Comprehensive Identification and Characterization of HML-9 Group in Chimpanzee Genome
Abstract
:1. Introduction
2. Materials and Methods
2.1. Identification, Localization, and Genomic Distribution of ERV-K HML-9 Proviruses and Solo LTR Elements
2.2. Structural Analyses
2.3. Phylogenetic Analyses
2.4. Estimated Time of Integration
2.5. Conserved Motif Analysis
2.6. Functional Prediction of cis-Regulatory Regions and Enrichment Analysis
2.7. PBS Type
3. Results
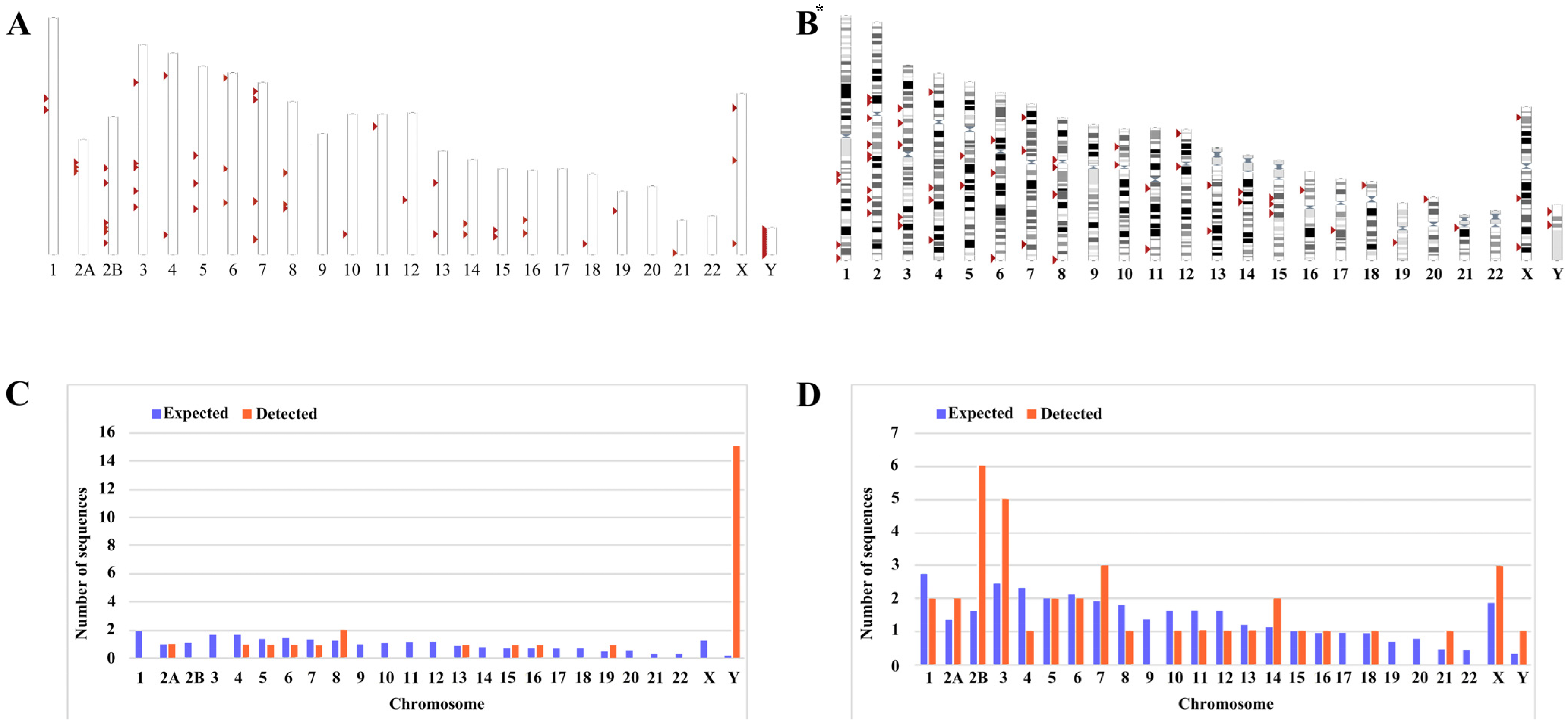
3.1. HML-9 Element Identification, Localization, and Distribution in Chimpanzee Genome Assembly January 2018 (Clint_PTRv2/panTro6)
3.2. Structural Characterization of HML-9 Proviral Sequences
3.3. Phylogenetic Analyses
3.4. Estimated Time of Integration
3.5. Conserved Motif Analysis
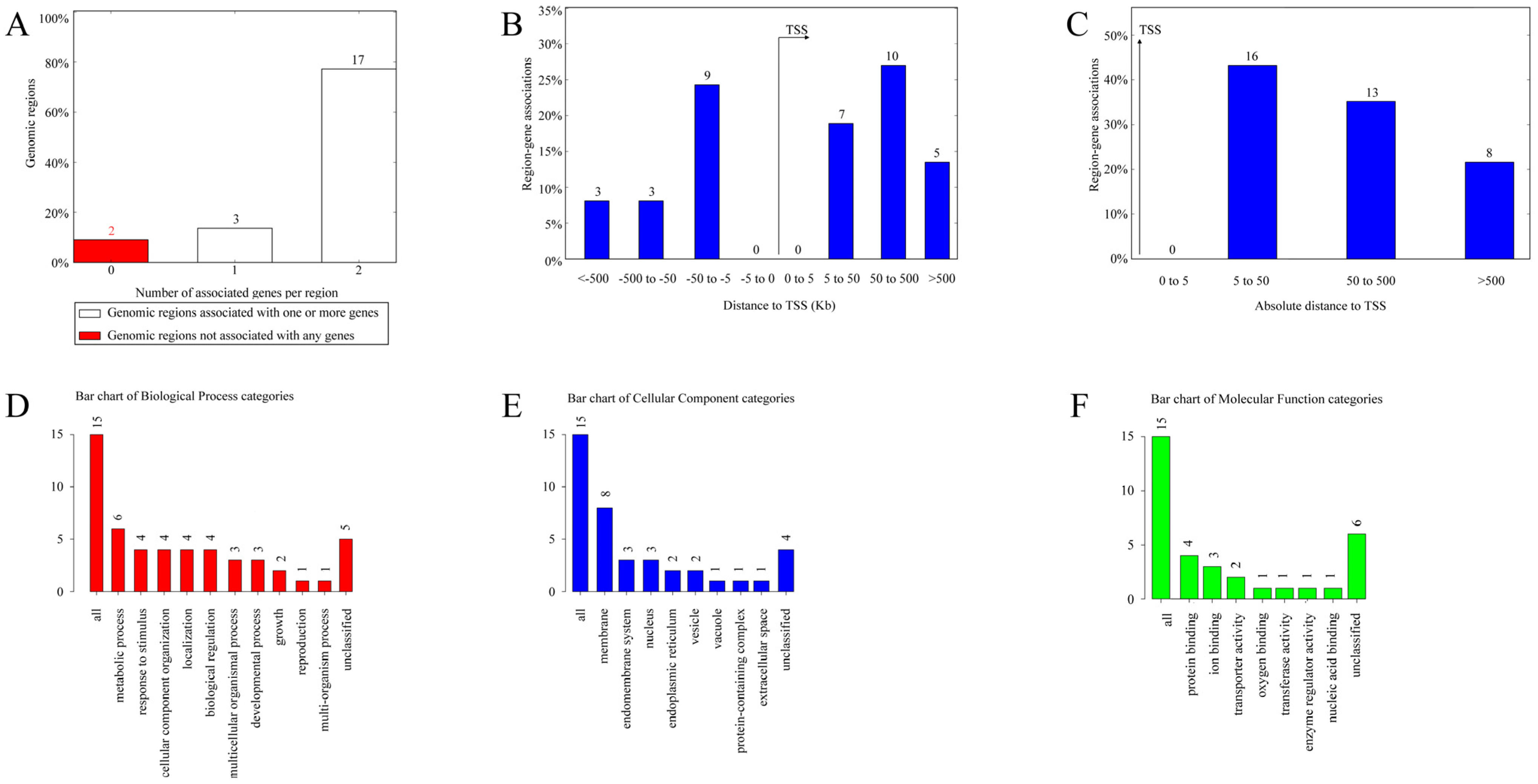
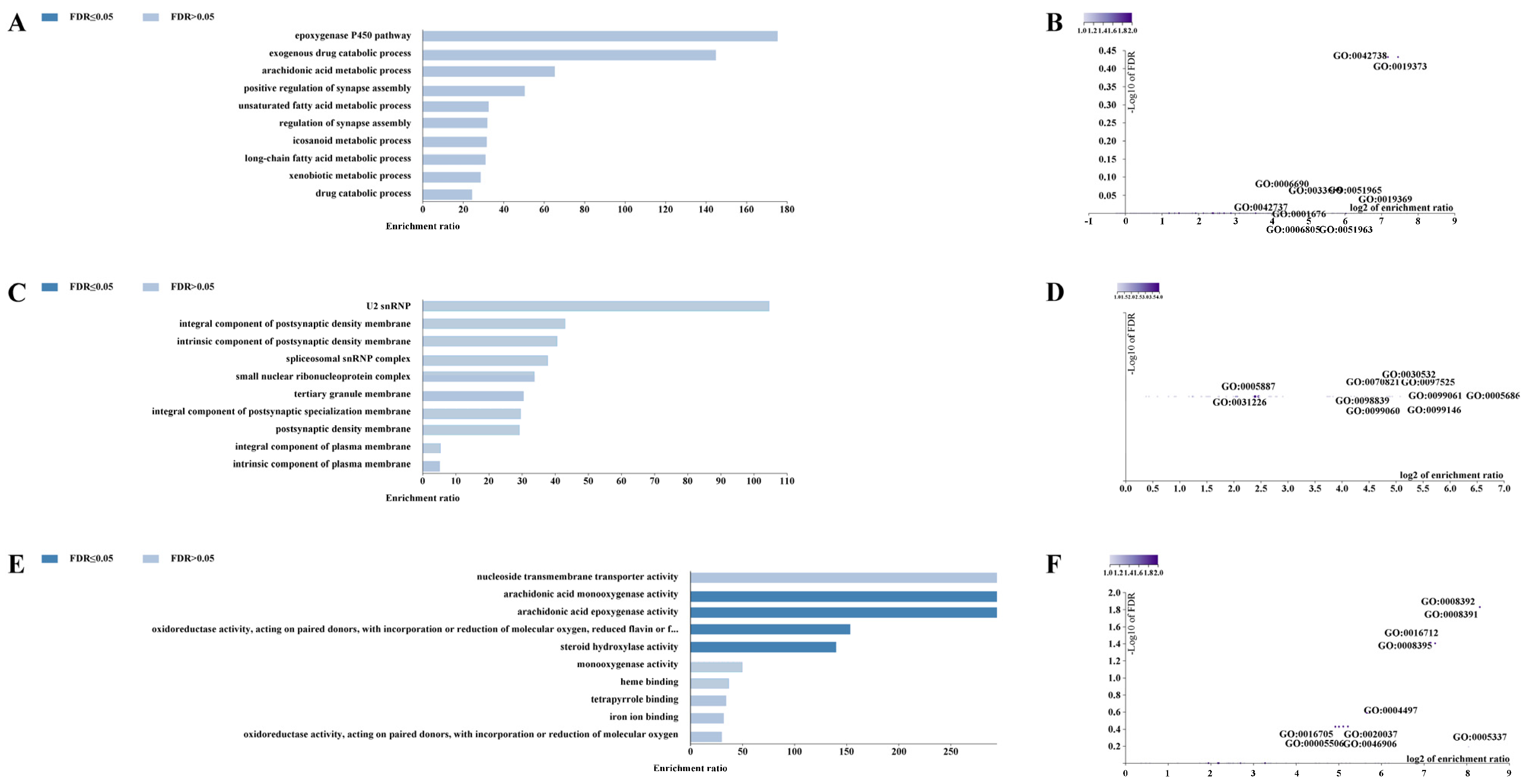
3.6. Functional Prediction of cis-Regulatory Regions and Enrichment Analysis
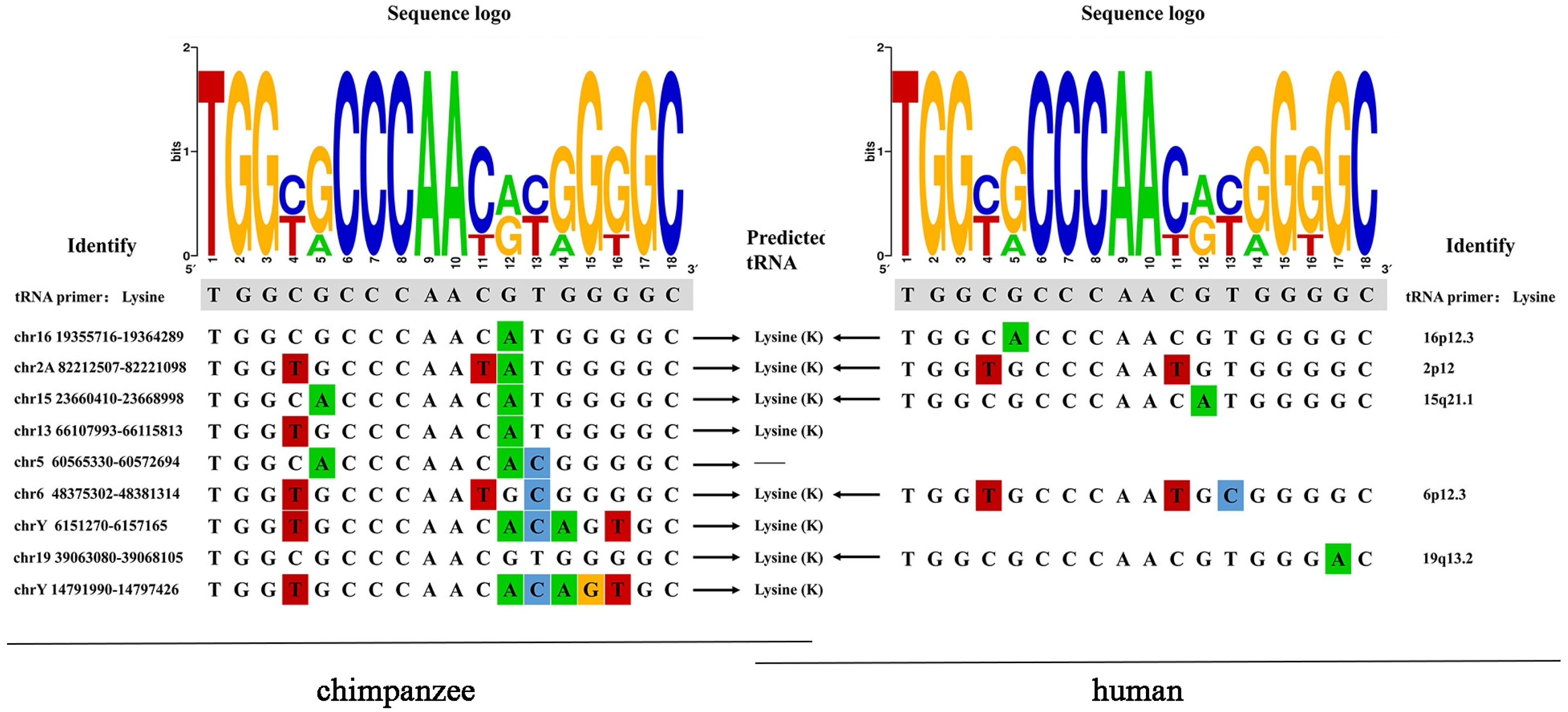
3.7. PBS Type
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Weiss, R.A. The discovery of endogenous retroviruses. Retrovirology 2006, 3, 67. [Google Scholar] [CrossRef] [PubMed]
- Johnson, W.E. Origins and evolutionary consequences of ancient endogenous retroviruses. Nat. Rev. Microbiol. 2019, 17, 355–370. [Google Scholar] [CrossRef] [PubMed]
- Attermann, A.; Bjerregaard, A.-M.; Saini, S.; Grønbæk, K.; Hadrup, S. Human endogenous retroviruses and their implication for immunotherapeutics of cancer. Ann. Oncol. 2018, 29, 2183–2191. [Google Scholar] [CrossRef] [PubMed]
- Michael Bock Stoye, J.P. Endogenous retroviruses and the human germline. Curr. Opin. Genet. Dev. 2000, 10, 651–655. [Google Scholar]
- Magiorkinis, G.; Gifford, R.J.; Katzourakis, A.; De Ranter, J.; Belshaw, R. Env-less endogenous retroviruses are genomic superspreaders. Proc. Natl. Acad. Sci. USA 2012, 109, 7385–7390. [Google Scholar] [CrossRef] [PubMed]
- Mager, D.L.; Stoye, J.P. Mammalian Endogenous Retroviruses. Microbiol. Spectr. 2015, 3, MDNA3-0009-2014. [Google Scholar]
- Vargiu, L.; Rodriguez-Tomé, P.; Sperber, G.O.; Cadeddu, M.; Grandi, N.; Blikstad, V.; Tramontano, E.; Blomberg, J. Classification and characterization of human endogenous retroviruses; mosaic forms are common. Retrovirology 2016, 13, 7. [Google Scholar] [CrossRef]
- Jern, P.; Sperber, G.O.; Blomberg, J. Use of endogenous retroviral sequences (ERVs) and structural markers for retroviral phylogenetic inference and taxonomy. Retrovirology 2005, 2, 50. [Google Scholar] [CrossRef]
- Jern, P.; Coffin, J.M. Effects of retroviruses on host genome function. Annu. Rev. Genet. 2008, 42, 709–732. [Google Scholar] [CrossRef]
- Leung, D.C.; Lorincz, M.C. Silencing of endogenous retroviruses: When and why do histone marks predominate? Trends Biochem. Sci. 2012, 37, 127–133. [Google Scholar] [CrossRef]
- Vargiu, L.; Rodriguez-Tomé, P.; Sperber, G.O.; Cadeddu, M.; Grandi, N.; Blikstad, V.; Tramontano, E.; Blomberg, J. Active endogenous retroviral elements in human pluripotent stem cells play a role in regulating host gene expression. Nucleic Acids Res. 2022, 50, 4959–4973. [Google Scholar]
- Brattås, P.L.; Jönsson, M.E.; Fasching, L.; Wahlestedt, J.N.; Shahsavani, M.; Falk, R.; Falk, A.; Jern, P.; Parmar, M.; Jakobsson, J. TRIM28 Controls a Gene Regulatory Network Based on Endogenous Retroviruses in Human Neural Progenitor Cells. Cell Rep. 2017, 18, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Fasching, L.; Kapopoulou, A.; Sachdeva, R.; Petri, R.; Jönsson, M.E.; Männe, C.; Turelli, P.; Jern, P.; Cammas, F.; Trono, D.; et al. TRIM28 represses transcription of endogenous retroviruses in neural progenitor cells. Cell Rep. 2015, 10, 20–28. [Google Scholar] [CrossRef] [PubMed]
- Zhuo, X.; Du, A.Y.; Pehrsson, E.C.; Li, D.; Wang, T. Epigenomic differences in the human and chimpanzee genomes are associated with structural variation. Genome Res. 2020, 31, 279–290. [Google Scholar] [CrossRef] [PubMed]
- Thomas, J.; Perron, H.; Feschotte, C. Variation in proviral content among human genomes mediated by LTR recombination. Mob. DNA 2018, 9, 36. [Google Scholar] [CrossRef] [PubMed]
- Copeland, N.G.; Hutchison, K.W.; Jenkins, N.A. Excision of the DBA ecotropic provirus in dilute coat-color revertants of mice occurs by homologous recombination involving the viral LTRs. Cell 1983, 33, 379–387. [Google Scholar] [CrossRef] [PubMed]
- Benachenhou, F.; Jern, P.; Oja, M.; Sperber, G.; Blikstad, V.; Somervuo, P.; Kaski, S.; Blomberg, J. Evolutionary conservation of orthoretroviral long terminal repeats (LTRs) and ab initio detection of single LTRs in genomic data. PLoS ONE 2009, 4, e5179. [Google Scholar] [CrossRef]
- Benachenhou, F.; O Sperber, G.; Bongcam-Rudloff, E.; Andersson, G.; Boeke, J.D.; Blomberg, J. Conserved structure and inferred evolutionary history of long terminal repeats (LTRs). Mob. DNA 2013, 4, 5. [Google Scholar] [CrossRef] [PubMed]
- Ruvolo, M. Molecular phylogeny of the hominoids: Inferences from multiple independent DNA sequence data sets. Mol. Biol. Evol. 1997, 14, 248–265. [Google Scholar] [CrossRef] [PubMed]
- Goodman, M.; Porter, C.A.; Czelusniak, J.; Page, S.L.; Schneider, H.; Shoshani, J.; Gunnell, G.; Groves, C.P. Toward a phylogenetic classification of Primates based on DNA evidence complemented by fossil evidence. Mol. Phylogenet. Evol. 1998, 9, 585–598. [Google Scholar] [CrossRef]
