HIV-1 Sub-Subtype A6: Settings for Normalised Identification and Molecular Epidemiology in the Southern Federal District, Russia
Abstract
:1. Introduction
2. Patients and Methods
Study Design and Participants
3. Results
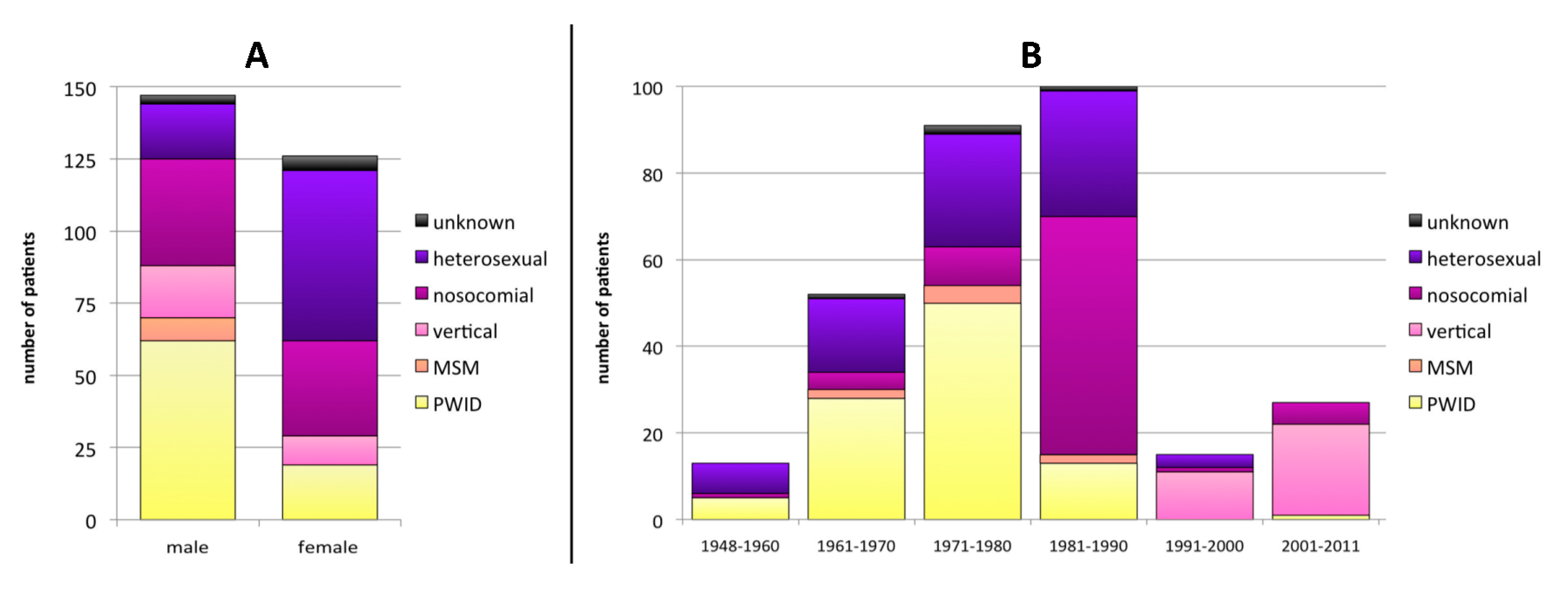
3.1. Baseline Characteristics
3.2. Viral Subtype Distribution
3.3. Resistance-Associated-Mutations (RAMs) and Drug-Susceptibility Profiles
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- UNAIDS DATA 2019. Available online: https://www.unaids.org/sites/default/files/media_asset/2019-UNAIDS-data_en.pdf (accessed on 2 February 2019).
