Comparison of Spaceborne and UAV-Borne Remote Sensing Spectral Data for Estimating Monsoon Crop Vegetation Parameters
Abstract
1. Introduction
- To build crop-specific parametric and non-parametric models to estimate crop vegetation parameters
- To evaluate the developed vegetation parameter estimation models against (a) the spectral sensitivity of the RS data (multispectral vs hyperspectral), (b) modelling method (parametric and non-parametric), and (c) crop type (finger millet, maize, and lablab)
- To explore how crop-wise vegetation parameter estimation is affected by agricultural treatment (irrigation and fertiliser)
2. Materials and Methods
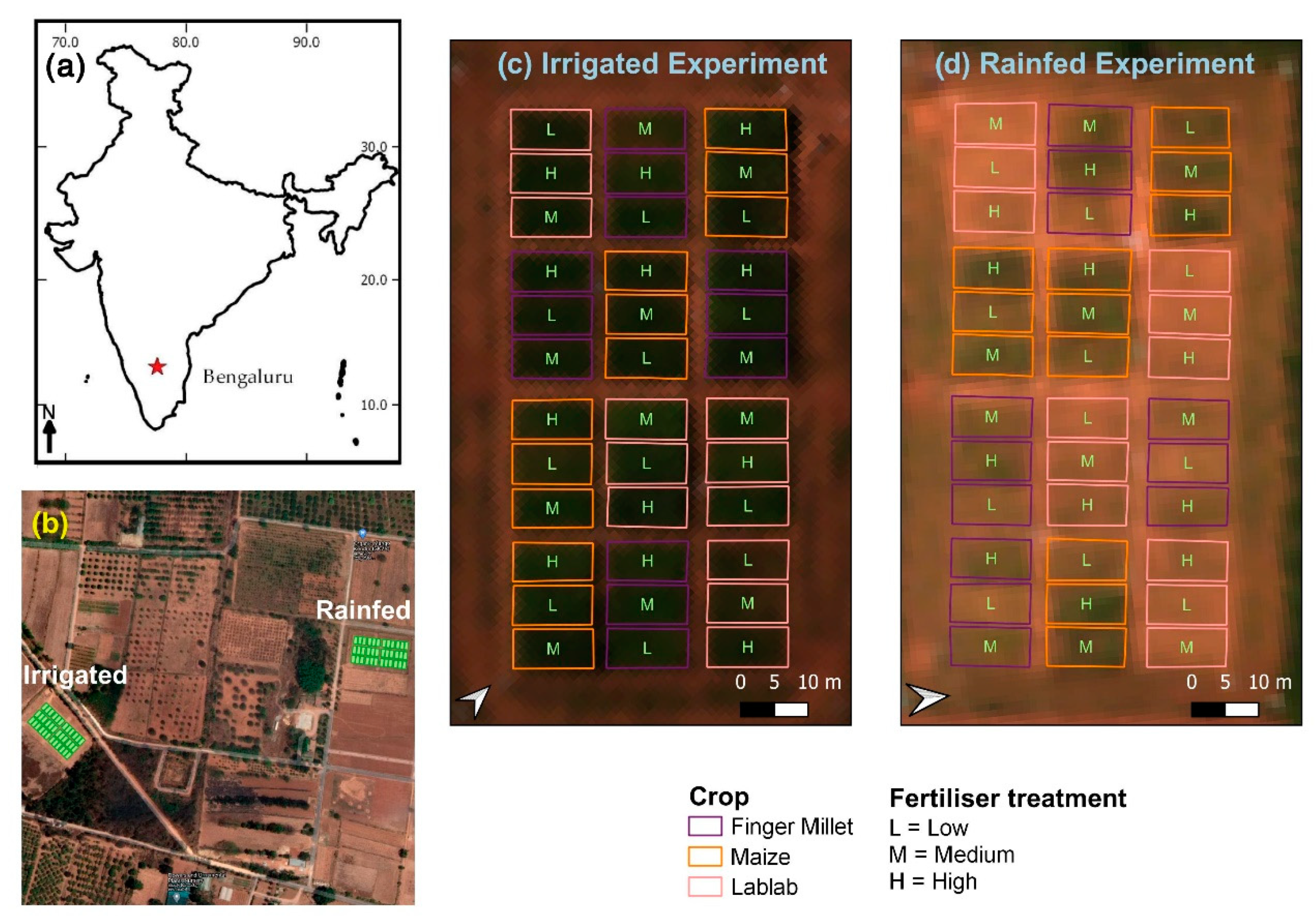
2.1. Study Site and Experimental Design
2.2. In-Situ Field Data
2.3. Remote Sensing Data
2.3.1. WorldView 3 Data
2.3.2. Cubert Hyperspectral Data
2.4. Model-Building Workflow for Crop Vegetation Parameter Estimation
3. Results
3.1. Crop Vegetation Parameter Data
3.2. Spectral and Vegetation Index Data
3.3. Crop Vegetation Parameter Estimation with Linear Regression
3.4. Crop Vegetation Parameter Estimation with Random Forest Regression
3.4.1. Key Wavebands
3.4.2. Model Performance
3.5. Best Models and Distribution of Residuals
4. Discussion
4.1. Finger Millet Vegetation Parameter Estimation
4.2. Lablab Vegetation Parameter Estimation
4.3. Maize Crop Vegetation Parameter Estimation
4.4. Overall Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
| Crop | Water | Min | Mean | SD | Max | CV |
|---|---|---|---|---|---|---|
| LAI (m2/m2) | ||||||
| Finger millet | Irrigated | 1.4 | 2.6 | 0.5 | 3.2 | 19.2% |