- Satta, Y.; Klein, J.; Takahata, N. DNA archives and our nearest relative: The trichotomy problem revisited. Mol. Phylogenet. Evol. 2000, 14, 259–275. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.C.; Li, W.H. Genomic divergences between humans and other hominoids and the effective population size of the common ancestor of humans and chimpanzees. Am. J. Hum. Genet. 2001, 68, 444–456. [Google Scholar] [CrossRef] [PubMed]
- Analysis Consortium. Initial sequence of the chimpanzee genome and comparison with the human genome. Nature 2005, 437, 69–87. [Google Scholar] [CrossRef] [PubMed]
- Kazazian, H.H.; Wong, C.; Youssoufian, H.; Scott, A.F.; Phillips, D.G.; Antonarakis, S.E. Haemophilia A resulting from de novo insertion of L1 sequences represents a novel mechanism for mutation in man. Nature 1988, 332, 164–166. [Google Scholar] [CrossRef] [PubMed]
- Brookfield, J.F. The ecology of the genome—Mobile DNA elements and their hosts. Nat. Rev. Genet. 2005, 6, 128–136. [Google Scholar] [CrossRef] [PubMed]
- Deniz, Ö.; Ahmed, M.; Todd, C.D.; Rio-Machin, A.; Dawson, M.A.; Branco, M.R. Endogenous retroviruses are a source of enhancers with oncogenic potential in acute myeloid leukaemia. Nat. Commun. 2020, 11, 3506. [Google Scholar] [CrossRef] [PubMed]
- Lukic, S.; Nicolas, J.C.; Levine, A.J. The diversity of zinc-finger genes on human chromosome 19 provides an evolutionary mechanism for defense against inherited endogenous retroviruses. Cell Death Differ. 2014, 21, 381–387. [Google Scholar] [CrossRef]
- Song, Y.; Hou, G.; Diep, J.; Ooi, Y.S.; Akopyants, N.S.; Beverley, S.M.; E Carette, J.; Greenberg, H.B.; Ding, S. Inhibitor of growth protein 3 epigenetically silences endogenous retroviral elements and prevents innate immune activation. Nucleic Acids Res. 2021, 49, 12706–12715. [Google Scholar] [CrossRef] [PubMed]
- Tokuyama, M.; Gunn, B.M.; Venkataraman, A.; Kong, Y.; Kang, I.; Rakib, T.; Townsend, M.J.; Costenbader, K.H.; Alter, G.; Iwasaki, A. Antibodies against human endogenous retrovirus K102 envelope activate neutrophils in systemic lupus erythematosus. J. Exp. Med. 2021, 218, e20191766. [Google Scholar] [CrossRef]
- Polavarapu, N.; Bowen, N.J.; Mcdonald, J.F. Identification, characterization and comparative genomics of chimpanzee endogenous retroviruses. Genome Biol. 2006, 7, R51. [Google Scholar] [CrossRef]
- Rj, B. Divergence between samples of chimpanzee and human DNA sequences is 5%, counting indels. Proc. Natl. Acad. Sci. USA 2002, 99, 13633–13635. [Google Scholar]
- Tsankov, A.M.; Thompson, D.A.; Socha, A.; Regev, A.; Rando, O.J. The role of nucleosome positioning in the evolution of gene regulation. PLoS Biol. 2010, 8, e1000414. [Google Scholar] [CrossRef] [PubMed]
- Ho, M.C.W.; Johnsen, H.; Goetz, S.E.; Schiller, B.J.; Bae, E.; Tran, D.A.; Shur, A.S.; Allen, J.M.; Rau, C.; Bender, W.; et al. Functional evolution of cis-regulatory modules at a homeotic gene in Drosophila. PLoS Genet. 2009, 5, e1000709. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.; Ludvik, A.E.; Arlotto, M.E.; Hayes, M.G.; Armstrong, L.L.; Scholtens, D.M.; Brown, C.D.; Newgard, C.B.; Becker, T.C.; Layden, B.T.; et al. Coordinated regulatory variation associated with gestational hyperglycaemia regulates expression of the novel hexokinase HKDC1. Nat. Commun. 2015, 6, 6069. [Google Scholar] [CrossRef]
- Zhang, L.; Reed, R.D. Genome editing in butterflies reveals that spalt promotes and Distal-less represses eyespot colour patterns. Nat. Commun. 2016, 7, 11769. [Google Scholar] [CrossRef] [PubMed]
- Landeen, E.L.; Muirhead, C.A.; Wright, L.; Meiklejohn, C.D.; Presgraves, D.C. Sex Chromosome-wide Transcriptional Suppression and Compensatory Cis-Regulatory Evolution Mediate Gene Expression in the Drosophila Male Germline. PLoS Biol. 2016, 14, e1002499. [Google Scholar] [CrossRef]
- Prescott, S.L.; Srinivasan, R.; Marchetto, M.C.; Grishina, I.; Narvaiza, I.; Selleri, L.; Gage, F.H.; Swigut, T.; Wysocka, J. Enhancer divergence and cis-regulatory evolution in the human and chimp neural crest. Cell 2015, 163, 68–83. [Google Scholar] [CrossRef]
- Trizzino, M.; Park, Y.; Holsbach-Beltrame, M.; Aracena, K.; Mika, K.; Caliskan, M.; Perry, G.H.; Lynch, V.J.; Brown, C.D. Transposable elements are the primary source of novelty in primate gene regulation. Genome Res. 2017, 27, 1623–1633. [Google Scholar] [CrossRef]
- Hughes, J.F.; Coffin, J.M. Human endogenous retroviral elements as indicators of ectopic recombination events in the primate genome. Genetics 2005, 171, 1183–1194. [Google Scholar] [CrossRef]
- Mun, S.; Lee, J.; Kim, Y.J.; Kim, H.S.; Han, K. Chimpanzee-specific endogenous retrovirus generates genomic variations in the chimpanzee genome. PLoS ONE 2014, 9, e101195. [Google Scholar] [CrossRef]
- Flockerzi, A.; Burkhardt, S.; Schempp, W.; Meese, E.; Mayer, J. Human endogenous retrovirus HERV-K14 families: Status, variants, evolution, and mobilization of other cellular sequences. J. Virol. 2005, 79, 2941–2949. [Google Scholar] [CrossRef] [PubMed]
- Sin, H.S.; Koh, E.; Kim, D.S.; Murayama, M.; Sugimoto, K.; Maeda, Y.; Yoshida, A.; Namiki, M. Human endogenous retrovirus K14C drove genomic diversification of the Y chromosome during primate evolution. J. Hum. Genet. 2010, 55, 717–725. [Google Scholar] [CrossRef] [PubMed]
