The Mechanism of Action of the Active Ingredients of Coptidis rhizoma against Porcine Epidemic Diarrhea Was Investigated Using Network Pharmacology and Molecular Docking Technology
Abstract
1. Introduction
2. Materials and Methods
3. Results
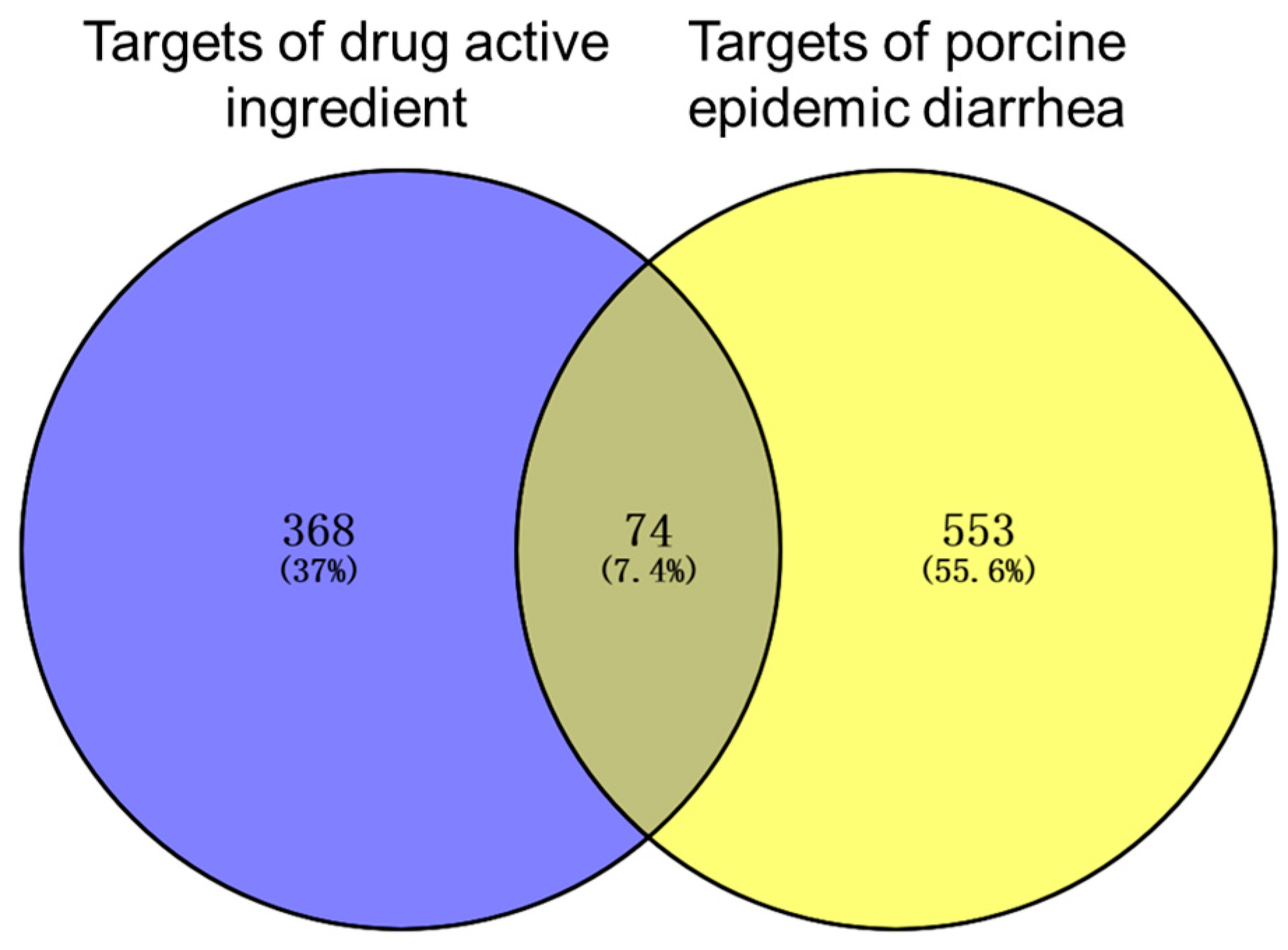
3.1. Drug Active Ingredients and Disease Targets
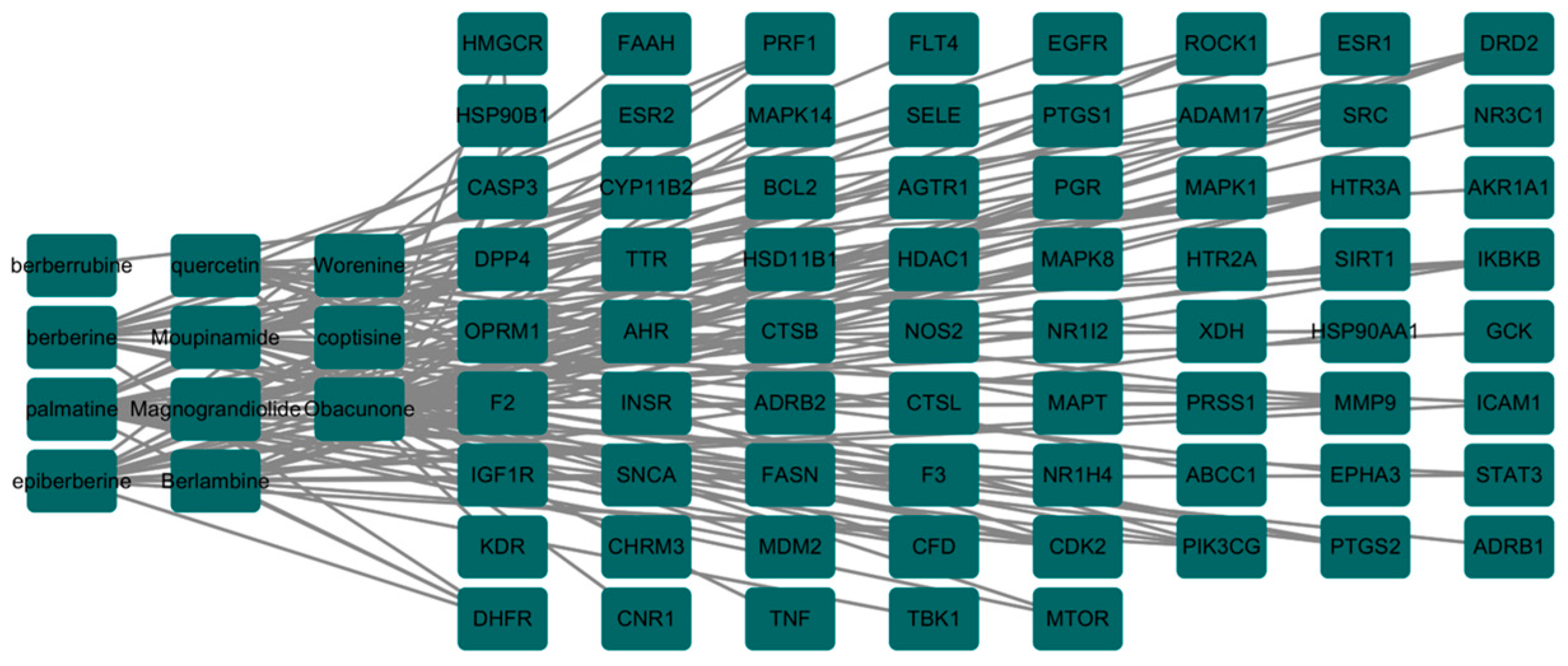
3.2. The Core Gene Targets of the Effective Active Components of the Drug against PED and Their Interaction Network Diagram
3.3. Enrichment Analysis of Anti-PED of Effective Active Ingredients of Drugs
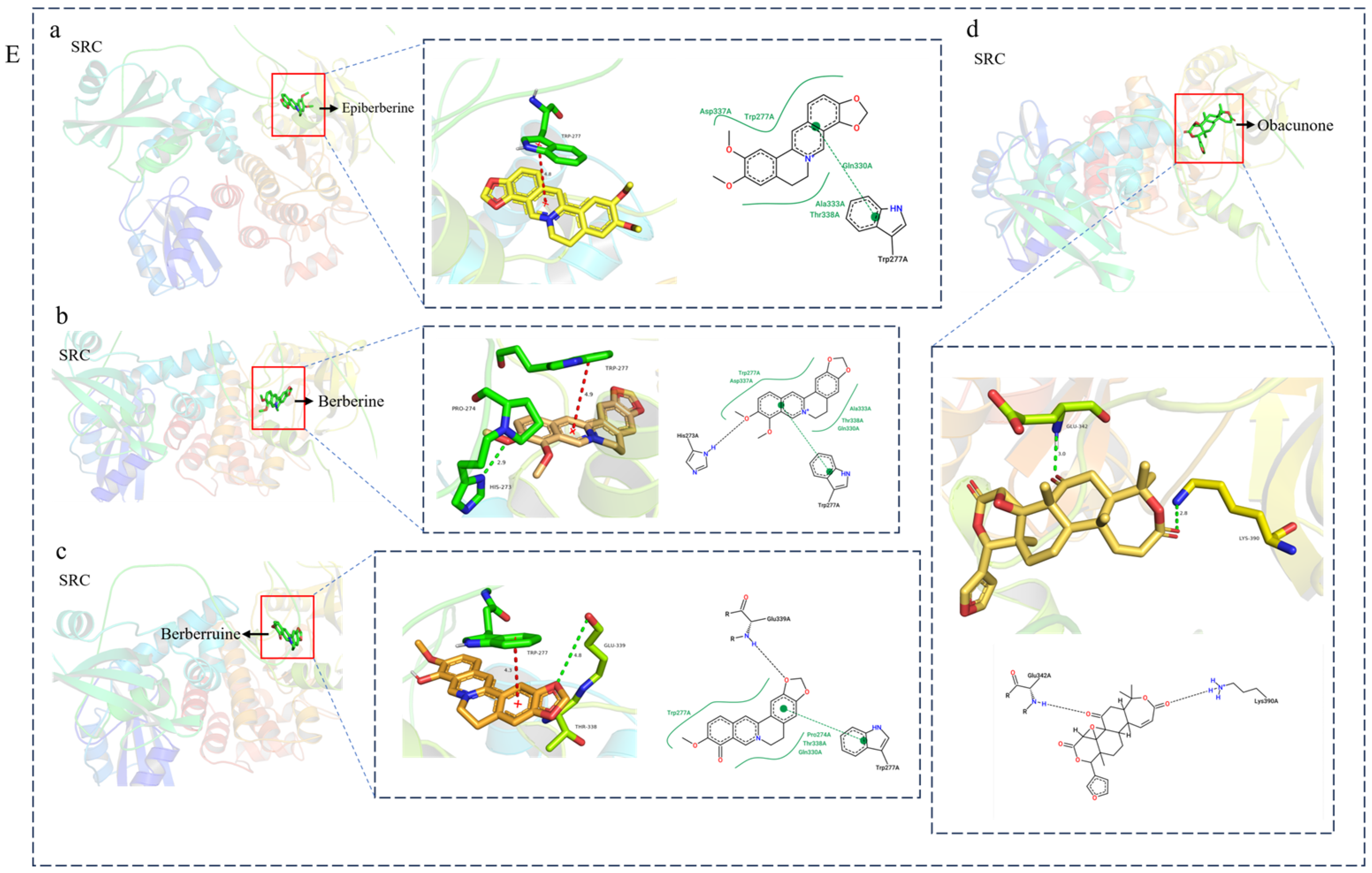
3.4. Molecular Docking Verification of the Main Genes Corresponding to the Active Components of the Drug and the Main Proteins Related to the Disease
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Lin, F.; Zhang, H.; Li, L.; Yang, Y.; Zou, X.; Chen, J.; Tang, X. PEDV: Insights and Advances into Types, Function, Structure, and Receptor Recognition. Viruses 2022, 14, 1744. [Google Scholar] [CrossRef]
- Jung, K.; Saif, L.J.; Wang, Q. Porcine epidemic diarrhea virus (PEDV): An update on etiology, transmission, pathogenesis, and prevention and control. Virus Res. 2020, 286, 198045. [Google Scholar] [CrossRef]
- Zhang, H.; Zou, C.; Peng, O.; Ashraf, U.; Xu, Q.; Gong, L.; Fan, B.; Zhang, Y.; Xu, Z.; Xue, C.; et al. Global Dynamics of Porcine Enteric Coronavirus PEDV Epidemiology, Evolution, and Transmission. Mol. Biol. Evol. 2023, 40, msad052. [Google Scholar] [CrossRef]
