1. Introduction
Lung cancer is the leading cause of cancer-related death worldwide, and non-small cell lung cancer (NSCLC) constitutes approximately 85% of lung cancer [
1,
2]. Accurate non-invasive diagnosis and early prognosis prediction based on biomarker detection help with precision medicine and prolong the survival of NSCLC. Traditional investigation of biomarkers in NSCLC is mainly based on immunohistochemistry (IHC) and fluorescent in situ hybridization (FISH) analysis of the tumor tissue. However, the difficulties in collecting tissue biopsies for repeated detection and the single biopsy bias due to intratumoral heterogeneity limit the application of the tissue-based assessment for accurate diagnosis, dynamic monitoring and prognosis prediction of NSCLC [
3,
4]. Recently, next generation sequencing (NGS) of the tumor tissue and plasma ctDNA has been increasingly accepted as an effective method to detect NSCLC mutation and to guide therapy in the clinical practice [
5,
6]. However, the detection cost is pretty high. Moreover, ctDNA is easy to be degraded, thus a large quantity of sample is needed to allow sufficient copy numbers of ctDNA derived mutations [
7]. Small extracellular vesicles (sEVs) are 30–150 nm cell-released vesicles that are widely present in various body fluids such as the blood, urine, saliva, etc. [
8,
9,
10]. They carry and transfer multiple information (e.g., proteins, nucleic acids, lipids) from their source cells [
11] and serve as desired liquid biopsy biomarkers for cancer detection and therefore provide opportunities for the precision treatment of NSCLC. Due to the complex mechanism under tumor initiation and progression, each biomarker has its distinct role and diagnostic significance. Single-marker analysis on the sEVs can hardly achieve high sensitivity and specificity in the diagnosis and prognosis prediction of NSCLC. Combinational marker analysis would help to improve the diagnostic and prognostic accuracy. The epidermal growth factor receptor (EGFR) is overexpressed in 40% to 80% of NSCLC and is associated with time to progression (TTP) and overall survival (OS) of NSCLC [
12,
13,
14]. C-X-C chemokine receptor 4 (CXCR4) is a chemokine receptor that promotes tumor progression and metastasis [
15]. It is overexpressed in various cancers including NSCLC, especially the advanced NSCLC (A/NSCLC) [
16,
17].
In this study, we established a machine learning-assisted dual-marker detection method based on microbead enrichment and signal amplification in flow cytometry to analyze the expression of EGFR and CXCR4 and in serum sEVs for the diagnosis and prognosis prediction of NSCLC. We have previously developed a microbead-based method in diagnosis and molecular phenotyping of breast cancer which overcame the problem that the nanoscale size of the sEVs exceeded the detection limit of the traditional flow cytometry, while the analysis approach is simple and traditional [
18]. In this study, we mainly focused on the intelligent and automated analysis of the detection results. A machine learning algorithm was developed based on EGFR and CXCR4 expression on serum sEVs to achieve automatic classification of healthy donors (HDs) and NSCLC patients with different malignancies. The dual biomarker analysis offered a high accuracy (97.4% for training cohort and 91.7% for validation cohort) for differentiating early stage NSCLC (E/NSCLC) from A/NSCLC and HDs, and showed potential in predicting the prognosis as early as three days after surgery. These results showed the application potential of this machine learning-assisted dual sEV marker analysis for the accurate diagnosis and prognosis prediction of NSCLC.
2. Materials and Methods
2.1. Cell Culture
All the cell lines were obtained from the Cell Resource Center, Peking Union Medical College (Beijing, China). H1975, A549, and H1650 were cultured in Gibco™ RPMI 1640 Medium (GIBCO-BRL, Gaithersburg, MD, USA) supplemented with 10% fetal bovine serum (FBS) (GIBCO-BRL, Gaithersburg, MD, USA) and 1% penicillin/streptomycin (GIBCO-BRL, Gaithersburg, MD, USA). SW620 was cultured in Dulbecco’s Modified Eagle Medium (DMEM) (GIBCO-BRL, Gaithersburg, MD, USA) supplemented with 10% FBS (GIBCO-BRL, Gaithersburg, MD, USA) and 1% penicillin/streptomycin (GIBCO-BRL, Gaithersburg, MD, USA). The cells were maintained in a humidified inhibitor at 37 °C with 5% CO2—95% air. Subcultivation of the cells was performed using 0.25% trypsin and 5 mM ethylenediaminetetraacetic acid (EDTA) (GIBCO-BRL, Gaithersburg, MD, USA).
