1. Introduction
Coronavirus infectious disease 2019 (COVID-19), caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), is essentially a human disease. However, the potential role of other mammalian species in the epidemiology of the disease either as competent hosts or as potential reservoirs has been in the spotlight [
1]. This interest spurred early epidemiological research aiming to predict and identify which animal species could be susceptible to SARS-CoV-2 infection and capable of transmitting the virus to either humans or other animals [
2,
3]. Experimental infections have attempted to identify potential SARS-CoV-2 hosts as well as animal models to recapitulate the COVID-19 immunopathological features observed in humans [
4]. Carnivores and primates have been identified as the most sensitive taxonomic groups to infection, with clinical disease reported in some instances [
4]. However, there is still limited information about the role of mammalian species in the epidemiology of COVID-19.
Despite the high incidence of SARS-CoV-2 human infection, the frequency of spillover events to other species has been relatively low [
5]. In most instances, cases of infection in non-human mammals have been largely asymptomatic and required extensive efforts to identify them. Pets from COVID-19-affected households have usually shown asymptomatic infections or mild respiratory/digestive clinical signs [
6,
7,
8,
9,
10]. Other domestic animals have shown less susceptibility to SARS-CoV-2 infection [
2,
5,
11]. In contrast, mink farms proved to be a singular exception with SARS-CoV-2 causing large outbreaks, resulting in massive culling in several countries. Infection in minks also resulted in the only documented transmission of SARS-CoV-2 from non-human mammals to persons [
12]. Overall, reports of natural infection and clinical disease in animals have been restricted to isolated cases, most likely identified due to extensive surveillance.
Zoos are singular regarding the epidemiology of human–animal interactions: they house multiple wildlife species of a wide array of taxa in relative proximity and the interactions between animals and humans are frequent, particularly for animal caretakers. The COVID-19 pandemic, therefore, triggered research for data and surveillance for suspicious cases of SARS-CoV-2 infection in zoo animals [
13]. Since April 2020, several cases of SARS-CoV-2 infections have been reported in zoo animals [
14]. Most reported outbreaks have involved large felids; curiously, only few cases of natural infection have been detected in non-human primates [
15].
The first SARS-CoV-2 outbreak identified and reported in a zoo took place at the Bronx Zoo, early in the COVID-19 epidemic in New York City, and resulted in a self-limiting disease affecting tigers (
Panthera tigris) and lions (
Panthera leo) [
16]. Unlike infections reported in domestic cats, which have been largely asymptomatic, tigers and lions displayed mild to moderate upper respiratory clinical signs (coughing and wheezing), lasting from 1 to 16 days. In addition, animals experienced transitory decreased appetite or anorexia, but all animals recovered spontaneously. Molecular and virological analyses identified the felid caretakers as their primary source of transmission [
16,
17]. Scarce information has been reported in other large felid infections occurring at zoos in South Africa, the Czech Republic, Sweden, and Sri Lanka [
18]. In addition, nine Asiatic lions kept at the Arignar Anna Zoological Park (India) were infected with the SARS-CoV-2 delta variant (pango B.1.617.2 lineage) in May 2021 [
19]. The source of infection could not be determined considering none of their caretakers were COVID-19 positive, but the sequenced viruses corresponded to the circulating variants in India at that time [
19]. Mild respiratory clinical signs were reported in some animals, while two of the lions died after succumbing to the infection [
19]. So far, no virological or pathological data has been reported but fatalities are presumably related to a COVID-19-like disease in these animals. While these studies provide important information on SARS-CoV-2 infection in large felids, many aspects remain unexplored.
In the present study, we report natural SARS-CoV-2 infection of the four lions at the Barcelona Zoo (Barcelona, Spain) and provide clinical, virological, and immunological data, as well as viral genomic comparison from animals and their caretakers.
2. Materials and Methods
2.1. Clinical Evaluation and Sample Collection
Animal handling and sampling were performed at the Barcelona Zoo (Catalonia, Spain) as part of the routine management and investigations performed when clinical signs are displayed and potential zoonoses are suspected.
