Susceptibility of Chickens to Low Pathogenic Avian Influenza (LPAI) Viruses of Wild Bird– and Poultry–Associated Subtypes
Abstract
:1. Introduction
2. Material and Methods
2.1. Ethical Statement
2.2. Virus Selection
2.3. Virus Isolation and Propagation
2.4. Animals and Housing
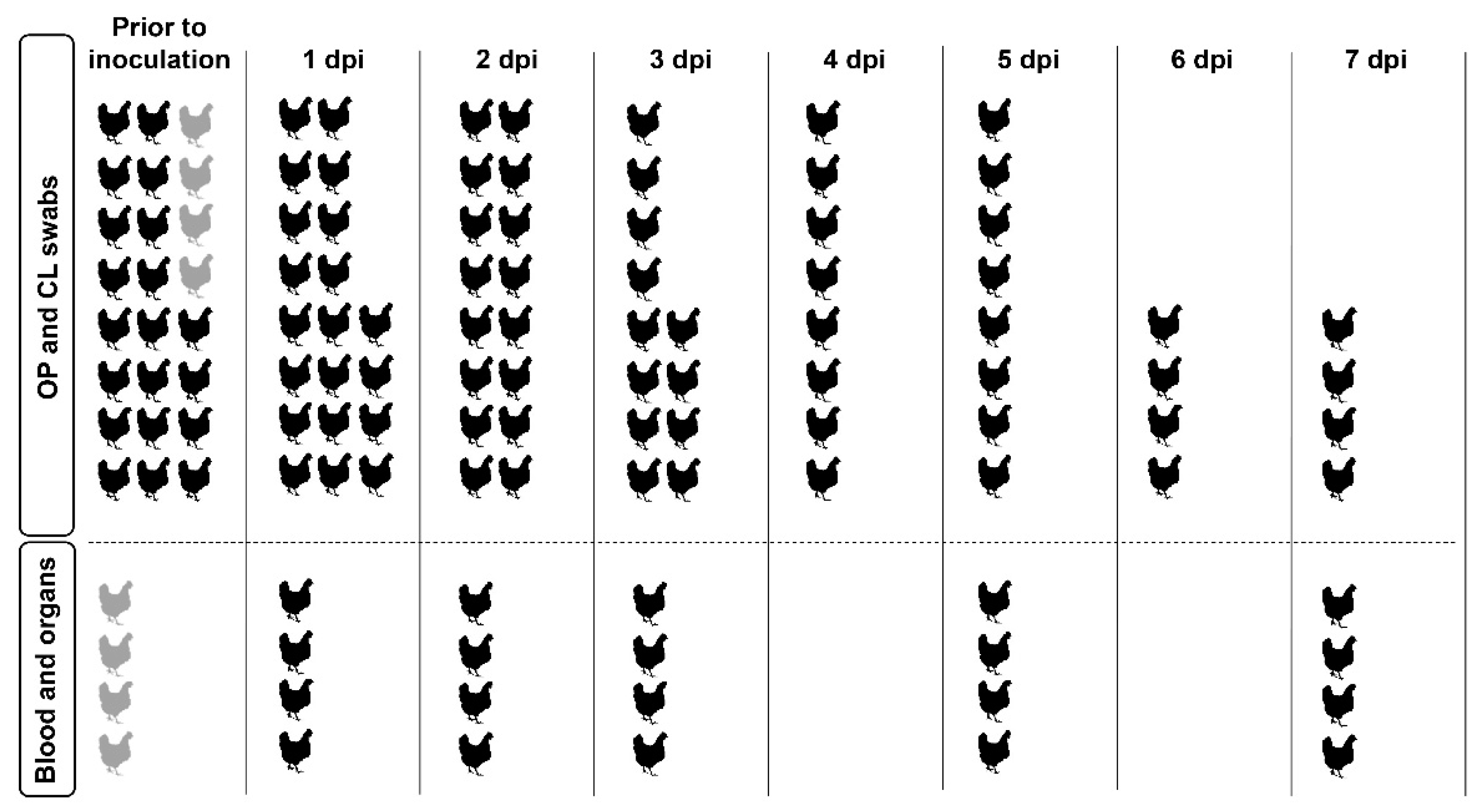
2.5. Experimental Design
2.6. Antibody Detection
2.7. Virus Detection in Swabs
2.8. Virus Detection in Tissue
2.9. Cytokine Expression in Tissue
2.10. Cluster Analysis of Internal Genes
3. Results
3.1. Identification of Host-Associations in the Internal Proteins
3.2. Clinical Signs and Seroconversion in LPAI Infected Chickens
3.3. Viral shedding of LPAI Infected Chickens
3.4. Viral Replication in the Respiratory and Intestinal Tract
3.5. Cytokine mRNA Expression in the Trachea and Ileum
3.6. Genetic Cluster Analysis of the Inoculated Viruses
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Stallknecht, D.E.; Shane, S.M. Host range of avian influenza virus in free-living birds. Vet. Res. Commun. 1988, 12, 125–141. [Google Scholar] [CrossRef] [PubMed]
- Webster, R.G.; Bean, W.J.; Gorman, O.T.; Chambers, T.M.; Kawaoka, Y. Evolution and ecology of influenza A viruses. Microbiol. Rev. 1992, 56, 152–179. [Google Scholar] [PubMed]
- Gonzales, J.L.; Elbers, A.R.W. Effective thresholds for reporting suspicions and improve early detection of avian influenza outbreaks in layer chickens. Sci. Rep. 2018, 8, 8533. [Google Scholar] [CrossRef] [PubMed]
- Webster, R.G.; Rott, R. Influenza virus A pathogenicity: The pivotal role of hemagglutinin. Cell 1987, 50, 665–666. [Google Scholar] [CrossRef]
- Capua, I.; Munoz, O. Emergence of influenza viruses with zoonotic potential: Open issues which need to be addressed. A review. Vet. Microbiol. 2013, 165, 7–12. [Google Scholar] [CrossRef]
- Olsen, B.; Munster, V.J.; Wallensten, A.; Waldenstrom, J.; Osterhaus, A.D.; Fouchier, R.A. Global patterns of influenza a virus in wild birds. Science 2006, 312, 384–388. [Google Scholar] [CrossRef]
- Fouchier, R.A.; Munster, V.; Wallensten, A.; Bestebroer, T.M.; Herfst, S.; Smith, D.; Rimmelzwaan, G.F.; Olsen, B.; Osterhaus, A.D. Characterization of a novel influenza A virus hemagglutinin subtype (H16) obtained from black-headed gulls. J. Virol. 2005, 79, 2814–2822. [Google Scholar] [CrossRef]
- Bergervoet, S.A.; Pritz-Verschuren, S.B.; Gonzales, J.L.; Bossers, A.; Poen, M.J.; Dutta, J.; Khan, Z.; Kriti, D.; van Bakel, H.; Bouwstra, R.; et al. Circulation of low pathogenic avian influenza (LPAI) viruses in wild birds and poultry in the Netherlands, 2006–2016. Sci. Rep. 2019, 9, 13681. [Google Scholar] [CrossRef]