2.2. Clinical Samples
Human peripheral blood samples collected from healthy donors and NSCLC patients and paraffin-embedded lung sections from NSCLC patients were all obtained from the department of thoracic surgery, Peking Union Medical College Hospital, China. The collection of human samples was approved by the Medical Ethical Committee of the Peking Union Medical College Hospital (JS-1263). All the participants, including 33 NSCLC patients and 18 healthy volunteers, were recruited with informed consent. The diagnostic criteria and the demographic details of the patients are described in the supplementary information (
Supplementary Tables S1 and S2). Serum samples and metastatic tumor specimens were collected after the lung cancer was pathologically confirmed, and before any chemo-/radio- therapies. The peripheral blood was collected in blood collection tubes and was allowed to clot for 30 min at room temperature. The serum was separated by centrifugation at 3000 rpm for 10 min, aliquoted and stored at −80 °C prior to use. Tumor specimens collected by surgical removal was embedded in 4% paraformaldehyde at room temperature for at least 24 h. After dehydration and paraffin embedding, the tumor specimens were sliced into paraffin sections using a rotary microtome (Leica RM2265, Nussloch, Germany) and IHC assessment was performed on the freshly prepared specimen sections.
All of the samples were randomly selected from larger cohorts and were analyzed in a blinded fashion. Unblinding of clinical parameters and corresponding experimental data was performed only after finishing all experiments.
2.3. sEV Purification
sEVs were purified by ultracentrifugation according to a previously described procedure with modifications [
19,
20]. To isolate sEVs from the cell culture supernatant, the cell culture medium was changed to a medium supplemented with 10% EV-free FBS (GIBCO-BRL, Gaithersburg, MD, USA) and 1% penicillin/streptomycin (GIBCO-BRL, Gaithersburg, MD, USA) for 48 h before sEV purification. To isolate sEVs from human sera, 500 μL of human sera was diluted with PBS solution to the final volume of 27 mL before sEV purification. The prepared cell culture medium or the diluted human sera was collected into 50-mL centrifuge tubes and centrifuged successively at 800×
g for 5 min and 2000×
g for 10 min, followed by filtration through a 0.22-μm filter (Merck Millipore, Darmstadt, Germany) to eliminate large dead cells and cell debris. The conditioned medium was then transferred to the 26.3-mL polycarbonate ultracentrifugation tubes matching the 70-Ti rotor and was ultracentrifuged at 100,000×
g for 2 h at 4 °C to purify sEVs from the cell culture medium and at 150,000 g overnight at 4 °C to purify sEVs from the human sera (OPTIMA XPN-100, Beckman Coulter Inc, Brea, CA, USA) due to the high viscosity of sera. The supernatant was removed completely and the pellet was washed with PBS and ultracentrifuged (at 100,000×
g to purify sEVs from the cell culture medium and at 150,000×
g to purify sEVs from the human sera) for 2 h at 4 °C for a second time. The purified sEV pellets were resuspended in 100 μL PBS.