- Beyrer, C.; Wirtz, A.L.; O’Hara, G.; Leon, N.; Kazatchkine, M. The expanding epidemic of HIV-1 in the Russian Federation. PLoS Med. 2017, 14, e1002462. [Google Scholar] [CrossRef] [Green Version]
- Bobkova, M. Current status of HIV-1 diversity and drug resistance monitoring in the former USSR. AIDS Rev. 2013, 15, 204–212. [Google Scholar] [PubMed]
- GBD 2017 HIV collaborators. Global, regional, and national incidence, prevalence, and mortality of HIV, 1980–2017, and forecasts to 2030, for 195 countries and territories: A systematic analysis for the Global Burden of Diseases, Injuries, and Risk Factors Study 2017. Lancet HIV 2019, 6, e831–e859. [Google Scholar] [CrossRef] [Green Version]
- Diez-Fuertes, F.; Cabello, M.; Thomson, M.M. Bayesian phylogeographic analyses clarify the origin of the HIV-1 subtype A variant circulating in former Soviet Union’s countries. Infect Genet. Evol. 2015, 33, 197–205. [Google Scholar] [CrossRef] [PubMed]
- Novitsky, V.A.; Montano, M.A.; Essex, M. Molecular epidemiology of an HIV-1 subtype A subcluster among injection drug users in the Southern Ukraine. AIDS Res. Hum. Retrovir. 1998, 14, 1079–1085. [Google Scholar] [CrossRef]
- Thomson, M.M.; de Parga, E.V.; Vinogradova, A.; Sierra, M.; Yakovlev, A.; Rakhmanova, A.; Delgado, E.; Casado, G.; Munoz, M.; Carmona, R.; et al. New insights into the origin of the HIV type 1 subtype A epidemic in former Soviet Union’s countries derived from sequence analyses of preepidemically transmitted viruses. AIDS Res. Hum. Retrovir. 2007, 23, 1599–1604. [Google Scholar] [CrossRef]
- Bobkov, A.; Cheingsong-Popov, R.; Selimova, L.; Ladnaya, N.; Kazennova, E.; Kravchenko, A.; Fedotov, E.; Saukhat, S.; Zverev, S.; Pokrovsky, V.; et al. An HIV type 1 epidemic among injecting drug users in the former Soviet Union caused by a homogeneous subtype a strain. AIDS Res. Hum. Retrovir. 1997, 13, 1195–1201. [Google Scholar] [CrossRef]
- Rumyantseva, O.A.; Olkhovskiy, I.A.; Malysheva, M.A.; Ruzaeva, L.A.; Vasiliev, A.V.; Kazennova, E.V.; Bobkova, M.R.; Lukashov, V.V. Epidemiological networks and drug resistance of HIV type 1 in Krasnoyarsk region, Russia. AIDS Res. Hum. Retrovir. 2009, 25, 931–936. [Google Scholar] [CrossRef]
- Karkashadze, E.; Dvali, N.; Bolokadze, N.; Sharvadze, L.; Gabunia, P.; Karchava, M.; Tchelidze, T.; Tsertsvadze, T.; DeHovitz, J.; Del Rio, C.; et al. Epidemiology of human immunodeficiency virus (HIV) drug resistance in HIV patients with virologic failure of first-line therapy in the country of Georgia. J. Med. Virol. 2019, 91, 235–240. [Google Scholar] [CrossRef]
- Foley, B.T.; Leitner, T.; Paraskevis, D.; Peeters, M. Primate immunodeficiency virus classification and nomenclature: Review. Infect Genet. Evol. 2016, 46, 150–158. [Google Scholar] [CrossRef] [Green Version]
