Evaluation of EPISEQ SARS-CoV-2 and a Fully Integrated Application to Identify SARS-CoV-2 Variants from Several Next-Generation Sequencing Approaches
Abstract
:1. Introduction
2. Materials and Methods
2.1. Patients and Samples
2.2. Sequencing
2.3. Sequencing Data Export and Analysis
2.4. Data Analysis
3. Results
3.1. EPISEQ® SARS-CoV-2 Application
3.2. Validation of EPISEQ SARS-CoV-2
3.2.1. SARS-CoV-2 Genome Coverage
3.2.2. SARS-CoV-2 Variant Call
3.2.3. SARS-CoV-2 Whole-Genome Consensus Sequence
3.2.4. SARS-CoV-2 Spike Protein Mutations
3.3. Comparative Performance of Sequencing Platforms and Kits Using EPISEQ SARS-CoV-2
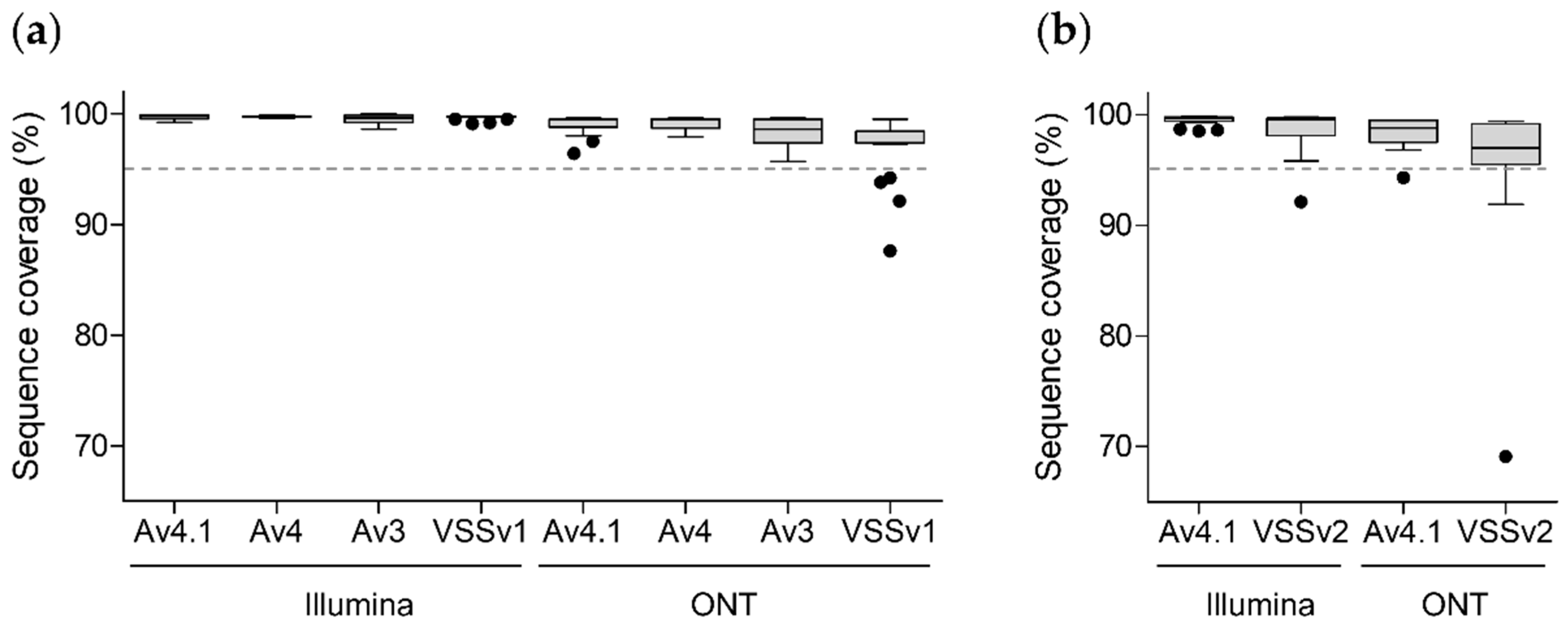
3.3.1. SARS-CoV-2 Genome Coverage
3.3.2. SARS-CoV-2 Variant Call
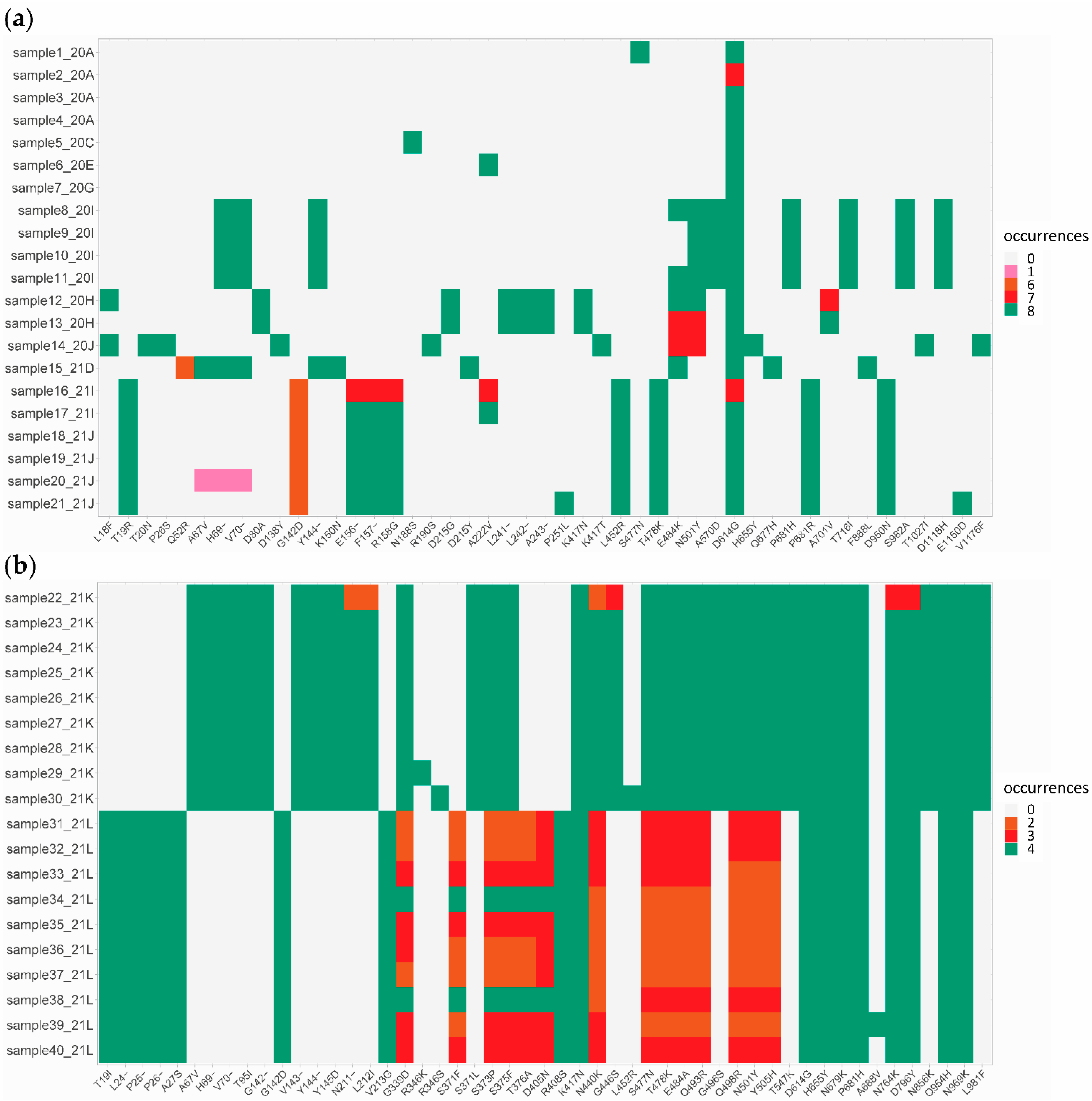
3.3.3. SARS-CoV-2 Amino Acid and Nucleotide Mutations
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Worobey, M.; Pekar, J.; Larsen, B.B.; Nelson, M.I.; Hill, V.; Joy, J.B.; Rambaut, A.; Suchard, M.A.; Wertheim, J.O.; Lemey, P. The Emergence of SARS-CoV-2 in Europe and North America. Science 2020, 370, 564–570. [Google Scholar] [CrossRef] [PubMed]
- Charre, C.; Ginevra, C.; Sabatier, M.; Regue, H.; Destras, G.; Brun, S.; Burfin, G.; Scholtes, C.; Morfin, F.; Valette, M.; et al. Evaluation of NGS-Based Approaches for SARS-CoV-2 Whole Genome Characterisation. Virus Evol. 2020, 6, veaa075. [Google Scholar] [CrossRef] [PubMed]
