Molecular Tools to Identify and Characterize Malignant Catarrhal Fever Viruses (MCFV) of Ruminants and Captive Artiodactyla
Abstract
:1. Introduction
1.1. Herpesviruses
1.2. Malignant Catarrhal Fever Viruses and MCF
2. Molecular Methodologies
2.1. Consensus Polymerase Chain Reaction Assays
2.2. Consensus Degenerate Hybrid Oligonucleotide Primers PCR Assays
2.3. Consensus Polymerase Chain Reaction Assay for Herpesvirus Detection
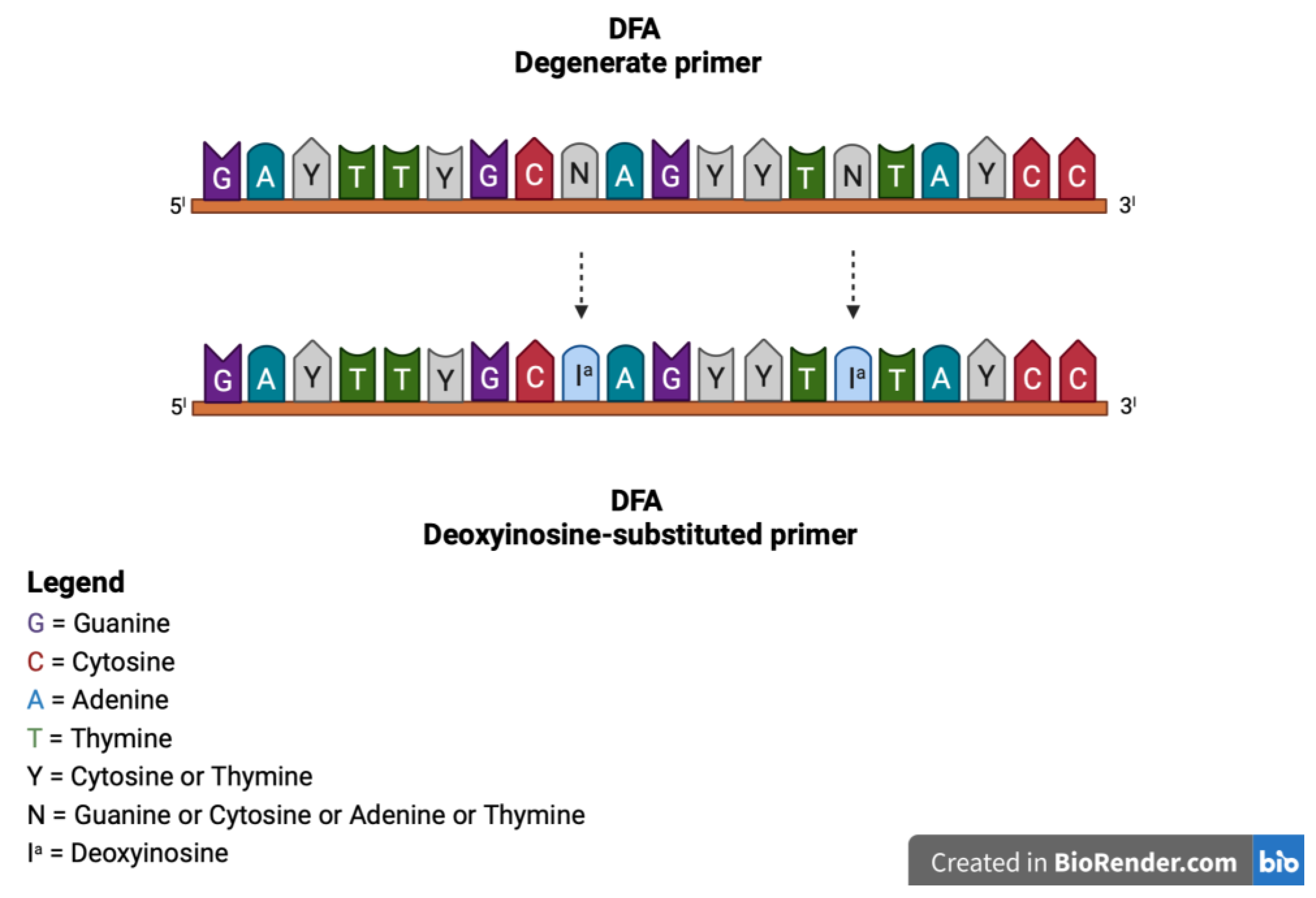
2.4. Consensus Panherpes PCR with Consensus/Degenerate and Deoxyinosine-Substituted Primers
2.5. Ovine Herpesvirus 2 Sequence Analysis
2.6. Long-Distance Polymerase Chain Reaction Assay and Primer Walking
2.7. Locked Nucleic Acids Approaches
2.8. Next Generation Sequencing
3. Conclusions and Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Malignant Catarrhal Fever. Available online: https://www.woah.org/app/uploads/2021/03/malignant-catharral-fever.pdf (accessed on 25 August 2022).
