In Silico Analysis of USP7 Inhibitors Based on Building QSAR Models and Fragment Design for Screening Marine Compound Libraries
Abstract
:1. Introduction
2. Results
2.1. The Analysis of Three Different QSAR Models
2.1.1. Construction and Verification of AutoQSAR Model
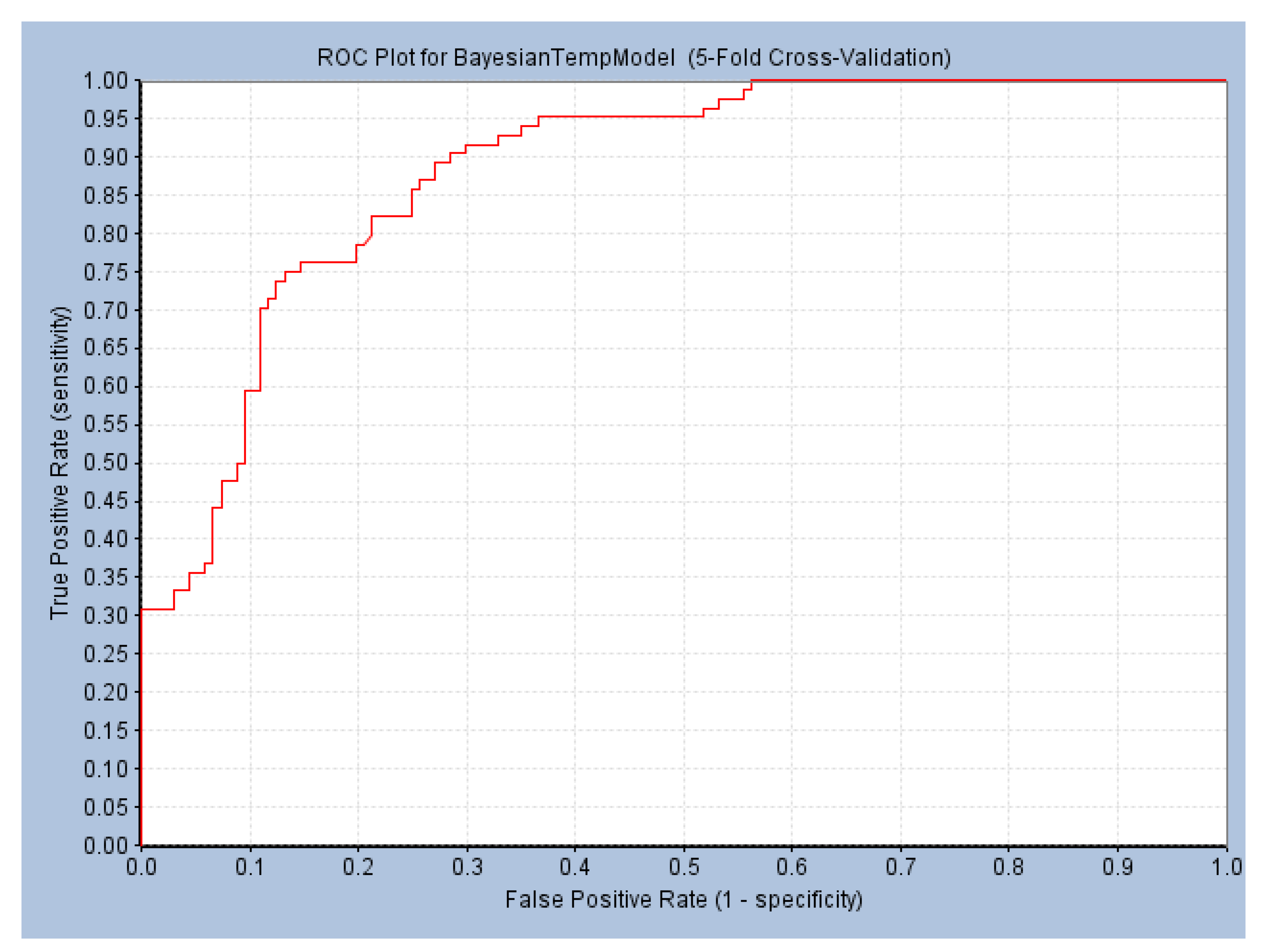
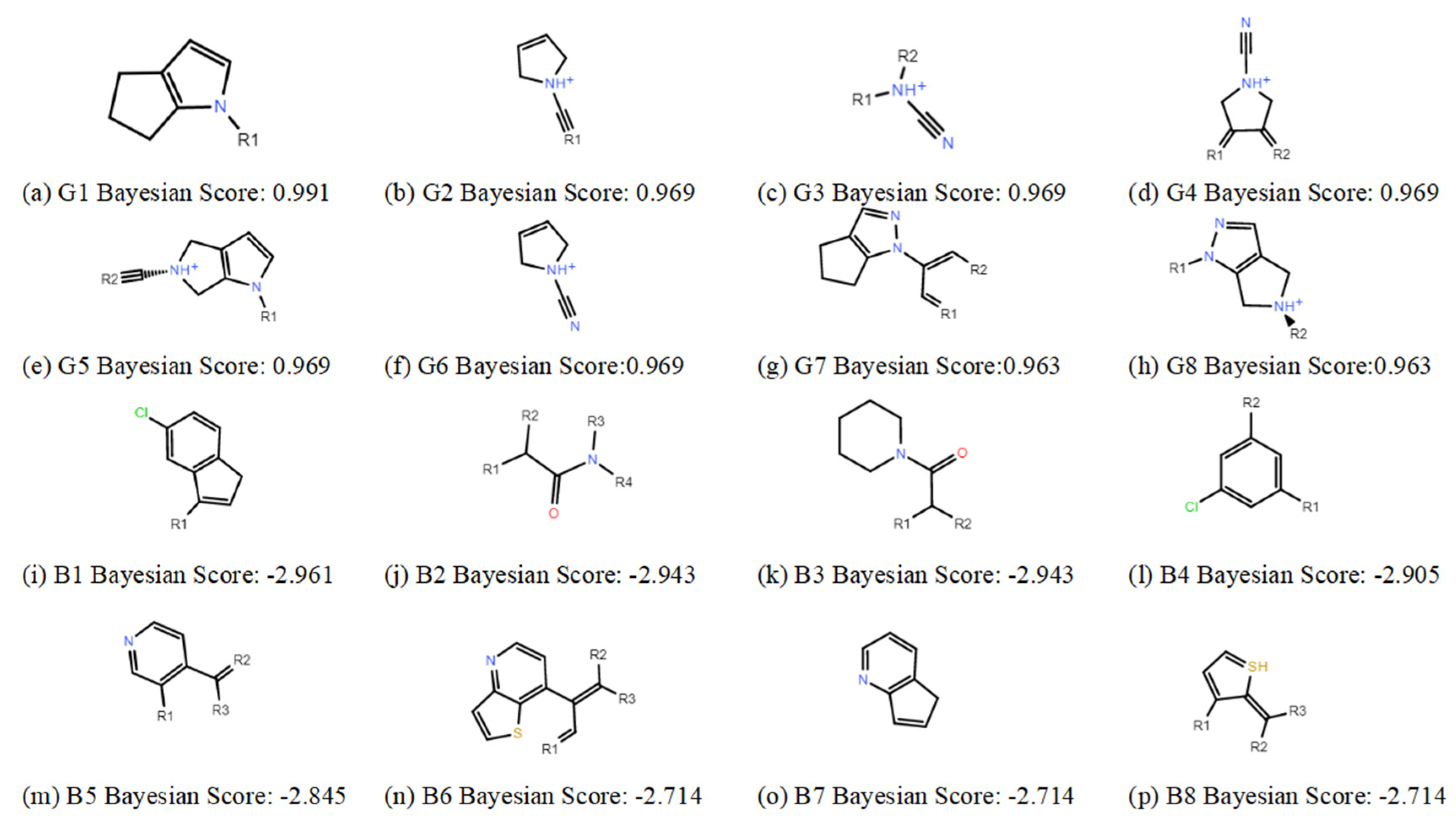
2.1.2. Construction and Verification of Naive Bayesian Model
2.1.3. Construction and Verification of Multiple Linear Regression Model
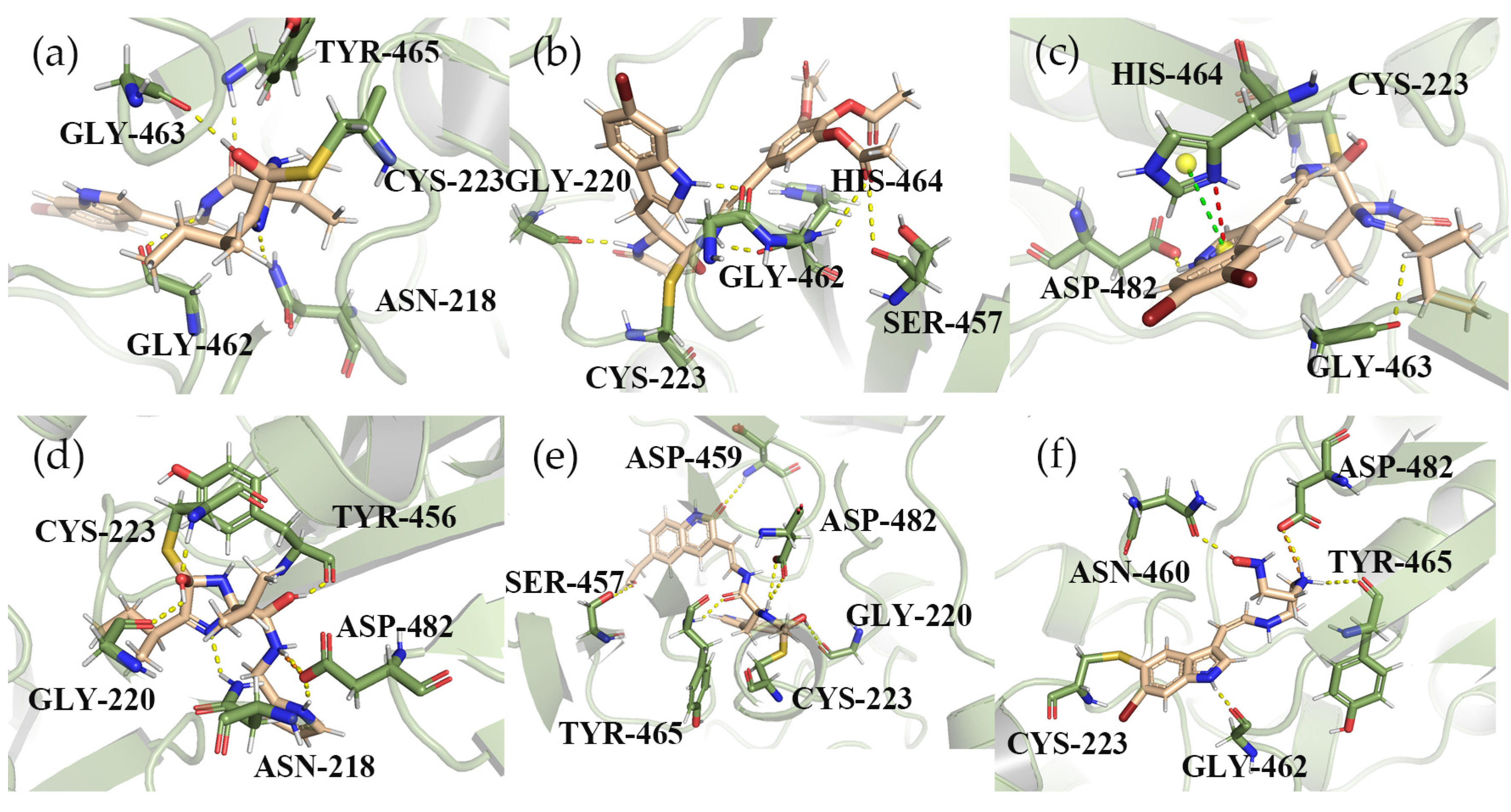
2.2. The Analysis of Molecular Docking
2.3. Covalent Docking
2.4. Scaffold Hopping
2.5. Analysis before and after Scaffold Hopping
2.6. Property Analysis of ADMET
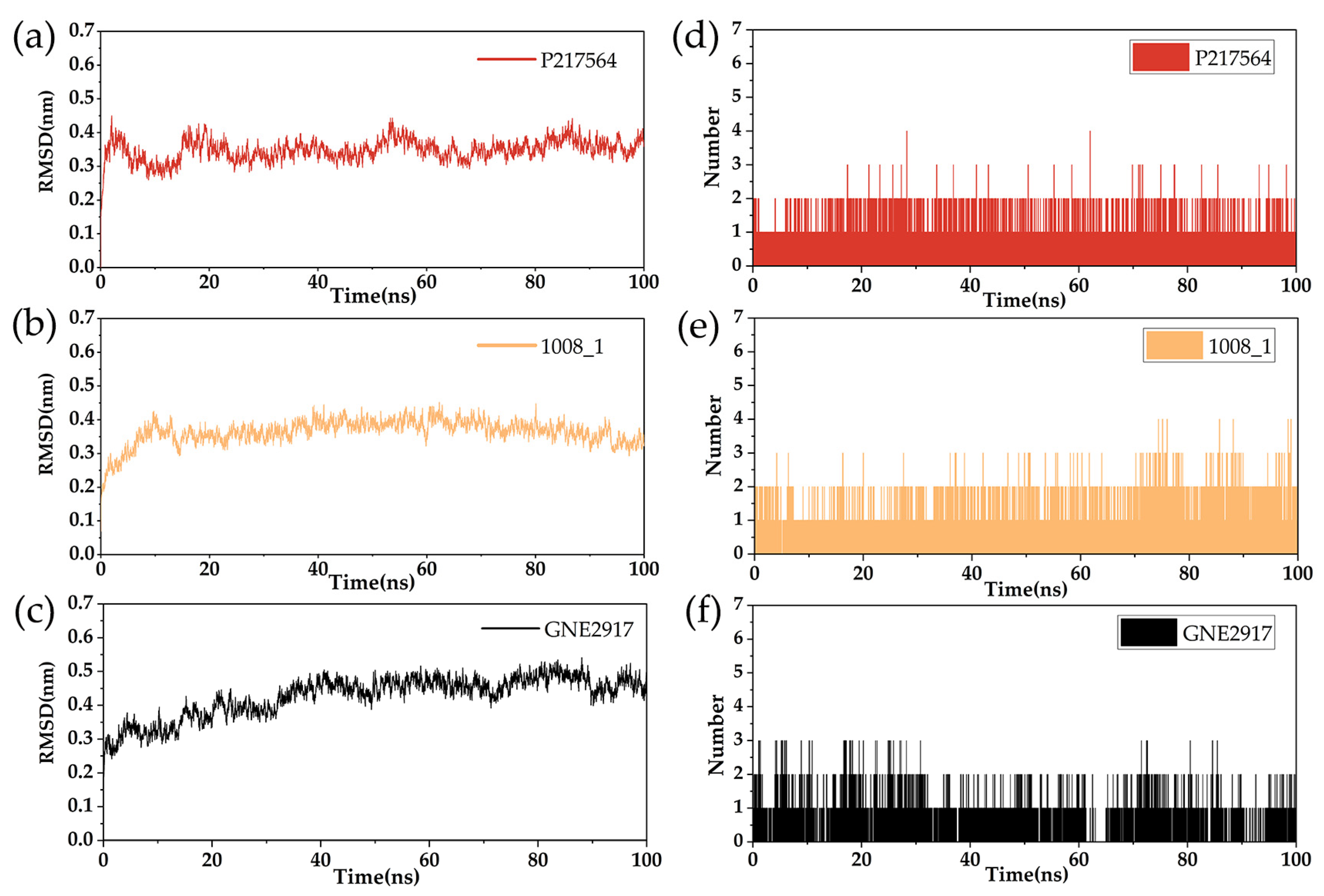
2.7. The Analysis of RMSD and RMSF
2.8. Hydrogen Bond Analysis
3. Discussion
4. Materials and Methods
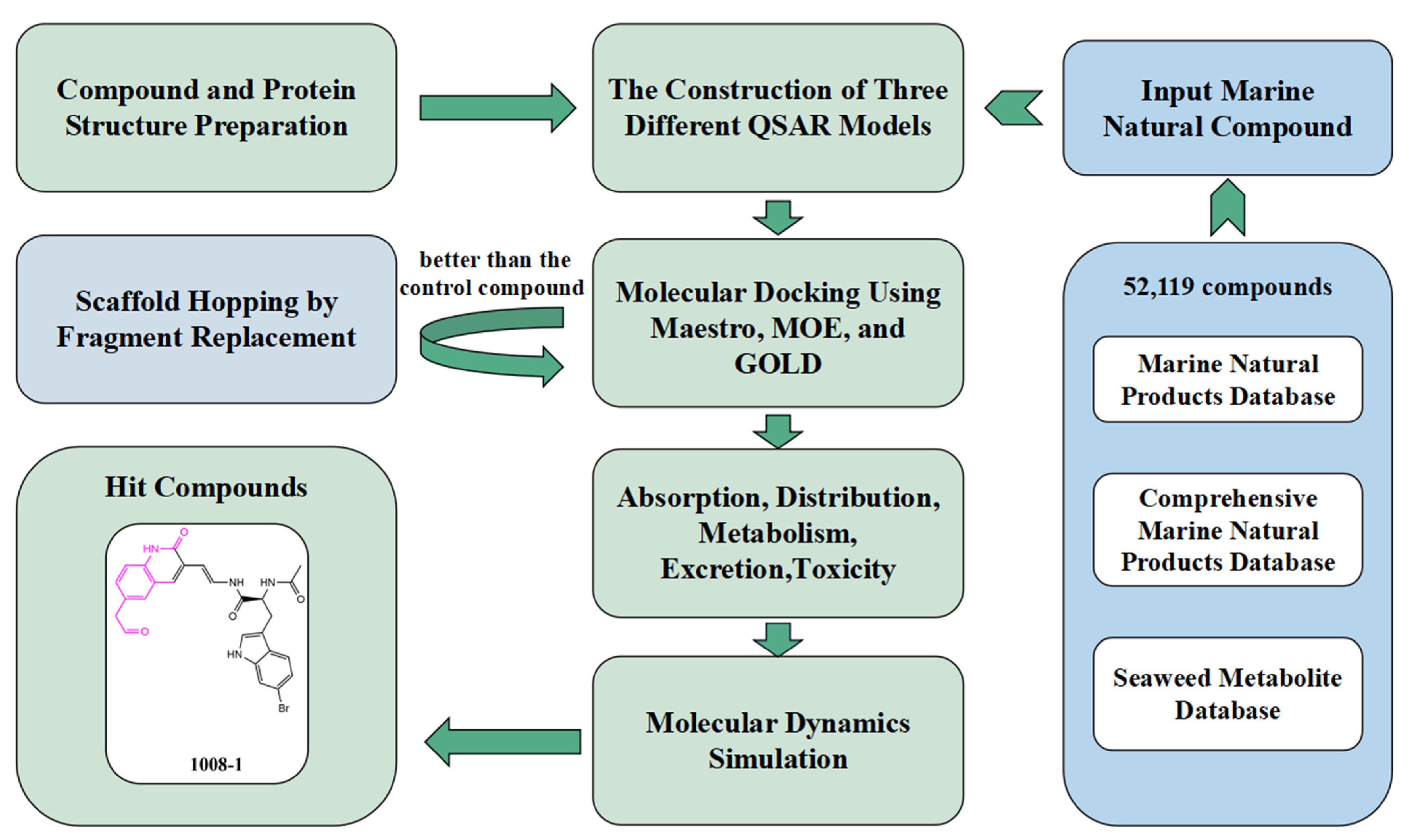
4.1. Compound Data Set Preparation
4.2. Protein Crystal Structure Preparation
4.3. The Construction of Three Different QSAR Models
4.3.1. Construction and Prediction of AutoQSAR Model
4.3.2. Construction and Prediction of Naive Bayesian Model
4.3.3. Construction and Prediction of Multiple Linear Regression Model
4.4. Structure-Based Virtual Screening
4.4.1. Molecular Docking Using Maestro
4.4.2. Molecular Docking Using MOE
4.4.3. Molecular Docking Using GOLD
4.5. Scaffold Hopping by Fragment Replacement
4.5.1. Molecular Processing
4.5.2. Optimization and Scoring
4.6. Prediction of ADMET Properties
4.7. Molecular Dynamics Simulations
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Imai, Y.; Soda, M.; Takahashi, R. Parkin Suppresses Unfolded Protein Stress-induced Cell Death through Its E3 Ubiquitin-protein Ligase Activity. J. Biol. Chem. 2000, 275, 35661–35664. [Google Scholar] [CrossRef] [PubMed]
- Karin, M.; Ben-Neriah, Y. Phosphorylation meets ubiquitination: The control of NF-κB activity. Annu. Rev. Immunol. 2000, 18, 621–663. [Google Scholar] [CrossRef] [PubMed]