- Sin, H.S.; Koh, E.; Taya, M.; IIjima, M.; Sugimoto, K.; Maeda, Y.; Yoshida, A.; Iwamoto, T.; Namiki, M. A novel Y chromosome microdeletion with the loss of an endogenous retrovirus related, testis specific transcript in AZFb region. J. Urol. 2011, 186, 1545–1552. [Google Scholar] [CrossRef]
- Jansz, N.; Faulkner, G.J. Endogenous retroviruses in the origins and treatment of cancer. Genome Biol. 2021, 22, 147. [Google Scholar] [CrossRef] [PubMed]
- Marchi, E.; Kanapin, A.; Magiorkinis, G.; Belshaw, R. Unfixed endogenous retroviral insertions in the human population. J. Virol. 2014, 88, 9529–9537. [Google Scholar] [CrossRef]
- Jia, L.; Liu, M.; Yang, C.; Li, H.; Liu, Y.; Han, J.; Zhai, X.; Wang, X.; Li, T.; Li, J.; et al. Comprehensive identification and characterization of the HERV-K (HML-9) group in the human genome. Retrovirology 2022, 19, 11. [Google Scholar] [CrossRef] [PubMed]
- Grandi, N.; Cadeddu, M.; Pisano, M.P.; Esposito, F.; Blomberg, J.; Tramontano, E. Identification of a novel HERV-K(HML10): Comprehensive characterization and comparative analysis in non-human primates provide insights about HML10 proviruses structure and diffusion. Mob. DNA 2017, 8, 15. [Google Scholar] [CrossRef]
- Pisano, M.P.; Grandi, N.; Cadeddu, M.; Blomberg, J.; Tramontano, E. Comprehensive Characterization of the Human Endogenous Retrovirus HERV-K(HML-6) Group: Overview of Structure, Phylogeny, and Contribution to the Human Genome. J. Virol. 2019, 93, e00110-19. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, R.P.; Wildschutte, J.H.; Russo, C.; Coffin, J.M. Identification, characterization, and comparative genomic distribution of the HERV-K (HML-2) group of human endogenous retroviruses. Retrovirology 2011, 8, 90. [Google Scholar] [CrossRef]
- Brady, T.; Lee, Y.N.; Ronen, K.; Malani, N.; Berry, C.C.; Bieniasz, P.D.; Bushman, F.D. Integration target site selection by a resurrected human endogenous retrovirus. Genes Dev. 2009, 23, 633–642. [Google Scholar] [CrossRef]
- Lebedev, Y.B.; Belonovitch, O.S.; Zybrova, N.V.; Khil, P.P.; Kurdyukov, S.G.; Vinogradova, T.V.; Hunsmann, G.; Sverdlov, E.D. Differences in HERV-K LTR insertions in orthologous loci of humans and great apes. Gene 2000, 247, 265–277. [Google Scholar] [CrossRef] [PubMed]
- Grandi, N.; Cadeddu, M.; Blomberg, J.; Tramontano, E. Contribution of type W human endogenous retroviruses to the human genome: Characterization of HERV-W proviral insertions and processed pseudogenes. Retrovirology 2016, 13, 67. [Google Scholar] [CrossRef] [PubMed]
- Grandi, N.; Pisano, M.P.; Pessiu, E.; Scognamiglio, S.; Tramontano, E. HERV-K(HML7) Integrations in the Human Genome: Comprehensive Characterization and Comparative Analysis in Non-Human Primates. Biology 2021, 10, 439. [Google Scholar] [CrossRef]
- Waterston, R.H.; Lindblad-Toh, K.; Birney, E.; Rogers, J.; Abril, J.F.; Agarwal, P.; Agarwala, R.; Ainscough, R.; Alexandersson, M.; An, P.; et al. Initial sequencing and comparative analysis of the mouse genome. Nature 2002, 420, 520–562. [Google Scholar] [PubMed]
- Venter, J.C.; Smith, H.O.; Adams, M.D. The sequence of the human genome. Science 2001, 291, 1304–1351. [Google Scholar] [CrossRef] [PubMed]
- Thompson, P.J.; Macfarlan, T.S.; Lorincz, M.C. Long Terminal Repeats: From Parasitic Elements to Building Blocks of the Transcriptional Regulatory Repertoire. Mol. Cell 2016, 62, 766–776. [Google Scholar] [CrossRef]
- Feschotte, C.; Gilbert, C. Endogenous viruses: Insights into viral evolution and impact on host biology. Nat. Rev. Genet. 2012, 13, 283–296. [Google Scholar] [CrossRef] [PubMed]
- Chuong, E.B.; Elde, N.C.; Feschotte, C. Regulatory activities of transposable elements: From conflicts to benefits. Nat. Rev. Genet. 2017, 18, 71–86. [Google Scholar] [CrossRef]
- Jacques, P.E.; Jeyakani, J.; Bourque, G. The majority of primate-specific regulatory sequences are derived from transposable elements. PLoS Genet. 2013, 9, e1003504. [Google Scholar] [CrossRef]
- Hancks, D.C.; Kazazian, H.H., Jr. Roles for retrotransposon insertions in human disease. Mob. DNA 2016, 7, 9. [Google Scholar] [CrossRef]
- Campos-Sánchez, R.; Kapusta, A.; Feschotte, C.; Chiaromonte, F.; Makova, K.D. Genomic landscape of human, bat, and ex vivo DNA transposon integrations. Mol. Biol. Evol. 2014, 31, 1816–1832. [Google Scholar] [CrossRef] [PubMed]
- Belshaw, R.; Pereira, V.; Katzourakis, A.; Talbot, G.; Pačes, J.; Burt, A.; Tristem, M. Long-term reinfection of the human genome by endogenous retroviruses. Proc. Natl. Acad. Sci. USA 2004, 101, 4894–4899. [Google Scholar] [CrossRef] [PubMed]
- Kuery, P.; Nath, A.; Creange, A.; Dolei, A.; Marche, P.; Gold, J.; Giovannoni, G.; Hartung, H.-P.; Perron, H. Human Endogenous Retroviruses in Neurological Diseases. Trends Mol. Med. 2018, 24, 379–394. [Google Scholar] [CrossRef] [PubMed]
- Löwer, R.; Löwer, J.; Kurth, R. The viruses in all of us characteristics and biological significance of human endogenous retrovirus sequences. Proc. Natl. Acad. Sci. USA 1996, 93, 5177–5184. [Google Scholar] [CrossRef] [PubMed]
- Young, G.R.; Eksmond, U.; Salcedo, R.; Alexopoulou, L.; Stoye, J.P.; Kassiotis, G. Resurrection of endogenous retroviruses in antibody-deficient mice. Nature 2012, 491, 774–778. [Google Scholar] [CrossRef] [PubMed]