- Treanor, J.J.; Snyder, M.H.; London, W.T.; Murphy, B.R. The B allele of the NS gene of avian influenza viruses, but not the A allele, attenuates a human influenza A virus for squirrel monkeys. Virology 1989, 171, 1–9. [Google Scholar] [CrossRef]
- Dou, D.; Revol, R.; Ostbye, H.; Wang, H.; Daniels, R. Influenza A Virus Cell Entry, Replication, Virion Assembly and Movement. Front. Immunol. 2018, 9, 1581. [Google Scholar] [CrossRef]
- Ayllon, J.; Garcia-Sastre, A. The NS1 protein: A multitasking virulence factor. Curr. Top. Microbiol. Immunol. 2015, 386, 73–107. [Google Scholar] [PubMed]
- Cauldwell, A.V.; Long, J.S.; Moncorge, O.; Barclay, W.S. Viral determinants of influenza A virus host range. J. Gen. Virol. 2014, 95, 1193–1210. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- EU. Directive 2010/63/EU of the European Parliament and of the Council of 22 September 2010 on the protection of animals used for scientific purposes. Off. J. Eur. Union 2010, 276, 33–79. [Google Scholar]
- Pickett, B.E.; Liu, M.; Sadat, E.L.; Squires, R.B.; Noronha, J.M.; He, S.; Jen, W.; Zaremba, S.; Gu, Z.; Zhou, L.; et al. Metadata-driven comparative analysis tool for sequences (meta-CATS): An automated process for identifying significant sequence variations that correlate with virus attributes. Virology 2013, 447, 45–51. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef]
- Spackman, E.; Killian, M.L. Avian influenza virus isolation, propagation, and titration in embryonated chicken eggs. Methods Mol. Biol. 2014, 1161, 125–140. [Google Scholar]
- Bergervoet, S.A.; Ho, C.K.Y.; Heutink, R.; Bossers, A.; Beerens, N. Spread of Highly Pathogenic Avian Influenza (HPAI) H5N5 Viruses in Europe in 2016–2017 Appears Related to the Timing of Reassortment Events. Viruses 2019, 11, 501. [Google Scholar] [CrossRef]
- CBS. Centraal Bureau voor de Statistiek; CBS: Den Haag, The Netherlands, 2016; Available online: https://opendata.cbs.nl/statline (accessed on 28 October 2019).
- Killian, M.L. Avian influenza virus sample types, collection, and handling. Methods Mol. Biol. 2008, 436, 7–12. [Google Scholar]
- Bouwstra, R.J.; Koch, G.; Heutink, R.; Harders, F.; Van der Spek, A.; Elbers, A.R.; Bossers, A. Phylogenetic analysis of highly pathogenic avian influenza A(H5N8) virus outbreak strains provides evidence for four separate introductions and one between-poultry farm transmission in the Netherlands, November 2014. Euro Surveill.Bull. Eur. Sur Les Mal. Transm. Eur. Commun. Dis. Bull. 2015, 20, 21174. [Google Scholar] [CrossRef]