- Pagano, M.; Tam, S.W.; Theodoras, A.M.; Beer-Romero, P.; Del Sal, G.; Chau, V.; Yew, P.R.; Draetta, G.F.; Rolfe, M. Role of the ubiquitin-proteasome pathway in regulating abundance of the cyclin-dependent kinase inhibitor p27. Science 1995, 269, 682–685. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Liu, H.-M. Recent advances in the development of ubiquitin-specific-processing protease 7 (USP7) inhibitors. Eur. J. Med. Chem. 2020, 191, 112107. [Google Scholar] [CrossRef] [PubMed]
- Turnbull, A.P.; Ioannidis, S.; Krajewski, W.W.; Pinto-Fernandez, A.; Heride, C.; Martin, A.C.L.; Tonkin, L.M.; Townsend, E.C.; Buker, S.M.; Lancia, D.R.; et al. Molecular basis of USP7 inhibition by selective small-molecule inhibitors. Nature 2017, 550, 481–486. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.-A.; Young, M.-J.; Jeng, W.-Y.; Liu, C.-Y.; Hung, J.-J. USP24 stabilizes bromodomain containing proteins to promote lung cancer malignancy. Sci. Rep. 2020, 10, 20870. [Google Scholar] [CrossRef] [PubMed]
- Nininahazwe, L.; Liu, B.; He, C.; Zhang, H.; Chen, Z.-S. The emerging nature of Ubiquitin-specific protease 7 (USP7): A new target in cancer therapy. Drug Discov. Today 2021, 26, 490–502. [Google Scholar] [CrossRef]
- Mussell, A.; Frangou, C.; Zhang, J. Regulation of the Hippo signaling pathway by deubiquitinating enzymes in cancer. Genes Dis. 2019, 6, 335–341. [Google Scholar] [CrossRef]
- Song, M.S.; Salmena, L.; Carracedo, A.; Egia, A.; Lo-Coco, F.; Teruya-Feldstein, J.; Pandolfi, P.P. The deubiquitinylation and localization of PTEN are regulated by a HAUSP–PML network. Nature 2008, 455, 813–817. [Google Scholar] [CrossRef]
- Wang, Q.; Ma, S.; Song, N.; Li, X.; Liu, L.; Yang, S.; Ding, X.; Shan, L.; Zhou, X.; Su, D.; et al. Stabilization of histone demethylase PHF8 by USP7 promotes breast carcinogenesis. J. Clin. Investig. 2016, 126, 2205–2220. [Google Scholar] [CrossRef]
- Georges, A.; Marcon, E.; Greenblatt, J.; Frappier, L. Identification and characterization of USP7 targets in cancer cells. Sci. Rep. 2018, 8, 15833. [Google Scholar] [CrossRef] [PubMed]
- An, T.; Gong, Y.; Li, X.; Kong, L.; Ma, P.; Gong, L.; Zhu, H.; Yu, C.; Liu, J.; Zhou, H.; et al. USP7 inhibitor P5091 inhibits Wnt signaling and colorectal tumor growth. Biochem. Pharmacol. 2017, 131, 29–39. [Google Scholar] [CrossRef] [PubMed]