- Cegolon, L.; Salata, C.; Weiderpass, E.; Vineis, P.; Palù, G.; Mastrangelo, G. Human endogenous retroviruses and cancer prevention evidence and prospects. BMC Cancer 2013, 13, 4. [Google Scholar] [CrossRef] [PubMed]
- Boso, G.; Fleck, K.; Carley, S.; Liu, Q.; Buckler-White, A.; A Kozak, C. The Oldest Co-opted gag Gene of a Human Endogenous Retrovirus Shows Placenta-Specific Expression and Is Upregulated in Diffuse Large B-Cell Lymphomas. Mol. Biol. Evol. 2021, 38, 5453–5471. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Cai, Q.; Pan, C.; Liu, W.; Li, L.; Liu, J.; Gao, M.; Li, X.; Wang, L.; Rao, Y.; et al. CDK2 Inhibition Enhances Antitumor Immunity by Increasing IFN Response to Endogenous Retroviruses. Cancer Immunol. Res. 2022, 10, 525–539. [Google Scholar] [CrossRef] [PubMed]
- Maze, E.A.; Agit, B.; Reeves, S.; Hilton, D.A.; Parkinson, D.B.; Laraba, L.; Ercolano, E.; Kurian, K.M.; Hanemann, C.O.; Belshaw, R.D.; et al. Human Endogenous Retrovirus Type K Promotes Proliferation and Confers Sensitivity to Antiretroviral Drugs in Merlin-Negative Schwannoma and Meningioma. Cancer Res. 2022, 82, 235–247. [Google Scholar]
- Fuentes, D.R.; Swigut, T.; Wysocka, J. Systematic perturbation of retroviral LTRs reveals widespread long-range effects on human gene regulation. Elife 2018, 7, e35989. [Google Scholar]
- Ono, M.; Yasunaga, T.; Miyata, T.; Ushikubo, H. Nucleotide sequence of human endogenous retrovirus genome related to the mouse mammary tumor virus genome. J. Virol. 1986, 60, 589–598. [Google Scholar] [CrossRef] [PubMed]
- Contreras-Galindo, R.; Kaplan, M.H.; Contreras-Galindo, A.C.; Gonzalez-Hernandez, M.J.; Ferlenghi, I.; Giusti, F.; Lorenzo, E.; Gitlin, S.D.; Dosik, M.H.; Yamamura, Y.; et al. Characterization of human endogenous retroviral elements in the blood of HIV-1-infected individuals. J. Virol. 2012, 86, 262–276. [Google Scholar] [CrossRef] [PubMed]
- Contreras-Galindo, R.; Kaplan, M.H.; Leissner, P.; Verjat, T.; Ferlenghi, I.; Bagnoli, F.; Giusti, F.; Dosik, M.H.; Hayes, D.F.; Gitlin, S.D.; et al. Human endogenous retrovirus K (HML-2) elements in the plasma of people with lymphoma and breast cancer. J. Virol. 2008, 82, 9329–9336. [Google Scholar] [CrossRef] [PubMed]
- Turner, G.; Barbulescu, M.; Su, M.; Jensen-Seaman, M.I.; Kidd, K.K.; Lenz, J. Insertional polymorphisms of full-length endogenous retroviruses in humans. Curr. Biol. 2001, 11, 1531–1535. [Google Scholar] [CrossRef] [PubMed]
- Medstrand, P.; Dl, M. Human-specific integrations of the HERV-K endogenous retrovirus family. J. Virol. 1998, 72, 9782–9787. [Google Scholar] [CrossRef] [PubMed]
- Barbulescu, M.; Turner, G.; Seaman, M.I.; Deinard, A.S.; Kidd, K.K.; Lenz, J. Many human endogenous retrovirus K (HERV-K) proviruses are unique to humans. Curr. Biol. 1999, 9, 861–868. [Google Scholar] [CrossRef] [PubMed]
- Macfarlane, C.M.; Badge, R.M. Genome-wide amplification of proviral sequences reveals new polymorphic HERV-K(HML-2) proviruses in humans and chimpanzees that are absent from genome assemblies. Retrovirology 2015, 12, 35. [Google Scholar] [CrossRef] [PubMed]
- Feschotte, C. Transposable elements and the evolution of regulatory networks. Nat. Rev. Genet. 2008, 9, 397–405. [Google Scholar] [CrossRef]
- Chuong, E.B.; Elde, N.C.; Feschotte, C. Regulatory evolution of innate immunity through co-option of endogenous retroviruses. Science 2016, 351, 1083–1087. [Google Scholar] [CrossRef]
- Tie, C.H.; Fernandes, L.; Conde, L.; Robbez-Masson, L.; Sumner, R.P.; Peacock, T.; Rodriguez-Plata, M.T.; Mickute, G.; Gifford, R.; Towers, G.J.; et al. KAP1 regulates endogenous retroviruses in adult human cells and contributes to innate immune control. EMBO Rep. 2018, 10, e45000. [Google Scholar]
- Hallast, P.; Ebert, P.; Loftus, M.; Yilmaz, F.; Audano, P.A.; Logsdon, G.A.; Bonder, M.J.; Zhou, W.; Hoeps, W.; Kim, K.; et al. Assembly of 43 human Y chromosomes reveals extensive complexity and variation. Nature 2023, 621, 355–364. [Google Scholar] [CrossRef] [PubMed]
- Collins, P.L.; Kyle, K.E.; Egawa, T.; Shinkai, Y.; Oltz, E.M. The histone methyltransferase SETDB1 represses endogenous and exogenous retroviruses in B lymphocytes. Proc. Natl. Acad. Sci. USA 2015, 112, 8367–8372. [Google Scholar] [CrossRef] [PubMed]
- Koide, S.; Oshima, M.; Takubo, K.; Yamazaki, S.; Nitta, E.; Saraya, A.; Aoyama, K.; Kato, Y.; Miyagi, S.; Nakajima-Takagi, Y.; et al. Setdb1 maintains hematopoietic stem and progenitor cells by restricting the ectopic activation of nonhematopoietic genes. Blood 2016, 128, 638–649. [Google Scholar] [CrossRef]
- Imbeault, M.; Helleboid, P.-Y.; Trono, D. KRAB zinc-finger proteins contribute to the evolution of gene regulatory networks. Nature 2017, 543, 550–554. [Google Scholar] [CrossRef] [PubMed]
- Johansson, E.M.; Bouchet, D.; Tamouza, R.; Ellul, P.; Morr, A.; Avignone, E.; Germi, R.; Leboyer, M.; Perron, H.; Groc, L. Human endogenous retroviral protein triggers deficit in glutamate synapse maturation and behaviors associated with psychosis. Sci. Adv. 2020, 6, eabc0708. [Google Scholar] [CrossRef]



| Number | Chromosome | Strand | Position Start | Position End | Length (bp) | Match + Mismatch (bp)/Full Length (bp) | Range | Q Gap Bases/(Match + Mismatch + Q Gap Bases) | Insertion or Deletion | Intergenic/Intron/Exon |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | chr16 | - | 19,355,716 | 19,364,289 | 8574 | 95.90% | (90–100%) | 1.14% | NA | intergenic |