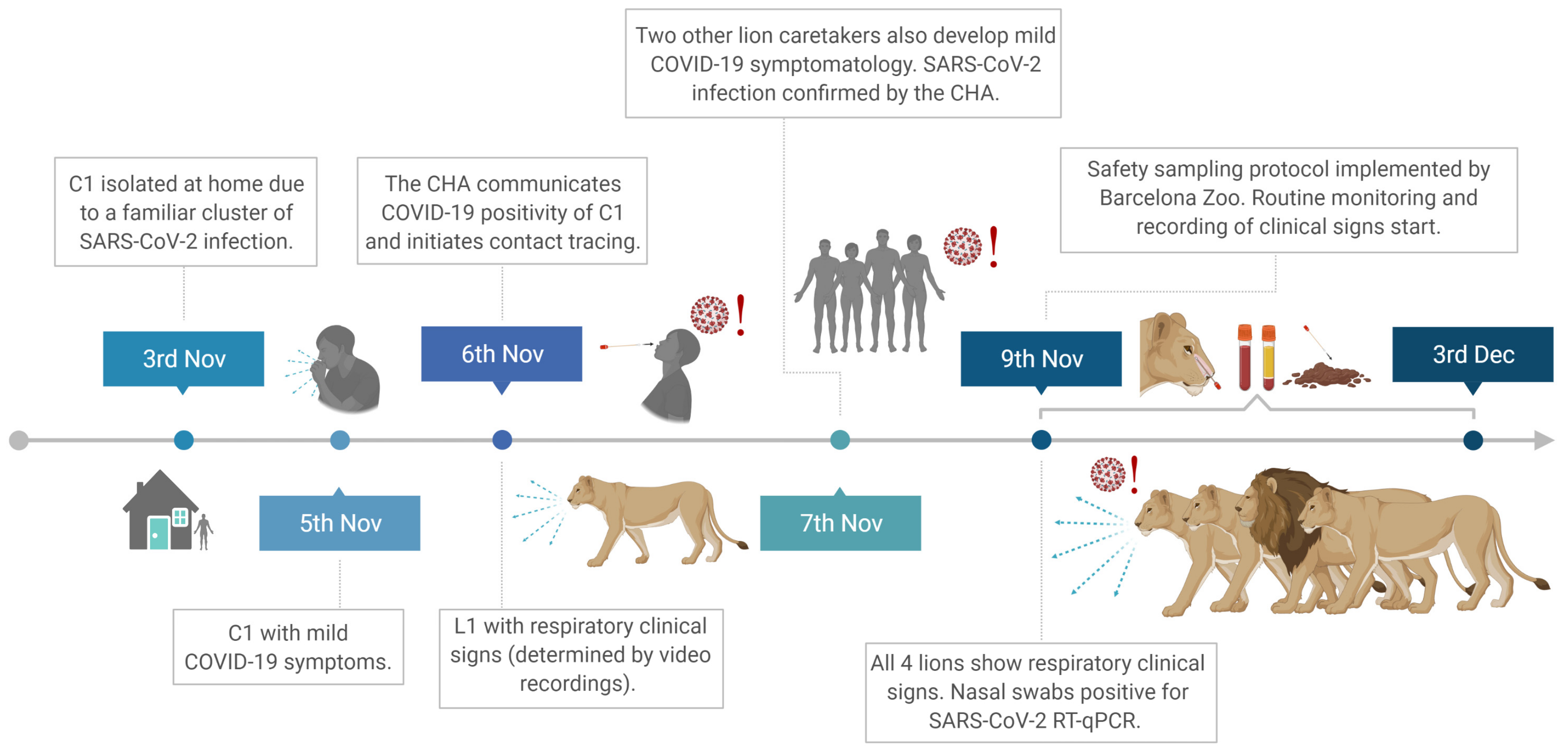
On 8 November 2020, four lions (Panthera leo bleyenberghi, three 16-year-old females and one 4-year-old male; L1, L2, L3, and L4, respectively) developed respiratory signs including sneezing, coughing, and nasal discharge. Additionally, the caretakers noticed changes in the behavior and attitude of the animals. Based on a previous report from the Wildlife Conservation Society describing natural infection with SARS-CoV-2 in large felids at the Bronx Zoo and the clinical suspicion for SARS-CoV-2 infection, a RADT was performed to confirm the infection in lions. Additionally, a safety sampling protocol was implemented together with routine animal management monitoring and more precise recording of clinical signs supported by video images captured continually at the indoor lion facilities. In addition, images between the 4 and 23 November were carefully reviewed to determine the potential onset of clinical signs.