- Rämet, M.; Manfruelli, P.; Pearson, A.; Mathey-Prevot, B.; Ezekowitz, R.A.B. Functional genomic analysis of phagocytosis and identification of a Drosophila receptor for E. coli. Nature 2002, 416, 644–648. [Google Scholar] [CrossRef] [PubMed]
- Kon, N.; Kobayashi, Y.; Li, M.; Brooks, C.L.; Ludwig, T.; Gu, W. Inactivation of HAUSP in vivo modulates p53 function. Oncogene 2009, 29, 1270–1279. [Google Scholar] [CrossRef] [PubMed]
- Marchenko, N.D.; Wolff, S.; Erster, S.; Becker, K.; Moll, U.M. Monoubiquitylation promotes mitochondrial p53 translocation. EMBO J. 2007, 26, 923–934. [Google Scholar] [CrossRef] [PubMed]
- Meulmeester, E.; Maurice, M.M.; Boutell, C.; Teunisse, A.F.A.S.; Ovaa, H.; Abraham, T.E.; Dirks, R.W.; Jochemsen, A.G. Loss of HAUSP-Mediated Deubiquitination Contributes to DNA Damage-Induced Destabilization of Hdmx and Hdm2. Mol. Cell 2005, 18, 565–576. [Google Scholar] [CrossRef]
- Zhou, Z.; Yao, X.; Li, S.; Xiong, Y.; Dong, X.; Zhao, Y.; Jiang, J.; Zhang, Q. Deubiquitination of Ci/Gli by Usp7/HAUSP Regulates Hedgehog Signaling. Dev. Cell 2015, 34, 58–72. [Google Scholar] [CrossRef] [PubMed]
- Tavana, O.; Li, D.; Dai, C.; Lopez, G.; Banerjee, D.; Kon, N.; Chen, C.; Califano, A.; Yamashiro, D.J.; Sun, H.; et al. HAUSP deubiquitinates and stabilizes N-Myc in neuroblastoma. Nat. Med. 2016, 22, 1180–1186. [Google Scholar] [CrossRef]
- van der Horst, A.; de Vries-Smits, A.M.M.; Brenkman, A.B.; van Triest, M.H.; van den Broek, N.; Colland, F.; Maurice, M.M.; Burgering, B.M.T. FOXO4 transcriptional activity is regulated by monoubiquitination and USP7/HAUSP. Nat. Cell Biol. 2006, 8, 1064–1073. [Google Scholar] [CrossRef]
- Wu, H.-T.; Kuo, Y.-C.; Hung, J.-J.; Huang, C.-H.; Chen, W.-Y.; Chou, T.-Y.; Chen, Y.; Chen, Y.-J.; Chen, Y.-J.; Cheng, W.-C. K63-polyubiquitinated HAUSP deubiquitinates HIF-1α and dictates H3K56 acetylation promoting hypoxia-induced tumour progression. Nat. Commun. 2016, 7, 13644. [Google Scholar] [CrossRef]
- Chauhan, D.; Tian, Z.; Nicholson, B.; Kumar, K.G.S.; Zhou, B.; Carrasco, R.; McDermott, J.L.; Leach, C.A.; Fulcinniti, M.; Kodrasov, M.P.; et al. A Small Molecule Inhibitor of Ubiquitin-Specific Protease-7 Induces Apoptosis in Multiple Myeloma Cells and Overcomes Bortezomib Resistance. Cancer Cell 2012, 22, 345–358. [Google Scholar] [CrossRef] [PubMed]