- Orono, S.A.; Gitao, G.C.; Mpatswenumugabo, J.P.; Chepkwony, M.; Mutisya, C.; Okoth, E.; Bronsvoort, B.M.C.; Russell, G.C.; Nene, V.; Cook, E.A.J. Field validation of clinical and laboratory diagnosis of wildebeest associated malignant catarrhal fever in cattle. BMC Vet. Res. 2019, 15, 69. [Google Scholar] [CrossRef] [PubMed]
- Meteyer, C.U.; Gonzales, B.J.; Heuschele, W.P.; Howard, E.B. Epidemiologic and pathologic aspects of an epizootic of malignant catarrhal fever in exotic hoofstock. J. Wildl. Dis. 1989, 25, 280–286. [Google Scholar] [CrossRef] [Green Version]
- Khudhair, Y.I.; Ayyez, H.N.; Hussain, M.H. Phylogenetic analysis of ovine herpes virus-2 (OHV-2) in malignant catarrhal fever infected cattle in Al-Qadisiyah governorate of Iraq. Iraqi J. Vet. Sci. 2019, 33, 51–58. [Google Scholar] [CrossRef]
- Cunha, C.W.; Slater, O.M.; Macbeth, B.; Duignan, P.J.; Warren, A.; Highland, M.A.; Li, H. Domestic sheep and bighorn sheep carry distinct gammaherpesviruses belonging to the genus Macavirus. Virus Res. 2019, 272, 197729. [Google Scholar] [CrossRef]
- Sharma, B.; Parul, S.; Basak, G.; Mishra, R. Malignant catarrhal fever (MCF): An emerging threat. J. Entomol. Zool. Stud. 2019, 7, 26–32. [Google Scholar]
- Azab, W.; Dayaram, A.; Greenwood, A.D.; Osterrieder, N. How host specific are herpesviruses? Lessons from herpesviruses infecting wild and endangered Mammals. Ann. Rev. Virol. 2018, 5, 53–68. [Google Scholar] [CrossRef]
- Davison, A.J.; Eberle, R.; Ehlers, B.; Hayward, G.S.; McGeoch, D.J.; Minson, A.C.; Pellett, P.E.; Roizman, B.; Studdert, M.J.; Thiry, E. The order Herpesvirales. Arch. Virol. 2009, 154, 171–177. [Google Scholar] [CrossRef]
- McGeoch, D.J.; Cook, S.; Dolan, A.; Jamieson, F.E.; Telford, E.A.R. Molecular phylogeny and evolutionary timescale for the family of mammalian herpesviruses. J. Mol. Biol. 1995, 247, 443–458. [Google Scholar] [CrossRef]
- McGeoch, D.J.; Rixon, F.J.; Davison, A.J. Topics in herpesvirus genomics and evolution. Virus Res. 2006, 117, 90–104. [Google Scholar] [CrossRef]
- International Committee on Taxonomy of Viruses. Virus Taxonomy: 2021 Release. EC 53, Online, July 2021. Email Ratification March 2022 (MSL #37). Available online: https://talk.ictvonline.org/taxonomy/ (accessed on 5 July 2022).
