Indoxyl Sulfate-Mediated Metabolic Alteration of Transcriptome Signatures in Monocytes of Patients with End-Stage Renal Disease (ESRD)
Abstract
:1. Introduction
2. Results
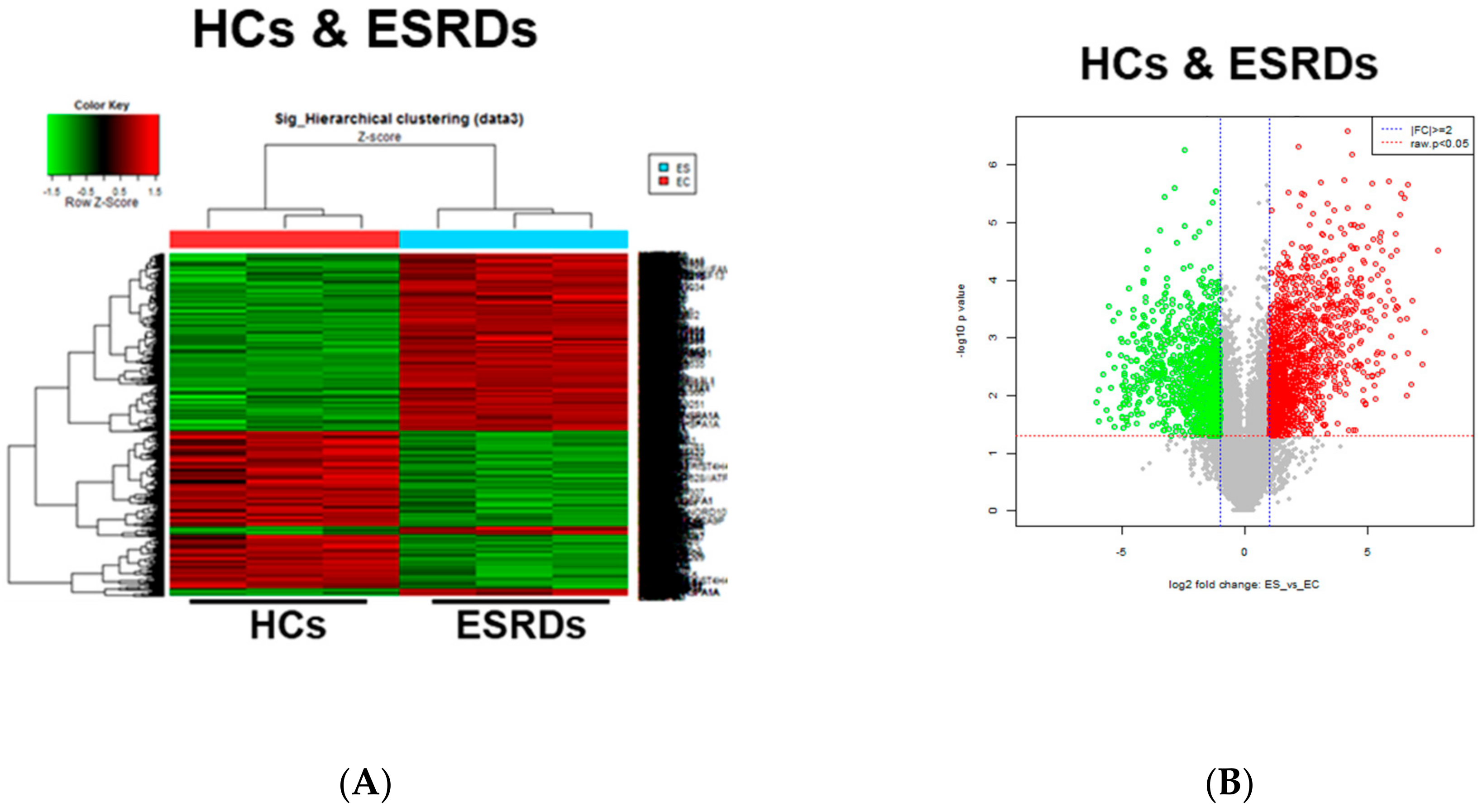
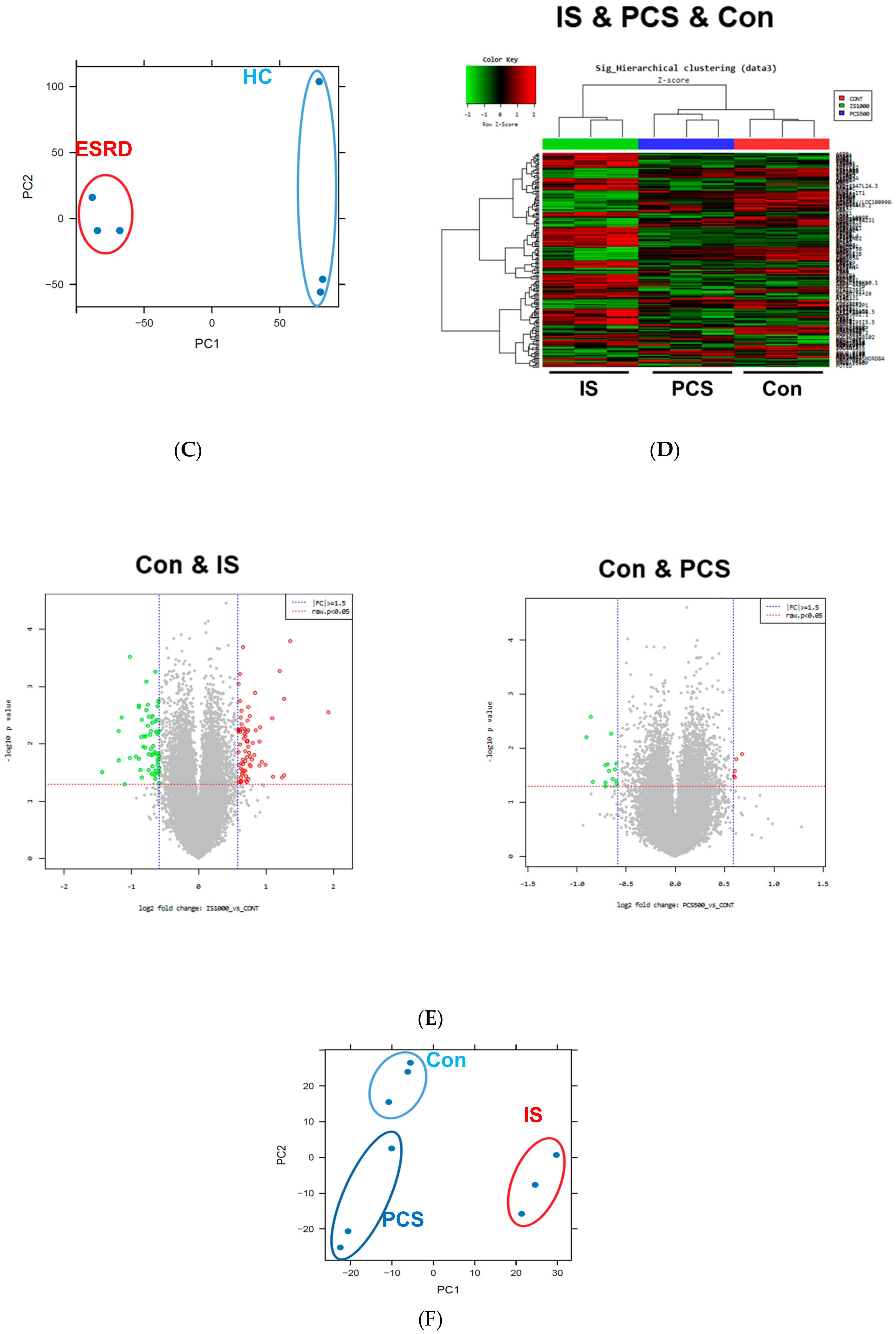
2.1. Transcriptomic Analysis of Monocytes Derived from ESRD Patients and Monocytes Stimulated with Major Uremic Toxins, IS and PCS
2.2. Common Differentially Expressed Genes (DEGs) between ESRD Patient-Derived Monocytes and IS-Stimulated Monocytes
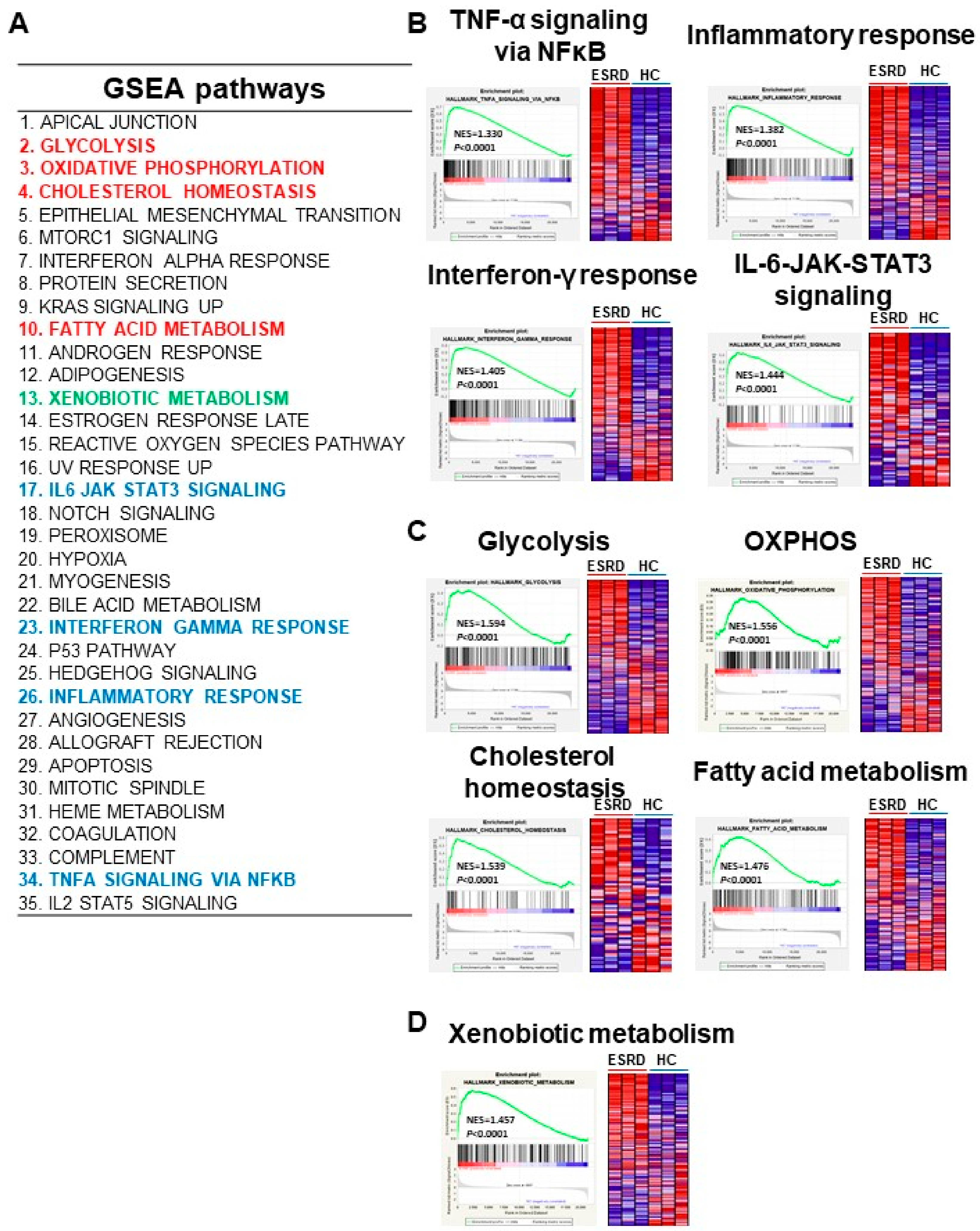
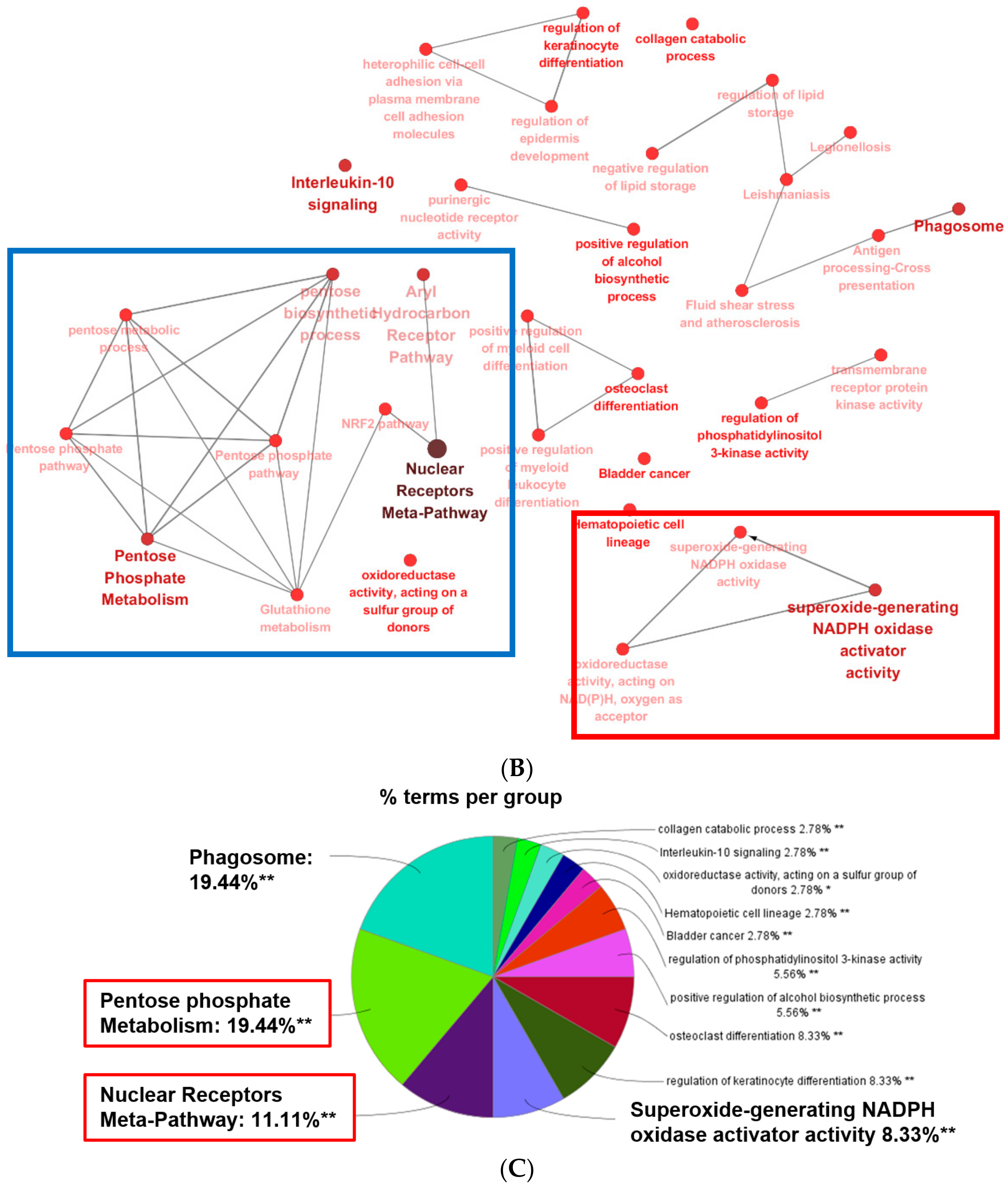
2.3. Principle Enriched Pathways in ESRD Patient-Derived Monocytes
2.4. Interaction Network Analysis of Common DEGs comparing ESRD Patient-Derived Monocytes and IS-Treated Monocytes
2.5. Experimental Validation of DEGs in IS-Treated Monocytes
3. Discussion
4. Materials and Methods
4.1. Peripheral Blood Mononuclear Cell (PBMC) Isolation
4.2. Sample Preparation for Microarray
4.3. Microarray Gene Expression Analysis
4.4. Data Preparation and Descriptive Statistics
4.5. Gene Expression Profiling and Analysis
4.6. Gene Set Enrichment Analysis
4.7. GO and Protein Pathway Analysis
4.8. Gene Functional Annotation and GO Annotation
4.9. Validation Microarray Genes-Quantitative RT-PCR
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Kanda, H.; Hirasaki, Y.; Iida, T.; Kanao-Kanda, M.; Toyama, Y.; Chiba, T.; Kunisawa, T. Perioperative Management of Patients with End-Stage Renal Disease. J. Cardiothorac. Vasc. Anesth. 2017, 31, 2251–2267. [Google Scholar] [CrossRef] [PubMed]