- Pozhidaeva, A.; Valles, G.; Wang, F.; Wu, J.; Sterner, D.E.; Nguyen, P.; Weinstock, J.; Kumar, K.G.S.; Kanyo, J.; Wright, D.; et al. USP7-Specific Inhibitors Target and Modify the Enzyme’s Active Site via Distinct Chemical Mechanisms. Cell Chem. Biol. 2017, 24, 1501–1512.e5. [Google Scholar] [CrossRef] [PubMed]
- Lamberto, I.; Liu, X.; Seo, H.-S.; Schauer, N.J.; Iacob, R.E.; Hu, W.; Das, D.; Mikhailova, T.; Weisberg, E.L.; Engen, J.R.; et al. Structure-Guided Development of a Potent and Selective Non-covalent Active-Site Inhibitor of USP7. Cell Chem. Biol. 2017, 24, 1490–1500.e11. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, S.; Chaugule, S.; More, S.; Rane, G.; Indap, M. Methanolic extract of Euchelus asper exhibits in-ovo anti-angiogenic and in vitro anti-proliferative activities. Biol. Res. 2017, 50, 41. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.-H.; Wang, Y.-J.; Sheng, J.; Wang, F.; Zheng, Y.; Lin, X.-K.; Sun, M. Antitumor Peptides from Marine Organisms. Mar. Drugs 2011, 9, 1840–1859. [Google Scholar] [CrossRef] [PubMed]
- Luo, L.; Wang, Q.; Liao, Y. The Inhibitors of CDK4/6 from a Library of Marine Compound Database: A Pharmacophore, ADMET, Molecular Docking and Molecular Dynamics Study. Mar. Drugs 2022, 20, 319. [Google Scholar] [CrossRef] [PubMed]
- Luo, L.; Zhong, A.; Wang, Q.; Zheng, T. Structure-based pharmacophore modeling, virtual screening, molecular docking, ADMET, and molecular dynamics (MD) simulation of potential inhibitors of PD-L1 from the library of marine natural products. Mar. Drugs 2021, 20, 29. [Google Scholar] [CrossRef]
- Singh, J.; Petter, R.C.; Baillie, T.A.; Whitty, A. The resurgence of covalent drugs. Nat. Rev. Drug Discov. 2011, 10, 307–317. [Google Scholar] [CrossRef]
- Wen, C.; Yan, X.; Gu, Q.; Du, J.; Wu, D.; Lu, Y.; Zhou, H.; Xu, J. Systematic studies on the protocol and criteria for selecting a covalent docking tool. Molecules 2019, 24, 2183. [Google Scholar] [CrossRef]