- Cornelissen, J.B.; Post, J.; Peeters, B.; Vervelde, L.; Rebel, J.M. Differential innate responses of chickens and ducks to low-pathogenic avian influenza. Avian Pathol. J. WVPA 2012, 41, 519–529. [Google Scholar] [CrossRef]
- Kap, M.; Oomen, M.; Arshad, S.; de Jong, B.; Riegman, P. Fit for purpose frozen tissue collections by RNA integrity number-based quality control assurance at the Erasmus MC tissue bank. Biopreservation Biobanking 2014, 12, 81–90. [Google Scholar] [CrossRef] [PubMed]
- Staines, K.; Batra, A.; Mwangi, W.; Maier, H.J.; Van Borm, S.; Young, J.R.; Fife, M.; Butter, C. A Versatile Panel of Reference Gene Assays for the Measurement of Chicken mRNA by Quantitative PCR. PLoS ONE 2016, 11, e0160173. [Google Scholar] [CrossRef] [PubMed]
- Andersen, C.L.; Jensen, J.L.; Orntoft, T.F. Normalization of real-time quantitative reverse transcription-PCR data: A model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 2004, 64, 5245–5250. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Shu, Y.; McCauley, J. GISAID: Global initiative on sharing all influenza data—From vision to reality. EURO Surveill. Bull. Eur. Sur Les Mal. Transm. Eur. Commun. Dis. Bull. 2017, 22, 30494. [Google Scholar] [CrossRef]
- Katoh, K.; Misawa, K.; Kuma, K.; Miyata, T. MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Larsson, A. AliView: A fast and lightweight alignment viewer and editor for large datasets. Bioinformatics 2014, 30, 3276–3278. [Google Scholar] [CrossRef]
- Fu, L.; Niu, B.; Zhu, Z.; Wu, S.; Li, W. CD-HIT: Accelerated for clustering the next-generation sequencing data. Bioinformatics 2012, 28, 3150–3152. [Google Scholar] [CrossRef]
- Li, W.; Godzik, A. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 2006, 22, 1658–1659. [Google Scholar] [CrossRef]
- Morales, A.C., Jr.; Hilt, D.A.; Williams, S.M.; Pantin-Jackwood, M.J.; Suarez, D.L.; Spackman, E.; Stallknecht, D.E.; Jackwood, M.W. Biologic characterization of H4, H6, and H9 type low pathogenicity avian influenza viruses from wild birds in chickens and turkeys. Avian Dis. 2009, 53, 552–562. [Google Scholar] [CrossRef] [PubMed]
- Pantin-Jackwood, M.J.; Smith, D.M.; Wasilenko, J.L.; Spackman, E. Low pathogenicity avian influenza viruses infect chicken layers by different routes of inoculation. Avian Dis. 2012, 56, 276–281. [Google Scholar] [CrossRef] [PubMed]
- Spickler, A.R.; Trampel, D.W.; Roth, J.A. The onset of virus shedding and clinical signs in chickens infected with high-pathogenicity and low-pathogenicity avian influenza viruses. Avian Pathol. J. WVPA 2008, 37, 555–577. [Google Scholar] [CrossRef]