Prior to 9 November, lion keepers wore personal protective equipment (PPE) including nitrile gloves and UNE-00065 certified cloth masks when working in the lion facility or preparing their food. From that day onwards, enhanced PPE included donning dedicated whole-body coveralls, FFP3 masks, face shields, and waterproof shoe covers to access the lion facility.
All four lions were target-trained and were compliant to deep nasal swabbing using sterile cotton swabs, which were inserted into the nasal cavity of the subject, briefly rotated, and extracted. Fecal samples were also collected from the facility, despite the fact that the samples could not be linked to a known individual in most instances. Additionally, saliva was obtained from each lion when possible during the week after respiratory clinical signs’ exhibition. All samples were stored at −80 °C and shipped to the IRTA-CReSA facilities (Catalonia, Spain) on cold packs. All samples were collected at different time-points during a 3-week period between 9 November and 2 December 2020, as shown in
Supplementary Table S1.
The lions were anesthetized for health examination, including blood extraction and pulmonary ultrasound between 2 and 5 weeks after clinical presentation, and three of them were checked-up again 4 months post-clinical presentation. L1 was not anesthetized 4 months later due to risk assessment; this animal was known for a specific reaction to anesthetic procedures and veterinarians were reticent regarding extra anesthesia. Anesthesia procedures were performed by remote darting following initial premedication with midazolam (0.1–0.12 mg/kg BW) using a combination of ketamine (1.5–2 mg/kg BW), butorphanol (0.1–0.15 mg/kg BW), and detomidine (0.075–0.08 mg/kg BW). Whole-blood samples were obtained by jugular venipuncture using standard procedures into vacuum serum collection tubes (Vacutainer®, Beckton Dickinson, San Diego, CA, USA). Blood samples from all lions collected in February 2020 were used as control samples before infection.
2.2. Additional Sample Collection
On initial case presentation, additional nasal swabs from other large carnivores held at the Barcelona Zoo were obtained, including hyenas (Crocutta crocutta), wolves (Canis lupus signatus), bears (Ursus arctos), tigers (Panthera tigris sumatrae), jaguars (Panthera onca), and leopards (Panthera pardus kotiya).
Additionally, two lion keepers who were at home on COVID-19 quarantine at the time when the lions showed clinical signs were asked to collect nasal swab self-samples during the week after clinical presentation of the animals. This period corresponded approximately to one week after their confirmation of COVID-19 positivity. The swabs were stored at −20 °C until the quarantine was lifted and the samples were then shipped to the laboratory.
2.3. SARS-CoV-2 Antigen Test
Nasal swabs obtained as previously described were tested for the presence of the SARS-CoV-2 nucleoprotein (N) antigen using a commercially available kit (COVID-19 Ag Test reference 243103N-20, nal von minden GmbH) following manufacturer’s instructions. Briefly, nasal swabs were immersed, stirred, and incubated at room temperature for 2 min in 10 drops sample buffer. Then, swabs were removed and 2 drops of the sample solution were transferred to the sample well of the rapid test cassette and incubated for 15 min at room temperature. Results were read by interpreting visible bands at the control line and test line.