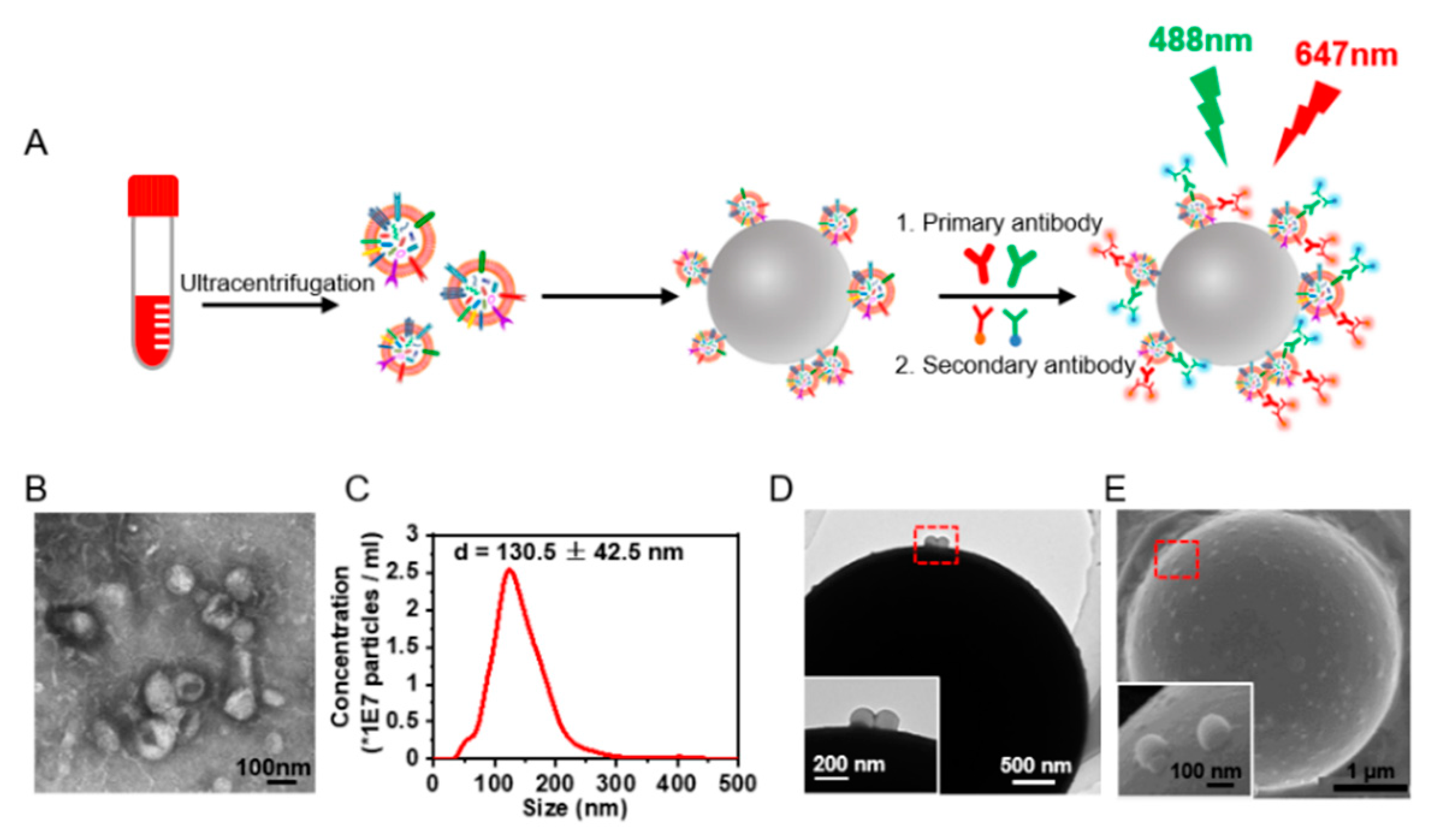
The size and morphology of the purified sEVs were characterized by nanoparticle tracking analysis (NTA), transmission electron microscopy (TEM) and scanning electron microscopy (SEM) as described below.
2.4. Nanoparticle Tracking Analysis
The number and size distribution of sEVs were measured using the NanoSight LM14 system with a 405 nm laser (NanoSight Technology, Malvern, UK). The sEVs derived from cell cultures and sera were diluted in PBS to keep the concentration at 108–109 particles/mL. Samples were injected into the sample chamber with a syringe, measured in triplicate with a high-sensitivity scientific complementary metal-oxide semiconductor (sCMOS) camera at camera setting 16 with an acquisition time of 60 s and a detection threshold setting of 7. The sample chamber was rinsed three times between measuring different samples. Finally, the data were analyzed using the nanoparticle tracking analysis software (NTA version 2.3; Malvern Instruments, Malvern, UK).