- Choi, J.G.; Kang, H.M.; Kim, M.C.; Paek, M.R.; Kim, H.R.; Kim, B.S.; Kwon, J.H.; Kim, J.H.; Lee, Y.J. Genetic relationship of H3 subtype avian influenza viruses isolated from domestic ducks and wild birds in Korea and their pathogenic potential in chickens and ducks. Vet. Microbiol. 2012, 155, 147–157. [Google Scholar] [CrossRef] [PubMed]
- Cui, H.; Shi, Y.; Ruan, T.; Li, X.; Teng, Q.; Chen, H.; Yang, J.; Liu, Q.; Li, Z. Phylogenetic analysis and pathogenicity of H3 subtype avian influenza viruses isolated from live poultry markets in China. Sci. Rep. 2016, 6, 27360. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shi, Y.; Cui, H.; Wang, J.; Chi, Q.; Li, X.; Teng, Q.; Chen, H.; Yang, J.; Liu, Q.; Li, Z. Characterizations of H4 avian influenza viruses isolated from ducks in live poultry markets and farm in Shanghai. Sci. Rep. 2016, 6, 37843. [Google Scholar] [CrossRef] [Green Version]
- FASFC. Federal Agency for the Safety of the Food Chain; FASFC: Leuven, Belgium, 2019.
- Trampel, D.W.; Zhou, E.M.; Yoon, K.J.; Koehler, K.J. Detection of antibodies in serum and egg yolk following infection of chickens with an H6N2 avian influenza virus. J. Vet. Diagn. Investig. Off. Publ. Am. Assoc. Vet. Lab. Diagn. Inc. 2006, 18, 437–442. [Google Scholar] [CrossRef]
- Wille, M.; Latorre-Margalef, N.; Tolf, C.; Halpin, R.; Wentworth, D.; Fouchier, R.A.; Raghwani, J.; Pybus, O.G.; Olsen, B.; Waldenström, J. Where do all the subtypes go? Temporal dynamics of H8-H12 influenza A viruses in waterfowl. Virus Evol. 2018, 4, vey025. [Google Scholar] [CrossRef]
- Turnbull, M.L.; Wise, H.M.; Nicol, M.Q.; Smith, N.; Dunfee, R.L.; Beard, P.M.; Jagger, B.W.; Ligertwood, Y.; Hardisty, G.R.; Xiao, H.; et al. Role of the B Allele of Influenza A Virus Segment 8 in Setting Mammalian Host Range and Pathogenicity. J. Virol. 2016, 90, 9263–9284. [Google Scholar] [CrossRef] [Green Version]
- Rajsbaum, R.; Albrecht, R.A.; Wang, M.K.; Maharaj, N.P.; Versteeg, G.A.; Nistal-Villán, E.; García-Sastre, A.; Gack, M.U. Species-specific inhibition of RIG-I ubiquitination and IFN induction by the influenza A virus NS1 protein. PLoS Pathog. 2012, 8, e1003059. [Google Scholar] [CrossRef]
- Evseev, D.; Magor, K.E. Innate Immune Responses to Avian Influenza Viruses in Ducks and Chickens. Vet. Sci. 2019, 6, 5. [Google Scholar] [CrossRef] [PubMed]
- Cauthen, A.N.; Swayne, D.E.; Sekellick, M.J.; Marcus, P.I.; Suarez, D.L. Amelioration of influenza virus pathogenesis in chickens attributed to the enhanced interferon-inducing capacity of a virus with a truncated NS1 gene. J. Virol. 2007, 81, 1838–1847. [Google Scholar] [CrossRef] [PubMed]