| Rainfed | 0.4 | 1.0 | 0.4 | 1.6 | 40.0% | |
| Lablab | Irrigated | 1.7 | 2.5 | 0.6 | 3.5 | 24.0% |
| Rainfed | 0.2 | 0.5 | 0.2 | 0.7 | 40.0% | |
| Maize | Irrigated | 2.1 | 2.7 | 0.2 | 3.0 | 7.4% |
| Rainfed | 1.0 | 1.6 | 0.4 | 2.2 | 25.0% | |
| LCC (µg/cm2) | ||||||
| Finger millet | Irrigated | 19.3 | 39.7 | 13.6 | 65.6 | 34.3% |
| Rainfed | 10.2 | 12.8 | 3.4 | 21.4 | 26.6% | |
| Lablab | Irrigated | 17.5 | 36.3 | 7.8 | 43.0 | 21.5% |
| Rainfed | 20.1 | 27.5 | 4.3 | 33.3 | 15.6% | |
| Maize | Irrigated | 15.7 | 42.1 | 19.6 | 76.4 | 46.6% |
| Rainfed | 11.9 | 20.3 | 5.3 | 30.6 | 26.1% | |
| CWC (kg/m2) | ||||||
| Finger millet | Irrigated | 0.4 | 1.4 | 0.7 | 2.7 | 46.5% |
| Rainfed | 0.1 | 0.5 | 0.2 | 1.0 | 52.1% | |
| Lablab | Irrigated | 0.4 | 0.7 | 0.3 | 1.6 | 48.5% |
| Rainfed | 0.03 | 0.08 | 0.04 | 0.1 | 50.0% | |
| Maize | Irrigated | 0.8 | 1.5 | 0.4 | 2.3 | 26.6% |
| Rainfed | 0.2 | 0.9 | 0.4 | 1.5 | 45.5% | |



References
- D’Amour, C.B.; Reitsma, F.; Baiocchi, G.; Barthel, S.; Güneralp, B.; Erb, K.H.; Haberl, H.; Creutzig, F.; Seto, K.C. Future urban land expansion and implications for global croplands. Proc. Natl. Acad. Sci. USA 2017, 114, 8939–8944. [Google Scholar] [CrossRef]
- Kübler, D.; Lefèvre, C. Megacity governance and the state. Urban Res. Pract. 2018, 11, 378–395. [Google Scholar] [CrossRef]
- Patil, S.; Dhanya, B.; Vanjari, R.S.; Purushothaman, S. Urbanisation and new agroecologies. Econ. Polit. Wkly. 2018, LIII, 71–77. [Google Scholar]
- Directorate of Economics and Statistics. Report on Area, Production, Productivity and Prices of Agriculture Crops in Karnataka, 2009–2010; DES: Bengaluru, India, 2012.