- Li, H.; Keller, J.; Knowles, D.P.; Crawford, T.B. Recognition of another member of the Malignant Catarrhal Fever Virus group: An endemic gammaherpesvirus in domestic goats. J. Gen. Virol. 2001, 82, 227–232. [Google Scholar] [CrossRef]
- Li, H.; Cunha, C.W.; Abbitt, B.; deMaar, T.W.; Lenz, S.D.; Hayes, J.R.; Taus, N.S. Goats are a potential reservoir for the herpesvirus (MCFV-WTD), causing malignant catarrhal fever in deer. J. Zoo Wildl. Med. 2013, 44, 484–486. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Cunha, C.W.; Taus, N.S.; Knowles, D.P. Malignant Catarrhal Fever: Inching toward understanding. Annu. Rev. Anim. Biosci. 2014, 2, 209–233. [Google Scholar] [CrossRef] [PubMed]
- Sood, R.; Kumar, N.; Bhatia, S. Malignant Catarrhal Fever. In Emerging and Re-Emerging Infectious Diseases of Livestock; Bayry, J., Ed.; Springer International Publishing: Berlin/Heidelberg, Germany, 2017; pp. 347–362. ISBN 978-3-319-47424-3. [Google Scholar]
- Russell, G.C.; Stewart, J.P.; Haig, D.M. Malignant Catarrhal Fever: A review. Vet. J. 2009, 179, 324–335. [Google Scholar] [CrossRef] [PubMed]
- Jia, J.; Delhon, G.; Tulman, E.R.; Diel, D.G.; Osorio, F.A.; Wen, X.; Kutish, G.F.; Rock, D.L. Novel gammaherpesvirus functions encoded by Bovine Herpesvirus 6 (Bovine Lymphotropic Virus). J. Gen. Virol 2014, 95, 1790–1798. [Google Scholar] [CrossRef]
- Franzo, G.; Drigo, M.; Legnardi, M.; Grassi, L.; Menandro, M.L.; Pasotto, D.; Cecchinato, M.; Tucciarone, C.M. Porcine gammaherpesviruses in Italian commercial swine population: Frequent but harmless. Pathogens 2021, 10, 47. [Google Scholar] [CrossRef]
- Carvallo, F.R.; Uzal, F.A.; Moore, J.D.; Jackson, K.; Nyaoke, A.C.; Naples, L.; Davis-Powell, J.; Stadler, C.K.; Boren, B.A.; Cunha, C.; et al. Ibex-associated malignant catarrhal fever in Duikers (Cephalophus spp.). Vet. Pathol. 2022, 57, 577–581. [Google Scholar] [CrossRef]
- Flach, E.J.; Reid, H.; Pow, I.; Klemt, A. gamma herpesvirus carrier status of captive artiodactyls. Res. Vet. Sci. 2002, 73, 93–99. [Google Scholar] [CrossRef] [PubMed]
- Wambua, L.; Wambua, P.N.; Ramogo, A.M.; Mijele, D.; Otiende, M.Y. Wildebeest-associated Malignant Catarrhal Fever: Perspectives for integrated control of a lymphoproliferative disease of cattle in sub-Saharan Africa. Arch. Virol. 2016, 161, 1–10. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Cunha, C.W.; Taus, N.S. Malignant Catarrhal Fever: Understanding molecular diagnostics in context of epidemiology. Int. J. Mol. Sci. 2011, 12, 6881–6893. [Google Scholar] [CrossRef]
- Campos, M.J.; Quesada, A. Strategies to improve efficiency and specificity of degenerate primers in PCR. In PCR; Domingues, L., Ed.; Methods in Molecular Biology; Springer New York: New York, NY, USA, 2017; Volume 1620, pp. 75–85. ISBN 978-1-4939-7059-9. [Google Scholar]
- Staheli, J.P.; Boyce, R.; Kovarik, D.; Rose, T.M. CODEHOP PCR and CODEHOP PCR primer design. In PCR Protocols; Park, D.J., Ed.; Methods in Molecular Biology; Humana Press: Totowa, NJ, USA, 2011; Volume 687, pp. 57–73. ISBN 978-1-60761-943-7. [Google Scholar]