2.4. Cell Culture and SARS-CoV-2
Vero E6 cells (ATCC
® repository, CRL-1586™) and SARS-CoV-2 stocks (passage 3; hCoV-19/Spain/CT-2020030095/2020; GISAID ID EPI_ISL_510689) were prepared as previously reported in [
20,
21] in Dulbecco’s modified Eagle Medium, DMEM (Lonza, Basel, Switzerland) supplemented with 5% fetal calf serum (FCS; EuroClone, Milan, Italy), 100 U/mL penicillin, 100 μg/mL streptomycin, and 2 mM glutamine (all from Gibco Life Technologies, Madrid, Spain).
SARS-CoV-2 was isolated from a nasopharyngeal swab collected from an 89-year-old person in Barcelona, Spain, and characterized by genomic sequencing (hCoV-19/Spain/CT-IrsiCaixa-04/2020; GISAID ID EPI_ISL_418268) as previously described [
20]. This isolate carries D614G and R682L amino acid substitutions in the spike protein compared to the previously identified Wuhan/Hu-1/2019 reference variant (NCBI accession number: NC_045512.2). The infectious titer of the SARS-CoV-2 stock was calculated by determining the dilution that caused CPE in 50% of the inoculated cell cultures (50% tissue culture infectious dose endpoint, TCID
50) in Vero E6 cells.
2.5. RNA Extraction and Detection by RT-qPCR
Nasal, saliva, and fecal swabs were transferred into cryotubes containing 500 µL DMEM supplemented with 100 U/mL penicillin, 100 μg/mL streptomycin, and 2 mM glutamine, vortexed and stored at −80 °C until use. Detection of SARS-CoV-2 RNA was performed by a reverse transcriptase-quantitative PCR (RT-qPCR) as previously described [
22]. Viral RNA was extracted using the Indimag Pathogen kit (Indical Biosciences, Leipzig, Germany) on a Biosprint 96 workstation (Qiagen, Hilden, Germany) according to the manufacturers’ instructions. Detection of SARS-CoV-2 in RNA extracts was performed following a previously described protocol targeting the E gene (REF), applying minor modifications. Briefly, RT-qPCR was carried out using AgPath-ID
TM One-Step RT-PCR Reagents (Applied Biosystems, Life Technologies), and amplification was done by using a 7500 Fast Real-Time PCR System (Applied Biosystems, Life Technologies, Waltham, MA, USA) with the following thermal profile: 10 min at 50 °C; 10 s at 95 °C; and 45 cycles of 15 s at 94 °C and 30 s at 58 °C. Samples with a Cq value ≤ 40 were considered positive for SARS-CoV-2 RNA.
2.6. SARS-CoV-2 Genome Sequencing and Phylogenetic Analyses
Viral RNA was extracted as described above and converted to cDNA with the PrimeScript™ RT reagent kit (Takara Bio Europe SAS, Saint-Germain-en-Laye, France) using a combination of oligo-dT and random hexamer methods according to the manufacturer’s protocol. cDNA was used for viral DNA enrichment using the ARTIC-Cov v3 PCR protocol and primer design with the Q5 Hot-start HF polymerase. Resulting PCR products were used for sequencing-ready library preparation with the Illumina DNA LibPrep kit (Illumina, San Diego, CA, USA). Sequencing-ready libraries were then loaded onto the Illumina MiSeq platform and a 150 bp paired-end sequencing kit (300 cycles). Sequence reads were quality-filtered and adapter primer sequences were trimmed using Trimmomatic. Sequencing reads were then mapped against the reference Wuhan/Hu-1/2019 variant (NCBI accession number: NC_045512.2) using the bowtie2 tool [
23]. Consensus genomic sequence were generated from the resulting alignments at average depth of coverages ranging from 85 to 1475x using iVar/samtools [
24,
25]. Genomic sequences from lion specimens were deposited at the GISAID repository with correlative accession IDs from EPI_ISL_1443170 to EPI_ISL_1443186, and EPI_ISL_1740499 for the lion caretaker.