- Chiara, M.; D’Erchia, A.M.; Gissi, C.; Manzari, C.; Parisi, A.; Resta, N.; Zambelli, F.; Picardi, E.; Pavesi, G.; Horner, D.S.; et al. Next Generation Sequencing of SARS-CoV-2 Genomes: Challenges, Applications and Opportunities. Brief. Bioinform. 2021, 22, 616–630. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.; Chen, Z.; Chen, W.; Chen, X.; Hosseini, M.; Yang, Z.; Li, J.; Ho, D.; Turay, D.; Gheorghe, C.P.; et al. A Benchmarking Study of SARS-CoV-2 Whole-Genome Sequencing Protocols Using COVID-19 Patient Samples. iScience 2021, 24, 102892. [Google Scholar] [CrossRef]
- Wang, M.; Fu, A.; Hu, B.; Tong, Y.; Liu, R.; Liu, Z.; Gu, J.; Xiang, B.; Liu, J.; Jiang, W.; et al. Nanopore Targeted Sequencing for the Accurate and Comprehensive Detection of SARS-CoV-2 and Other Respiratory Viruses. Small 2020, 16, 2002169. [Google Scholar] [CrossRef]
- De Maio, N.; Walker, C.; Borges, R.; Weilguny, L.; Slodkowicz, G.; Goldman, N. Issues with SARS-CoV-2 Sequencing Data. Available online: https://virological.org/t/issues-with-sars-cov-2-sequencing-data/473 (accessed on 15 March 2022).
- Yutao, F. SARS-CoV-2 Samples from Same Early COVID-19 Patients Were Sequenced Repeatedly with Errors Distorting Phylogenetic Trees. Available online: https://virological.org/t/sars-cov-2-samples-from-same-early-covid-19-patients-were-sequenced-repeatedly-with-errors-distorting-phylogenetic-trees/434 (accessed on 15 March 2022).