- Ren, T.; Feng, H.; Xu, Y.; Ling, Y. Revealing the mechanism of Dahuang Huanglian Xiexin Decoction attenuates dysbiosis via IL-17 signaling pathway based on network pharmacology and experimental validation. J. Ethnopharmacol. 2024, 331, 118267. [Google Scholar] [CrossRef]
- Zhang, C.; Chen, H.; Sun, L.; Zhao, P.; Qi, C.; Yang, Y.; Si, A.; Qian, Y.; Jung, Y.-S. Bis-Benzylisoquinoline Alkaloids Inhibit Porcine Epidemic Diarrhea Virus by Disrupting Virus Entry. Pathogens 2023, 12, 845. [Google Scholar] [CrossRef]
- Xiang, G.; Yang, L.; Qin, J.; Wang, S.; Zhang, Y.; Yang, S. Revealing the potential bioactive components and mechanism of Qianhua Gout Capsules in the treatment of gouty arthritis through network pharmacology, molecular docking and pharmacodynamic study strategies. Heliyon 2024, 10, e30983. [Google Scholar] [CrossRef]
- Xue, Y.; Chen, Y.; Sun, S.; Tong, X.; Chen, Y.; Tang, S.; Wang, X.; Bi, S.; Qiu, Y.; Zhao, Q.; et al. TET2-STAT3-CXCL5 nexus promotes neutrophil lipid transfer to fuel lung adeno-to-squamous transition. J. Exp. Med. 2024, 221, e20240111. [Google Scholar] [CrossRef]
- Huang, H.; Li, Y.; Li, D.; Wang, L.; Jiao, W.; Bai, Y.; Zhang, G. The tyrosine phosphatase PTPN14 inhibits the activation of STAT3 in PEDV infected Vero cells. Vet. Microbiol. 2022, 267, 109391. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Chen, D.; Tian, G.; Mao, X.; He, J.; Zheng, P.; Yu, J.; Luo, Y.; Luo, J.; Huang, Z.; et al. 1,25-Dihydroxyvitamin D3 Negatively Regulates the Inflammatory Response to Porcine Epidemic Diarrhea Virus Infection by Inhibiting NF-κB and JAK/STAT Signaling Pathway in IPEC-J2 Porcine Epithelial Cells. Int. J. Mol. Sci. 2022, 23, 10603. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Sun, J.; Prinz, R.A.; Liu, X.; Xu, X. Inhibition of porcine epidemic diarrhea virus (PEDV) replication by A77 1726 through targeting JAK and Src tyrosine kinases. Virology 2020, 551, 75–83. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Zhang, Z.; Li, Y.; Zhao, Y.; Ren, Y.; Tian, Y.; Hou, M.; Guo, Y.; Li, Q.; Tian, W.; et al. Estrogen promotes gonadotropin-releasing hormone expression by regulating tachykinin 3 and prodynorphin systems in chicken. Poult. Sci. 2024, 103, 103820. [Google Scholar] [CrossRef]
- Jasiński, T.; Zdrojkowski, Ł.; Ferreira-Dias, G.; Kautz, E.; Juszczuk-Kubiak, E.; Domino, M. Molecular Mechanism of Equine Endometrosis: The NF-κB-Dependent Pathway Underlies the Ovarian Steroid Receptors’ Dysfunction. Int. J. Mol. Sci. 2022, 23, 7360. [Google Scholar] [CrossRef]
- Qian, F.; Zhong, Q.; Chen, Z. Role of mitochondrial dysfunction in acute traumatic brain injury: Evidence from bioinformatics analysis. Heliyon 2024, 10, e31121. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Zhang, Y.; Wang, J.; Yan, Y.; Liu, Y.; Shi, X.; Zhang, Q.; Xu, X. The CREB and AP-1-Dependent Cell Communication Network Factor 1 Regulates Porcine Epidemic Diarrhea Virus-Induced Cell Apoptosis Inhibiting Virus Replication Through the p53 Pathway. Front. Microbiol. 2022, 13, 831852. [Google Scholar] [CrossRef]