To compare the sequenced genomes obtained from the lion respiratory tract and fecal specimens, as well as the caretaker nasopharyngeal samples, evolutionary analysis was inferred by the maximum likelihood method and the Hasegawa–Kishino–Yano model [
26]. The percentage of trees in which the associated taxa clustered together is shown next to the branches. Initial tree(s) for the heuristic search were obtained automatically by applying neighbor-joining and BioNJ algorithms to a matrix of pairwise distances estimated using the maximum composite likelihood approach and then by selecting the topology with the superior log likelihood value. The tree is drawn to scale with branch lengths measured in the number of substitutions per site. The analysis involved 20 nucleotide sequences. Codon positions included were 1st + 2nd + 3rd + Non-coding. All positions with less than 80% site coverage were eliminated, i.e., fewer than 20% alignment gaps, missing data, and ambiguous bases were allowed at any position (partial deletion option). There was a total of 25,132 positions in the final dataset. Evolutionary analyses were conducted in MEGA X [
27].
2.7. Virus Isolation in Vero E6 Cells
Nasal and fecal swabs positive for SARS-CoV-2 as determined by RT-qPCR were evaluated for the presence of the infectious virus by titration in Vero E6 cells, as previously reported [
21,
28]. Direct samples and ten-fold dilutions were transferred to Vero E6 monolayers. Plates were monitored daily under light microscope and wells were assessed for the presence of cytopathic effects for 6 days. The amount of infectious virus in the swabs was calculated by determining the TCID
50.
2.8. SARS-CoV-2 RBD and N-ELISA
Serological assays were performed to detect antibodies against the S and N proteins of SARS-CoV-2. Prior to ELISA tests, serum samples were heat inactivated at 56 °C for 30 min. Afterwards, specific nAbs against the RBD of the S protein were determined in serum samples from all collected time-points using the SARS-CoV-2 sVNT (Genescript) following the manufacturer’s instructions. The specific percentage of inhibition for anti-RBD antibodies was calculated after subtracting the background signal obtained for each sample in antigen-free wells and using the mean OD of the negative control samples. Samples with an activity of ≥30% inhibition were considered positive.
In addition, antibodies targeting the N protein were detected using an in-house double-antigen sandwich ELISA (DAS-N-ELISA). The method uses a recombinant N protein antigen coated to the plate and a recombinant N protein conjugated with HRP (N-HRP), expressed in
Escherichia coli and purified by immobilized metal affinity chromatography. Briefly, 0.2 µg/mL of the unconjugated recombinant SARS-CoV-2 N protein (50 µL per well) were coated onto ELISA microplates. Tested sera were diluted 1/2.5 in diluent buffer in a final volume of 50 µL, added to the coated plate, and incubated for 60 min at room temperature. Negative and positive controls were included in each plate at the same serum dilution. After three washing cycles, 50 µL/well of the HRP-conjugated recombinant SARS-CoV-2 N protein was added and the plate was incubated for another 60 min at room temperature. After washing, TMB substrate (Surmodics) was added and the ELISA microplates were incubated in the dark at room temperature for 20 min. The colorimetric reaction was then stopped by the addition of 2N sulphuric acid and absorbance values were read at 450 nm using an ELISA reader. Results were expressed as a percentage of reactivity compared with the positive control included in each plate (%S/P), calculated as (OD sample − average OD neg control/average OD pos control − average OD neg control) × 100. More details of the development and analytical performance of the in-house DAS-N-ELISA can be found in the
Supplementary Materials (
Appendix A).
2.9. SARS-CoV-2 Neutralization Assay
The seroneutralization assay using replicating SARS-CoV-2 was performed as previously described [
21,
29]. Serum samples were inactivated for 30 min at 56 °C. Then, sera were first diluted 1:50 and then were 3-fold serially diluted in supplemented DMEM, mixed 1:1 with the SARS-CoV-2 isolate (EPI_ISL_418268), and incubated 1 h at 37 °C. Then, each dilution mixture (in duplicates) was transferred to Vero E6 monolayers containing 100 TCID50 of SARS-CoV-2 per well and were cultured for 3 days at 37 °C and 5% CO
2. CPE was measured after 3 days using the CellTiter-Glo luminescent cell viability assay (Promega, Madison, WI, USA) according to the manufacturer’s protocol. Luminescence was measured as luminescence units in a Fluoroskan Ascent FL luminometer (ThermoFisher Scientific, Waltham, MA, USA). Serum virus neutralization titer (SNT50) was defined as the reciprocal value of the sample dilution that showed 50% protection of virus growth. Sera with titers ≥ 1/100 were considered positive for SARS-CoV-2 nAbs.
