Nationwide Harmonization Effort for Semi-Quantitative Reporting of SARS-CoV-2 PCR Test Results in Belgium
Abstract
:1. Introduction
2. Materials and Methods
2.1. Preparation of SARS-CoV-2 Control Material
2.2. Distribution of SARS-CoV-2 Control Material and Setup of Standard Curves
2.3. Data Processing: Data Requirements
2.4. Data Processing: Approach for Each PCR Assay
3. Results
3.1. Preparation of the SARS-CoV-2 Control Material
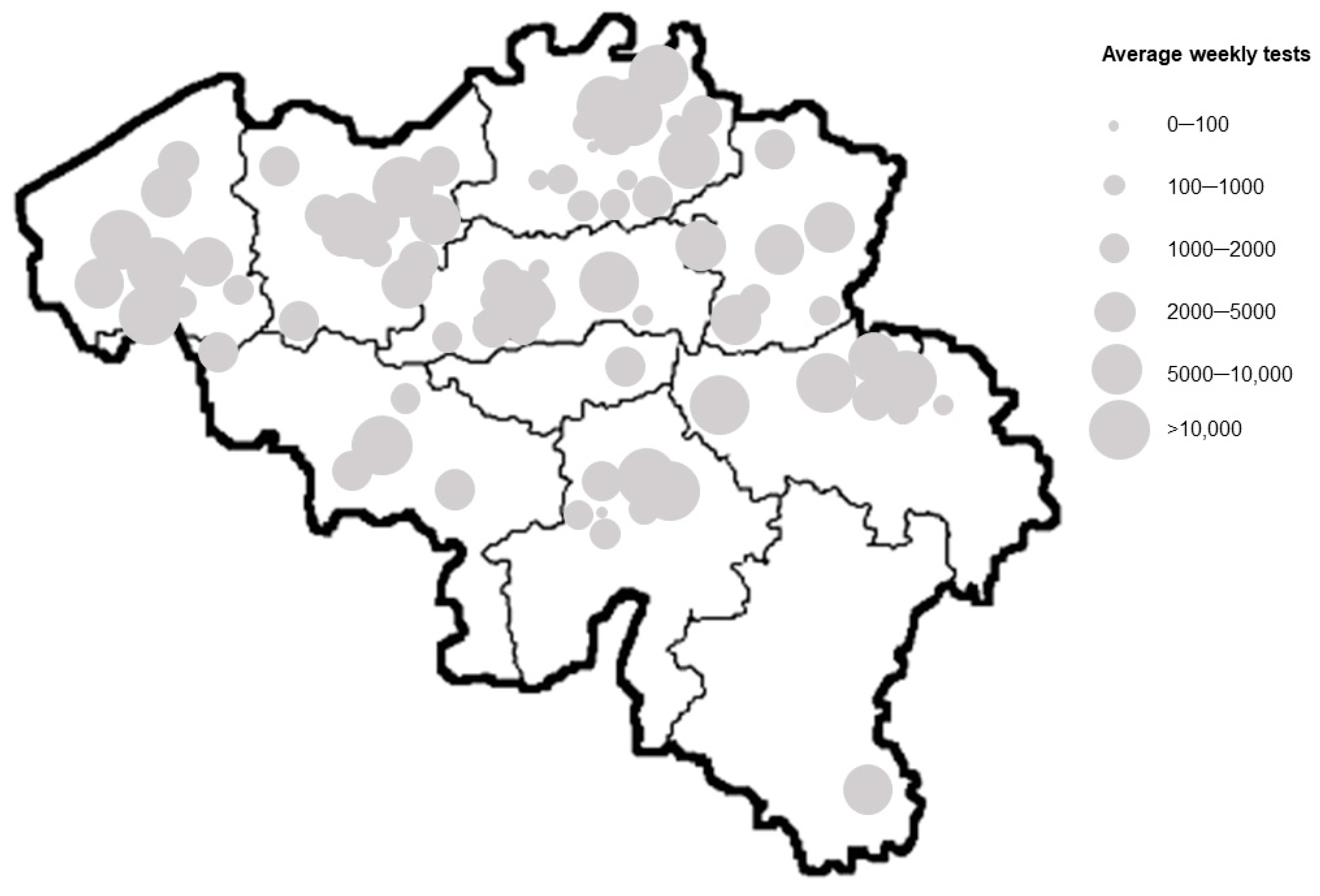
3.2. Distribution of the SARS-CoV-2 Control Material to the Testing Laboratories in Belgium
3.3. The SARS-CoV-2 Nucleocapsid Gene Is Most Often Targeted in PCR Assays Used in Routine Diagnostics in Belgium
3.4. The Difference in Cycle Threshold Values between the Wide Arsenal of SARS-CoV-2 Assays
4. Discussion
4.1. Harmonized Semi-Quantitative Reporting of 85% of the PCR Tests in Belgium
4.2. Limitations When Reporting Test Results Based on Ct Values
4.3. Impact of the Target Composition of PCR Assays on SARS-CoV-2 Genomic Surveillance
4.4. Interpretation of SARS-CoV-2 PCR Test Results
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mardian, Y.; Kosasih, H.; Karyana, M.; Neal, A.; Lau, C.-Y. Review of current COVID-19 diagnostics and opportunities for further development. Front Med. 2021, 8, 615099. [Google Scholar] [CrossRef] [PubMed]
- La Scola, B.; Le Bideau, M.; Andreani, J.; Hoang, V.T.; Grimaldier, C.; Colson, P.; Gautret, P.; Raoult, D. Viral RNA load as determined by cell culture as a management tool for discharge of SARS-CoV-2 patients from infectious disease wards. Eur. J. Clin. Microbiol. 2020, 39, 1059–1061. [Google Scholar] [CrossRef] [PubMed]
- Bullard, J.; Dust, K.; Funk, D.; Strong, J.E.; Alexander, D.; Garnett, L.; Boodman, C.; Bello, A.; Hedley, A.; Schiffman, Z.; et al. Predicting infectious severe acute respiratory syndrome coronavirus 2 from diagnostic samples. Clin. Infect. Dis. 2020, 71, 2663–2666. [Google Scholar] [CrossRef] [PubMed]
- Jefferson, T.; Spencer, E.A.; Brassey, J.; Heneghan, C. Viral cultures for coronavirus disease 2019 infectivity assessment: A systematic review. Clin. Infect. Dis. 2020, 73, e3884–e3899. [Google Scholar] [CrossRef]
- Arons, M.M.; Hatfield, K.M.; Reddy, S.C.; Kimball, A.; James, A.; Jacobs, J.R.; Taylor, J.; Spicer, K.; Bardossy, A.C.; Oakley, L.P.; et al. Presymptomatic SARS-CoV-2 infections and transmission in a skilled nursing facility. N. Engl. J. Med. 2020, 382, 2081–2090. [Google Scholar] [CrossRef]
- Brown, C.S.; Clare, K.; Chand, M.; Andrews, J.; Auckland, C.; Beshir, S.; Choudhry, S.; Davies, K.; Freeman, J.; Gallini, A.; et al. Snapshot PCR surveillance for SARS-CoV-2 in hospital staff in England. J. Infect. 2020, 81, 427–434. [Google Scholar] [CrossRef]