- Xu, X.; Ma, M.; Shi, X.; Yan, Y.; Liu, Y.; Yang, N.; Wang, Q.; Zhang, S.; Zhang, Q. The novel Nsp9-interacting host factor H2BE promotes PEDV replication by inhibiting endoplasmic reticulum stress-mediated apoptosis. Vet. Res. 2023, 54, 27. [Google Scholar] [CrossRef] [PubMed]
- Luk, I.S.; Bridgwater, C.M.; Yu, A.; Boila, L.D.; Yáñez-Bartolomé, M.; Lampano, A.E.; Hulahan, T.S.; Boukhali, M.; Kathiresan, M.; Macarulla, T.; et al. SRC inhibition enables formation of a growth suppressive MAGI1-PP2A complex in isocitrate dehydrogenase-mutant cholangiocarcinoma. Sci. Transl. Med. 2024, 16, eadj7685. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Cao, Y.; Yang, Q. Transferrin receptor 1 levels at the cell surface influence the susceptibility of newborn piglets to PEDV infection. PLoS Pathog. 2020, 16, e1008682. [Google Scholar] [CrossRef]
- Jian, J.; Yang, Q.; Huang, X. Src regulates Tyr(20) phosphorylation of transferrin receptor-1 and potentiates breast cancer cell survival. J. Biol. Chem. 2011, 286, 35708–35715. [Google Scholar] [CrossRef]
- Lunney, J.K. Advances in swine biomedical model genomics. Int. J. Biol. Sci. 2007, 3, 179–184. [Google Scholar] [CrossRef]
- Schook, L.B.; Collares, T.V.; Darfour-Oduro, K.A.; De, A.K.; Rund, L.A.; Schachtschneider, K.M.; Seixas, F.K. Unraveling the swine genome: Implications for human health. Annu. Rev. Anim. Biosci. 2015, 3, 219–244. [Google Scholar] [CrossRef]