- Saran, R.; Robinson, B.; Abbott, K.C.; Agodoa, L.Y.C.; Bhave, N.; Bragg-Gresham, J.; Balkrishnan, R.; Dietrich, X.; Eckard, A.; Eggers, P.W.; et al. US Renal Data System 2017 Annual Data Report: Epidemiology of Kidney Disease in the United States. Am. J. Kidney Dis. Off. J. Natl. Kidney Found. 2018, 71, A7. [Google Scholar] [CrossRef] [PubMed]
- Foley, R.N.; Parfrey, P.S.; Sarnak, M.J. Clinical epidemiology of cardiovascular disease in chronic renal disease. Am. J. Kidney Dis. Off. J. Natl. Kidney Found. 1998, 32, S112–S119. [Google Scholar] [CrossRef]
- Sarnak, M.J.; Jaber, B.L. Mortality caused by sepsis in patients with end-stage renal disease compared with the general population. Kidney Int. 2000, 58, 1758–1764. [Google Scholar] [CrossRef] [PubMed]
- Collins, A.J.; Li, S.; Ma, J.Z.; Herzog, C. Cardiovascular disease in end-stage renal disease patients. Am. J. Kidney Dis. Off. J. Natl. Kidney Found. 2001, 38, S26–S29. [Google Scholar] [CrossRef] [PubMed]
- Cozzolino, M.; Mangano, M.; Stucchi, A.; Ciceri, P.; Conte, F.; Galassi, A. Cardiovascular disease in dialysis patients. Nephrol. Dial. Transplant. Off. Publ. Eur. Dial. Transplant. Assoc. Eur. Ren. Assoc. 2018, 33, iii28–iii34. [Google Scholar] [CrossRef]
- Ercan, E. Statin treatment in dialysis patients after acute myocardial infarction improves overall mortality. Atherosclerosis 2017, 267, 156–157. [Google Scholar] [CrossRef]