- Folgueira, M.D.; Luczkowiak, J.; Lasala, F.; Pérez-Rivilla, A.; Delgado, R. Prolonged SARS-CoV-2 cell culture replication in respiratory samples from patients with severe COVID-19. Clin. Microbiol. Infect. 2021, 27, 886–891. [Google Scholar] [CrossRef]
- Basile, K.; McPhie, K.; Carter, I.; Alderson, S.; Rahman, H.; Donovan, L.; Kumar, S.; Tran, T.; Ko, D.; Sivaruban, T.; et al. Cell-based Culture Informs Infectivity and Safe De-Isolation Assessments in Patients with Coronavirus Disease 2019. Clin. Infect. Dis. 2020, 73, e2952–e2959. [Google Scholar] [CrossRef]
- Singanayagam, A.; Patel, M.; Charlett, A.; Bernal, J.L.; Saliba, V.; Ellis, J.; Ladhani, S.; Zambon, M.; Gopal, R. Duration of infectiousness and correlation with RT-PCR cycle threshold values in cases of COVID-19, England, January to May 2020. Eurosurveillance 2020, 25, 2001483. [Google Scholar] [CrossRef]
- Gniazdowski, V.; Paul Morris, C.; Wohl, S.; Mehoke, T.; Ramakrishnan, S.; Thielen, P.; Powell, H.; Smith, B.; Armstrong, D.T.; Herrera, M.; et al. Repeated Coronavirus Disease 2019 Molecular Testing: Correlation of Severe Acute Respiratory Syndrome Coronavirus 2 Culture With Molecular Assays and Cycle Thresholds. Clin. Infect. Dis. 2021, 73, e860–e869. [Google Scholar] [CrossRef]
- Young, B.E.; Ong, S.W.X.; Ng, L.F.P.; Anderson, D.E.; Chia, W.N.; Chia, P.Y.; Ang, L.W.; Mak, T.-M.; Kalimuddin, S.; Chai, L.Y.A.; et al. Viral Dynamics and Immune Correlates of Coronavirus Disease 2019 (COVID-19) Severity. Clin. Infect. Dis. 2021, 73, e2932–e2942. [Google Scholar] [CrossRef] [PubMed]
- Rhee, C.; Kanjilal, S.; Baker, M.; Klompas, M. Duration of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Infectivity: When Is It Safe to Discontinue Isolation? Clin. Infect. Dis. 2020, 72, 1467–1474. [Google Scholar] [CrossRef] [PubMed]
- Ladhani, S.N.; Chow, J.Y.; Janarthanan, R.; Fok, J.; Crawley-Boevey, E.; Vusirikala, A.; Fernandez, E.; Perez, M.S.; Tang, S.; Dun-Campbell, K.; et al. Investigation of SARS-CoV-2 outbreaks in six care homes in London, April 2020. EClinicalMedicine 2020, 26, 100533. [Google Scholar] [CrossRef] [PubMed]
- Jaafar, R.; Aherfi, S.; Wurtz, N.; Grimaldier, C.; Van Hoang, T.; Colson, P.; Raoult, D.; La Scola, B. Correlation between 3790 Quantitative Polymerase Chain Reaction–Positives Samples and Positive Cell Cultures, Including 1941 Severe Acute Respiratory Syndrome Coronavirus 2 Isolates. Clin. Infect. Dis. 2020, 72, e921. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Fu, D.; Yang, Q.; Geng, Z.; Xia, J.; Wang, Z.; Wang, L. Low transmission risk of 9 asymptomatic carriers tested positive for both SARS-CoV-2 nucleic acid and serum IgG. J. Infect. 2020, 81, 452–482. [Google Scholar] [CrossRef]
