; Ghorbaniasl, G. A hyperparameters selection technique for support vector regression models. Appl. Soft Comput. 2017, 61, 139–148. [Google Scholar] [CrossRef]
- Kaneko, H.; Funatsu, K. Fast optimization of hyperparameters for support vector regression models with highly predictive ability. Chemom. Intell. Lab. 2015, 142, 64–69. [Google Scholar] [CrossRef]
- Cherkassky, V.; Ma, Y. Practical selection of SVM parameters and noise estimation for SVM regression. Neural Netw. 2004, 17, 113–126. [Google Scholar] [CrossRef]
- Tang, Y.; Guo, W.; Gao, J. Efficient model selection for Support Vector Machine with Gaussian kernel function. In Proceedings of the 2009 IEEE Symposium on Computational Intelligence and Data Mining, Nashville, TN, USA, 30 March–2 April 2009; pp. 40–45. [Google Scholar] [CrossRef]
- Heaton, J. Introduction to Neural Networks with JAVA; Heaton Research: St. Louis, MI, USA, 2008. [Google Scholar]
- Kingma, D.P.; Ba, J. Adam: A Method for Stochastic Optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar]
- Li, H.; Liang, Y.; Xu, Q.; Cao, D. Key wavelengths screening using competitive adaptive reweighted sampling method for multivariate calibration. Anal. Chim. Acta 2009, 648, 77–84. [Google Scholar] [CrossRef] [PubMed]
- Malek, S.; Melgani, F.; Bazi, Y. One-dimensional convolutional neural networks for spectroscopic signal regression. J. Chemom. 2017, 32. [Google Scholar] [CrossRef]
- Cui, C.; Fearn, T. Modern practical convolutional neural networks for multivariate regression: Applications to NIR calibration. Chemom. Intell. Lab. 2018, 182, 9–20. [Google Scholar] [CrossRef]
- Ni, C.; Wang, D.; Tao, Y. Variable weighted convolutional neural network for the nitrogen content quantization of Masson pine seedling leaves with near-infrared spectroscopy. Spectrochim. Acta A 2019, 209, 32–39. [Google Scholar] [CrossRef]









| abs | abs_snv | abs1d | abs1d_snv | abs2d | abs2d_snv | |
|---|---|---|---|---|---|---|
| ABCD (full spec.) | 419 (19) | 575 (17) | 542 (22) | 728 (25) | 585 (25) | 608 (25) |
| ABCE (full spec.) | 439 (18) | 536 (17) | 637 (25) | 698 (25) | 534 (25) | 587 (24) |
| ABDE (full spec.) | 472 (18) | 604 (18) | 540 (25) | 692 (25) | 633 (24) | 650 (22) |
| ACDE (full spec.) | 459 (18) | 523 (23) | 576 (25) | 578 (25) | 598 (25) | 597 (25) |
| BCDE (full spec.) | 489 (17) | 528 (18) | 598 (23) | 581 (24) | 593 (19) | 552 (25) |
| Big train (full spec.) | 413 (19) | 581 (19) | 595 (25) | 622 (25) | 623 (22) | 612 (23) |
| Small train (full spec.) | 579 (22) | 543 (23) | 560 (23) | 798 (13) | 651 (23) | 691 (24) |
| ABCD (no Chl) | 311 (13) | 309 (11) | 430 (25) | 378 (24) | 328 (22) | 429 (24) |
| ABCE (no Chl) | 263 (12) | 333 (9) | 438 (25) | 418 (24) | 341 (25) | 417 (24) |
| ABDE (no Chl) | 245 (13) | 365 (10) | 384 (24) | 374 (25) | 356 (24) | 381 (24) |
| ACDE (no Chl) | 256 (13) | 289 (10) | 458 (22) | 427 (25) | 356 (25) | 414 (18) |
| BCDE (no Chl) | 227 (14) | 306 (10) | 410 (24) | 399 (24) | 348 (25) | 373 (22) |
| Big train(no Chl) | 250 (13) | 276 (14) | 406 (23) | 423 (25) | 374 (24) | 393 (23) |