| 2 | chr2A | + | 82,212,507 | 82,221,098 | 8592 | 95.75% | (90–100%) | 1.32% | NA | intergenic |
| 3 | chr8 | - | 43,516,710 | 43,525,262 | 8553 | 93.95% | (90–100%) | 3.39% | NA | intergenic |
| 4 | chr15 | - | 23,660,410 | 23,668,998 | 8589 | 89.68% | (80–90%) | 7.89% | NA | exonic&intronic |
| 5 | chr13 | + | 66,107,993 | 66,115,813 | 7821 | 85.42% | (80–90%) | 8.35% | NA | intergenic |
| 6 | chr5 | - | 60,565,330 | 60,572,694 | 7365 | 75.51% | (70–80%) | 22.56% | deletion | intron |
| 7 | chr6 | + | 48,375,302 | 48,381,314 | 6013 | 64.13% | (60–70%) | 35.21% | deletion | intergenic |
| 8 | chrY | - | 2,019,892 | 2,025,806 | 5915 | 60.86% | (60–70%) | 37.99% | deletion | intergenic |
| 9 | chrY | + | 6,151,270 | 6,157,165 | 5896 | 60.78% | (60–70%) | 37.88% | deletion | intergenic |
| 10 | chr19 | + | 39,063,080 | 39,068,105 | 5026 | 56.83% | (50–60%) | 41.91% | deletion | intergenic |
| 11 | chrY | + | 1,786,750 | 1,792,226 | 5477 | 55.32% | (50–60%) | 40.62% | deletion | intergenic |
| 12 | chrY | + | 14,791,990 | 14,797,426 | 5437 | 55.17% | (50–60%) | 40.45% | deletion | intergenic |
| 13 | chrY | - | 16,748,108 | 16,753,037 | 4930 | 54.55% | (50–60%) | 39.48% | deletion | intergenic |
| 14 | chr4 | - | 168,066,681 | 168,071,528 | 4848 | 54.03% | (50–60%) | 0.96% | NA | intergenic |
| 15 | chrY | + | 16,516,529 | 16,520,265 | 3737 | 35.85% | (30–40%) | 52.96% | deletion;insertion | genic&intergenic |
| 16 | chr7 | - | 147,138,951 | 147,141,648 | 2698 | 27.43% | (20–30%) | 11.17% | deletion | intergenic |
| 17 | chr8 | + | 76,757,424 | 76,759,791 | 2368 | 26.57% | (20–30%) | 0.48% | NA | intron |
| 18 | chrY | + | 10,590,060 | 10,592,276 | 2217 | 24.95% | (20–30%) | 8.91% | NA | intergenic |
| 19 | chrY | + | 8,716,313 | 8,718,506 | 2194 | 22.85% | (20–30%) | 6.51% | NA | intergenic |
| 20 | chrY | - | 10,822,682 | 10,824,473 | 1792 | 19.48% | (10–20%) | 3.45% | NA | intergenic |
| 21 | chrY | + | 13,999,593 | 14,001,168 | 1576 | 17.24% | (10–20%) | 12.03% | deletion | intergenic |
| 22 | chrY | + | 5,360,268 | 5,361,838 | 1571 | 17.18% | (10–20%) | 12.07% | deletion | intergenic |
| 23 | chrY | - | 13,423,428 | 13,425,003 | 1576 | 17.03% | (10–20%) | 13.10% | deletion | intergenic |
| 24 | chrY | - | 4,753,348 | 4,754,918 | 1571 | 16.97% | (10–20%) | 13.14% | deletion | intergenic |
| 25 | chrY | - | 10,496,235 | 10,498,083 | 1849 | 6.83% | (0–10%) | 67.62% | deletion;insertion | intergenic |
| 26 | chrY | - | 9,765,071 | 9,765,511 | 441 | 4.72% | (0–10%) | 3.10% | NA | intergenic |
| Number | Chromosome | Strand | Position Start | Position End | Length (bp) | Percentage of LTR14C in Length | Match + Mismatch/Full Length | Range | Qgap(bp)/(Match + Mismatch + Qgap(bp)) | Insertion or Deletion | Intergenic/Intron/Exon |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | chr1 | + | 137,442,976 | 137,443,751 | 776 | 101.19% | 6.90% | (0–10%) | 4.81% | NA | intergenic |
| 2 | chrX | - | 88,705,817 | 88,706,830 | 1014 | 100.85% | 6.88% | (0–10%) | 2.47% | NA | intergenic |
| 3 | chr2B | + | 66,792,632 | 66,793,222 | 591 | 100.34% | 6.84% | (0–10%) | 0.34% | NA | intergenic |
| 4 | chr5 | - | 84,066,356 | 84,066,944 | 589 | 100.00% | 6.82% | (0–10%) | 0.00% | NA | intergenic |
| 5 | chr7 | - | 48,429,668 | 48,430,256 | 589 | 100.00% | 6.82% | (0–10%) | 0.34% | NA | intergenic |
| 6 | chr15 | + | 17,411,474 | 17,412,062 | 589 | 100.00% | 6.82% | (0–10%) | 0.00% | NA | intergenic |
| 7 | chr2B | - | 80,853,578 | 80,854,165 | 588 | 99.83% | 6.81% | (0–10%) | 0.00% | NA | intergenic |
| 8 | chr3 | - | 161,301,797 | 161,302,385 | 589 | 99.83% | 6.81% | (0–10%) | 0.00% | NA | intron |
| 9 | chr4 | + | 19,038,454 | 19,039,041 | 588 | 99.83% | 6.81% | (0–10%) | 0.17% | NA | intergenic |
| 10 | chr8 | - | 49,114,425 | 49,115,012 | 588 | 99.83% | 6.81% | (0–10%) | 0.00% | NA | intergenic |
| 11 | chr10 | - | 19,329,882 | 19,330,469 | 588 | 99.83% | 6.81% | (0–10%) | 0.00% | NA | intergenic |
| 12 | chr14 | + | 19,027,337 | 19028300 | 964 | 99.83% | 6.81% | (0–10%) | 35.96% | deletion;insertion | intron |
| 13 | chr2B | + | 22,085,802 | 22,086,389 | 588 | 99.66% | 6.80% | (0–10%) | 0.17% | NA | intron |
| 14 | chr3 | - | 81,878,200 | 81,878,786 | 587 | 99.66% | 6.80% | (0–10%) | 0.17% | NA | intergenic |
| 15 | chr3 | - | 84,155,763 | 84,156,349 | 587 | 99.66% | 6.80% | (0–10%) | 0.17% | NA | intergenic |
| 16 | chr6 | - | 79,799,818 | 79,800,406 | 589 | 99.66% | 6.80% | (0–10%) | 0.34% | NA | intergenic |
| 17 | chr7 | + | 14,600,052 | 14,600,637 | 586 | 99.49% | 6.78% | (0–10%) | 0.34% | NA | intron |
| 18 | chrY | - | 18,630,426 | 18,631,014 | 589 | 99.49% | 6.78% | (0–10%) | 0.51% | NA | intergenic |
| 19 | chr2A | - | 81,498,383 | 81,498,970 | 588 | 99.32% | 6.77% | (0–10%) | 0.51% | NA | intergenic |
| 20 | chrX | + | 10,991,573 | 10,992,157 | 585 | 99.32% | 6.77% | (0–10%) | 0.51% | NA | intergenic |
| 21 | chr2A | + | 78,001,602 | 78,002,185 | 584 | 99.15% | 6.76% | (0–10%) | 0.68% | NA | intergenic |
| 22 | chr18 | + | 10,395,647 | 10,396,231 | 585 | 99.15% | 6.76% | (0–10%) | 0.68% | NA | intergenic |
| 23 | chrX | - | 138,003,751 | 138,004,333 | 583 | 98.98% | 6.75% | (0–10%) | 0.00% | NA | intergenic |