- Masharsky, A.E.; Klimov, N.A.; Kozlov, A.P. Molecular cloning and analysis of full-length genome of HIV type 1 strains prevalent in countries of the former Soviet Union. AIDS Res. Hum. Retrovir. 2003, 19, 933–939. [Google Scholar] [CrossRef] [PubMed]
- Lapovok, I.; Laga, V.; Kazennova, E.; Bobkova, M. HIV Type 1 Integrase Natural Polymorphisms in Viral Variants Circulating in FSU Countries. Curr. HIV Res. 2017, 15, 318–326. [Google Scholar] [CrossRef] [PubMed]
- Riva, C.; Romano, L.; Saladini, F.; Lai, A.; Carr, J.K.; Francisci, D.; Balotta, C.; Zazzi, M. Identification of a possible ancestor of the subtype A1 HIV Type 1 variant circulating in the former Soviet Union. AIDS Res. Hum. Retrovir. 2008, 24, 1319–1325. [Google Scholar] [CrossRef] [PubMed]
- Lapovok, I.A.; Lopatukhin, A.E.; Kireev, D.E.; Kazennova, E.V.; Lebedev, A.V.; Bobkova, M.R.; Kolomeets, A.N.; Turbina, G.I.; Shipulin, G.A.; Ladnaya, N.N.; et al. Molecular epidemiological analysis of HIV-1 variants circulating in Russia in 1987-2015. Ter. Arkh. 2017, 89, 44–49. [Google Scholar] [CrossRef]
- Pokrovskii, V.V.; Eramova, I.; Deulina, M.O.; Lipetikov, V.V.; Iashkulov, K.B.; Sliusareva, L.A.; Chemizova, N.M.; Savchenko, S.P. An intrahospital outbreak of HIV infection in Elista. Zhurnal Mikrobiol. Epidemiol. Immunobiol. 1990, 17–23. [Google Scholar]
- Bobkov, A.; Cheingsong-Popov, R.; Garaev, M.; Rzhaninova, A.; Kaleebu, P.; Beddows, S.; Bachmann, M.H.; Mullins, J.I.; Louwagie, J.; Janssens, W.; et al. Identification of an env G subtype and heterogeneity of HIV-1 strains in the Russian Federation and Belarus. AIDS 1994, 8, 1649–1655. [Google Scholar] [CrossRef]
- Bobkov, A.; Garaev, M.M.; Rzhaninova, A.; Kaleebu, P.; Pitman, R.; Weber, J.N.; Cheingsong-Popov, R. Molecular epidemiology of HIV-1 in the former Soviet Union: Analysis of env V3 sequences and their correlation with epidemiologic data. AIDS 1994, 8, 619–624. [Google Scholar] [CrossRef]
- Murzakova, A.; Kireev, D.; Baryshev, P.; Lopatukhin, A.; Serova, E.; Shemshura, A.; Saukhat, S.; Kolpakov, D.; Matuzkova, A.; Suladze, A.; et al. Molecular Epidemiology of HIV-1 Subtype G in the Russian Federation. Viruses 2019, 11, 348. [Google Scholar] [CrossRef] [Green Version]
- Sierra, S.; Kaiser, R.; Lubke, N.; Thielen, A.; Schuelter, E.; Heger, E.; Daumer, M.; Reuter, S.; Esser, S.; Fatkenheuer, G.; et al. Prediction of HIV-1 coreceptor usage (tropism) by sequence analysis using a genotypic approach. J. Vis. Exp. 2011, 58, e3264. [Google Scholar] [CrossRef]