- Rose, T.M.; Schultz, E.R.; Henikoff, J.G.; Pietrokovski, S.; McCallum, C.M.; Henikoff, S. Consensus-degenerate hybrid oligonucleotide primers for amplification of distantly related sequences. Nucleic Acids Res. 1998, 26, 1628–1635. [Google Scholar] [CrossRef]
- Rose, T. CODEHOP (COnsensus-DEgenerate Hybrid Oligonucleotide Primer) PCR primer design. Nucleic Acids Res. 2003, 31, 3763–3766. [Google Scholar] [CrossRef] [PubMed]
- Boyce, R.; Chilana, P.; Rose, T.M. ICODEHOP: A new interactive program for designing COnsensus-DEgenerate hybrid oligonucleotide primers from multiply aligned protein sequences. Nucleic Acids Res. 2009, 37, W222–W228. [Google Scholar] [CrossRef] [Green Version]
- Rose, T.M. CODEHOP-Mediated PCR–A powerful technique for the identification and characterization of viral genomes. Virol. J. 2005, 2, 20. [Google Scholar] [CrossRef] [PubMed]
- Partin, T.G.; Schrenzel, M.D.; Braun, J.; Witte, C.L.; Kubiski, S.V.; Lee, J.; Rideout, B.A. Herpesvirus survellance and discovery in zoo-housed ruminants. PLoS ONE 2021, 16, e0246162. [Google Scholar] [CrossRef] [PubMed]
- Davison, A.J. Evolution of the herpesviruses. Vet. Microbiol. 2002, 86, 69–88. [Google Scholar] [CrossRef] [PubMed]
- McGeoch, D.J. Molecular Evolution of the γ–Herpesvirinae. Philos. Trans. Royal Soc. Lond. Ser. B 2001, 356, 421–435. [Google Scholar] [CrossRef] [Green Version]
- Ito, J.; Braithwaite, D.K. Compilation and alignment of DNA polymerase sequences. Nucleic Acids Res. 1991, 19, 4045–4057. [Google Scholar] [CrossRef] [Green Version]
- VanDevanter, D.R.; Warrener, P.; Bennett, L.; Schultz, E.R.; Coulter, S.; Garber, R.L.; Rose, T.M. Detection and analysis of diverse herpesviral species by consensus primer PCR. J. Clin. Microbiol. 1996, 34, 1666–1671. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Dyer, N.; Keller, J.; Crawford, T.B. Newly recognized herpesvirus causing Malignant Catarrhal Fever in White-Tailed deer (Odocoileus Virginianus). J. Clin. Microbiol. 2000, 38, 1313–1318. [Google Scholar] [CrossRef] [Green Version]
- Li, H.; Gailbreath, K.; Flach, E.J.; Taus, N.S.; Cooley, J.; Keller, J.; Russell, G.C.; Knowles, D.P.; Haig, D.M.; Oaks, J.L.; et al. A novel subgroup of Rhadinoviruses in ruminants. J. Gen. Virol. 2005, 86, 3021–3026. [Google Scholar] [CrossRef]
- Li, H.; Gailbreath, K.; Bender, L.C.; West, K.; Keller, J.; Crawford, T.B. Evidence of three new members of Malignant Catarrhal Fever Virus group in Muskox (Ovibos Moschatus), Nubian Ibex (Capra Nubiana), and Gemsbok (Oryx Gazella). J. Wildl. Dis. 2003, 39, 875–880. [Google Scholar] [CrossRef] [PubMed]
- Ehlers, B.; Borchers, K.; Grund, C.; Fro¨lich, K.; Ludwig, H.; Buhk, H.-J. Detection of new DNA polymerase genes of known and potentially novel herpesviruses by PCR with degenerate and deoxyinosine-substituted primers. Virus Genes 1999, 18, 211–220. [Google Scholar] [CrossRef] [PubMed]