- Fellstrom, B.C.; Jardine, A.G.; Schmieder, R.E.; Holdaas, H.; Bannister, K.; Beutler, J.; Chae, D.W.; Chevaile, A.; Cobbe, S.M.; Gronhagen-Riska, C.; et al. Rosuvastatin and cardiovascular events in patients undergoing hemodialysis. N. Engl. J. Med. 2009, 360, 1395–1407. [Google Scholar] [CrossRef] [Green Version]
- Ferro, C.J.; Mark, P.B.; Kanbay, M.; Sarafidis, P.; Heine, G.H.; Rossignol, P.; Massy, Z.A.; Mallamaci, F.; Valdivielso, J.M.; Malyszko, J.; et al. Lipid management in patients with chronic kidney disease. Nat. Rev. Nephrol. 2018, 14, 727–749. [Google Scholar] [CrossRef] [Green Version]
- Cohen, G.; Glorieux, G.; Thornalley, P.; Schepers, E.; Meert, N.; Jankowski, J.; Jankowski, V.; Argiles, A.; Anderstam, B.; Brunet, P.; et al. Review on uraemic toxins III: Recommendations for handling uraemic retention solutes in vitro--towards a standardized approach for research on uraemia. Nephrol. Dial. Transplant. Off. Publ. Eur. Dial. Transplant. Assoc. Eur. Ren. Assoc. 2007, 22, 3381–3390. [Google Scholar] [CrossRef] [Green Version]
- Rossi, M.; Campbell, K.L.; Johnson, D.W.; Stanton, T.; Vesey, D.A.; Coombes, J.S.; Weston, K.S.; Hawley, C.M.; McWhinney, B.C.; Ungerer, J.P.; et al. Protein-bound uremic toxins, inflammation and oxidative stress: A cross-sectional study in stage 3-4 chronic kidney disease. Arch. Med Res. 2014, 45, 309–317. [Google Scholar] [CrossRef] [PubMed]