- Wölfel, R.; Corman, V.M.; Guggemos, W.; Seilmaier, M.; Zange, S.; Müller, M.A.; Niemeyer, D.; Jones, T.C.; Vollmar, P.; Rothe, C.; et al. Virological assessment of hospitalized patients with COVID-2019. Nature 2020, 581, 465–469. [Google Scholar] [CrossRef] [Green Version]
- Huang, C.-G.; Lee, K.-M.; Hsiao, M.-J.; Yang, S.-L.; Huang, P.-N.; Gong, Y.-N.; Hsieh, T.-H.; Huang, P.-W.; Lin, Y.-J.; Liu, Y.-C.; et al. Culture-Based Virus Isolation To Evaluate Potential Infectivity of Clinical Specimens Tested for COVID-19. J. Clin. Microbiol. 2020, 58, e01068-20. [Google Scholar] [CrossRef]
- Perera, R.A.; Tso, E.; Tsang, O.T.; Tsang, D.N.; Fung, K.; Leung, Y.W.; Chin, A.W.; Chu, D.K.; Cheng, S.M.; Poon, L.L.; et al. SARS-CoV-2 Virus Culture and Subgenomic RNA for Respiratory Specimens from Patients with Mild Coronavirus Disease. Emerg. Infect. Dis. 2020, 26, 2701–2704. [Google Scholar] [CrossRef]
- Pekosz, A.; Parvu, V.; Li, M.; Andrews, J.C.; Manabe, Y.C.; Kodsi, S.; Gary, D.S.; Roger-Dalbert, C.; Leitch, J.; Cooper, C.K. Antigen-Based Testing but Not Real-Time Polymerase Chain Reaction Correlates With Severe Acute Respiratory Syndrome Coronavirus 2 Viral Culture. Clin. Infect. Dis. 2021, 73, e2861–e2866. [Google Scholar] [CrossRef]
- L’Huillier, A.G.; Torriani, G.; Pigny, F.; Kaiser, L.; Eckerle, I. Culture-Competent SARS-CoV-2 in Nasopharynx of Symptomatic Neonates, Children, and Adolescents. Emerg. Infect. Dis. 2020, 26, 2494–2497. [Google Scholar] [CrossRef]
- Van Kampen, J.J.A.; Van De Vijver, D.A.M.C.; Fraaij, P.L.A.; Haagmans, B.L.; Lamers, M.M.; Okba, N.; Van Den Akker, J.P.C.; Endeman, H.; Gommers, D.A.M.P.J.; Cornelissen, J.J.; et al. Duration and key determinants of infectious virus shedding in hospitalized patients with coronavirus disease-2019 (COVID-19). Nat. Commun. 2021, 12, 267. [Google Scholar] [CrossRef] [PubMed]
- Goyal, A.; Reeves, D.B.; Cardozo-Ojeda, E.F.; Schiffer, J.T.; Mayer, B.T. Wrong person, place and time: Viral load and contact network structure predict SARS-CoV-2 transmission and super-spreading events. Elife 2021, 10, e63537. [Google Scholar] [CrossRef] [PubMed]
- Jones, T.C.; Biele, G.; Mühlemann, B.; Veith, T.; Schneider, J.; Beheim-Schwarzbach, J.; Bleicker, T.; Tesch, J.; Schmidt, M.L.; Sander, L.E.; et al. Estimating infectiousness throughout SARS-CoV-2 infection course. Science 2021, 373, 6551. [Google Scholar] [CrossRef] [PubMed]