| 24 | chr5 | - | 38,971,726 | 38,972,307 | 582 | 98.64% | 6.73% | (0–10%) | 0.00% | NA | intergenic |
| 25 | chr12 | - | 50,879,963 | 50,880,545 | 583 | 98.47% | 6.71% | (0–10%) | 0.69% | NA | intergenic |
| 26 | chr14 | + | 28,857,266 | 28,857,843 | 578 | 97.96% | 6.68% | (0–10%) | 2.04% | NA | intergenic |
| 27 | chr21 | - | 979,256 | 979,845 | 590 | 97.79% | 6.67% | (0–10%) | 2.05% | NA | intron |
| 28 | chr6 | + | 166,662,836 | 166,663,420 | 585 | 97.27% | 6.63% | (0–10%) | 2.56% | NA | intergenic |
| 29 | chr16 | - | 31,333,554 | 31,334,129 | 576 | 96.76% | 6.60% | (0–10%) | 2.91% | NA | intergenic |
| 30 | chr3 | + | 44,753,258 | 44,753,829 | 572 | 96.42% | 6.58% | (0–10%) | 3.41% | NA | intergenic |
| 31 | chr2B | + | 27,967,992 | 27,968,572 | 581 | 96.25% | 6.56% | (0–10%) | 3.58% | NA | intron |
| 32 | chr7 | + | 140,435,975 | 140,436,547 | 573 | 96.25% | 6.56% | (0–10%) | 3.58% | NA | intergenic |
| 33 | chr2B | - | 25,439,654 | 25,440,248 | 595 | 96.08% | 6.55% | (0–10%) | 2.76% | NA | intergenic |
| 34 | chr13 | - | 19,353,563 | 19,354,142 | 580 | 95.91% | 6.54% | (0–10%) | 4.09% | NA | intron |
| 35 | chr2B | - | 11,387,988 | 11,388,572 | 585 | 95.40% | 6.51% | (0–10%) | 4.44% | NA | intergenic |
| 36 | chr3 | - | 59,703,426 | 59704196 | 771 | 94.21% | 6.42% | (0–10%) | 5.63% | NA | intergenic |
| 37 | chr11 | + | 120,039,148 | 120,039,717 | 570 | 94.04% | 6.41% | (0–10%) | 2.99% | NA | intergenic |
| 38 | chr1 | + | 143,492,646 | 143,493,186 | 541 | 88.07% | 6.01% | (0–10%) | 3.36% | NA | intron |
| Species | Chromosome | Strand | Position Start | Position End |
|---|---|---|---|---|
| chimp * | chr2A | + | 82,212,507 | 82,221,098 |
| human * | chr2 | + | 82,022,660 | 82,031,279 |
| chimp | chr4 | - | 168,066,681 | 168,071,528 |
| human | chr4 | - | 170,126,345 | 170,133,883 |
| chimp | chr6 | + | 48,375,302 | 48,381,314 |
| human | chr6 | + | 48,873,675 | 48,879,725 |
| chimp | chr7 | - | 147,138,951 | 147,141,648 |
| human | chr7 | - | 150,561,277 | 150,563,994 |
| chimp | chr8 | - | 43,516,710 | 43,525,262 |
| human | chr8 | - | 43,694,016 | 43,702,583 |
| chimp | chr8 | + | 76,757,424 | 76,759,791 |
| human | chr8 | + | 78,652,302 | 78,654,820 |
| chimp | chr13 | + | 66,107,993 | 66,115,813 |
| human | chr13 | + | 84,869,526 | 84,877,320 |
| chimp | chr15 | - | 23,660,410 | 23,668,998 |
| human | chr15 | - | 45,234,477 | 45,243,073 |
| chimp | chr16 | - | 19,355,716 | 19,364,289 |
| human | chr16 | - | 19,393,581 | 19,402,152 |
| chimp | chr19 | + | 39,063,080 | 39,068,105 |
| human | chr19 | + | 40,954,172 | 40,959,178 |
| chimp # | chr5 | - | 60,565,330 | 60,572,694 |
| chimp | chrY | + | 1,786,750 | 1,792,226 |
| chimp | chrY | - | 2,019,892 | 2,025,806 |
| chimp | chrY | - | 4,753,348 | 4,754,918 |
| chimp | chrY | + | 5,360,268 | 5,361,838 |
| chimp | chrY | + | 6,151,270 | 61,571,65 |
| chimp | chrY | + | 8,716,313 | 8,718,506 |
| chimp | chrY | - | 9,765,071 | 9,765,511 |
| chimp | chrY | - | 10,496,235 | 10,498,083 |
| chimp | chrY | + | 10,590,060 | 10,592,276 |
| chimp | chrY | - | 10,822,682 | 10,824,473 |
| chimp | chrY | - | 13,423,428 | 13,425,003 |
| chimp | chrY | + | 13,999,593 | 14,001,168 |
| chimp | chrY | + | 14,791,990 | 14,797,426 |
| chimp | chrY | + | 16,516,529 | 16,520,265 |
| chimp | chrY | - | 16,748,108 | 16,753,037 |
| human | chr1 | - | 155,629,408 | 155,632,775 |
| human | chr5 | - | 156,660,448 | 156,663,815 |
| human | chr8 | + | 145,019,974 | 145,032,719 |
| human | chr10 | - | 99,822,511 | 99,825,532 |
| human | chr12 | + | 48,509,228 | 48,511,681 |
| human | chr21 | - | 18,563,368 | 18,566,735 |
| human | chrY | - | 8,121,821 | 8,126,768 |
| human | chrY | + | 8,996,062 | 9,000,755 |
| human | chrY | - | 9,273,707 | 9,279,611 |
| human | chrY | - | 17,669,948 | 17,671,523 |
| human | chrY | - | 18,622,534 | 18,626,952 |
| human | chrY | + | 21,580,120 | 21,585,551 |
| human | chrY | - | 21,845,475 | 21,850,069 |
| Species | Chromosome | Strand | Position Start | Position End |
|---|---|---|---|---|
| chimp * | chr1 | + | 143,492,646 | 143,493,186 |
| human * | chr1 | + | 168,457,190 | 168,457,732 |
| chimp | chr2A | + | 78,001,602 | 78,002,185 |
| human | chr2 | + | 77,807,602 | 77,808,185 |
| chimp | chr2A | − | 81,498,383 | 81,498,970 |
| human | chr2 | − | 81,304,430 | 81,305,068 |
| chimp | chr3 | + | 44,753,258 | 44,753,829 |
| human | chr3 | + | 44,534,488 | 44,535,059 |
| chimp | chr3 | − | 59,703,426 | 59,704,196 |
| human | chr3 | − | 59,469,489 | 59,470,030 |
| chimp | chr3 | − | 81,878,200 | 81,878,786 |
| human | chr3 | − | 81,329,902 | 81,330,488 |
| chimp | chr3 | − | 84,155,763 | 84,156,349 |
| human | chr3 | − | 83,618,409 | 83,618,995 |
| chimp | chr3 | − | 161,301,797 | 161,302,385 |
| human | chr3 | − | 163,283,189 | 163,283,777 |
| chimp | chr4 | + | 19,038,454 | 19,039,041 |
| human | chr4 | + | 19,556,097 | 19,556,684 |
| chimp | chr5 | − | 38,971,726 | 38,972,307 |
| human | chr5 | + | 75,859,521 | 75,860,102 |
| chimp | chr5 | − | 84,066,356 | 84,066,944 |
| human | chr5 | − | 105,998,962 | 105,999,549 |
| chimp | chr6 | − | 79,799,818 | 79,800,406 |
| human | chr6 | − | 82,297,755 | 82,298,498 |
| chimp | chr6 | + | 166,662,836 | 166,663,420 |
| human | chr6 | + | 169,084,226 | 169,084,808 |
| chimp | chr7 | + | 14,600,052 | 14,600,637 |
| human | chr7 | + | 14,509,240 | 14,509,827 |