- Kreier, F. Deltacron: The Story of the Variant That Wasn’t. Nature 2022, 602, 19. [Google Scholar] [CrossRef]
- Maxmen, A. Omicron Blindspots: Why It’s Hard to Track Coronavirus Variants. Nature 2021, 600, 579. [Google Scholar] [CrossRef]
- Pekar, J.; Parker, E.; Havens, J.L.; Suchard, M.A.; Andersen, K.G.; Moshiri, N.; Worobey, M.; Rambaut, A.; Wertheim, J.O. Evidence Against the Veracity of SARS-CoV-2 Genomes Intermediate between Lineages A and B. Available online: https://virological.org/t/evidence-against-the-veracity-of-sars-cov-2-genomes-intermediate-between-lineages-a-and-b/754 (accessed on 15 March 2022).
- Sanderson, T.; Barrett, J.C. Variation at Spike Position 142 in SARS-CoV-2 Delta Genomes Is a Technical Artifact Caused by Dropout of a Sequencing Amplicon. Wellcome Open Res. 2021, 6, 305. [Google Scholar] [CrossRef]
- Davis, J.J.; Long, S.W.; Christensen, P.A.; Olsen, R.J.; Olson, R.; Shukla, M.; Subedi, S.; Stevens, R.; Musser, J.M. Analysis of the ARTIC Version 3 and Version 4 SARS-CoV-2 Primers and Their Impact on the Detection of the G142D Amino Acid Substitution in the Spike Protein. Microbiol. Spectr. 2021, 9, e0180321. [Google Scholar] [CrossRef]
- GISAID—HCov19 Variants. Available online: https://www.gisaid.org/hcov19-variants// (accessed on 29 March 2022).