- Food and Agriculture Organization of the United Nations. The Future of Food and Agriculture: Trends and Challenges; FAO: Rome, Italy, 2017. [Google Scholar]
- Mulla, D.J. Twenty five years of remote sensing in precision agriculture: Key advances and remaining knowledge gaps. Biosyst. Eng. 2013, 114, 358–371. [Google Scholar] [CrossRef]
- Sishodia, R.P.; Ray, R.L.; Singh, S.K. Applications of remote sensing in precision agriculture: A review. Remote Sens. 2020, 12, 3136. [Google Scholar] [CrossRef]
- Jones, H.G.; Vaughan, R.A. Remote Sensing of Vegetation—Principles, Techniques, and Applications; Oxford University Press: New York, NY, USA, 2010; ISBN 978-0-19-920779-4. [Google Scholar]
- Thenkabail, P.S.; Lyon, J.G.; Huete, A. Hyperspectral Remote Sensing of Vegetation; Prasad, S.T., John, G.L., Alfredo, H., Eds.; CRC Press: Boca Raton, FL, USA, 2011; ISBN 1439845387. [Google Scholar]
- Mananze, S.; Pôças, I.; Cunha, M. Retrieval of maize leaf area index using hyperspectral and multispectral data. Remote Sens. 2018, 10, 1942. [Google Scholar] [CrossRef]
- Verrelst, J.; Malenovský, Z.; Van der Tol, C.; Camps-Valls, G.; Gastellu-Etchegorry, J.P.; Lewis, P.; North, P.; Moreno, J. Quantifying Vegetation Biophysical Variables from Imaging Spectroscopy Data: A Review on Retrieval Methods. Surv. Geophys. 2019, 40, 589–629. [Google Scholar] [CrossRef]
- Rivera, J.P.; Verrelst, J.; Delegido, J.; Veroustraete, F.; Moreno, J. On the semi-automatic retrieval of biophysical parameters based on spectral index optimization. Remote Sens. 2014, 6, 4927–4951. [Google Scholar] [CrossRef]
- Verrelst, J.; Camps-Valls, G.; Muñoz-Marí, J.; Rivera, J.P.; Veroustraete, F.; Clevers, J.G.P.W.; Moreno, J. Optical remote sensing and the retrieval of terrestrial vegetation bio-geophysical properties—A review. ISPRS J. Photogramm. Remote Sens. 2015, 108, 273–290. [Google Scholar] [CrossRef]
- Caicedo, J.P.R. Optimized and Automated Estimation of Vegetation Properties: Opportunities for Sentinel-2. Ph.D. Thesis, Universitat De València, Valencia, Spain, 2014. [Google Scholar]
- Chen, J.M.; Black, T.A. Defining leaf area index for non-flat leaves. Plant. Cell Environ. 1992, 15, 421–429. [Google Scholar] [CrossRef]
- Xu, J.; Quackenbush, L.J.; Volk, T.A.; Im, J. Forest and crop leaf area index estimation using remote sensing: Research trends and future directions. Remote Sens. 2020, 12, 2934. [Google Scholar] [CrossRef]
- Houborg, R.; McCabe, M.F.; Cescatti, A.; Gitelson, A.A. Leaf chlorophyll constraint on model simulated gross primary productivity in agricultural systems. Int. J. Appl. Earth Obs. Geoinf. 2015, 43, 160–176. [Google Scholar] [CrossRef]
- Croft, H.; Chen, J.M.; Wang, R.; Mo, G.; Luo, S.; Luo, X.; He, L.; Gonsamo, A.; Arabian, J.; Zhang, Y.; et al. The global distribution of leaf chlorophyll content. Remote Sens. Environ. 2020, 236. [Google Scholar] [CrossRef]