- RAG Meeting 08/12/2020: Interpretation and Reporting of SARS-CoV-2 PCR Results. Available online: https://covid-19.sciensano.be/sites/default/files/Covid19/20201208_Advice%20RAG%20Interpretation%20and%20reporting%20of%20COVID%20PCR%20results.pdf (accessed on 15 March 2022).
- RAG Meeting 05/31/2021: Interpretation of RT-PCR Results of SARS-CoV-2 Traces. Available online: https://covid-19.sciensano.be/sites/default/files/Covid19/20210531_Advice_RAG_Interpretation%20of%20traces%20of%20SARS-CoV-2_NL.pdf (accessed on 9 May 2022).
- Laboratory Test Result Healthdata.be Sciensano. Available online: https://covid19lab.healthdata.be/data-collection/laboratorytestresult (accessed on 22 April 2022).
- Meurisse, M.; Lajot, A.; Dupont, Y.; Lesenfants, M.; Klamer, S.; Rebolledo, J.; Lernout, T.; Leroy, M.; Capron, A.; Van Bussel, J.; et al. One year of laboratory-based COVID-19 surveillance system in Belgium: Main indicators and performance of the laboratories (March 2020–2021). Arch. Public Health 2021, 79, 188. [Google Scholar] [CrossRef]
- Planas, D.; Saunders, N.; Maes, P.; Guivel-Benhassine, F.; Planchais, C.; Buchrieser, J.; Bolland, W.H.; Porrot, F.; Staropoli, I.; Lemoine, F.; et al. Considerable escape of SARS-CoV-2 Omicron to antibody neutralization. Nature 2021, 602, 671–675. [Google Scholar] [CrossRef]
- Bruel, T.; Hadjadj, J.; Maes, P.; Planas, D.; Seve, A.; Staropoli, I.; Guivel-Benhassine, F.; Porrot, F.; Bolland, W.-H.; Nguyen, Y.; et al. Serum neutralization of SARS-CoV-2 Omicron sublineages BA.1 and BA.2 in patients receiving monoclonal antibodies. Nat. Med. 2022, 2022, 1–6. [Google Scholar] [CrossRef]
- Wawina-Bokalanga, T.; Martí-Carreras, J.; Vanmechelen, B.; Bloemen, M.; Wollants, E.; Laenen, L.; Cuypers, L.; Beuselinck, K.; Lagrou, K.; André, E.; et al. Genetic diversity and evolution of SARS-CoV-2 in Belgium during the first wave outbreak. bioRxiv 2021. [Google Scholar] [CrossRef]
- Vanmechelen, B.; Logist, A.-S.; Wawina-Bokalanga, T.; Verlinden, J.; Martí-Carreras, J.; Geenen, C.; Slechten, B.; Cuypers, L.; André, E.; Baele, G.; et al. Identification of the First SARS-CoV-2 Lineage B.1.1.529 Virus Detected in Europe. Microbiol. Resour. Announc. 2022, 11, e0116121. [Google Scholar] [CrossRef]
- Janssen, R.; Laenen, L.; Cuypers, L.; Capron, A.; Beuselinck, K.; COVID-19 Genomics Belgium Consortium; Van Ranst, M.; Lagrou, K.; André, E.; Dequeker, E. Achieving quality assurance of high-throughput diagnostic molecular testing during the SARS-CoV-2 pandemic—role of the Belgian national reference laboratory for SARS-CoV-2. 2022; Manuscript in preparation. [Google Scholar]
- Van Vooren, S.; Grayson, J.; Van Ranst, M.; Dequeker, E.; Laenen, L.; Janssen, R.; Gillet, L.; Bureau, F.; Coppieters, W.; Devos, N.; et al. Reliable and Scalable SARS-CoV-2 qPCR Testing at a High Sample Throughput: Lessons Learned from the Belgian Initiative. Life 2022, 12, 159. [Google Scholar] [CrossRef]