| chimp | chr7 | + | 140,435,975 | 140,436,547 |
| human | chr7 | + | 143,472,173 | 143,472,744 |
| chimp | chr8 | − | 49,114,425 | 49,115,012 |
| human | chr8 | − | 51,178,592 | 51,179,179 |
| chimp | chr10 | − | 19,329,882 | 19,330,469 |
| human | chr10 | − | 18,856,645 | 18,857,233 |
| chimp | chr11 | + | 120,039,148 | 120,039,717 |
| human | chr11 | + | 124,270,705 | 124,271,275 |
| chimp | chr12 | − | 50,879,963 | 50,880,545 |
| human | chr12 | + | 38,144,469 | 38,145,052 |
| chimp | chr13 | − | 19,353,563 | 19,354,142 |
| human | chr13 | − | 38,319,721 | 38,320,300 |
| chimp | chr14 | + | 19,027,337 | 19,028,300 |
| human | chr14 | + | 38,011,040 | 38,012,012 |
| chimp | chr14 | + | 28,857,266 | 28,857,843 |
| human | chr14 | + | 48,011,215 | 48,011,780 |
| chimp | chr15 | + | 17,411,474 | 17,412,062 |
| human | chr15 | + | 39,011,033 | 39,011,621 |
| chimp | chr18 | + | 10,395,647 | 10,396,231 |
| human | chr18 | − | 4,527,618 | 4,528,209 |
| chimp | chr21 | − | 979,256 | 979,845 |
| human | chr21 | − | 14,080,466 | 14,081,052 |
| chimp | chrX | + | 10,991,573 | 10,992,157 |
| human | chrX | + | 11,033,746 | 11,034,330 |
| chimp | chrX | − | 88,705,817 | 88,706,830 |
| human | chrX | − | 93,273,183 | 93,274,197 |
| chimp | chrX | − | 138,003,751 | 138,004,333 |
| human | chrX | − | 142,767,872 | 142,768,454 |
| chimp # | chr1 | + | 137,442,976 | 137,443,751 |
| chimp | chr2B | − | 11,387,988 | 11,388,572 |
| chimp | chr2B | + | 22,085,802 | 22,086,389 |
| chimp | chr2B | − | 25,439,654 | 25,440,248 |
| chimp | chr2B | + | 27,967,992 | 27,968,572 |
| chimp | chr2B | + | 66,792,632 | 66,793,222 |
| chimp | chr2B | − | 80,853,578 | 80,854,165 |
| chimp | chr7 | − | 48,429,668 | 48,430,256 |
| chimp | chr16 | − | 31,333,554 | 31,334,129 |
| chimp | chrY | − | 18,630,426 | 18,631,014 |
| human | chr1 | + | 162,419,359 | 162,419,942 |
| human | chr2 | + | 97,964,920 | 97,965,508 |
| human | chr2 | − | 125,024,208 | 125,024,792 |
| human | chr2 | + | 135,521,883 | 135,522,470 |
| human | chr2 | − | 138,860,917 | 138,861,512 |
| human | chr2 | + | 171,365,032 | 171,365,617 |
| human | chr2 | + | 180,236,847 | 180,237,437 |
| human | chr2 | − | 194,256,159 | 194,256,746 |
| human | chr3 | − | 154,944,330 | 154,944,911 |
| human | chr4 | + | 116,980,222 | 116,980,809 |
| human | chr4 | + | 129,080,872 | 129,081,454 |
| human | chr11 | − | 62,185,237 | 62,185,824 |
| human | chr12 | − | 4,720,007 | 4,720,593 |
| human | chr15 | − | 54,594,796 | 54,595,373 |
| human | chr17 | + | 52,961,655 | 52,962,071 |
| human | chr18 | + | 63,648,105 | 63,648,555 |
| human | chr20 | − | 2,809,052 | 2,809,886 |
| human | chrX | − | 144,791,258 | 144,791,846 |
| human | chrY | − | 25,974,734 | 25,975,320 |
| Number | Provirus Regions | 5′LTR (%) | Gag (%) | Pro (%) | Pol (%) | Env (%) | 3′LTR (%) |
|---|---|---|---|---|---|---|---|
| 1 | chr16 19,355,716 19,364,289 | 100.00% | 99.61% | 99.89% | 99.43% | 99.17% | 99.66% |
| 2 | chr2A 82,212,507 82,221,098 | 98.98% | 99.66% | 100.00% | 99.47% | 99.90% | 99.15% |
| 3 | chr8 43,516,710 43,525,262 | 99.66% | 99.83% | 98.20% | 99.40% | 98.83% | 99.83% |
| 4 | chr15 23,660,410 23,668,998 | 99.83% | 99.22% | 99.77% | 99.51% | 99.46% | 99.83% |
| 5 | chr13 66,107,993 66,115,813 | 35.95% | 98.49% | 52.25% | 99.66% | 99.76% | 99.66% |
| 6 | chr5 60,565,330 60,572,694 | 73.42% | 99.72% | 99.55% | 75.25% | 48.48% | 98.64% |
| 7 | chr6 48,375,302 48,381,314 | 98.47% | 92.91% | 61.71% | 6.04% | 99.85% | 98.98% |
| 8 | chrY 2,019,892 2,025,806 | 88.25% | 91.74% | 64.86% | 6.19% | 99.56% | 99.15% |
| 9 | chrY 6,151,270 6,157,165 | 83.82% | 91.68% | 65.32% | 6.04% | 99.56% | 99.83% |
| 10 | chr19 39,063,080 39,068,105 | 98.98% | 99.22% | 64.30% | 6.11% | 67.35% | 79.56% |
| 11 | chrY 1,786,750 1,792,226 | 88.59% | 91.74% | 64.86% | 6.19% | 99.56% | 22.83% |
| 12 | chrY 14,791,990 14,797,426 | 88.59% | 91.74% | 64.86% | 6.19% | 99.56% | 16.52% |
| 13 | chrY 16,748,108 16,753,037 | 0.00% | 91.35% | 64.86% | 6.19% | 99.56% | 99.83% |
| 14 | chr4 168,066,681 168,071,528 | 0.00% | 92.69% | 99.77% | 87.66% | 0.00% | 0.00% |
| 15 | chrY 16,516,529 16,520,265 | 0.00% | 64.99% | 64.86% | 6.19% | 93.70% | 0.00% |
| 16 | chr7 147,138,951 147,141,648 | 0.00% | 0.00% | 0.00% | 16.64% | 99.12% | 51.45% |
| 17 | chr8 76,757,424 76,759,791 | 0.00% | 0.00% | 0.00% | 0.00% | 86.95% | 100.00% |
| 18 | chrY 10,590,060 10,592,276 | 56.90% | 91.74% | 7.77% | 0.00% | 0.00% | 0.00% |
| 19 | chrY 8,716,313 8,718,506 | 0.00% | 0.00% | 0.00% | 6.11% | 99.32% | 10.22% |
| 20 | chrY 10,822,682 10,824,473 | 0.00% | 0.00% | 0.00% | 0.00% | 58.26% | 99.66% |
| 21 | chrY 13,999,593 14,001,168 | 52.81% | 62.09% | 0.00% | 0.00% | 0.00% | 0.00% |
| 22 | chrY 5,360,268 5,361,838 | 52.47% | 61.92% | 0.00% | 0.00% | 0.00% | 0.00% |
| 23 | chrY 13,423,428 13,425,003 | 52.64% | 62.14% | 0.00% | 0.00% | 0.00% | 0.00% |
| 24 | chrY 4,753,348 4,754,918 | 51.96% | 62.09% | 0.00% | 0.00% | 0.00% | 0.00% |
| 25 | chrY 10,496,235 10,498,083 | 0.00% | 0.00% | 0.00% | 0.00% | 86.41% | 9.20% |
| 26 | chrY 9,765,071 9,765,511 | 0.00% | 0.00% | 0.00% | 0.00% | 21.31% | 0.00% |
| Species | Provirus Regions | Divergence from Consensus Sequence | Mean Divergences | T = D/0.2 | Age/Million Years (Gene vs. Consensus) | Divergence between 2 LTRs | T = D/0.2/2 | Age/Million Years (LTR vs. LTR) | |||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Gag | Pro | Pol | Env | ||||||||