- Caicedo, J.P.R.; Verrelst, J.; Munoz-Mari, J.; Moreno, J.; Camps-Valls, G. Toward a semiautomatic machine learning retrieval of biophysical parameters. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2014, 7, 1249–1259. [Google Scholar] [CrossRef]
- Liang, L.; Qin, Z.; Zhao, S.; Di, L.; Zhang, C.; Deng, M.; Lin, H.; Zhang, L.; Wang, L.; Liu, Z. Estimating crop chlorophyll content with hyperspectral vegetation indices and the hybrid inversion method. Int. J. Remote Sens. 2016, 37, 2923–2949. [Google Scholar] [CrossRef]
- Penuelas, J.; Filella, I.; Biel, C.; Serrano, L.; Save, R. The reflectance at the 950–970 nm region as an indicator of plant water status. Int. J. Remote Sens. 1993, 14, 1887–1905. [Google Scholar] [CrossRef]
- Clevers, J.G.P.W.; Kooistra, L.; Schaepman, M.E. Estimating canopy water content using hyperspectral remote sensing data. Int. J. Appl. Earth Obs. Geoinf. 2010, 12, 119–125. [Google Scholar] [CrossRef]
- Penuelas, J.; Pinol, J.; Ogaya, R.; Filella, I. Estimation of plant water concentration by the reflectance Water Index WI (R900/R970). Int. J. Remote Sens. 1997, 18, 2869–2875. [Google Scholar] [CrossRef]
- Zhang, F.; Zhou, G. Estimation of vegetation water content using hyperspectral vegetation indices: A comparison of crop water indicators in response to water stress treatments for summer maize. BMC Ecol. 2019, 19, 1–12. [Google Scholar] [CrossRef]
- Zhang, F.; Zhou, G. Estimation of canopy water content by means of hyperspectral indices based on drought stress gradient experiments of maize in the north plain China. Remote Sens. 2015, 7, 15203–15223. [Google Scholar] [CrossRef]
- Weiss, M.; Jacob, F.; Duveiller, G. Remote sensing for agricultural applications: A meta-review. Remote Sens. Environ. 2020, 236, 111402. [Google Scholar] [CrossRef]
- Tsouros, D.C.; Bibi, S.; Sarigiannidis, P.G. A Review on UAV-Based Applications for Precision Agriculture. Information 2019, 10, 349. [Google Scholar] [CrossRef]
- Lu, B.; He, Y.; Dao, P.D. Comparing the performance of multispectral and hyperspectral images for estimating vegetation properties. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2019, 12, 1784–1797. [Google Scholar] [CrossRef]
- Dayananda, S.; Astor, T.; Wijesingha, J.; Chickadibburahalli Thimappa, S.; Dimba Chowdappa, H.; Mudalagiriyappa; Nidamanuri, R.R.; Nautiyal, S.; Wachendorf, M. Multi-Temporal Monsoon Crop Biomass Estimation Using Hyperspectral Imaging. Remote Sens. 2019, 11, 1771. [Google Scholar] [CrossRef]
- Danner, M.; Locherer, M.; Hank, T.; Richter, K. Measuring Leaf Area Index (LAI) with the LI-Cor LAI 2200C or LAI-2200; EnMAP Field Guide Technical Report; GFZ Data Services: Potsdam, Germany, 2015. [Google Scholar]
- Cerovic, Z.G.; Masdoumier, G.; Ghozlen, N.B.; Latouche, G. A new optical leaf-clip meter for simultaneous non-destructive assessment of leaf chlorophyll and epidermal flavonoids. Physiol. Plant. 2012, 146, 251–260. [Google Scholar] [CrossRef]