- Shu, Y.; McCauley, J. GISAID: Global Initiative on Sharing All Influenza Data—From Vision to Reality. Eurosurveill 2017, 22, 30494. [Google Scholar] [CrossRef] [Green Version]
- Tilloy, V.; Cuzin, P.; Leroi, L.; Guérin, E.; Durand, P.; Alain, S. ASPICov: An Automated Pipeline for Identification of SARS-Cov2 Nucleotidic Variants. PLoS ONE 2022, 17, e0262953. [Google Scholar] [CrossRef]
- Wagner, D.D.; Marine, R.L.; Ramos, E.; Ng, T.F.F.; Castro, C.J.; Okomo-Adhiambo, M.; Harvey, K.; Doho, G.; Kelly, R.; Jain, Y.; et al. VPipe: An Automated Bioinformatics Platform for Assembly and Management of Viral Next-Generation Sequencing Data. Microbiol. Spectr. 2022, 10, e0256421. [Google Scholar] [CrossRef]
- Rueca, M.; Giombini, E.; Messina, F.; Bartolini, B.; Di Caro, A.; Capobianchi, M.R.; Gruber, C.E. The Easy-to-Use SARS-CoV-2 Assembler for Genome Sequencing: Development Study. JMIR Bioinform. Biotech. 2022, 3, e31536. [Google Scholar] [CrossRef]
- Farkas, C.; Mella, A.; Turgeon, M.; Haigh, J.J. A Novel SARS-CoV-2 Viral Sequence Bioinformatic Pipeline Has Found Genetic Evidence That the Viral 3′ Untranslated Region (UTR) Is Evolving and Generating Increased Viral Diversity. Front. Microbiol. 2021, 12, 665041. [Google Scholar] [CrossRef]
- Advancing Real-Time Infection Control (ARTIC) Network—ARTIC Network. Available online: https://artic.network/ (accessed on 29 March 2022).
- Gautreau, I. NEBNext® ARTIC Protocols Collection. Available online: https://www.protocols.io/view/nebnext-artic-protocols-collection-bw2apgae (accessed on 29 March 2022).
- Quick, J. NCoV-2019 Sequencing Protocol v2 (GunIt). Available online: https://www.protocols.io/view/ncov-2019-sequencing-protocol-v2-bdp7i5rn (accessed on 29 March 2022).
- Genepii Seqmet. Available online: https://github.com/genepii/seqmet (accessed on 7 April 2022).
- Bal, A.; Simon, B.; Destras, G.; Chalvignac, R.; Semanas, Q.; Oblette, A.; Queromes, G.; Fanget, R.; Regue, H.; Morfin, F.; et al. Detection and Prevalence of SARS-CoV-2 Co-Infections during the Omicron Variant Circulation, France, December 2021–February 2022. Medrxiv 2022. [Google Scholar] [CrossRef]
- Martin, M. Cutadapt Removes Adapter Sequences from High-Throughput Sequencing Reads. EMBnet. J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Li, H. Minimap2: Pairwise Alignment for Nucleotide Sequences. Bioinform. 2018, 34, 3094–3100. [Google Scholar] [CrossRef]
- Picard Tools—By Broad Institute. Available online: http://broadinstitute.github.io/picard/ (accessed on 21 July 2022).
- Mose, L.E.; Perou, C.M.; Parker, J.S. Improved Indel Detection in DNA and RNA via Realignment with ABRA2. Bioinform. 2019, 35, 2966–2973. [Google Scholar] [CrossRef] [Green Version]
- Danecek, P.; Bonfield, J.K.; Liddle, J.; Marshall, J.; Ohan, V.; Pollard, M.O.; Whitwham, A.; Keane, T.; McCarthy, S.A.; Davies, R.M.; et al. Twelve Years of SAMtools and BCFtools. GigaScience 2021, 10, giab008. [Google Scholar] [CrossRef]
- Garrison, E.; Marth, G. Haplotype-Based Variant Detection from Short-Read Sequencing. arXiv 2012, arXiv:1207.3907. [Google Scholar] [CrossRef]
- Tan, A.; Abecasis, G.R.; Kang, H.M. Unified Representation of Genetic Variants. Bioinform 2015, 31, 2202–2204. [Google Scholar] [CrossRef]
- Nextstrain—Nextclade. Available online: https://github.com/nextstrain/nextclade (accessed on 30 March 2022).
- Centre for Genomic Pathogen Surveillance—Phylogenetic Assignment of Named Global Outbreak LINeages (Pangolin). Available online: https://github.com/cov-lineages/pangolin (accessed on 30 March 2022).