- Munir, M.; Zohari, S.; Metreveli, G.; Baule, C.; Belak, S.; Berg, M. Alleles A and B of non-structural protein 1 of avian influenza A viruses differentially inhibit beta interferon production in human and mink lung cells. J. Gen. Virol. 2011, 92 Pt 9, 2111–2121. [Google Scholar] [CrossRef]
- Zohari, S.; Munir, M.; Metreveli, G.; Belak, S.; Berg, M. Differences in the ability to suppress interferon beta production between allele A and allele B NS1 proteins from H10 influenza A viruses. Virol. J. 2010, 7, 376. [Google Scholar] [CrossRef] [PubMed]
- Adams, S.; Xing, Z.; Li, J.; Mendoza, K.; Perez, D.; Reed, K.; Cardona, C. The effect of avian influenza virus NS1 allele on virus replication and innate gene expression in avian cells. Mol. Immunol. 2013, 56, 358–368. [Google Scholar] [CrossRef] [PubMed]
- Suarez, D.L.; Garcia, M.; Latimer, J.; Senne, D.; Perdue, M. Phylogenetic analysis of H7 avian influenza viruses isolated from the live bird markets of the Northeast United States. J. Virol. 1999, 73, 3567–3573. [Google Scholar] [PubMed]
- Lund, J.M.; Alexopoulou, L.; Sato, A.; Karow, M.; Adams, N.C.; Gale, N.W.; Iwasaki, A.; Flavell, R.A. Recognition of single-stranded RNA viruses by Toll-like receptor 7. Proc. Natl. Acad. Sci. USA 2004, 101, 5598–5603. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nang, N.T.; Lee, J.S.; Song, B.M.; Kang, Y.M.; Kim, H.S.; Seo, S.H. Induction of inflammatory cytokines and Toll-like receptors in chickens infected with avian H9N2 influenza virus. Vet. Res. 2011, 42, 64. [Google Scholar] [CrossRef]
- Bergervoet, S.A. Analysis of Cytokine Response in Virus-Positive Samples of Chickens Inoculated with Low Pathogenic Avian Influenza (LPAI) Viruses of Wild Bird– and Poultry-Associated Subtypes; Department of Virology, Wageningen Bioveterinary Research: Lelystad, The Netherlands, 2019. [Google Scholar]
- Lloren, K.K.S.; Lee, T.; Kwon, J.J.; Song, M.S. Molecular Markers for Interspecies Transmission of Avian Influenza Viruses in Mammalian Hosts. Int. J. Mol. Sci. 2017, 18, 2706. [Google Scholar] [CrossRef]
- Manz, B.; Schwemmle, M.; Brunotte, L. Adaptation of avian influenza A virus polymerase in mammals to overcome the host species barrier. J. Virol. 2013, 87, 7200–7209. [Google Scholar] [CrossRef]
- Obenauer, J.C.; Denson, J.; Mehta, P.K.; Su, X.; Mukatira, S.; Finkelstein, D.B.; Xu, X.; Wang, J.; Ma, J.; Fan, Y.; et al. Large-scale sequence analysis of avian influenza isolates. Science 2006, 311, 1576–1580. [Google Scholar] [CrossRef] [PubMed]