- Leitner, T.; Korber, B.; Daniels, M.; Calef, C.; Foley, B. HIV-1 Subtype and Circulating Recombinant Form (CRF) Reference Sequences. HIV Seq. Compend. 2005, 2005, 41–48. [Google Scholar]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, T.F.; Shafer, R.W. Web Resources for HIV type 1 Genotypic-Resistance Test Interpretation. Clin. Infect. Dis. 2006, 42, 1608–1618. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aibekova, L.; Foley, B.; Hortelano, G.; Raees, M.; Abdraimov, S.; Toichuev, R.; Ali, S. Molecular epidemiology of HIV-1 subtype A in former Soviet Union countries. PLoS ONE 2018, 13, e0191891. [Google Scholar] [CrossRef] [PubMed]
- Kazennova, E.; Laga, V.; Gromov, K.; Lebedeva, N.; Zhukova, E.; Pronin, A.; Grezina, L.; Dement’eva, N.; Shemshura, A.; Bobkova, M. Genetic Variants of HIV Type 1 in Men Who Have Sex with Men in Russia. AIDS Res. Hum. Retrovir. 2017, 33, 1061–1064. [Google Scholar] [CrossRef] [PubMed]
- Foley, B.; Leitner, T.; Apetrei, C.; Hahn, B.; Mizrachi, I.; Mullins, J.; Rambaut, A.; Wolinsky, S.; Korber, B. HIV Sequence Compendium 2018; Theoretical Biology and Biophysics Group T-6, Los Alamos National Laboratory: New Mexico, NM, USA, 2018. [Google Scholar]
- Miri, L.; Wakrim, L.; Kassar, H.; Hemminki, K.; Khyatti, M. Impact of immigration on HIV-1 molecular epidemiology in West Africa, Maghreb and Southern Europe. AIDS Rev. 2014, 16, 109–116. [Google Scholar]
- Perez-Parra, S.; Chueca, N.; Alvarez, M.; Pasquau, J.; Omar, M.; Collado, A.; Vinuesa, D.; Lozano, A.B.; Yebra, G.; Garcia, F. High prevalence and diversity of HIV-1 non-B genetic forms due to immigration in southern Spain: A phylogeographic approach. PLoS ONE 2017, 12, e0186928. [Google Scholar] [CrossRef]
- Pineda-Peña, A.C.; Pingarilho, M.; Li, G.; Vrancken, B.; Libin, P.; Gomes, P.; Camacho, R.J.; Theys, K.; Barroso Abecasis, A.; Portuguese, H.I.V.R.S.G. Drivers of HIV-1 transmission: The Portuguese case. PLoS ONE 2019, 14, e0218226. [Google Scholar] [CrossRef] [Green Version]
- Heipertz, R.A., Jr.; Ayemoba, O.; Sanders-Buell, E.; Poltavee, K.; Pham, P.; Kijak, G.H.; Lei, E.; Bose, M.; Howell, S.; O’Sullivan, A.M.; et al. Significant contribution of subtype G to HIV-1 genetic complexity in Nigeria identified by a newly developed subtyping assay specific for subtype G and CRF02_AG. Medicine (Baltimore) 2016, 95, e4346. [Google Scholar] [CrossRef]
- Gonzalez-Alba, J.M.; Holguin, A.; Garcia, R.; Garcia-Bujalance, S.; Alonso, R.; Suarez, A.; Delgado, R.; Cardenoso, L.; Gonzalez, R.; Garcia-Bermejo, I.; et al. Molecular surveillance of HIV-1 in Madrid, Spain: A phylogeographic analysis. J. Virol. 2011, 85, 10755–10763. [Google Scholar] [CrossRef] [Green Version]
- Karamov, E.; Epremyan, K.; Siniavin, A.; Zhernov, Y.; Cuevas, M.T.; Delgado, E.; Sanchez-Martinez, M.; Carrera, C.; Kornilaeva, G.; Turgiev, A.; et al. HIV-1 Genetic Diversity in Recently Diagnosed Infections in Moscow: Predominance of AFSU, Frequent Branching in Clusters, and Circulation of the Iberian Subtype G Variant. AIDS Res. Hum. Retrovir. 2018, 34, 629–634. [Google Scholar] [CrossRef]