2.5. Transmission Electron Microscopy
An optical concentration of sEVs or sEVs-bound beads were loaded onto 200-mesh carbon/formvar coated grids (Beijing Zhongjingkeyi Technology Co., Ltd., Beijing, China) and were allowed to absorb on the grids for 20 min, followed by negative staining with uranyl acetate for 10 min. After rinsing with PBS, the grids were air-dried and subsequently observed with a Hitachi transmission electron microscope.
2.6. Scanning Electron Microscopy
Isolated sEVs were loaded onto silicon wafers and dried in a drying oven, followed by sputter-coating with a thin layer of gold. SEM images were obtained using a Hitachi S-3400N scanning electron microscope (Hitachi High-Tech, Tokyo, Japan) at an acceleration potential of 15 kV.
2.7. Flow Cytometry Analysis
For flow cytometry analysis based on EV-bound beads, 4 μg sEVs were attached to 1 μL 4-μm aldehyde/sulphate latex beads (Invitrogen, Waltham, MA, USA) for 1 h at room temperature with continuous rotation (the sEV/beads ratio is determined by the saturation assay in
Supplementary Figure S2). The input of sEVs was normalized by total protein content on the sEVs according to relative protein quantification using a bicinchoninic acid (BCA) kit (Solarbio, Beijing, China). The reaction was stopped with 100 μM glycine and left rotating for 30 min at room temperature. EV-bound beads were washed once in 0.5% Bovine Serum Albumin (BSA)/PBS and blocked with 5% BSA/PBS with rotation at room temperature for 1 h, then washed a second time in 0.5% BSA/PBS and incubated with anti-EGFR (rabbit mAb, Cell Signaling Technology (CST), #4267) and anti- CXCR4 (goat mAb, Abcam, ab1670, Cambridge, UK) when rotating at 4 °C for 1 h. Beads were centrifuged for 3 min at 14,800×
g, the supernatant was discarded and beads were washed in 0.5% BSA/PBS, then incubated with Alexa-647 (Abcam, anti-rabbit, ab150107) or Alexa-488 (Abcam, anti-goat, ab150073) tagged secondary antibodies with 30 min rotation at 4 °C. After blocking with 5% BSA/PBS, secondary antibodies were incubated with the EV-bound beads as controls as described in the previous studies [
19,
21,
22]. The samples were finally washed by 0.5% BSA/PBS three times and re-suspended in 200 μL PBS. Flow cytometry analysis was performed on a BD Accuri
TM C6 Flow Cytometer (BD Bioscience, Franklin Lakes, NJ, USA).
2.8. Protein Separation and Western Blot Analysis
Cells or sEVs were lysed with lysis buffer, supplemented with protease inhibitor cocktail and phenylmenthysulfonyl fluoride (Thermo Scientific, Waltham, MA, USA) on ice for 60 min. Protein fractions were collected by centrifugation and were normalized according to relative protein quantification using a BCA protein assay kit (Solarbio). Proteins were separated in NuPAGE 10% Bis-Tris Gels (Thermo Scientific, Waltham, MA, USA) under reducing condition, and transferred onto polyvinylidene difluoride (PVDF) membranes (0.45 μm, Millipore, Bedford, MA, USA). The membranes were blocked with 5% non-fat milk (BD Bioscience, Franklin Lakes, NJ, USA) in Tris-buffered saline with 0.1% Tween (TBST) for 1 h and then incubated with anti-EGFR (rabbit mAb, CST, #4267, Boston, MA, USA), anti-CXCR4 (goat mAb, Abcam, ab1670, Cambridge, UK), anti-CD44 (rabbit mAb, Abcam, ab51037, Cambridge, UK), anti-ALDH1A1 (rabbit mAb, CST, #36671, Boston, MA, USA), anti-β actin (mouse mAb, EASYBIO, BE0037, Beijing, China), anti-CD63 (mouse mAb, Santa Cruz, sc-5275, Dallas, TX, USA), anti-CD81 (mouse mAb, Santa Cruz, sc-166029, Dallas, TX, USA), anti-flotillin (rabbit mAb, Abcam, 133497, Cambridge, UK), anti-APOA1(rabbit pAb, Sino biological, 10686-T52, Beijing, China) or anti-calnexin (rabbit pAb, Sino biological, 102109-T32, Beijing, China) overnight at 4 °C. Horse-radish-peroxidase (HRP) conjugated goat anti-rabbit IgG (CST, #7074, Boston, MA, USA), goat anti-mouse IgG (CST #7076, Boston, MA, USA) or donkey anti-goat IgG (Santa Cruz, sc-2020, Dallas, TX, USA) were used as secondary antibodies. The blots were visualized by Image Lab (BIO-RAD, Hercules, CA, USA) with an Enhanced Chemiluminescence Kit (Thermo Pierce, Waltham, MA, USA).