- Korenev, G.; Yakukhnov, S.; Druk, A.; Golovina, A.; Chasov, V.; Mirgayazova, R.; Ivanov, R.; Bulatov, E. USP7 inhibitors in cancer immunotherapy: Current status and perspective. Cancers 2022, 14, 5539. [Google Scholar] [CrossRef]
- Babaoglu, K.; Shoichet, B.K. Deconstructing fragment-based inhibitor discovery. Nat. Chem. Biol. 2006, 2, 720–723. [Google Scholar] [CrossRef] [PubMed]
- Pfoh, R.; Lacdao, I.K.; Saridakis, V. Deubiquitinases and the new therapeutic opportunities offered to cancer. Endocr.-Relat. Cancer 2015, 22, T35–T54. [Google Scholar] [CrossRef] [PubMed]
- Sarkari, F.; Sanchez-Alcaraz, T.; Wang, S.; Holowaty, M.N.; Sheng, Y.; Frappier, L. EBNA1-mediated recruitment of a histone H2B deubiquitylating complex to the Epstein-Barr virus latent origin of DNA replication. PLoS Pathog. 2009, 5, e1000624. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Brooks, C.L.; Kon, N.; Gu, W. A dynamic role of HAUSP in the p53-Mdm2 pathway. Mol. Cell 2004, 13, 879–886. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Kang, W.; You, Y.; Pang, J.; Ren, H.; Suo, Z.; Liu, H.; Zheng, Y. USP7: Novel drug target in cancer therapy. Front. Pharmacol. 2019, 10, 427. [Google Scholar] [CrossRef] [PubMed]
- Jing, B.; Liu, M.; Yang, L.; Cai, H.Y.; Chen, J.B.; Li, Z.X.; Kou, X.; Wu, Y.Z.; Qin, D.J.; Zhou, L.; et al. Characterization of naturally occurring pentacyclic triterpenes as novel inhibitors of deubiquitinating protease USP7 with anticancer activity in vitro. Acta Pharmacol. Sin. 2018, 39, 492–498. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Zhou, X.; Li, M.; Zhao, W.; Zhou, S.; Cheng, K.; Xu, Q.; Chen, C.; Wen, X.; Sun, H.; et al. Discovery of Ubiquitin-Specific Protease 7 (USP7) Inhibitors with Novel Scaffold Structures by Virtual Screening, Molecular Dynamics Simulation, and Biological Evaluation. J. Chem. Inf. Model 2020, 60, 3255–3264. [Google Scholar] [CrossRef] [PubMed]
- Singh, J. The Ascension of Targeted Covalent Inhibitors. J. Med. Chem. 2022, 65, 5886–5901. [Google Scholar] [CrossRef]
- Bashore, C.; Jaishankar, P.; Skelton, N.J.; Fuhrmann, J.; Hearn, B.R.; Liu, P.S.; Renslo, A.R.; Dueber, E.C. Cyanopyrrolidine Inhibitors of Ubiquitin Specific Protease 7 Mediate Desulfhydration of the Active-Site Cysteine. ACS Chem. Biol. 2020, 15, 1392–1400. [Google Scholar] [CrossRef]