- Barisione, C.; Ghigliotti, G.; Canepa, M.; Balbi, M.; Brunelli, C.; Ameri, P. Indoxyl sulfate: A candidate target for the prevention and treatment of cardiovascular disease in chronic kidney disease. Curr. Drug Targets 2015, 16, 366–372. [Google Scholar] [CrossRef]
- Jing, Y.J.; Ni, J.W.; Ding, F.H.; Fang, Y.H.; Wang, X.Q.; Wang, H.B.; Chen, X.N.; Chen, N.; Zhan, W.W.; Lu, L.; et al. P-Cresyl sulfate is associated with carotid arteriosclerosis in hemodialysis patients and promotes atherogenesis in apoE-/- mice. Kidney Int. 2016, 89, 439–449. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Opdebeeck, B.; D’Haese, P.C.; Verhulst, A. Molecular and Cellular Mechanisms that Induce Arterial Calcification by Indoxyl Sulfate and P-Cresyl Sulfate. Toxins 2020, 12, 58. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kato, S.; Chmielewski, M.; Honda, H.; Pecoits-Filho, R.; Matsuo, S.; Yuzawa, Y.; Tranaeus, A.; Stenvinkel, P.; Lindholm, B. Aspects of immune dysfunction in end-stage renal disease. Clin. J. Am. Soc. Nephrol. 2008, 3, 1526–1533. [Google Scholar] [CrossRef] [Green Version]
- Kim, H.Y.; Yoo, T.H.; Hwang, Y.; Lee, G.H.; Kim, B.; Jang, J.; Yu, H.T.; Kim, M.C.; Cho, J.Y.; Lee, C.J.; et al. Indoxyl sulfate (IS)-mediated immune dysfunction provokes endothelial damage in patients with end-stage renal disease (ESRD). Sci. Rep. 2017, 7, 3057. [Google Scholar] [CrossRef] [Green Version]