- Sarabia, I.; Bosque, A. HIV-1 Latency and Latency Reversal: Does Subtype Matter? Viruses 2019, 11, 1104. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Daumer, M.; Awerkiew, S.; Aragon, S.S.; Kartashev, V.; Poplavskaja, T.; Klein, R.; Sichtig, N.; Thiele, B.; Lengauer, T.; Roomp, K.; et al. Short communication: Selection of thymidine analogue resistance mutational patterns in children infected from a common HIV type 1 subtype G source. AIDS Res. Hum. Retrovir. 2010, 26, 275–278. [Google Scholar] [CrossRef] [PubMed]
- Wainberg, M.A.; Brenner, B.G. Role of HIV Subtype Diversity in the Development of Resistance to Antiviral Drugs. Viruses 2010, 2, 2493–2508. [Google Scholar] [CrossRef] [Green Version]
- Maldonado, J.O.; Mansky, L.M. The HIV-1 Reverse Transcriptase A62V Mutation Influences Replication Fidelity and Viral Fitness in the Context of Multi-Drug-Resistant Mutations. Viruses 2018, 10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dvali, N.; Parker, M.M.; Chkhartishvili, N.; Sharvadze, L.; Gochitashvili, N.; Abutidze, A.; Karchava, M.; DeHovitz, J.A.; Tsertsvadze, T. Characterization of HIV-1 subtypes and drug resistance mutations among individuals infected with HIV in Georgia. J. Med. Virol. 2012, 84, 1002–1008. [Google Scholar] [CrossRef]
- Kolomeets, A.N.; Varghese, V.; Lemey, P.; Bobkova, M.R.; Shafer, R.W. A uniquely prevalent nonnucleoside reverse transcriptase inhibitor resistance mutation in Russian subtype A HIV-1 viruses. AIDS 2014, 28, F1–F8. [Google Scholar] [CrossRef] [Green Version]
- Bacheler, L.T.; Anton, E.D.; Kudish, P.; Baker, D.; Bunville, J.; Krakowski, K.; Bolling, L.; Aujay, M.; Wang, X.V.; Ellis, D.; et al. Human immunodeficiency virus type 1 mutations selected in patients failing efavirenz combination therapy. Antimicrob. Agents Chemother. 2000, 44, 2475–2484. [Google Scholar] [CrossRef] [Green Version]
- Huang, W.; Gamarnik, A.; Limoli, K.; Petropoulos, C.J.; Whitcomb, J.M. Amino acid substitutions at position 190 of human immunodeficiency virus type 1 reverse transcriptase increase susceptibility to delavirdine and impair virus replication. J. Virol. 2003, 77, 1512–1523. [Google Scholar] [CrossRef] [Green Version]
- Brenner, B.G. Selective acquisition of G190S in HIV-1 subtype A from Russia leading to efavirenz and nevirapine treatment failure. AIDS 2014, 28, 2619–2621. [Google Scholar] [CrossRef]
- Avidor, B.; Turner, D.; Mor, Z.; Chalom, S.; Riesenberg, K.; Shahar, E.; Pollack, S.; Elbirt, D.; Sthoeger, Z.; Maayan, S.; et al. Transmission patterns of HIV-subtypes A/AE versus B: Inferring risk-behavior trends and treatment-efficacy limitations from viral genotypic data obtained prior to and during antiretroviral therapy. PLoS ONE 2013, 8, e57789. [Google Scholar] [CrossRef] [PubMed]