- Rossolini, G.M.; Cresti, S.; Ingianni, A.; Cattani, P.; Riccio, M.L.; Satta, G. Use of Deoxyinosine-containing primers vs degenerate primers for polymerase chain reaction based on ambiguous sequence information. Mol. Cell. Probes 1994, 8, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Ehlers, B.; Ulrich, S.; Goltz, M. Detection of two novel porcine herpesviruses with high similarity to gammaherpesviruses. J. Gen. Virol. 1999, 80, 971–978. [Google Scholar] [CrossRef]
- Prepens, S.; Kreuzer, K.-A.; Leendertz, F.; Nitsche, A.; Ehlers, B. Discovery of herpesviruses in multi-infected primates using locked nucleic acids (LNA) and a bigenic PCR approach. Virol. J. 2007, 4, 84. [Google Scholar] [CrossRef] [Green Version]
- Chmielewicz, B.; Goltz, M.; Franz, T.; Bauer, C.; Brema, S.; Ellerbrok, H.; Beckmann, S.; Rziha, H.-J.; Lahrmann, K.-H.; Romero, C.; et al. A novel porcine gammaherpesvirus. Virology 2003, 308, 317–329. [Google Scholar] [CrossRef] [Green Version]
- Russell, G.C.; Scholes, S.F.; Twomey, D.F.; Courtenay, A.E.; Grant, D.M.; Lamond, B.; Norris, D.; Willoughby, K.; Haig, D.M.; Stewart, J.P. Analysis of the genetic diversity of ovine herpesvirus 2 in samples from livestock with malignant catarrhal fever. Vet. Microbiol. 2014, 172, 63–71. [Google Scholar] [CrossRef] [Green Version]
- Chmielewicz, B.; Goltz, M.; Ehlers, B. Detection and multigenic characterization of a novel gammaherpesvirus in goats. Virus Res. 2001, 75, 87–94. [Google Scholar] [CrossRef]
- Ehlers, B.; Küchler, J.; Yasmum, N.; Dural, G.; Voigt, S.; Schmidt-Chanasit, J.; Jäkel, T.; Matuschka, F.-R.; Richter, D.; Essbauer, S.; et al. Identification of novel rodent herpesviruses, including the first gammaherpesvirus of Mus Musculus. J. Virol 2007, 81, 8091–8100. [Google Scholar] [CrossRef] [Green Version]
- Sverdlov, E.; Azhikina, T. Primer walking. Wiley Online Library. 2005. Available online: https://doi.org/10.1038/npg.els.0005382 (accessed on 10 November 2022).
- Ehlers, B.; Dural, G.; Yasmum, N.; Lembo, T.; de Thoisy, B.; Ryser-Degiorgis, M.-P.; Ulrich, R.G.; McGeoch, D.J. Novel mammalian herpesviruses and lineages within the Gammaherpesvirinae: Cospeciation and interspecies transfer. J. Virol. 2008, 82, 3509–3516. [Google Scholar] [CrossRef] [Green Version]
- Mana, T.; Bhattacharya, B.; Lahiri, H.; Mukhopadhyay, R. XNAs: A troubleshooter for nucleic acid sensing. ACS Omega. 2022, 7, 15296–15307. [Google Scholar] [CrossRef] [PubMed]
- Rose, K.M.; Alves Ferreira-Bravo, I.; Li, M.; Craigie, R.; Ditzler, M.A.; Holliger, P.; DeStefano, J.J. Selection of 2′-Deoxy-2′-fluoroarabino nucleic acid (FANA) aptamers that bind HIV-1 integrase with picomolar affinity. ACS Chem. Biol. 2019, 14, 2166–2175. [Google Scholar] [CrossRef] [PubMed]
- Rodda, A.E.; Parker, B.J.; Spencer, A.; Corrie, S.R. Extending circulating tumor DNA analysis to ultralow abundance mutations: Techniques and challenges. ACS Sens. 2018, 3, 540–560. [Google Scholar] [CrossRef] [PubMed]