- Nakano, T.; Katsuki, S.; Chen, M.; Decano, J.L.; Halu, A.; Lee, L.H.; Pestana, D.V.S.; Kum, A.S.T.; Kuromoto, R.K.; Golden, W.S.; et al. Uremic Toxin Indoxyl Sulfate Promotes Proinflammatory Macrophage Activation Via the Interplay of OATP2B1 and Dll4-Notch Signaling. Circulation 2019, 139, 78–96. [Google Scholar] [CrossRef]
- Luo, T.; Zheng, F.; Wang, K.; Xu, Y.; Xu, H.; Shen, W.; Zhu, C.; Zhang, X.; Sui, W.; Tang, D.; et al. A single-cell map for the transcriptomic signatures of peripheral blood mononuclear cells in end-stage renal disease. Nephrol. Dial. Transplant. Off. Publ. Eur. Dial. Transplant. Assoc. Eur. Ren. Assoc. 2019. [Google Scholar] [CrossRef]
- Al-Chaqmaqchi, H.A.; Moshfegh, A.; Dadfar, E.; Paulsson, J.; Hassan, M.; Jacobson, S.H.; Lundahl, J. Activation of Wnt/beta-catenin pathway in monocytes derived from chronic kidney disease patients. PLoS ONE 2013, 8, e68937. [Google Scholar] [CrossRef] [Green Version]
- Zhang, R.; Saredy, J.; Shao, Y.; Yao, T.; Liu, L.; Saaoud, F.; Yang, W.Y.; Sun, Y.; Johnson, C.; Drummer, C.T.; et al. End-stage renal disease is different from chronic kidney disease in upregulating ROS-modulated proinflammatory secretome in PBMCs—A novel multiple-hit model for disease progression. Redox Biol. 2020, 20, 101460. [Google Scholar] [CrossRef]