- Aksamentov, I.; Roemer, C.; Hodcroft, E.B.; Neher, R.A. Nextclade: Clade assignment, mutation calling and quality control for viral genomes. J. Open Source Softw. 2021, 6, 3773. [Google Scholar] [CrossRef]
- O’Toole, Á.; Scher, E.; Underwood, A.; Jackson, B.; Hill, V.; McCrone, J.T.; Colquhoun, R.; Ruis, C.; Abu-Dahab, K.; Taylor, B.; et al. Assignment of Epidemiological Lineages in an Emerging Pandemic Using the Pangolin Tool. Virus Evol. 2021, 7, veab064. [Google Scholar] [CrossRef] [PubMed]
- Dahdouh, E.; Lázaro-Perona, F.; Romero-Gómez, M.P.; Mingorance, J.; García-Rodriguez, J. Ct values from SARS-CoV-2 diagnostic PCR assays should not be used as direct estimates of viral load. J. Infect. 2021, 82, 414–451. [Google Scholar] [CrossRef] [PubMed]
- Rhoads, D.; Peaper, D.R.; She, R.C.; Nolte, F.S.; Wojewoda, C.M.; Anderson, N.W.; Pritt, B.S. College of American Pathologists (CAP) Microbiology Committee Perspective: Caution Must Be Used in Interpreting the Cycle Threshold (Ct) Value. Clin. Infect. Dis. 2021, 72, e685–e686. [Google Scholar] [CrossRef] [PubMed]
- van Kasteren, P.B.; van Der Veer, B.; van den Brink, S.; Wijsman, L.; de Jonge, J.; van den Brandt, A.; Molenkamp, R.; Reusken, C.B.E.M.; Meijer, A. Comparison of seven commercial RT-PCR diagnostic kits for COVID-19. J. Clin. Virol. 2020, 128, 104412. [Google Scholar] [CrossRef] [PubMed]
- Kanwar, N.; Banerjee, D.; Sasidharan, A.; Abdulhamid, A.; Larson, M.; Lee, B.; Selvarangan, R.; Liesman, R.M. Comparison of Diagnostic Performance of Five Molecular Assays for Detection of SARS-CoV-2. Diagn. Microbiol. Infect. Dis. 2021, 101, 115518. [Google Scholar] [CrossRef]
- Ehret, R.; Breuer, S.; Dhein, J.; Reinhardt, B.; Obermeier, M. Clinical evaluation of the automated Abbott RealTime SARS-CoV-2, Alinity m SARS-CoV-2, and Alinity m Resp-4-Plex assays. J. Virol. Methods 2022, 299, 114338. [Google Scholar] [CrossRef]
- Bentley, E.; Mee, E.T.; Routley, S.; Mate, R.; Fritzsche, M.; Hurley, M.; Le Duff, Y.; Anderson, R.; Hockley, J.; Rigsby, P.; et al. Collaborative Study for the Establishment of a WHO International Standard for SARS-CoV-2 RNA. Expert Committee on Biological Standardization; World Health Organization: Geneva, Switzerland, 2020. [Google Scholar]
- AACC Recommendation for Reporting SARS-CoV-2 Cycle Threshold (Ct) Values. Available online: https://www.aacc.org/science-and-research/covid-19-resources/statements-on-covid-19-testing/aacc-recommendation-for-reporting-sars-cov-2-cycle-threshold-ct-values (accessed on 2 May 2022).