- Lechmann, J.; Ackermann, M.; Kaiser, V.; Bachofen, C. Viral infections shared between water buffaloes and small ruminants in Switzerland. J. Vet. Diagn. Investig. 2021, 33, 894–905. [Google Scholar] [CrossRef]
- Gagnon, C.A.; Traesel, C.K.; Music, N.; Laroche, J.; Tison, N.; Auger, J.P.; Music, S.; Provost, C.; Bellehumeur, C.; Abrahamyan, L.; et al. Whole genome sequencing of a Canadian bovine Gammaherpesvirus 4 strain and the possible link between the viral infection and respiratory and reproductive clinical manifestations in dairy cattle. Front. Vet. Sci. 2017, 4, 92. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cornejo Castro, E.M.; Marshall, V.; Lack, J.; Lurain, K.; Immonen, T.; Labo, N.; Fisher, N.C.; Ramaswami, R.; Polizzotto, M.N.; Keele, B.F.; et al. Dual infection and recombination of Kaposi sarcoma herpesvirus revealed by whole-genome sequence analysis of effusion samples. Virus Evol. 2020, 6, veaa047. [Google Scholar] [CrossRef]
- Liu, P.; Fang, X.; Feng, Z.; Guo, Y.M.; Peng, R.J.; Liu, T.; Huang, Z.; Feng, Y.; Sun, X.; Xiong, Z.; et al. Direct sequencing and characterization of a clinical isolate of Epstein-Barr virus from nasopharyngeal carcinoma tissue by using next-generation sequencing technology. J Virol. 2011, 85, 11291–11299. [Google Scholar] [CrossRef] [Green Version]
- Bovo, S.; Schiavo, G.; Bolner, M.; Ballan, M.; Fontanesi, L. Mining livestock genome datasets for an unconventional characterization of animal DNA viromes. Genomics 2022, 114, 110312. [Google Scholar] [CrossRef]


| Virus | Clinical MCF | Reservoir Species | Susceptible Species | Reference |
|---|---|---|---|---|
| Alcelaphine γ- herpesvirus-1 | Yes | Wildebeest | Cattle Deer | [14] |
| Alcelaphine γ- herpesvirus-2 | Yes | Hartebeest Topi | Barbary red deer Bison | [14] |
| Ovine γ-herpesvirus-2 | Yes | Sheep Mouflon | Cattle Water buffalo Moose Domestic goats Deer Bison Pig Giraffe | [1,14] |
| Caprine γ-herpesvirus-2 | Yes | Goat | Sika deer White-tailed deer | [14] |
| Hippotragine γ-herpesvirus-1 | No | Roan antelope | No susceptible species identified | [14] |
| Bovine γ-herpesvirus-6 | No | Cattle | No susceptible species identified | [17] |
| Suid γ-herpesvirus-3 | No | Pig | No susceptible species identified | [18] |
| Suid γ-herpesvirus-4 | No | Pig | No susceptible species identified | [18] |
| Suid γ-herpesvirus-5 | No | Pig | No susceptible species identified | [18] |
| MCFV–white-tailed deer/Caprine γ-herpesvirus-3 | Yes | Goat | White-tailed deer Red brocket deer Reindeer | [14] |
| Ibex-MCFV | Yes | Ibex | Bongo antelopes Anoa Pronghorn Duikers | [14,19] |
| 2a. Consensus Panherpes Polymerase Chain Reaction Primers | |
|---|---|
| First PCR | |
| Forward | Reverse |
| DFA: 5′-GAY TTY GCN AGY YTN TAY CC-3′ | KG1: 5′-GTC TTG CTC ACC AGN TCN ACN CCY TT-3′ |
| ILK: 5′-TCC TGG ACA AGC AGC ARN YSG CNM TNA A-3I’ | |
| Second PCR | |
| Forward | Reverse |