| Virus | Trachea | Lung | Ileum | ||||
|---|---|---|---|---|---|---|---|
| 1 dpi | 3 dpi | 1 dpi | 5 dpi | 1 dpi | 5 dpi | 7 dpi | |
| H3N8 NS allele A | 3/4 (32.6 ± 1.8) | 1/4 (34.3) | 4/4 (31.6 ± 3.4) | 0/4 | 0/4 | 0/4 | 1/4 (26.3) |
| H3N8 NS allele B | 1/4 (29.5) | 0/4 | 4/4 (34.6 ± 2.8) | 0/4 | 0/4 | 1/4 (23.2) | 2/4 (23.4 ± 0.2) |
| H4N6 NS allele A | 3/4 (27.5 ± 0.8) | 0/4 | 3/4 (26.9 ± 2.9) | 0/4 | 0/4 | 0/4 | 0/4 |
| H4N6 NS allele B | 1/4 (36.6) | 0/4 | 3/4 (32.5 ± 8.3) | 0/4 | 0/4 | 0/4 | 0/4 |
| H8N4 NS allele A | 2/4 (32.3 ± 0.2) | 0/4 | 4/4 (30.2 ± 0.5) | 0/4 | 0/4 | 0/4 | 4/4 (34.2 ± 4.9) |
| H8N4 NS allele B | 0/4 | 0/4 | 1/4 (35.7) | 1/4 (30.0) | 0/4 | 0/4 | 0/4 |
| H9N2 NS allele A | 2/4 (30.9 ± 0.9) | 0/4 | 0/4 | 0/4 | 0/4 | 0/4 | 0/4 |
| H9N2 NS allele B | 2/4 (30.1 ± 2.3) | 0/4 | 2/4 (33.5 ± 2.0) | 0/4 | 0/4 | 0/4 | 0/4 |
| Virus | PB1 | PB2 | PA | NP | MP | NS |
|---|---|---|---|---|---|---|
| H3N8 NS allele A | 3/15 | 0/91 | 0/23 | 2/37 | 0/49 | 0/67 |
| H3N8 NS allele B | 0/4 | 2/6 | 0/7 | 0/32 | 7/101 | 0/103 |
| H4N6 NS allele A | 4/17 | 0/59 | 1/5 | 1/27 | 6/304 | 13/557 |
| H4N6 NS allele B | 6/17 | 0/97 | 0/3 | 0/95 | 34/568 | 0/108 |
| H8N4 NS allele A | 0/160 | 2/33 | 0/21 | 0/92 | 38/698 | 33/886 |
| H8N4 NS allele B | 0/61 | 0/20 | 0/20 | 4/56 | 93/1034 | 0/79 |
| H9N2 NS allele A | 0/59 | 0/64 | 0/58 | 0/108 | 14/510 | 2/86 |
| H9N2 NS allele B | 0/15 | 0/16 | 0/56 | 0/1 | 0/149 | 8/35 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bergervoet, S.A.; Germeraad, E.A.; Alders, M.; Roose, M.M.; Engelsma, M.Y.; Heutink, R.; Bouwstra, R.; Fouchier, R.A.M.; Beerens, N. Susceptibility of Chickens to Low Pathogenic Avian Influenza (LPAI) Viruses of Wild Bird– and Poultry–Associated Subtypes. Viruses 2019, 11, 1010. https://doi.org/10.3390/v11111010
Bergervoet SA, Germeraad EA, Alders M, Roose MM, Engelsma MY, Heutink R, Bouwstra R, Fouchier RAM, Beerens N. Susceptibility of Chickens to Low Pathogenic Avian Influenza (LPAI) Viruses of Wild Bird– and Poultry–Associated Subtypes. Viruses. 2019; 11(11):1010. https://doi.org/10.3390/v11111010
Chicago/Turabian StyleBergervoet, Saskia A., Evelien A. Germeraad, Marc Alders, Marit M. Roose, Marc Y. Engelsma, Rene Heutink, Ruth Bouwstra, Ron A.M. Fouchier, and Nancy Beerens. 2019. "Susceptibility of Chickens to Low Pathogenic Avian Influenza (LPAI) Viruses of Wild Bird– and Poultry–Associated Subtypes" Viruses 11, no. 11: 1010. https://doi.org/10.3390/v11111010