- Dvali, N.; Chkhartishvili, N.; Karchava, M.; Sharvadze, L.; Tsertsvadze, T. Distinct drug resistance profile of HIV-1 subtype A strain circulating in Georgia. Georgian Med. News 2015, 19–24. [Google Scholar]
- Davy-Mendez, T.; Eron, J.J.; Brunet, L.; Zakharova, O.; Dennis, A.M.; Napravnik, S. New antiretroviral agent use affects prevalence of HIV drug resistance in clinical care populations. AIDS 2018, 32, 2593–2603. [Google Scholar] [CrossRef] [PubMed]
- Schultze, A.; Phillips, A.N.; Paredes, R.; Battegay, M.; Rockstroh, J.K.; Machala, L.; Tomazic, J.; Girard, P.M.; Januskevica, I.; Gronborg-Laut, K.; et al. HIV resistance testing and detected drug resistance in Europe. AIDS 2015, 29, 1379–1389. [Google Scholar] [CrossRef]
- Knops, E.; Schuülter, E.; Luübke, N.; Oette, M.; Faätkenheuer, G.; Hower, M.; Knechten, H.; Mutz, A.; Esser, S.; Scholten, S.; et al. The RESINA data support the individualized therapy based on primary resistance testing. Eur. Resist. Abstr. 67 2015, 2015, 66–67. [Google Scholar]
- Franzetti, M.; De Luca, A.; Ceccherini-Silberstein, F.; Spagnuolo, V.; Nicastri, E.; Mussini, C.; Antinori, A.; Monno, L.; Vecchiet, J.; Fanti, I.; et al. Evolution of HIV-1 transmitted drug resistance in Italy in the 2007-2014 period: A weighted analysis. J. Clin. Virol. 2018, 106, 49–52. [Google Scholar] [CrossRef]
- Hauser, A.; Hofmann, A.; Hanke, K.; Bremer, V.; Bartmeyer, B.; Kuecherer, C.; Bannert, N. National molecular surveillance of recently acquired HIV infections in Germany, 2013 to 2014. Euro Surveill 2017, 22, 30436. [Google Scholar] [CrossRef]
- Rossetti, B.; Di Giambenedetto, S.; Torti, C.; Postorino, M.C.; Punzi, G.; Saladini, F.; Gennari, W.; Borghi, V.; Monno, L.; Pignataro, A.R.; et al. Evolution of transmitted HIV-1 drug resistance and viral subtypes circulation in Italy from 2006 to 2016. HIV Med. 2018, 19, 619–628. [Google Scholar] [CrossRef]


| RAMs | Patients | ||||
|---|---|---|---|---|---|
| Total 1 | TN 2 | TE 3 | No Data 4 | p Value 5 | |
| PI | 261 | 102 | 110 | 49 | |
| D30N | 1 (0.4%) | 0 (0.0%) | 1 (0.9%) | 0 (0.0%) | ≥0.05 |
| M46IL | 13 (5.0%) | 2 (2.0%) | 9 (8.2%) | 2 (4.1%) | ≥0.05 |
| G48V | 2 (0.8%) | 2 (2.0%) | 0 (0.0%) | 0 (0.0%) | ≥0.05 |
| I50VL | 4 (1.5%) | 0 (0.0%) | 3 (2.7%) | 1 (2.0%) | ≥0.05 |
| I54AV | 8 (3.1%) | 1 (1.0%) | 6 (5.5%) | 1 (2.0%) | ≥0.05 |
| L76V | 2 (0.8%) | 1 (1.0%) | 1 (0.9%) | 0 (0.0%) | ≥0.05 |
| V82AFMST | 10 (3.8%) | 1 (1.0%) | 8 (7.3%) | 1 (2.0%) | 0,0363 |
| N88S | 1 (0.4%) | 0 (0.0%) | 0 (0.0%) | 1 (2.0%) | ≥0.05 |
| L90M | 10 (3.8%) | 1 (1.0%) | 8 (7.3%) | 1 (2.0%) | 0,0363 |
| NRTIs | 277 | 109 | 120 | 48 | |
| M41L | 22 (7.9%) | 1 (0.9%) | 19 (15.8%) | 2 (4.2%) | <0.0001 |