- Kuester, M. Radiometric Use of WorldView-3 Imagery; Digital Globe: Longmont, CO, USA, 2016. [Google Scholar]
- Digital Globe. WorldView-3; Digital Globe: Longmont, CO, USA, 2014. [Google Scholar]
- Davaadorj, A. Evaluating Atmospheric Correction Methods Using Worldview-3 Image. Master’s Thesis, University of Twente, Enschede, The Netherlands, 2019. [Google Scholar]
- Liu, K.; Zhou, Q.B.; Wu, W.B.; Xia, T.; Tang, H.J. Estimating the crop leaf area index using hyperspectral remote sensing. J. Integr. Agric. 2016, 15, 475–491. [Google Scholar] [CrossRef]
- Datt, B. Remote Sensing of Chlorophyll a, Chlorophyll b, Chlorophyll a+b, and Total Carotenoid Content in Eucalyptus Leaves. Remote Sens. Environ. 1998, 66, 111–121. [Google Scholar] [CrossRef]
- Rouse, J.W.; Haas, R.H.; Schell, J.A.; Deering, D.W. Monitoring the vernal advancement and retrogradation (green wave effect) of natural vegetation. Prog. Rep. RSC 1978-1 1973, 112. [Google Scholar]
- Gitelson, A.A.; Kaufman, Y.J.; Merzlyak, M.N. Use of a green channel in remote sensing of global vegetation from EOS- MODIS. Remote Sens. Environ. 1996, 58, 289–298. [Google Scholar] [CrossRef]
- Haboudane, D.; Miller, J.R.; Pattey, E.; Zarco-Tejada, P.J.; Strachan, I.B. Hyperspectral vegetation indices and novel algorithms for predicting green LAI of crop canopies: Modeling and validation in the context of precision agriculture. Remote Sens. Environ. 2004, 90, 337–352. [Google Scholar] [CrossRef]
- Clevers, J.G.P.W. Imaging Spectrometry in Agriculture—Plant Vitality And Yield Indicators. In Imaging Spectrometry—A Tool for Environmental Observations; Hill, J., Mégier, J., Eds.; Springer: Dordrecht, The Netherlands, 1994; pp. 193–219. ISBN 978-0-585-33173-7. [Google Scholar]
- Aasen, H.; Burkart, A.; Bolten, A.; Bareth, G. Generating 3D hyperspectral information with lightweight UAV snapshot cameras for vegetation monitoring: From camera calibration to quality assurance. ISPRS J. Photogramm. Remote Sens. 2015, 108, 245–259. [Google Scholar] [CrossRef]
- Cubert GmbH. Cubert S185; Cubert GmbH: Ulm, Germany, 2016. [Google Scholar]
- Wijesingha, J.; Astor, T.; Schulze-Brüninghoff, D.; Wengert, M.; Wachendorf, M. Predicting forage quality of grasslands using UAV-borne imaging spectroscopy. Remote Sens. 2020, 12, 126. [Google Scholar] [CrossRef]
- Yang, G.; Li, C.; Wang, Y.; Yuan, H.; Feng, H.; Xu, B.; Yang, X. The DOM generation and precise radiometric calibration of a UAV-mounted miniature snapshot hyperspectral imager. Remote Sens. 2017, 9, 642. [Google Scholar] [CrossRef]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Soukhavong, D. (Laurae) Ensembles of tree-based models: Why correlated features do not trip them—And why NA matters. Available online: https://medium.com/data-design/ensembles-of-tree-based-models-why-correlated-features-do-not-trip-them-and-why-na-matters-7658f4752e1b (accessed on 19 November 2020).