- Aksamentov, I.; Roemer, C.; Hodcroft, E.B.; Neher, R.A. Nextclade: Clade Assignment, Mutation Calling and Quality Control for Viral Genomes. J. Open Source Softw. 2021, 6, 3773. [Google Scholar] [CrossRef]
- Krzywinski, M.; Altman, N. Visualizing Samples with Box Plots. Nat. Methods 2014, 11, 119–120. [Google Scholar] [CrossRef]
- Chen, S.; He, C.; Li, Y.; Li, Z.; Melançon, C.E., III. A Computational Toolset for Rapid Identification of SARS-CoV-2, Other Viruses and Microorganisms from Sequencing Data. Brief. Bioinform. 2021, 22, 924–935. [Google Scholar] [CrossRef]
- Li, H. Aligning Sequence Reads, Clone Sequences and Assembly Contigs with BWA-MEM. arXiv 2013, arXiv:1303.3997. [Google Scholar] [CrossRef]
- Grubaugh, N.D.; Gangavarapu, K.; Quick, J.; Matteson, N.L.; De Jesus, J.G.; Main, B.J.; Tan, A.L.; Paul, L.M.; Brackney, D.E.; Grewal, S.; et al. An Amplicon-Based Sequencing Framework for Accurately Measuring Intrahost Virus Diversity Using PrimalSeq and IVar. Genome. Biol. 2019, 20, 8. [Google Scholar] [CrossRef] [Green Version]
- Loman, N.; Rowe, W.; Rambaut, A. ARTIC-NCoV-BioinformaticsSOP-v1.1.0. Available online: https://artic.network/ncov-2019/ncov2019-bioinformatics-sop.html (accessed on 28 April 2022).
- ECDC SARS-CoV-2 Variants of Concern (VOC). Available online: https://www.ecdc.europa.eu/en/covid-19/variants-concern (accessed on 28 April 2022).
- Sanderson, T.; De Maio, N.; Hinrichs, A.S.; de Bernardi Schneider, A.; Walker, C.; Goldman, N.; Turakhia, Y.; Lanfear, R.; Corbett-Detig, R. Issues with SARS-CoV-2 Sequencing Data—Systematic Errors Associated with Some Implementations of ARTIC V4 and a Fast Workflow to Prescreen Samples for New Problematic Sites. Available online: https://virological.org/t/issues-with-sars-cov-2-sequencing-data/473 (accessed on 15 March 2022).
- New England Biolabs SARS-CoV-2 Lineage Variant Summary. Available online: https://primer-monitor.neb.com/lineages (accessed on 29 March 2022).
- Wilkinson, S.; Groves, N.; Quick, J. Loman Erroneous Mutations Associated with 64_L-60_R Primer-Dimer in ARTIC 4/4.1—Laboratory. Available online: https://community.artic.network/t/erroneous-mutations-associated-with-64-l-60-r-primer-dimer-in-artic-4-4-1/419 (accessed on 29 April 2022).
- Lambisia, A.W.; Mohammed, K.S.; Makori, T.O.; Ndwiga, L.; Mburu, M.W.; Morobe, J.M.; Moraa, E.O.; Musyoki, J.; Murunga, N.; Mwangi, J.N.; et al. Optimization of the SARS-CoV-2 ARTIC Network V4 Primers and Whole Genome Sequencing Protocol. Front. Med. 2022, 9, 836728. [Google Scholar] [CrossRef]
- W-L ProblematicSites_SARS-CoV2—Human-Friendly Version of the Vcf File. Available online: https://github.com/W-L/ProblematicSites_SARS-CoV2 (accessed on 17 May 2022).
- ARTIC Network SARS-CoV-2 Version 4 Scheme Release—Laboratory. Available online: https://community.artic.network/t/sars-cov-2-version-4-scheme-release/312 (accessed on 29 April 2022).
- ARTIC Network SARS-CoV-2 V4.1 Update for Omicron Variant—Laboratory. Available online: https://community.artic.network/t/sars-cov-2-v4-1-update-for-omicron-variant/342 (accessed on 29 April 2022).