- Sethuraman, N.; Jeremiah, S.S.; Ryo, A. Interpreting Diagnostic Tests for SARS-CoV-2. JAMA 2020, 323, 2249–2251. [Google Scholar] [CrossRef]
- Payne, D.; Newton, D.; Evans, P.; Osman, H.; Baretto, R. Preanalytical issues affecting the diagnosis of COVID-19. J. Clin. Pathol. 2020, 74, 207–208. [Google Scholar] [CrossRef]
- Rahbari, R.; Moradi, N.; Abdi, M. rRT-PCR for SARS-CoV-2: Analytical considerations. Clin. Chim. Acta 2021, 516, 1–7. [Google Scholar] [CrossRef]
- Gand, M.; Vanneste, K.; Thomas, I.; Van Gucht, S.; Capron, A.; Herman, P.; Roosens, N.; De Keersmaecker, S. Deepening of in Silico Evaluation of SARS-CoV-2 Detection RT-qPCR Assays in the Context of New Variants. Genes 2021, 12, 565. [Google Scholar] [CrossRef]
- Wollschläger, P.; Todt, D.; Gerlitz, N.; Pfaender, S.; Bollinger, T.; Sing, A.; Dangel, A.; Ackermann, N.; Korn, K.; Ensser, A.; et al. SARS-CoV-2 N gene dropout and N gene Ct value shift as indicator for the presence of B.1.1.7 lineage in a commercial multiplex PCR assay. Clin. Microbiol. Infect. 2021, 27, 1353.e1–1353.e5. [Google Scholar] [CrossRef] [PubMed]
- Brown, K.A.; Gubbay, J.; Hopkins, J.; Patel, S.; Buchan, S.A.; Daneman, N.; Goneau, L.W. S-Gene Target Failure as a Marker of Variant B.1.1.7 among SARS-CoV-2 Isolates in the Greater Toronto Area, December 2020 to March 2021. JAMA 2021, 325, 2115. [Google Scholar] [CrossRef] [PubMed]
- Clark, C.; Schrecker, J.; Hardison, M.; Taitel, M.S. Validation of reduced S-gene target performance and failure for rapid surveillance of SARS-CoV-2 variants. medRxiv 2022. [Google Scholar] [CrossRef]
- Mendes, R.A. We need increased targeted measures now to slow the spread of omicron. BMJ 2021, 375, n3133. [Google Scholar] [CrossRef]
- UK Health Security Agency. SARS-CoV-2 Variants of Concern and Variants Under Investigation in England, Technical Briefing 32. 17 December 2021. Available online: https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/1042688/RA_Technical_Briefing_32_DRAFT_17_December_2021_2021_12_17.pdf (accessed on 3 May 2022).
- Walsh, K.A.; Jordan, K.; Clyne, B.; Rohde, D.; Drummond, L.; Byrne, P.; Ahern, S.; Carty, P.G.; O’Brien, K.K.; O’Murchu, E.; et al. SARS-CoV-2 detection, viral load and infectivity over the course of an infection. J. Infect. 2020, 81, 357–371. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.-D.; Chang, S.-Y.; Wang, J.-T.; Tsai, M.-J.; Hung, C.-C.; Hsu, C.-L.; Chang, S.-C. Prolonged virus shedding even after seroconversion in a patient with COVID-19. J. Infect. 2020, 81, 318–356. [Google Scholar] [CrossRef]
- Quicke, K.; Gallichotte, E.; Sexton, N.; Young, M.; Janich, A.; Gahm, G.; Carlton, E.J.; Ehrhart, N.; Ebel, G.D. Longitudinal surveillance for SARS-CoV-2 RNA among asymptomatic staff in five Colorado skilled nursing home facilities: Epidemiologic, virologic and sequence analysis. medRxiv 2020. [Google Scholar] [CrossRef]