- Kursa, M.B.; Rudnicki, W.R. Feature selection with the boruta package. J. Stat. Softw. 2010, 36, 1–13. [Google Scholar] [CrossRef]
- Probst, P.; Wright, M.N.; Boulesteix, A.L. Hyperparameters and tuning strategies for random forest. Wiley Interdiscip. Rev. Data Min. Knowl. Discov. 2019, 9, 1–19. [Google Scholar] [CrossRef]
- Nembrini, S.; König, I.R.; Wright, M.N. The revival of the Gini importance? Bioinformatics 2018, 34, 3711–3718. [Google Scholar] [CrossRef]
- Lang, M.; Binder, M.; Richter, J.; Schratz, P.; Pfisterer, F.; Coors, S.; Au, Q.; Casalicchio, G.; Kotthoff, L.; Bischl, B. mlr3: A modern object-oriented machine learning framework in R. J. Open Source Softw. 2019, 4, 1903. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2020. [Google Scholar]
- Wright, M.N.; Ziegler, A. Ranger: A fast implementation of random forests for high dimensional data in C++ and R. J. Stat. Softw. 2017, 77. [Google Scholar] [CrossRef]
- Kvalseth, T.O. Cautionary note about R2. Am. Stat. 1985, 39, 279–285. [Google Scholar]
- Afrasiabian, Y.; Noory, H.; Mokhtari, A.; Nikoo, M.R.; Pourshakouri, F.; Haghighatmehr, P. Effects of spatial, temporal, and spectral resolutions on the estimation of wheat and barley leaf area index using multi- and hyper-spectral data (case study: Karaj, Iran). Precis. Agric. 2020. [Google Scholar] [CrossRef]
- Lambert, M.J.; Traoré, P.C.S.; Blaes, X.; Baret, P.; Defourny, P. Estimating smallholder crops production at village level from Sentinel-2 time series in Mali’s cotton belt. Remote Sens. Environ. 2018, 216, 647–657. [Google Scholar] [CrossRef]
- Taylor, J.R.N.; Kruger, J. Millets. Encycl. Food Heal. 2015, 748–757. [Google Scholar] [CrossRef]
- Shafian, S.; Rajan, N.; Schnell, R.; Bagavathiannan, M.; Valasek, J.; Shi, Y.; Olsenholler, J. Unmanned aerial systems-based remote sensing for monitoring sorghum growth and development. PLoS ONE 2018, 13, e0196605. [Google Scholar] [CrossRef] [PubMed]
- Bhadra, S.; Sagan, V.; Maimaitijiang, M.; Maimaitiyiming, M.; Newcomb, M.; Shakoor, N.; Mockler, T.C. Quantifying leaf chlorophyll concentration of sorghum from hyperspectral data using derivative calculus and machine learning. Remote Sens. 2020, 12, 2082. [Google Scholar] [CrossRef]
- Li, J.; Shi, Y.; Veeranampalayam-Sivakumar, A.N.; Schachtman, D.P. Elucidating sorghum biomass, nitrogen and chlorophyll contents with spectral and morphological traits derived from unmanned aircraft system. Front. Plant Sci. 2018, 9, 1–12. [Google Scholar] [CrossRef]
- Allen, L.H. Legumes. Encycl. Hum. Nutr. 2012, 3–4, 74–79. [Google Scholar]
- Haboudane, D.; Miller, J.R.; Tremblay, N.; Pattey, E.; Vigneault, P. Estimation of leaf area index using ground spectral measurements over agriculture crops: Prediction capability assessment of optical indices. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. ISPRS Arch. 2004, 35, 108–113. [Google Scholar]