| Sequencer | Primer Pool | Kits |
|---|---|---|
| MiSeq (Illumina) | ARTIC v3 | NEBNext® ARTIC SARS-CoV-2 Library Prep Kit (Illumina) (NEB, E7650) |
| ARTIC v4 | NEBNext® ARTIC SARS-CoV-2 Library Prep Kit (Illumina) (NEB, E7650); ARTIC V4 NCOV-2019 Panel (IDT, 10008554) | |
| ARTIC v4.1 | NEBNext® ARTIC SARS-CoV-2 FS Library Prep Kit (Illumina) (NEB, E7658); ARTIC V4.1 NCOV-2019 Panel (IDT, 10011442) | |
| VSS v1 | NEBNext® ARTIC SARS-CoV-2 FS Library Prep Kit (Illumina) (NEB, E7658) | |
| VSS v2 | NEBNext® ARTIC SARS-CoV-2 FS Library Prep Kit (Illumina) (NEB, E7658) | |
| GridION Mk1 (Oxford Nanopore Technologies) | ARTIC v3 | NEBNext® ARTIC SARS-CoV-2 Companion Kit (ONT) (NEB, E7660) |
| ARTIC v4 | NEBNext® ARTIC SARS-CoV-2 Companion Kit (ONT) (NEB, E7660); ARTIC V4 NCOV-2019 Panel (IDT, 10008554) | |
| ARTIC v4.1 | NEBNext® ARTIC SARS-CoV-2 Companion Kit (ONT) (NEB, E7660); ARTIC V4.1 NCOV-2019 Panel (IDT, 10011442) | |
| VSS v1 | NEBNext® ARTIC SARS-CoV-2 Companion Kit (ONT) (NEB, E7660) | |
| VSS v2 | NEBNext® ARTIC SARS-CoV-2 Companion Kit (ONT) (NEB, E7660) |
| Nextstrain Clade | Pango Lineage | |||
|---|---|---|---|---|
| Sequencing Kit | n/N 1 | % [95% CI] | n/N 1 | % [95% CI] |
| ARTIC v3 | 527/527 2 | 100.0% [99.3–100.0] | 525/527 | 99.6% [98.6–99.9] |
| ARTIC v4 | 316/316 3 | 100.0% [98.8–100.0] | 315/316 | 99.7% [98.3–99.9] |
| ARTIC v4.1 | 517/519 4 | 99.6% [98.6–99.9] | 512/519 | 98.7% [97.2–99.5] |
| Total | 1360/1362 | 99.9% [99.5–100.0] | 1352/1362 | 99.3% [98.7–99.7] |
| Sequencing Kit | 0 SNP n/N 1 (%) | 1 SNP n/N 1 (%) | 2 SNPs n/N 1 (%) | >2 SNPs n/N 1 (%) |
|---|---|---|---|---|
| ARTIC v3 | 524/527 (99.4%) | 3/527 (0.6%) | 0/527 (0.0%) | 0/527 (0.0%) |
| ARTIC v4 | 253/316 (80.1%) | 55/316 (17.4%) | 8/316 (2.5%) | 0/316 (0.0%) |
| ARTIC v4.1 | 363/519 (69.9%) | 137/519 (26.4%) | 15/519 (2.9%) | 4/519 (0.8%) |
| Total | 1140/1362 (83.7%) | 195/1362 (14.3%) | 23/1362 (1.7%) | 4/1362 (0.3%) |
| Sequencing Kit | Spike Mutations, n/N 1 (%) |
|---|---|
| ARTIC v3 | 527/527 (100.0%) |
| ARTIC v4 | 315/316 (99.7%) |
| ARTIC v4.1 | 510/519 (98.3%) |
| Total | 1352/1362 (99.3%) |
| SARS-CoV-2 Samples | Nextstrain Clade | Pango Lineage |
|---|---|---|
| Pre-omicron variants 1 | 21/21 (100.0%) | 21/21 (100.0%) |
| Omicron variants 2 | 19/19 (100.0%) | 19/19 (100.0%) |
| Total | 40/40 (100.0%) | 40/40 (100.0%) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mugnier, N.; Griffon, A.; Simon, B.; Rambaud, M.; Regue, H.; Bal, A.; Destras, G.; Tournoud, M.; Jaillard, M.; Betraoui, A.; et al. Evaluation of EPISEQ SARS-CoV-2 and a Fully Integrated Application to Identify SARS-CoV-2 Variants from Several Next-Generation Sequencing Approaches. Viruses 2022, 14, 1674. https://doi.org/10.3390/v14081674
Mugnier N, Griffon A, Simon B, Rambaud M, Regue H, Bal A, Destras G, Tournoud M, Jaillard M, Betraoui A, et al. Evaluation of EPISEQ SARS-CoV-2 and a Fully Integrated Application to Identify SARS-CoV-2 Variants from Several Next-Generation Sequencing Approaches. Viruses. 2022; 14(8):1674. https://doi.org/10.3390/v14081674
Chicago/Turabian StyleMugnier, Nathalie, Aurélien Griffon, Bruno Simon, Maxence Rambaud, Hadrien Regue, Antonin Bal, Gregory Destras, Maud Tournoud, Magali Jaillard, Abel Betraoui, and et al. 2022. "Evaluation of EPISEQ SARS-CoV-2 and a Fully Integrated Application to Identify SARS-CoV-2 Variants from Several Next-Generation Sequencing Approaches" Viruses 14, no. 8: 1674. https://doi.org/10.3390/v14081674