- Niu, Z.; Zhang, S.; Xu, S.; Wang, J.; Wang, S.; Hu, X.; Zhang, L.; Ren, L.; Zhang, J.; Liu, X.; et al. Porcine Epidemic Diarrhea Virus Replication in Human Intestinal Cells Reveals Potential Sus ceptibility to Cross-Species Infection. Viruses 2023, 15, 956. [Google Scholar] [CrossRef]



| Molecule Name | Chemical Structural | Molecule ID | Molecular Weight | Oral Bioavailability (%) | Drug-Likeness | Targets |
|---|---|---|---|---|---|---|
| Berberine | 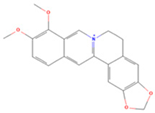 | MOL001454 | 336.39 | 36.86 | 0.78 | 100 |
| Obacunone | 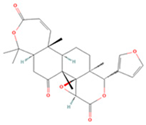 | MOL013352 | 454.56 | 43.29 | 0.77 | 100 |
| Berberrubine | 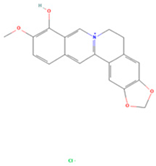 | MOL002894 | 322.36 | 35.74 | 0.73 | 22 |
| Epiberberine |  | MOL002897 | 336.39 | 43.09 | 0.78 | 100 |
| (R)-Canadine | 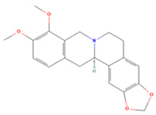 | MOL002903 | 339.42 | 55.37 | 0.77 | 100 |
| Berlambine | 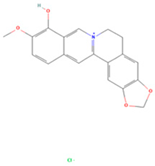 | MOL002904 | 351.38 | 36.68 | 0.82 | 100 |
| Corchoroside A_qt | 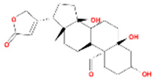 | MOL002907 | 404.55 | 104.95 | 0.78 | 2 |
| Magnograndiolide | 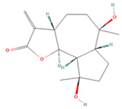 | MOL000622 | 266.37 | 63.71 | 0.19 | 49 |
| Palmidin A | 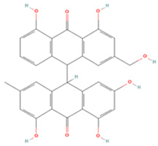 | MOL000762 | 510.52 | 35.36 | 0.65 | 0 |
| Palmatine | 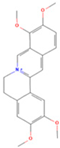 | MOL000785 | 352.44 | 64.60 | 0.65 | 100 |
| Quercetin | 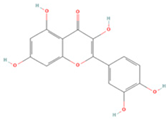 | MOL000098 | 302.55 | 46.43 | 0.28 | 154 |
| Coptisine |  | MOL001458 | 320.34 | 30.67 | 0.86 | 23 |