- Schlemmera, M.; Gitelson, A.; Schepersa, J.; Fergusona, R.; Peng, Y.; Shanahana, J.; Rundquist, D. Remote estimation of nitrogen and chlorophyll contents in maize at leaf and canopy levels. Int. J. Appl. Earth Obs. Geoinf. 2013, 25, 47–54. [Google Scholar] [CrossRef]
- Lichtenthaler, H.K. Chlorophylls and Carotenoids: Pigments of Photosynthetic Biomembranes. Methods Enzymol. 1987, 148, 350–382. [Google Scholar]
- Zhu, W.; Sun, Z.; Yang, T.; Li, J.; Peng, J.; Zhu, K.; Li, S.; Gong, H.; Lyu, Y.; Li, B.; et al. Estimating leaf chlorophyll content of crops via optimal unmanned aerial vehicle hyperspectral data at multi-scales. Comput. Electron. Agric. 2020, 178, 105786. [Google Scholar] [CrossRef]
- Dawson, T.P.; Curran, P.J. Technical note A new technique for interpolating the reflectance red edge position. Int. J. Remote Sens. 1998, 19, 2133–2139. [Google Scholar] [CrossRef]



| Crop | Phenological Stage (Days after Sowing) | |
|---|---|---|
| Irrigated Experiment | Rainfed Experiment | |
| Finger millet | Inflorescence emergence (87) | Inflorescence emergence (79) |
| Lablab | Ripening (83) | Development of fruit (78) |
| Maize | Development of fruit (87) | Development of fruit (79) |
| Band Name | Centre Wavelength (nm) | Effective Bandwidth (nm) |
|---|---|---|
| Coastal blue (CB) | 427.4 | 40.5 |
| Blue (BL) | 481.9 | 54.0 |
| Green (GR) | 547.1 | 61.8 |
| Yellow (YE) | 604.3 | 38.1 |
| Red (RD) | 660.1 | 58.5 |
| Red-edge (RE) | 722.7 | 38.7 |
| Near-infrared 1 (N1) | 824.0 | 100.4 |
| Near-infrared 2 (N2) | 913.6 | 88.9 |
| VI | Formula for WV3 Bands | Formula for CUB Bands | Reference |
|---|---|---|---|
| NDVI800,670 | [37] | ||
| NDVI750,550 | [38] | ||
| DATT4 | [36] | ||
| MTVI | [39] | ||
| REIP | [40] | ||
| WI | [23] |
| Parameter | Crop | RS Data | LR Model with VIs | |||
|---|---|---|---|---|---|---|
| Best Vegetation Index | r | R2cv | nRMSEcv (%) | |||
| LAI (m2/m2) | Finger millet | CUB | NDVI800_670 | 0.88 | 0.74 | 15.7 |
| WV3 | REIP | 0.88 | 0.74 | 16.1 | ||
| Lablab | CUB | NDVI800_670 | 0.90 | 0.77 | 15.6 | |
| WV3 | REIP | 0.90 | 0.77 | 15.9 | ||
| Maize | CUB | NDVI800_670 | 0.90 | 0.77 | 14.9 | |
| WV3 | NDVI800_670 | 0.89 | 0.73 | 16.0 | ||
| LCC (µg/cm2) | Finger millet | CUB | DATT4 | 0.83 | 0.63 | 18.0 |
| WV3 | NDVI750_550 | 0.76 | 0.50 | 21.0 | ||
| Lablab | CUB | NDVI750_550 | 0.67 | 0.37 | 23.3 | |
| WV3 | NDVI750_550 | 0.66 | 0.36 | 23.4 | ||
| Maize | CUB | NDVI800_670 | 0.59 | 0.21 | 24.1 | |
| WV3 | DATT4 | 0.61 | 0.26 | 23.3 | ||
| CWC (kg/m2) | Finger millet | CUB | NDVI750_550 | 0.73 | 0.44 | 19.5 |
| WV3 | NDVI750_550 | 0.73 | 0.43 | 19.9 | ||
| Lablab | CUB | NDVI800_670 | 0.77 | 0.53 | 16.3 | |
| WV3 | REIP | 0.81 | 0.58 | 15.6 | ||
| Maize | CUB | NDVI800_670 | 0.68 | 0.36 | 19.9 | |
| WV3 | DATT4 | 0.76 | 0.51 | 17.0 | ||
| Parameter | Crop | Selected Wavebands from Cubert Data | Selected Wavebands from WorldView3 Data |
|---|---|---|---|