- Wang, F.; Wang, L.; Wu, J.; Sokirniy, I.; Nguyen, P.; Bregnard, T.; Weinstock, J.; Mattern, M.; Bezsonova, I.; Hancock, W.W. Active site-targeted covalent irreversible inhibitors of USP7 impair the functions of Foxp3+ T-regulatory cells by promoting ubiquitination of Tip60. PLoS ONE 2017, 12, e0189744. [Google Scholar] [CrossRef]
- Dixon, S.L.; Duan, J.; Smith, E.; Von Bargen, C.D.; Sherman, W.; Repasky, M.P. AutoQSAR: An automated machine learning tool for best-practice quantitative structure–activity relationship modeling. Future Med. Chem. 2016, 8, 1825–1839. [Google Scholar] [CrossRef] [PubMed]
- Elekofehinti, O.O.; Iwaloye, O.; Olawale, F.; Chukwuemeka, P.O.; Folorunso, I.M. Newly designed compounds from scaffolds of known actives as inhibitors of survivin: Computational analysis from the perspective of fragment-based drug design. In Silico Pharmacol. 2021, 9, 47. [Google Scholar] [CrossRef] [PubMed]
- Sledz, P.; Silvestre, H.L.; Hung, A.W.; Ciulli, A.; Blundell, T.L.; Abell, C. Optimization of the interligand Overhauser effect for fragment linking: Application to inhibitor discovery against Mycobacterium tuberculosis pantothenate synthetase. J. Am. Chem. Soc. 2010, 132, 4544–4545. [Google Scholar] [CrossRef] [PubMed]
- Boudreau, P.D.; Miller, B.W.; McCall, L.-I.; Almaliti, J.; Reher, R.; Hirata, K.; Le, T.; Siqueira-Neto, J.L.; Hook, V.; Gerwick, W.H. Design of gallinamide A analogs as potent inhibitors of the cysteine proteases human cathepsin L and Trypanosoma cruzi cruzain. J. Med. Chem. 2019, 62, 9026–9044. [Google Scholar] [CrossRef] [PubMed]
- Xiong, G.; Wu, Z.; Yi, J.; Fu, L.; Yang, Z.; Hsieh, C.; Yin, M.; Zeng, X.; Wu, C.; Lu, A.; et al. ADMETlab 2.0: An integrated online platform for accurate and comprehensive predictions of ADMET properties. Nucleic Acids Res. 2021, 49, W5–W14. [Google Scholar] [CrossRef] [PubMed]
- Van Der Spoel, D.; Lindahl, E.; Hess, B.; Groenhof, G.; Mark, A.E.; Berendsen, H.J. GROMACS: Fast, flexible, and free. J. Comput. Chem. 2005, 26, 1701–1718. [Google Scholar] [CrossRef] [PubMed]