- Kitajima, S.; Iwata, Y.; Furuichi, K.; Sagara, A.; Shinozaki, Y.; Toyama, T.; Sakai, N.; Shimizu, M.; Sakurai, T.; Kaneko, S.; et al. Messenger RNA expression profile of sleep-related genes in peripheral blood cells in patients with chronic kidney disease. Clin. Exp. Nephrol. 2016, 20, 218–225. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chan, C.T.; Liu, P.P.; Arab, S.; Jamal, N.; Messner, H.A. Nocturnal hemodialysis improves erythropoietin responsiveness and growth of hematopoietic stem cells. J. Am. Soc. Nephrol. 2009, 20, 665–671. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lano, G.; Burtey, S.; Sallée, M. Indoxyl Sulfate, a Uremic Endotheliotoxin. Toxins 2020, 12, 229. [Google Scholar] [CrossRef] [Green Version]
- Lin, C.J.; Chuang, C.K.; Jayakumar, T.; Liu, H.L.; Pan, C.F.; Wang, T.J.; Chen, H.H.; Wu, C.J. Serum p-cresyl sulfate predicts cardiovascular disease and mortality in elderly hemodialysis patients. Arch. Med. Sci. 2013, 9, 662–668. [Google Scholar] [CrossRef]
- Lin, C.J.; Liu, H.L.; Pan, C.F.; Chuang, C.K.; Jayakumar, T.; Wang, T.J.; Chen, H.H.; Wu, C.J. Indoxyl sulfate predicts cardiovascular disease and renal function deterioration in advanced chronic kidney disease. Arch. Med. Res. 2012, 43, 451–456. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, W.C.; Tomino, Y.; Lu, K.C. Impacts of Indoxyl Sulfate and p-Cresol Sulfate on Chronic Kidney Disease and Mitigating Effects of AST-120. Toxins 2018, 10, 367. [Google Scholar] [CrossRef] [Green Version]
- Lin, C.J.; Pan, C.F.; Liu, H.L.; Chuang, C.K.; Jayakumar, T.; Wang, T.J.; Chen, H.H.; Wu, C.J. The role of protein-bound uremic toxins on peripheral artery disease and vascular access failure in patients on hemodialysis. Atherosclerosis 2012, 225, 173–179. [Google Scholar] [CrossRef]