| E44D | 1 (0.4%) | 0 (0.0%) | 1 (0.8%) | 0 (0.0%) | ≥0.05 |
| A62V | 61 (22.0%) | 30 (27.5%) | 22 (18.3%) | 9 (18.8%) | ≥0.05 |
| D67G | 3 (1.1%) | 1 (0.9%) | 2 (1.7%) | 0 (0.0%) | ≥0.05 |
| D67N | 17 (6.1%) | 1 (0.9%) | 13 (10.8%) | 3 (6.3%) | 0.0016 |
| K65DEN | 4 (1.4%) | 1 (0.9%) | 3 (2.5%) | 0 (0.0%) | ≥0.05 |
| T69DN | 3 (1.1%) | 0 (0.0%) | 3 (2.5%) | 0 (0.0%) | ≥0.05 |
| T69G_SG | 1 (0.4%) | 0 (0.0%) | 1 (0.8%) | 0 (0.0%) | ≥0.05 |
| K70ER | 14 (5.1%) | 1 (0.9%) | 11 (9.2%) | 2 (4.2%) | 0.0058 |
| L74IV | 9 (3.2%) | 0 (0.0%) | 6 (5.0%) | 3 (6.3%) | ≥0.05 |
| V75AIM | 5 (1.8%) | 0 (0.0%) | 5 (4.2%) | 0 (0.0%) | ≥0.05 |
| Y115F | 2 (0.7%) | 0 (0.0%) | 1 (0.8%) | 1 (2.1%) | ≥0.05 |
| M184IV | 60 (21.7%) | 5 (4.6%) | 44 (36.7%) | 11 (22.9%) | <0.0001 |
| L210W | 8 (2.9%) | 0 (0.0%) | 8 (6.7%) | 0 (0.0%) | 0.0073 |
| T215CFILNSY | 38 (13.7%) | 1 (0.9%) | 33 (27.5%) | 4 (8.3%) | <0.0001 |
| K219ENQ | 16 (5.8%) | 1 (0.9%) | 12 (10.0%) | 3 (6.3%) | 0.0030 |
| NNRTI | 277 | 109 | 120 | 48 | |
| A98G | 2 (0.7%) | 0 (0.0%) | 2 (1.7%) | 0 (0.0%) | ≥0.05 |
| K101EHPQ | 11 (4.0%) | 0 (0.0%) | 9 (7.5%) | 2 (4.2%) | 0.0036 |
| L100F | 2 (0.7%) | 0 (0.0%) | 2 (1.7%) | 0 (0.0%) | ≥0.05 |
| K103NS | 28 (10.1%) | 0 (0.0%) | 24 (20.0%) | 4 (8.3%) | <0.0001 |
| V106A | 1 (0.4%) | 0 (0.0%) | 1 (0.8%) | 0 (0.0%) | ≥0.05 |
| V108I | 6 (2.2%) | 0 (0.0%) | 3 (2.5%) | 3 (6.3%) | ≥0.05 |
| E138AGHKQR | 20 (7.2%) | 5 (4.6%) | 13 (10.8%) | 2 (4.2%) | ≥0.05 |
| V179DE | 6 (2.2%) | 2 (1.8%) | 4 (3.3%) | 0 (0.0%) | ≥0.05 |
| Y181CFIV | 13 (4.7%) | 0 (0.0%) | 11 (9.2%) | 2 (4.2%) | 0.0009 |
| Y188LS | 4 (1.4%) | 1 (0.9%) | 3 (2.5%) | 0 (0.0%) | ≥0.05 |
| G190AS | 21 (7.6%) | 0 (0.0%) | 18 (15.0%) | 3 (6.3%) | <0.0001 |
| H221Y | 2 (0.7%) | 0 (0.0%) | 2 (1.7%) | 0 (0.0%) | ≥0.05 |
| P225H | 4 (1.4%) | 0 (0.0%) | 3 (2.5%) | 1 (2.1%) | ≥0.05 |
| F227L | 1 (0.4%) | 0 (0.0%) | 1 (0.8%) | 0 (0.0%) | ≥0.05 |
| M230L | 1 (0.4%) | 0 (0.0%) | 1 (0.8%) | 0 (0.0%) | ≥0.05 |
| K238T | 1 (0.4%) | 0 (0.0%) | 0 (0.0%) | 1 (2.1%) | ≥0.05 |
| Y318F | 1 (0.4%) | 0 (0.0%) | 1 (0.8%) | 0 (0.0%) | ≥0.05 |
| INSTI | 61 | 19 | 21 | 21 | |
| T6TK | 1 (1.6%) | 0 (0.0%) | 1 (4.8%) | 0 (0.0%) | ≥0.05 |
| G140A | 1 (1.6%) | 0 (0.0%) | 1 (4.8%) | 0 (0.0%) | ≥0.05 |
| Q148R | 1 (1.6%) | 0 (0.0%) | 1 (4.8%) | 0 (0.0%) | ≥0.05 |
| N155H | 1 (1.6%) | 0 (0.0%) | 1 (4.8%) | 0 (0.0%) | ≥0.05 |
| Patients | |||||
|---|---|---|---|---|---|
| Total 1 | TN 2 | TE 3 | No Data 4 | ||
| Available PR Sequences | 261 | 102 | 110 | 49 | |
| PI | IR | 11 (4.2%) | 6 (5.9%) | 5 (4.5%) | 0 (0.0%) |
| FR | 21 (8.0%) | 3 (2.9%) | 14 (12.7%) | 4 (8.2%) | |
| CR-resistance | 32 (12.3%) | 9 (8.8%) | 19 (17.3%) | 4 (8.2%) | |
| Available RT Sequences | 277 | 109 | 120 | 48 | |