- Baang, J.H.; Smith, C.; Mirabelli, C.; Valesano, A.L.; Manthei, D.M.; Bachman, M.A.; Wobus, C.E.; Adams, M.; Washer, L.; Martin, E.T.; et al. Prolonged Severe Acute Respiratory Syndrome Coronavirus 2 Replication in an Immunocompromised Patient. J. Infect. Dis. 2020, 223, 23–27. [Google Scholar] [CrossRef]
| Category of SARS-CoV-2 Positivity | SARS-CoV-2 Viral Load (RNA Copies/mL) | Interpretation with Respect to Infectivity |
|---|---|---|
| Very strongly positive | ≥107 | patient is contagious |
| Strongly positive | ≥105–<107 | patient is probably contagious |
| Moderate positive | ≥103–<105 | patient is potentially contagious, unless there is clinical and/or serological evidence of an old, cleared infection |
| Weakly positive | <103 | patient is probably not or no longer contagious if there is also clinical and/or serological evidence of an old, cleared infection |
| SARS-CoV-2 PCR Assay | Number of Laboratories | Targeted SARS-CoV-2 Genes |
|---|---|---|
| Abbott Realtime SARS-CoV-2 assay (m2000) * | 3 | RdRp and N |
| Alinity m SARS-CoV-2 assay * | 6 | RdRp and N |
| Allplex SARS-CoV-2 assay * | 21 | E, RdRp, S and N |
| Allplex SARS-CoV-2/FluA/FluB/RSV assay * | 8 | S, RdRp and N |
| Aries SARS-CoV-2 LDT | 4 | E |
| Aries SARS-CoV-2 RUO * | 10 | ORF1ab and N |
| BD SARS-CoV-2 for BD Max * | 3 | N1 and N2 |
| CE-IVD Perkin Elmer SARS-CoV-2 RT-qPCR * | 2 | ORF1ab and N |
| Cobas SARS-CoV-2 * | 4 | ORF1ab and E |
| GeneFinderTM COVID-19 Plus RealAmp kit * | 12 | RdRp, E and N |
| Ingenius | 2 | RdRp, E and N |
| Liat Cobas SARS-CoV-2 and influenza A/B 1 | 4 | RdRp and N |
| LightMix Modular SARS and Wuhan CoV E-gene kit 2 | 5 | E |
| N1 CDC PCR | 2 | N1 |
| NeuMoDx SARS-CoV-2 * | 2 | N and Nsp2 |
| Powerchek SARS-CoV-2 Real-time PCR kit 1 | 2 | ORF1ab and E |
| R-DiaSARS-CoV-2 * | 3 | E |
| RealStar SARS-CoV-2 RT-PCR kit 1.0 * | 5 | E and S |
| SARS-CoV-2 ELITe MGB kit * | 2 | RdRp and ORF8 |
| SARS-CoV-2 plus ELITe MGB * | 2 | RdRp, ORF8 |
| SARS-CoV-2 N1 + N2 Qiagen 2 | 3 | N1 and N2 |
| TaqPath COVID-19 CE-IVD RT-PCR kit * | 22 | N, S and ORF1ab |
| Viasure SARS-CoV-2 RT PCR 1 | 2 | ORF1ab and N |
| Xpert Xpress SARS-CoV-2 * | 14 | N2 and E |
| Xpert Xpress SARS-CoV-2/Flu/RSV * | 11 | N2 and E |
| PCR Assay | Included (Contributed) Laboratories | SARS-CoV-2 Gene | Mean Ct Value or Equivalent Metric (SD) | ||
|---|---|---|---|---|---|
| 107 RNA Copies/mL * | 105 RNA Copies/mL * | 103 RNA Copies/mL * | |||
| Abbott RealTime SARS-CoV-2 | 3 (3) | RdRp/N | 6.2 (0.34) | 12.7 (0.37) | 19.1 (0.50) |
| Alinity m SARS-CoV-2 | 6 (6) | RdRp/N | 15.0 (0.65) | 21.9 (0.68) | 28.9 (0.75) |
| Allplex SARS-CoV-2 | 15 (19) | E | 17.0 (0.49) | 24.3 (0.72) | 31.5 (1.02) |
| RdRp/S | 17.9 (0.76) | 25.1 (0.96) | 32.2 (1.28) | ||
| N | 17.9 (0.67) | 24.9 (0.80) | 31.9 (1.04) | ||
| Allplex SARS-CoV-2/FluA/FluB/RSV | 7 (8) | S | 15.7 (0.44) | 22.6 (0.55) | 29.6 (0.82) |
| RdRp | 15.3 (0.47) | 22.5 (0.63) | 29.7 (0.89) | ||
| N | 15.4 (0.54) | 22.1 (0.71) | 28.8 (0.99) | ||
| Aries SARS-CoV-2 RUO | 7 (9) | N | 20.3 (0.68) | 27.4 (0.78) | 34.4 (0.96) |
| ORF1ab | 18.3 (0.54) | 25.3 (0.73) | 32.4 (1.06) | ||