2.9. Immunohistochemical Analysis
Tissues were sectioned into 5 μm thick slices using a microtome and transferred into adhesive slides, dried, deparaffinized in xylene and rehydrated in graded alcohol. Antigen retrieval was performed in a citrate buffer (pH 6) for 15 min after. After blocking with 5% normal goat serum (Solarbio), following staining using EGFR antibody (rabbit mAb, CST, #4267), or CXCR4 antibody (goat mAb, Abcam, ab51037). IHC staining was done using a Vectastain Elite avidin-biotin complex detection kit (Vector Laboratories), and sections were developed by DAB (Sigma-Aldrich, Darmstadt, Germany) according to the manufacturer’s recommendations. Sections were rinsed in tap water, counterstained, cleared and mounted. The image screening and photography of sections were performed using a EVOS® XL Core Imaging System (Thermo Fisher Scientific, Waltham, MA, USA).
2.10. RNA Extraction and Real-Time Polymerase Chain Reaction (PCR)
Total RNA was extracted from sEVs using TRIzol (Life Technologies, Waltham, MA, USA) according to the manufacturer’s instructions. The first-strand cDNA was synthetized by RNA reverse-transcription using QuantScript RT Kit (TIANGEN) before Quantitative Realtime PCR (qPCR) was performed on a Realtime PCR System (Eppendorf), using SuperReal PreMix Plus (SYBR Green) (TIANGEN) according to the manufacturer’s directions. All of the reactions were run in triplicate. The mRNA levels were normalized to glyceraldehyde 3-phosphate dehydrogenase (GAPDH). The relative mRNA expression normalized to control was calculated with the equation 2^(−∆Ct), in which ∆Ct = Ct − Ct(control).
2.11. Logistic Regression
The logistic regression algorithm is a generalized linear model which was used in this work to compute a weighted sum of the expression of EGFR and CXCR4 of sera sEVs. We used the binary logistic regression in SPSS (Statistical Product and Service Solutions) statistical software to weigh the combination of the two biomarkers. Receiver operating characteristic (ROC) analysis was used to evaluate the specificity and sensitivity of EGFR, CXCR4 and the biomarker combination in distinguishing HDs from NSCLC patients, E/NSCLC from A/NSCLC and HDs from E/NSCLC. The area under the ROC curve (AUC) was estimated for each biomarker. All ROC analyses were performed using SPSS statistical software, and the cut-off value was determined using the Youden index.
2.12. Machine Learning
To choose an appropriate classification algorithm for the combinational biomarker of sera sEVs, the cross validation was performed using the whole 51 sEVs samples from HDs and NSCLCs. By comparing the classification performance with different algorithms, Random Forest, which is one of the most powerful machine learning algorithms, was finally chosen as the classification model. Comparing with some “single algorithm” such as SVM (Support Vector Machine) and Decision Tree, Random Forest is an ensemble learning algorithm containing many decision trees, which means better classification and prediction efficacy. The program was written in the Scikit-Learn library in the Python language. In the program, 51 sera samples, including 18 HDs, 16 E/NSCLC and 17 A/NSCLC patients were randomly divided into two groups. One group is a training cohort including 39 samples (14 HDs, 12 E/NSCLC and 13 A/NSCLC patients) and the other group is a validation cohort including 12 samples (4 HDs, 4 E/NSCLC and 4 A/NSCLC patients). EGFR and CXCR4 expression are imported as two independent variables, and the HDs, E/NSCLC and A/NSCLC patients were divided into three classes, termed as 0, 1, and 2, respectively. After learning using the training set database, the efficacy of the classification algorithm was validated by the validation cohort. The performance of classification in both the training and validation sets was evaluated by the accuracy.