| NRTI | IR | 9 (3.2%) | 2 (1.8%) | 6 (5.0%) | 1 (2.1%) |
| FR | 72 (26.0%) | 5 (4.6%) | 54 (45.0%) | 13 (27.1%) | |
| CR-resistance | 81 (29.2%) | 7 (6.4%) | 60 (50.0%) | 14 (29.2%) | |
| NNRTI | IR | 2 (0.7%) | 0 (0.0%) | 1 (0.8%) | 1 (2.1%) |
| FR | 54 (19.5%) | 0 (0.0%) | 46 (38.3%) | 8 (16.7%) | |
| CR-resistance | 56 (20.2%) | 0 (0.0%) | 47 (39.2%) | 9 (18.8%) | |
| NRTI + NNRTI | IR | 3 (1.1%) | 0 (0.0%) | 2 (1.7%) | 1 (2.1%) |
| FR | 30 (10.83%) | 0 (0.0%) | 26 (21.7%) | 4 (8.3%) | |
| CR-resistance | 33 (11.9%) | 0 (0.0%) | 28 (23.3%) | 5 (10.4%) | |
| Available PR + RT Sequences | 260 | 102 | 109 | 49 | |
| PI + NRTI | IR | 1 (0.4%) | 1 (1.0%) | 0 (0.0%) | 0 (0.0%) |
| FR | 9 (3.5%) | 2 (2.0%) | 5 (4.6%) | 2 (4.1%) | |
| CR-resistance | 10 (3.8%) | 3 (2.9%) | 5 (4.6%) | 2 (4.1%) | |
| PI + NNRTI | IR | 1 (0.4%) | 0 (0.0%) | 1 (0.9%) | 0 (0.0%) |
| FR | 1 (0.4%) | 0 (0.0%) | 1 (0.9%) | 0 (0.0%) | |
| CR-resistance | 2 (0.8%) | 0 (0.0%) | 2 (1.8%) | 0 (0.0%) | |
| PI + NRTI + NNRTI | IR | 0 (0.0%) | 0 (0.0%) | 0 (0.0%) | 0 (0.0%) |
| FR | 5 (1.9%) | 0 (0.0%) | 3 (2.8%) | 2 (4.1%) | |
| CR-resistance | 5 (1.9%) | 0 (0.0%) | 3 (2.8%) | 2 (4.1%) | |
| Available IN Sequences | 61 | 19 | 21 | 21 | |
| INSTI | IR | 0 (0.0%) | 0 (0.0%) | 0 (0.0%) | 0 (0.0%) |
| FR | 2 (3.3%) | 0 (0.0%) | 2 (9.5%) | 2 (4.1%) | |
| CR-resistance | 3 (3.3%) | 0 (0.0%) | 2 (9.5%) | 2 (4.1%) | |
| Available RT + IN Sequences | 52 | 18 | 17 | 17 | |
| NRTI + NNRTI + INSTI | IR | 0 (0.0%) | 0 (0.0%) | 0 (0.0%) | 0 (0.0%) |
| FR | 2 (3.8%) | 0 (0.0%) | 2 (11.8%) | 0 (0.0%) | |
| CR-resistance | 2 (3.8%) | 0 (0.0%) | 2 (11.8%) | 0 (0.0%) | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Schlösser, M.; Kartashev, V.V.; Mikkola, V.H.; Shemshura, A.; Saukhat, S.; Kolpakov, D.; Suladze, A.; Tverdokhlebova, T.; Hutt, K.; Heger, E.; et al. HIV-1 Sub-Subtype A6: Settings for Normalised Identification and Molecular Epidemiology in the Southern Federal District, Russia. Viruses 2020, 12, 475. https://doi.org/10.3390/v12040475
Schlösser M, Kartashev VV, Mikkola VH, Shemshura A, Saukhat S, Kolpakov D, Suladze A, Tverdokhlebova T, Hutt K, Heger E, et al. HIV-1 Sub-Subtype A6: Settings for Normalised Identification and Molecular Epidemiology in the Southern Federal District, Russia. Viruses. 2020; 12(4):475. https://doi.org/10.3390/v12040475
Chicago/Turabian StyleSchlösser, Madita, Vladimir V. Kartashev, Visa H. Mikkola, Andrey Shemshura, Sergey Saukhat, Dmitriy Kolpakov, Alexandr Suladze, Tatiana Tverdokhlebova, Katharina Hutt, Eva Heger, and et al. 2020. "HIV-1 Sub-Subtype A6: Settings for Normalised Identification and Molecular Epidemiology in the Southern Federal District, Russia" Viruses 12, no. 4: 475. https://doi.org/10.3390/v12040475