| BioGX SARS-CoV-2 BD MAX | 3 (3) | N1 | 16.4 (0.71) | 23.4 (0.56) | 30.4 (0.42) |
| N2 | 16.8 (0.97) | 23.8 (0.99) | 30.9 (1.03) | ||
| Cobas SARS-CoV-2 | 4 (4) | ORF1ab | 17.3 (0.50) | 23.6 (0.52) | 29.8 (0.56) |
| E | 17.6 (0.50) | 24.0 (0.51) | 30.4 (0.55) | ||
| GeneFinder COVID-19 Plus RealAmp | 6 (13) | E | 18.9 (0.37) | 26.1 (0.61) | 33.3 (0.85) |
| RdRp | 18.6 (0.65) | 25.3 (0.74) | 32.1 (0.88) | ||
| N | 19.4 (0.99) | 26.2 (1.00) | 33.0 (1.06) | ||
| NeuMoDx SARS-CoV-2 | 2 (2) | N | 15.7 (0.09) | 22.6 (0.10) | 29.4 (0.29) |
| Nsp2 | 16.4 (0.01) | 23.3 (0.31) | 30.2 (0.62) | ||
| PerkinElmer SARS-CoV-2 RT-PCR | 2 (2) | N | 18.8 (2.34) | 25.1 (1.89) | 31.5 (1.45) |
| ORF1ab | 16.4 (1.82) | 22.7 (1.63) | 28.9 (1.44) | ||
| R-DiaSARS-CoV-2 | 3 (3) | E | 17.6 (0.39) | 23.9 (0.44) | 30.3 (0.68) |
| RealStar SARS-CoV-2 RT-PCR kit 1.0 | 2 (4) | S | 16.3 (1.38) | 23.1 (1.69) | 30.0 (2.00) |
| E | 19.1 (1.10) | 25.6 (1.49) | 32.2 (1.87) | ||
| SARS-CoV-2 ELITe MGB | 2 (2) | RdRp | 18.8 (0.84) | 25.4 (0.84) | 32.0 (0.83) |
| ORF8 | 17.8 (0.86) | 24.6 (0.94) | 31.3 (1.02) | ||
| SARS-CoV-2 plus ELITe MGB | 2 (2) | RdRp/ORF8 | 16.8 (0.66) | 23.8 (0.71) | 30.9 (0.76) |
| TaqPath COVID-19 CE-IVD RT-PCR | 17 (21) | N | 14.5 (0.69) | 21.3 (0.78) | 28.1 (0.95) |
| ORF1ab | 14.1 (0.71) | 20.9 (0.75) | 27.8 (0.95) | ||
| S | 14.1 (0.70) | 20.9 (0.74) | 27.8 (0.92) | ||
| Xpert Xpress SARS-CoV-2 | 7 (13) | N2 | 19.1 (0.48) | 26.2 (0.69) | 33.2 (1.04) |
| E | 16.7 (0.49) | 23.4 (0.51) | 30.1 (0.54) | ||
| Xpert Xpress SARS-CoV-2/Flu/RSV | 9 (11) | N2/E | 16.5 (0.57) | 23.2 (0.92) | 29.9 (1.35) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cuypers, L.; Bode, J.; Beuselinck, K.; Laenen, L.; Dewaele, K.; Janssen, R.; Capron, A.; Lafort, Y.; Paridaens, H.; Bearzatto, B.; et al. Nationwide Harmonization Effort for Semi-Quantitative Reporting of SARS-CoV-2 PCR Test Results in Belgium. Viruses 2022, 14, 1294. https://doi.org/10.3390/v14061294
Cuypers L, Bode J, Beuselinck K, Laenen L, Dewaele K, Janssen R, Capron A, Lafort Y, Paridaens H, Bearzatto B, et al. Nationwide Harmonization Effort for Semi-Quantitative Reporting of SARS-CoV-2 PCR Test Results in Belgium. Viruses. 2022; 14(6):1294. https://doi.org/10.3390/v14061294
Chicago/Turabian StyleCuypers, Lize, Jannes Bode, Kurt Beuselinck, Lies Laenen, Klaas Dewaele, Reile Janssen, Arnaud Capron, Yves Lafort, Henry Paridaens, Bertrand Bearzatto, and et al. 2022. "Nationwide Harmonization Effort for Semi-Quantitative Reporting of SARS-CoV-2 PCR Test Results in Belgium" Viruses 14, no. 6: 1294. https://doi.org/10.3390/v14061294
APA StyleCuypers, L., Bode, J., Beuselinck, K., Laenen, L., Dewaele, K., Janssen, R., Capron, A., Lafort, Y., Paridaens, H., Bearzatto, B., Cauchie, M., Huwart, A., Degosserie, J., Fagnart, O., Overmeire, Y., Rouffiange, A., Vandecandelaere, I., Deffontaine, M., Pilate, T., ... André, E. (2022). Nationwide Harmonization Effort for Semi-Quantitative Reporting of SARS-CoV-2 PCR Test Results in Belgium. Viruses, 14(6), 1294. https://doi.org/10.3390/v14061294