2.13. Statistical Analysis
The GraphPad Prism version 6.0 (GraphPad Software) was used for the analysis of flow cytometry results, and data were presented as the mean ± SD in the scatter plots. Comparisons between two groups were made using a Student’s
t-test. An ROC analysis was used to evaluate the specificity and sensitivity of EGFR, CXCR4 and combinational marker of sEVs in differentiating E/NSCLC, A/NSCLC and HDs. The area under the ROC curve (AUC) was estimated for each biomarker. All ROC analyses were performed using SPSS statistical software. The cut-off value was determined using the Youden index. We have outlined the methods of our experiments on EV-TRACK (evtrack.org). The resulting link is
http://evtrack.org/review.php (accessed on 18 January 2022), and the EV-TRACK ID is EV190065.
4. Discussion
sEVs have attracted increasing attention with regard to liquid biopsies that are part of the cancer diagnosis. However, given the complex tumor initiation and progression mechanisms, single-marker analysis on sEVs can hardly achieve high diagnostic and prognostic accuracy. Here we established a dual-marker detection method to analyze the expression of EGFR and CXCR4 on serum sEVs for the diagnosis and prognosis prediction of NSCLS. sEVs were enriched on microbeads and stained with fluorescent antibodies against EGFR and CXCR4 to facilitate signal amplification of these two proteins on the sEVs in flow cytometry analysis, overcoming the problem whereby the nano-scaled size of the sEVs exceeds the detection limit of the traditional flow cytometry. We demonstrated that the expression levels of EGFR and CXCR4 on the sEVs well represented the ones in the source lung cancer cells. We compared serum sEVs from the HDs, E/NSCLCs and A/NSCLCs, and found that the expressions of EGFR and CXCR4 on serum sEVs were significantly higher in A/NSCLCs compared to HDs and E/NSCLCs, suggesting the capability of serum sEV EGFR and CXCR4 for the diagnosis of NSCLC. Moreover, the expression level of EGFR and CXCR4 in serum sEVs correlated well with the ones in the primary tumor tissues as assessed by IHC, suggesting that sEV-based assessments could be used as a noninvasive surrogate to the tissue-based examination by IHC and therefore had potential in clinical application. Considering the various significance of EGFR and CXCR4 for NSCLC progression, logistic regression analysis was used to obtain an unweighted sum of EGFR and CXCR4, which was used as the combinational marker. ROC analysis revealed that the combinational marker had better performance than the single marker in discriminating NSCLC patients from HDs, especially in discriminating A/NSCLC patients from HDs, demonstrating the potential of the combined protein marker in acting as an independent detection biomarker for NSCLC. We further established an intelligent and automated sEV-based method for the accurate detection of NSCLC based on a machine learning algorithm. We found that EGFR and CXCR4 expression identified by machine learning showed an accuracy of 97.4% for the training cohort and 91.7% for the validation cohort in diagnosing and staging NSCLC patients. Moreover, utilizing this machine learning algorithm, we have successfully predicted the possibility of tumor relapse in three patients by classifying their serum sEVs before and three days after surgery. Machine learning-based prognostic classification correlated well with the real clinical prognosis, indicating the capability of the machine learning-assisted serum sEV dual-marker detection method for the early prediction of tumor recurrence after surgery.
In conclusion, the current study demonstrated that the combination of EGFR and CXCR4 in serum sEVs can act as efficient liquid biopsy biomarkers and that machine learning applied in EGFR and CXCR4 expression of serum sEVs improved the diagnostic effectiveness. This study could shed light on clinical applications of this detection method with machine learning analysis for NSCLC diagnosis and early prediction of relapse after surgery. Because of the high accuracy and intelligent characteristics, the detection platform shows clinical potential in monitoring the development of NSCLC, evaluating the prognosis, predicting the possibility of tumor recurrence and facilitating precision therapy.