- Sousa da Silva, A.W.; Vranken, W.F. ACPYPE-Antechamber python parser interface. BMC Res. Notes 2012, 5, 367. [Google Scholar] [CrossRef]
- Mark, P.; Nilsson, L. Structure and dynamics of the TIP3P, SPC, and SPC/E water models at 298 K. J. Phys. Chem. A 2001, 105, 9954–9960. [Google Scholar] [CrossRef]
- Lindorff-Larsen, K.; Piana, S.; Palmo, K.; Maragakis, P.; Klepeis, J.L.; Dror, R.O.; Shaw, D.E. Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins Struct. Funct. Bioinform. 2010, 78, 1950–1958. [Google Scholar] [CrossRef]




| Model Code | Score | S.D. | R2 | RMSE | Q2 |
|---|---|---|---|---|---|
| kpls_radial_20 | 0.7847 | 0.5998 | 0.8077 | 0.6184 | 0.8016 |
| kpls_radial_5 | 0.7827 | 0.5797 | 0.8224 | 0.6101 | 0.8018 |
| kpls_linear_20 | 0.7776 | 0.4559 | 0.8889 | 0.5662 | 0.8337 |
| kpls_dendritic_20 | 0.7765 | 0.4491 | 0.8922 | 0.5648 | 0.8345 |
| kpls_dendritic_34 | 0.7700 | 0.5324 | 0.8501 | 0.6018 | 0.8048 |
| kpls_linear_5 | 0.7641 | 0.5264 | 0.8528 | 0.6050 | 0.8051 |
| kpls_linear_34 | 0.7613 | 0.5377 | 0.8471 | 0.6116 | 0.7984 |
| kpls_radial_13 | 0.7598 | 0.5320 | 0.8498 | 0.6110 | 0.8050 |
| kpls_molprint2D_13 | 0.7501 | 0.5263 | 0.8545 | 0.6161 | 0.8017 |
| kpls_dendritic_5 | 0.7500 | 0.5276 | 0.8522 | 0.6178 | 0.7967 |
| Name | Pharmacophore Limitation | Filter Criteria | Before Scaffold Replacement | After Scaffold Replacement | Score |
|---|---|---|---|---|---|
| 1008-1 | No | Weight < 600, SlogP [–4, 8], TPSA [40, 140], Score less than −12 | 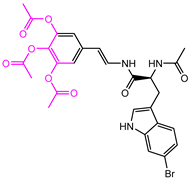 | 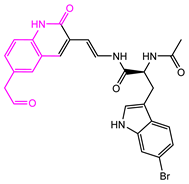 | −18.4846 |
| 24428-35 | 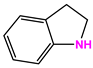 | Weight<500, SlogP [–4, 8], TPSA [40, 140] |  |  | −13.1977 |
| 13058-2 | 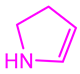 | Weight < 500, SlogP [–4, 8], TPSA [40, 140], Score less than −10 | 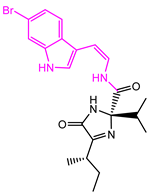 |  | −10.7885 |
| 13058-3 | 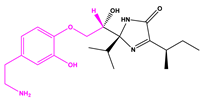 | −13.0436 | |||
| 13057-1 |  | Weight < 500, SlogP [–4, 8], TPSA [40, 140] | 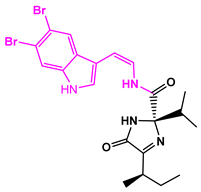 | 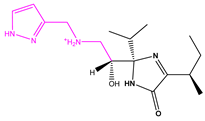 | −11.2398 |
| 13057-2 | 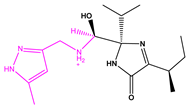 | −11.4333 | |||