| chimp * | chr2A 82,212,507 82,221,098 | 0.057 | 0.060 | 0.063 | 0.085 | 0.066 | 0.33 | 33.13 | 0.057 | 0.14 | 14.25 |
| human * | chr2 82,022,660 82,031,279 | 0.061 | 0.182 | 0.204 | 0.101 | 0.137 | 0.69 | 68.50 | 0.070 | 0.18 | 17.50 |
| chimp | chr4 168,066,681 168,071,528 | 0.051 | 0.063 | NA | NA | 0.057 | 0.29 | 28.50 | NA | NA | NA |
| human | chr4 170,126,345 170,133,883 | 0.058 | 0.172 | 0.214 | NA | 0.148 | 0.74 | 74.00 | NA | NA | NA |
| chimp | chr6 48,375,302 48,381,314 | 0.091 | NA | NA | 0.078 | 0.085 | 0.42 | 42.25 | 0.106 | 0.27 | 26.00 |
| human | chr6 48,873,675 48,879,725 | 0.063 | NA | NA | 0.106 | 0.085 | 0.42 | 42.25 | 0.110 | 0.28 | 27.50 |
| chimp | chr7 147,138,951 147,141,648 | NA | NA | NA | 0.235 | 0.235 | 1.18 | 117.50 | NA | NA | NA |
| human | chr7 150,561,277 150,563,994 | NA | NA | NA | 0.197 | 0.197 | 0.99 | 98.50 | NA | NA | NA |
| chimp | chr8 43,516,710 43,525,262 | 0.072 | 0.076 | 0.071 | 0.086 | 0.076 | 0.38 | 38.13 | 0.103 | 0.26 | 25.75 |
| human | chr8 43,694,016 43,702,583 | 0.091 | 0.177 | 0.231 | 0.121 | 0.155 | 0.78 | 77.50 | 0.107 | 0.27 | 26.75 |
| chimp | chr13 66,107,993 66,115,813 | 0.058 | NA | 0.055 | 0.070 | 0.061 | 0.31 | 30.50 | NA | NA | NA |
| human | chr13 84,869,526 84,877,320 | 0.054 | NA | 0.208 | 0.103 | 0.122 | 0.61 | 60.83 | NA | NA | NA |
| chimp | chr15 23,660,410 23,668,998 | 0.056 | 0.054 | 0.046 | 0.063 | 0.055 | 0.27 | 27.38 | 0.076 | 0.19 | 19.00 |
| human | chr15 45,234,477 45,243,073 | 0.051 | 0.126 | 0.206 | 0.099 | 0.121 | 0.60 | 60.25 | 0.080 | 0.20 | 20.00 |
| chimp | chr16 19,355,716 19,364,289 | 0.053 | 0.038 | 0.053 | 0.063 | 0.052 | 0.26 | 25.88 | 0.090 | 0.23 | 22.00 |
| human | chr16 19,393,581 19,402,152 | 0.059 | 0.158 | 0.206 | 0.089 | 0.128 | 0.64 | 64.00 | 0.082 | 0.21 | 20.50 |
| chimp | chr19 39,063,080 39,068,105 | 0.045 | NA | NA | NA | 0.045 | 0.23 | 22.50 | 0.109 | 0.27 | 27.25 |
| human | chr19 40,954,172 40,959,178 | 0.075 | NA | NA | NA | 0.075 | 0.38 | 37.50 | 0.097 | 0.24 | 24.25 |
| chimp # | chr5 60,565,330 60,572,694 | 0.061 | 0.056 | NA | NA | 0.059 | 0.29 | 29.25 | 0.089 | 0.22 | 22.25 |
| chimp | chrY 1,786,750 1,792,226 | 0.115 | NA | NA | 0.099 | 0.107 | 0.54 | 53.50 | NA | NA | NA |
| chimp | chrY 2,019,892 2,025,806 | 0.113 | NA | NA | 0.100 | 0.107 | 0.53 | 53.25 | 0.144 | 0.36 | 36.00 |
| chimp | chrY 6,151,270 6,157,165 | 0.114 | NA | NA | 0.098 | 0.106 | 0.53 | 53.00 | 0.144 | 0.36 | 36.00 |
| chimp | chrY 8,716,313 8,718,506 | NA | NA | NA | 0.100 | 0.100 | 0.50 | 50.00 | NA | NA | NA |
| chimp | chrY 10,590,060 10,592,276 | 0.114 | NA | NA | NA | 0.114 | 0.57 | 57.00 | NA | NA | NA |
| chimp | chrY 14,791,990 14,797,426 | 0.114 | NA | NA | 0.099 | 0.107 | 0.53 | 53.25 | NA | NA | NA |
| chimp | chrY 16,516,529 16,520,265 | NA | NA | NA | 0.097 | 0.097 | 0.49 | 48.50 | NA | NA | NA |
| chimp | chrY 16,748,108 16,753,037 | 0.112 | NA | NA | 0.098 | 0.105 | 0.53 | 52.50 | NA | NA | NA |
| human | chr1 155,629,408 155,632,775 | NA | 0.212 | 0.156 | NA | 0.184 | 0.92 | 92.00 | NA | NA | NA |
| human | chr5 156,660,448 156,663,815 | NA | 0.215 | 0.160 | NA | 0.188 | 0.94 | 93.75 | NA | NA | NA |
| human | chr8 145,019,974 145,032,719 | NA | NA | 0.513 | 0.093 | 0.303 | 1.52 | 151.50 | NA | NA | NA |
| human | chr10 99,822,511 99,825,532 | NA | NA | 0.169 | NA | 0.169 | 0.85 | 84.50 | NA | NA | NA |
| human | chr21 18,563,368 18,566,735 | NA | 0.266 | 0.190 | NA | 0.228 | 1.14 | 114.00 | NA | NA | NA |
| human | chrY 8,121,821 8,126,768 | NA | NA | NA | 0.125 | 0.125 | 0.63 | 62.50 | 0.157 | 0.39 | 39.25 |
| human | chrY 8,996,062 9,000,755 | NA | NA | NA | 0.133 | 0.133 | 0.67 | 66.50 | NA | NA | NA |
| human | chrY 9,273,707 9,279,611 | 0.114 | NA | NA | 0.139 | 0.127 | 0.63 | 63.25 | 0.141 | 0.35 | 35.25 |
| human | chrY 18,622,534 18,626,952 | NA | NA | NA | NA | NA | NA | NA | 0.194 | 0.49 | 48.50 |
| human | chrY 21,580,120 21,585,551 | 0.105 | NA | NA | 0.140 | 0.123 | 0.61 | 61.25 | NA | NA | NA |
| human | chrY 21,845,475 21,850,069 | NA | NA | NA | 0.143 | 0.143 | 0.72 | 71.50 | NA | NA | NA |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, M.; Yang, C.; Zhai, X.; Wang, C.; Liu, M.; Zhang, B.; Guo, X.; Wang, Y.; Li, H.; Liu, Y.; et al. Comprehensive Identification and Characterization of HML-9 Group in Chimpanzee Genome. Viruses 2024, 16, 892. https://doi.org/10.3390/v16060892
Chen M, Yang C, Zhai X, Wang C, Liu M, Zhang B, Guo X, Wang Y, Li H, Liu Y, et al. Comprehensive Identification and Characterization of HML-9 Group in Chimpanzee Genome. Viruses. 2024; 16(6):892. https://doi.org/10.3390/v16060892
Chicago/Turabian StyleChen, Mingyue, Caiqin Yang, Xiuli Zhai, Chunlei Wang, Mengying Liu, Bohan Zhang, Xing Guo, Yanglan Wang, Hanping Li, Yongjian Liu, and et al. 2024. "Comprehensive Identification and Characterization of HML-9 Group in Chimpanzee Genome" Viruses 16, no. 6: 892. https://doi.org/10.3390/v16060892
APA StyleChen, M., Yang, C., Zhai, X., Wang, C., Liu, M., Zhang, B., Guo, X., Wang, Y., Li, H., Liu, Y., Han, J., Wang, X., Li, J., Jia, L., & Li, L. (2024). Comprehensive Identification and Characterization of HML-9 Group in Chimpanzee Genome. Viruses, 16(6), 892. https://doi.org/10.3390/v16060892