| Small train (no Chl) | 310 (18) | 263 (18) | 455 (25) | 418 (25) | 349 (25) | 386 (25) |
| Best Preproc. | RMSEC | RMSEP | R | % | PG | |
|---|---|---|---|---|---|---|
| EV (full spec.) | abs1d_snv2 | 0.96 | 1.15 | 0.56 | 8.86 | 1.55 |
| Big (full spec.) | abs1d2 | 0.86 | 0.90 | 0.73 | 6.91 | 1.91 |
| Small (full spec.) | abs1d2 | 0.46 | 0.68 | 0.62 | 4.95 | 1.57 |
| EV (no Chl) | abs1d_snv2 | 0.97 | 1.11 | 0.58 | 8.57 | 1.60 |
| Big (no Chl) | abs1d2 | 0.90 | 0.98 | 0.68 | 7.54 | 1.75 |
| Small (no Chl) | abs1d2 | 0.48 | 0.76 | 0.55 | 5.60 | 1.38 |
| Best Preproc. | RMSEC | RMSEP | R | % | PG | |
|---|---|---|---|---|---|---|
| EV (full spec.) | abs1d_snv2 | 0.89 | 1.15 | 0.55 | 8.9 | 1.54 |
| Big (full spec.) | abs1d2 | 0.78 | 0.91 | 0.72 | 7.0 | 1.88 |
| EV (no Chl) | abs1d_snv2 | 0.91 | 1.11 | 0.57 | 8.63 | 1.59 |
| Big (no Chl) | abs1d2 | 0.85 | 0.99 | 0.67 | 7.56 | 1.74 |
| Best Preproc. | RMSEC | RMSEP | R | % | PG | |
|---|---|---|---|---|---|---|
| EV (full spec.) | abs1d_snv2 | 0.93 | 1.16 | 0.57 | 8.94 | 1.56 |
| Big (full spec.) | abs1d2 | 0.84 | 0.82 | 0.77 | 6.31 | 2.09 |
| Small (full spec.) | abs1d_snv2 | 0.59 | 0.62 | 0.66 | 4.59 | 1.71 |
| EV (no Chl) | abs1d_snv2 | 0.92 | 1.09 | 0.60 | 8.45 | 1.63 |
| Big (no Chl) | abs1d2 | 0.84 | 0.82 | 0.77 | 6.33 | 2.09 |
| Small (no Chl) | abs1d | 0.63 | 0.63 | 0.68 | 4.56 | 1.70 |
| Best Preproc. | RMSEC | RMSEP | R | % | PG | |
|---|---|---|---|---|---|---|
| EV (full spec.) | abs1d_snv2 | 0.60 | 1.15 | 0.57 | 8.89 | 1.55 |
| Big (full spec.) | abs1d2 | 0.54 | 0.88 | 0.75 | 6.78 | 1.96 |
| Small (full spec.) | abs2 | 0.39 | 0.70 | 0.58 | 5.15 | 1.53 |
| EV (no Chl) | abs_snv2 | 0.49 | 1.21 | 0.51 | 9.64 | 1.46 |
| Big (no Chl) | abs1d2 | 0.58 | 0.92 | 0.73 | 7.09 | 1.86 |
| Small (no Chl) | abs | 0.49 | 0.75 | 0.51 | 5.52 | 1.41 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Passos, D.; Rodrigues, D.; Cavaco, A.M.; Antunes, M.D.; Guerra, R. Non-Destructive Soluble Solids Content Determination for ‘Rocha’ Pear Based on VIS-SWNIR Spectroscopy under ‘Real World’ Sorting Facility Conditions. Sensors 2019, 19, 5165. https://doi.org/10.3390/s19235165
Passos D, Rodrigues D, Cavaco AM, Antunes MD, Guerra R. Non-Destructive Soluble Solids Content Determination for ‘Rocha’ Pear Based on VIS-SWNIR Spectroscopy under ‘Real World’ Sorting Facility Conditions. Sensors. 2019; 19(23):5165. https://doi.org/10.3390/s19235165
Chicago/Turabian StylePassos, Dário, Daniela Rodrigues, Ana Margarida Cavaco, Maria Dulce Antunes, and Rui Guerra. 2019. "Non-Destructive Soluble Solids Content Determination for ‘Rocha’ Pear Based on VIS-SWNIR Spectroscopy under ‘Real World’ Sorting Facility Conditions" Sensors 19, no. 23: 5165. https://doi.org/10.3390/s19235165
APA StylePassos, D., Rodrigues, D., Cavaco, A. M., Antunes, M. D., & Guerra, R. (2019). Non-Destructive Soluble Solids Content Determination for ‘Rocha’ Pear Based on VIS-SWNIR Spectroscopy under ‘Real World’ Sorting Facility Conditions. Sensors, 19(23), 5165. https://doi.org/10.3390/s19235165