| 13057-3 | 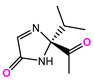 |  | 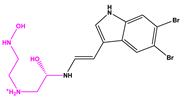 | −8.5368 | |
| 8171-3 |  | Weight < 500, SlogP [–4, 8], TPSA [40, 140] |  |  | −14.2391 |
| 8171-6 | 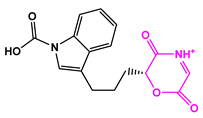 | −12.9634 | |||
| 8171-7 | 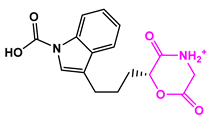 | −12.4163 |
| Compound | Hydrogen Bonds | π-π | Cation-π | Salt Bridge | Covalent Docking Score (kcal/mol) |
|---|---|---|---|---|---|
| P217564 | CYS223, GLY462 | - | - | ASP482, HIS464 | −5.583 |
| 13058 | TYR465, GLY463, GLY462, ASN218 | - | - | - | −5.866 |
| 13058-2 | CYS223, TYR465, GLY220, ASN218, ASP482 | - | - | ASP482 | −7.865 |
| 1008 | GLY220, GLY462, GLY483, SER457, HIS464 | - | - | - | −7.852 |
| 1008-1 | ASP459, ASP482, GLY220, TYR465, SER457 | - | - | - | −7.868 |
| 13057 | ASP482, GLY463 | HIS464 | HIS464 | - | −5.820 |
| 13057-3 | ASN460, ASP482, TYR465, GLY462 | - | - | ASP482 | −6.549 |
| Compound | MDCK | BBB | VD | Fu | CYP2C19-Sub | H-HT | AMES | EC | EI |
|---|---|---|---|---|---|---|---|---|---|
| 1008-1 | 8.34 × 10−6 | 0.071 | 1.813 | 29.79% | 0.053 | 0.116 | 0.442 | 0.003 | 0.006 |
| 13057-3 | 2.48 × 10−5 | 0.063 | 0.954 | 65.57% | 0.065 | 0.134 | 0.007 | 0.006 | 0.012 |
| 13058-2 | 3.48 × 10−6 | 0.355 | 1.011 | 67.65% | 0.064 | 0.062 | 0.007 | 0.003 | 0.019 |
| P217564 | 3.15 × 10−5 | 0.016 | 1.196 | 1.65% | 0.423 | 0.547 | 0.947 | 0.003 | 0.043 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tan, H.; Li, C.; Lai, T.; Luo, L. In Silico Analysis of USP7 Inhibitors Based on Building QSAR Models and Fragment Design for Screening Marine Compound Libraries. Mar. Drugs 2024, 22, 1. https://doi.org/10.3390/md22010001
Tan H, Li C, Lai T, Luo L. In Silico Analysis of USP7 Inhibitors Based on Building QSAR Models and Fragment Design for Screening Marine Compound Libraries. Marine Drugs. 2024; 22(1):1. https://doi.org/10.3390/md22010001
Chicago/Turabian StyleTan, Huiting, Chenying Li, Tianli Lai, and Lianxiang Luo. 2024. "In Silico Analysis of USP7 Inhibitors Based on Building QSAR Models and Fragment Design for Screening Marine Compound Libraries" Marine Drugs 22, no. 1: 1. https://doi.org/10.3390/md22010001