| TGV: 5′-TGT AAC TCG GTG TAY GGN TTY ACN GGN GT-3′ | IYG: 5′-CAC AGA GTC CGT RTC NCC RTA DAT-3′. |
| Approximate product: 215- to 315-bp | |
| Sanger Sequencing | |
| Forward | Reverse |
| TGVseq: 5′-CAT CTG ATG TAA CTC GGT GTA-3′ bottom line | IYGseq: 5′-GAC AAA CAC AGA GTC CGT-3′ |
| 2b. Consensus Panherpes PCR with Consensus/Degenerate and Deoxyinosine-Substituted Primers | |
| First PCR | |
| Forward | Reverse |
| DFA: 5′-GAY TTY GCN AGY YTN TAY CC-3′ | KG1: 5′-GTC TTG CTC ACC AGN TCN ACN CCY TT-3′ |
| ILK: 5′-TCC TGG ACA AGC AGC ARN YSG CNM TNA A-3′ | |
| Deoxyinosine-substituted equivalent | Deoxyinosine-substituted equivalent |
| DFA: 5′-GAY TTY GCIa AGY YTIa TAY CC-3I′ | KG1: 5′-GTC TTG CTC ACC AGIa TCIa ACIa CCY TT-3′ |
| ILK: 5′-TCC TGG ACA AGC AGC ARIa YSG CIaM TIaA-3′ | |
| Second PCR | |
| Forward | Reverse |
| TGV: 5′-TGT AAC TCG GTG TAY GGN TTY ACN GGN GT-3′ | IYG: 5′-CAC AGA GTC CGT RTC NCC RTA DAT-3′ |
| Deoxyinosine-substituted equivalent TGV: 5′-TGT AAC TCG GTG TAY GGIa TTY ACIa GGIa GT-3′ | Deoxyinosine-substituted equivalent IYG: 5′-CAC AGA GTC CGT RTC IaCC RTA IaAT-3′ |
| 3a. Semi-Nested PCR for the Glycoprotein B (gB) Gene | |
|---|---|
| First PCR | |
| Forward | Reverse |
| 702 Gb: 5′-CAR IaTIa CAR TWT GCM TAY GAC-3′ | 702 Gb: 5′-GTA RTA RTT RTA YTC YCT RAA-3′ |
| Second PCR | |
| Forward | Reverse |
| 734 gB: 5′-GCA AAA TCA ACC CTA CVA GYG TNA TG-3′ | 702 gB: 5′-GTA RTA RTT RTA YTC YCT RAA-3′ |
| Approximate product: 515bp | |
| 3b. Nested PCR for the Glycoprotein B (gB) Gene | |
| First PCR | |
| Forward | Reverse |
| GH1 2759: 5′-CCT CCC AGG TTC ART WYG CMT AYG A-3I | GH1 2762: 5′-CCG TTG AGG TTC TGA GTG TAR TAR TTR TAY TC-3′ |
| GH2 3029: 5′-CCC AGT TGC ART WYG GC(N/Ia) TAY GA-3′ | GH2 3033: 5′-GCC AGG CGT TGC GT(N/Ia) TAR TAR TTR TA-3′ |
| Second PCR | |
| Forward | Reverse |
| GH1 2760: 5′-AAG ATC AAC CCC AC(N/Ia) AG(N/Ia) GT(N/Ia) ATG-3′ | GH1 2761: 5′-GTG TAG TAG TTG TAC TCC CTR AAC AT(N/Ia) GTY TC-3′ |
| GH2 3031: 5′-CAA GAT TAA CCC CAC (N/Ia)AG (N/Ia)GT (N/Ia)AT G-3′ | GH2 3032: 5′-TTG CGT GTA GTA GTT GTA YTC (N/Ia)CT RAA CAT-3′ |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bianchessi, L.; Rocchi, M.S.; Maley, M.; Piccinini, R.; Turin, L. Molecular Tools to Identify and Characterize Malignant Catarrhal Fever Viruses (MCFV) of Ruminants and Captive Artiodactyla. Viruses 2022, 14, 2697. https://doi.org/10.3390/v14122697
Bianchessi L, Rocchi MS, Maley M, Piccinini R, Turin L. Molecular Tools to Identify and Characterize Malignant Catarrhal Fever Viruses (MCFV) of Ruminants and Captive Artiodactyla. Viruses. 2022; 14(12):2697. https://doi.org/10.3390/v14122697
Chicago/Turabian StyleBianchessi, Laura, Mara Silvia Rocchi, Madeleine Maley, Renata Piccinini, and Lauretta Turin. 2022. "Molecular Tools to Identify and Characterize Malignant Catarrhal Fever Viruses (MCFV) of Ruminants and Captive Artiodactyla" Viruses 14, no. 12: 2697. https://doi.org/10.3390/v14122697