2.10. Data Analysis and Statistics
Mean of binding or nAb data were analyzed using the parametric two-way ANOVA test and paired t-test for comparison of 2 groups. The normality of each data set was calculated using the Shapiro–Wilk test and p-values lower than 0.05 were considered statistically significant.
Dose–response curves of serum samples were adjusted to a non-linear fit regression model calculated with a normalized logistic curve with variable slope. For data normalization, uninfected cells and untreated virus-infected cells were used as negative and positive control of infection (% Neutralization = (RLUmax − RLUexperimental)/(RLUmax − RLUmin) × 100), respectively. All SNT50 values were determined as the concentration of sera blocking 50% of the SARS-CoV-2-induced CPE in Vero E6 cells and were expressed as reciprocal dilution. All statistical analyses were performed with GraphPad Prism 8.4.3 (GraphPad Software, Inc., San Diego, CA, USA).
4. Discussion
In the present study we reported extensive clinical, virological, and immunological insights of natural SARS-CoV-2 infection in lions. The investigated outbreak occurring in four lions from the Barcelona Zoo is consistent with previous reports of non-domestic large felids being infected with SARS-CoV-2 (
Panthera tigris,
Panthera leo, and
Puma concolor) [
16,
17,
18,
19,
30]. The chronological events and further genomic characterization fully support human-to-animal transmission of SARS-CoV-2. Moreover, viral genomic sequences retrieved from the respiratory specimens of the lions and their direct keepers were identical, confirming SARS-CoV-2 transmission. Although the transmission route could not be determined (lion keepers wore cloth facial masks inside the lion facility or when preparing their food), the main hypothesis points to close contact transmission between lions and their caretakers; however, human-to-animal indirect transmission or potential transmission events among lions cannot be excluded.
After displaying mild respiratory clinical signs (sneezing, coughing, nasal discharge, and mild apathy), all four lions were rapidly confirmed as infected by SARS-CoV-2. In fact, infection was monitored for a 3-week period at both RNA and protein levels, as determined by RT-qPCR and RADT, respectively, recognizing the N protein of SARS-CoV-2. Although diagnostic sensitivity of RADT is limited, it appears to be a useful tool to monitor SARS-CoV-2 infection in zoo animals. The animals shed viral RNA for approximately 2 weeks after the onset of clinical signs and confirmation of the viral infection. Although the peak of infection occurred slightly before the first sampling and viral shedding varied among individuals, infectious SARS-CoV-2 could be isolated from the nasal cavity of two out of four animals. Additionally, viral RNA was determined in other specimens such as saliva and feces, the latter containing limited loads of viral RNA and no infectious virus could be cultured from any of the collected fecal samples during the study. Although fecal sampling has been proposed as a valid surrogate to monitor infection in these large carnivores, it may be less sensitive than respiratory sampling and may not parallel the described respiratory disease. This is especially relevant for lion management and safe sample collection at zoo facilities.
Furthermore, whole-viral genome sequencing showed that all lions were infected with the same virus. The SARS-CoV-2 variant infecting lions belonged to the pango lineage B.1.177, which indeed was circulating in Europe (including Barcelona, Spain) in November 2020 when the human-to-lion transmission occurred. This lineage contains A222V, E583V, and D614G mutations in the S protein, which altered SARS-CoV-2 fitness and were fixed in the human population during 2020, with an increased transmissibility compared to other viral lineages [
22,
31,
32]. Distinct viral variants have also infected large felids in different zoos worldwide [
17,
19,
30], which can potentially create new niches for SARS-CoV-2 natural reservoirs. More studies are required to assess the impact of these mutations towards a potential broadening of the susceptible host range as already observed with other SARS-CoV-2 variants [
33].