| Worenine | 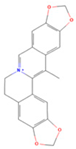 | MOL002668 | 334.37 | 45.83 | 0.87 | 23 |
| Moupinamide | 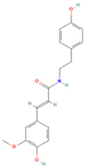 | MOL008647 | 313.38 | 86.71 | 0.26 | 100 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zou, H.; Niu, Z.; Tang, Z.; Cheng, P.; Yin, Y.; Luo, G.; Huang, S. The Mechanism of Action of the Active Ingredients of Coptidis rhizoma against Porcine Epidemic Diarrhea Was Investigated Using Network Pharmacology and Molecular Docking Technology. Viruses 2024, 16, 1229. https://doi.org/10.3390/v16081229
Zou H, Niu Z, Tang Z, Cheng P, Yin Y, Luo G, Huang S. The Mechanism of Action of the Active Ingredients of Coptidis rhizoma against Porcine Epidemic Diarrhea Was Investigated Using Network Pharmacology and Molecular Docking Technology. Viruses. 2024; 16(8):1229. https://doi.org/10.3390/v16081229
Chicago/Turabian StyleZou, Hong, Zheng Niu, Zhangchen Tang, Peng Cheng, Yanling Yin, Gan Luo, and Shilei Huang. 2024. "The Mechanism of Action of the Active Ingredients of Coptidis rhizoma against Porcine Epidemic Diarrhea Was Investigated Using Network Pharmacology and Molecular Docking Technology" Viruses 16, no. 8: 1229. https://doi.org/10.3390/v16081229
APA StyleZou, H., Niu, Z., Tang, Z., Cheng, P., Yin, Y., Luo, G., & Huang, S. (2024). The Mechanism of Action of the Active Ingredients of Coptidis rhizoma against Porcine Epidemic Diarrhea Was Investigated Using Network Pharmacology and Molecular Docking Technology. Viruses, 16(8), 1229. https://doi.org/10.3390/v16081229