| LAI (m2/m2) | Finger Millet | ρ522, ρ526, ρ582, ρ642, ρ694, ρ702, ρ706, ρ722, ρ730, ρ738, ρ750, ρ762, ρ946 | Blue, Green, Yellow, Red, Red-edge, Near-infrared 2 |
| Lablab | ρ690, ρ698, ρ706, ρ722, ρ726, ρ734, ρ750, ρ826, ρ918, ρ930, ρ946, ρ950, ρ954, ρ958 | Blue, Green, Yellow, Red, Red edge | |
| Maize | ρ474, ρ478, ρ674, ρ682, ρ690, ρ694, ρ794, ρ802, ρ806, ρ822, ρ870, ρ874, ρ890, ρ898, ρ906, ρ930, ρ954 | Blue, Green, Yellow, Red, Red edge, Near-infrared 2 | |
| LCC (µg/cm2) | Finger Millet | ρ746, ρ750, ρ754, ρ758, ρ762, ρ766 | Blue, Green, Yellow, Red, Red edge, Near-infrared 1, Near-infrared 2 |
| Lablab | ρ574, ρ638, ρ718, ρ742, ρ750 | Blue, Green, Yellow, Red, Red edge | |
| Maize | ρ682, ρ690, ρ698, ρ702 | Blue, Green, Yellow, Red, Red edge | |
| CWC (kg/m2) | Finger Millet | ρ470, ρ478, ρ522, ρ526, ρ694, ρ706, ρ710, ρ722, ρ742, ρ746 | Blue, Green, Yellow, Red, Red edge, Near-infrared 2 |
| Lablab | ρ502, ρ606, ρ614, ρ618, ρ630, ρ666, ρ678, ρ682, ρ742, ρ802, ρ834 | Blue, Green, Yellow, Red, Red edge | |
| Maize | ρ866, ρ878, ρ886, ρ918, ρ966, ρ970 | Blue, Green, Yellow, Red, Red edge |
| Parameter | Crop | RS Data | RFR Model with Selected Wavebands | ||
|---|---|---|---|---|---|
| No. of Wavebands | R2cv | nRMSEcv (%) | |||
| LAI (m2/m2) | Finger millet | CUB | 13 | 0.74 | 16.1 |
| WV3 | 6 | 0.70 | 17.1 | ||
| Lablab | CUB | 14 | 0.84 | 12.9 | |
| WV3 | 5 | 0.87 | 12.0 | ||
| Maize | CUB | 18 | 0.79 | 13.9 | |
| WV3 | 6 | 0.80 | 13.9 | ||
| LCC (µg/cm2) | Finger millet | CUB | 6 | 0.45 | 22.1 |
| WV3 | 7 | 0.51 | 20.8 | ||
| Lablab | CUB | 5 | 0.23 | 25.8 | |
| WV3 | 5 | 0.13 | 27.4 | ||
| Maize | CUB | 4 | 0.16 | 24.9 | |
| WV3 | 5 | 0.01 | 31.5 | ||
| CWC (kg/m2) | Finger millet | CUB | 10 | 0.43 | 19.9 |
| WV3 | 6 | 0.23 | 22.9 | ||
| Lablab | CUB | 11 | 0.51 | 16.9 | |
| WV3 | 5 | 0.42 | 18.2 | ||
| Maize | CUB | 4 | 0.24 | 21.4 | |
| WV3 | 5 | 0.26 | 21.4 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wijesingha, J.; Dayananda, S.; Wachendorf, M.; Astor, T. Comparison of Spaceborne and UAV-Borne Remote Sensing Spectral Data for Estimating Monsoon Crop Vegetation Parameters. Sensors 2021, 21, 2886. https://doi.org/10.3390/s21082886
Wijesingha J, Dayananda S, Wachendorf M, Astor T. Comparison of Spaceborne and UAV-Borne Remote Sensing Spectral Data for Estimating Monsoon Crop Vegetation Parameters. Sensors. 2021; 21(8):2886. https://doi.org/10.3390/s21082886
Chicago/Turabian StyleWijesingha, Jayan, Supriya Dayananda, Michael Wachendorf, and Thomas Astor. 2021. "Comparison of Spaceborne and UAV-Borne Remote Sensing Spectral Data for Estimating Monsoon Crop Vegetation Parameters" Sensors 21, no. 8: 2886. https://doi.org/10.3390/s21082886
APA StyleWijesingha, J., Dayananda, S., Wachendorf, M., & Astor, T. (2021). Comparison of Spaceborne and UAV-Borne Remote Sensing Spectral Data for Estimating Monsoon Crop Vegetation Parameters. Sensors, 21(8), 2886. https://doi.org/10.3390/s21082886