Importantly, the infection of lions resulted in high-genomic stability and no amino acid mutations were found in high-quality genomic sequences from respiratory specimens monitored for up to 2 weeks. However, we identified two punctual mutations (S50L and G496D) in the genomes of all fecal samples. S50L mutation specifically occurred in the N-terminal domain of the SARS-CoV-2 S protein, while G496D is found in the receptor-binding motif of the RBD domain. These mutations have also spontaneously emerged in infected humans around the globe but at a low prevalence and it has not been established within the currently circulating SARS-CoV-2 variants. Notably, both S50L and G496D mutations identified in feces from naturally SARS-CoV-2-infected lions were also found in fecal samples from different
Rhinolophus bat species (
R. affinis,
R. malayanus,
R. shameli,
R. sinicus,
R. stheno,
R. pusillus, and other several
Rhinolophus species) sampled during the 2010–2020 period in different geographical locations such as the Chinese province of Yunnan as well as Cambodia [
34]. Curiously, the closest SARSr-CoV known to date (RaTG13/2013), which shares 96.1% homology to SARS-CoV-2, as well as other SARSr-CoVs characterized in Malayan Pangolins sampled during 2017–2019 [
35], also possesses a leucine in the amino acid position 50 of the S protein. In addition, an aspartic acid at position 496 in the S protein (D496) has been continuously found in SARSr-CoVs containing the L50 residue recovered from bat feces sampled during 2018–2019 [
34]. Worthy to note, the viral evolution results presented here are in line with the only previous study providing genomic SARS-CoV-2 sequences from naturally infected large felids [
17,
19]. McAloose et al. sequenced the SARS-CoV-2 genomes from respiratory specimens of a naturally infected tiger at the Bronx Zoo, along with others from the fecal samples of four tigers and three lions. In addition, Mishra and collaborators sequenced the viruses from nasal swabs of four lions. Interestingly, none of these mutations were found in SARS-CoV-2 sequences obtained from the respiratory tract of tigers and lions [
17,
19], which also showed high-viral genomic stability and coincided with our studies performed in the upper respiratory tract. However, the G496D mutation was present in the SARS-CoV-2 sequences also obtained from feces of three naturally infected tigers at the Bronx Zoo [
17]. The presence of the G496D mutation exclusively in the two reported outbreaks providing SARS-CoV-2 sequences in feces from large felids might not be just incidental. Further studies are needed to elucidate the capabilities of SARS-CoV-2 to replicate in different anatomic compartments of large feline species and whether they impose different bottleneck pressures to the virus replicating inside the same host.
Our results also showed that naturally infected lions cleared the infection and recovered from respiratory clinical signs 2–3 weeks after infection, developing robust humoral responses against SARS-CoV-2 already at 2–5 weeks after infection. In line with previous studies in one tiger [
17], all lions developed high titers of nAbs against the S protein and specifically the RBD domain of SARS-CoV-2. Nonetheless, as similarly reported in domestic felines [
2,
6,
36,
37], limited levels of antibodies against the nucleoprotein were observed in three of the animals. Moreover, the levels of nAbs decreased four months after infection in all analyzed lions. Due to the lack of specific reagents, it was not possible to determine the isotype of the reactive antibodies against SARS-CoV-2, although IgGs are probably the most abundant ones in serum. More studies are needed to determine the duration of immunity developed by the lions and whether the animals develop sterilizing immunity or, conversely, they could be susceptible to re-infections.
In summary, SARS-CoV-2 infection dynamics in large felids from the Barcelona Zoo were monitored in respiratory and fecal samples for a period of 4 months. The viral variant belonging to the B.1.177 lineage was confirmed to jump from humans to lions and caused respiratory disease in both species without the need of specific mutations. Lions cleared viral infection within three weeks and developed high levels of neutralizing responses to SARS-CoV-2 that lasted at least for four months.