- Barreto, F.C.; Barreto, D.V.; Liabeuf, S.; Meert, N.; Glorieux, G.; Temmar, M.; Choukroun, G.; Vanholder, R.; Massy, Z.A. Serum indoxyl sulfate is associated with vascular disease and mortality in chronic kidney disease patients. Clin. J. Am. Soc. Nephrol. 2009, 4, 1551–1558. [Google Scholar] [CrossRef] [Green Version]
- Vanholder, R.; Glorieux, G.; De Smet, R.; Lameire, N.; European Uremic Toxin Work Group. New insights in uremic toxins. Kidney Int. 2003, 63 (Suppl. 84), S6–S10. [Google Scholar] [CrossRef] [Green Version]
- Ito, S.; Osaka, M.; Higuchi, Y.; Nishijima, F.; Ishii, H.; Yoshida, M. Indoxyl sulfate induces leukocyte-endothelial interactions through up-regulation of E-selectin. J. Biol. Chem. 2010, 285, 38869–38875. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tumur, Z.; Shimizu, H.; Enomoto, A.; Miyazaki, H.; Niwa, T. Indoxyl sulfate upregulates expression of ICAM-1 and MCP-1 by oxidative stress-induced NF-kappaB activation. Am. J. Nephrol. 2010, 31, 435–441. [Google Scholar] [CrossRef]
- Gao, H.; Liu, S. Role of uremic toxin indoxyl sulfate in the progression of cardiovascular disease. Life Sci. 2017, 185, 23–29. [Google Scholar] [CrossRef] [PubMed]
- Espi, M.; Koppe, L.; Fouque, D.; Thaunat, O. Chronic Kidney Disease-Associated Immune Dysfunctions: Impact of Protein-Bound Uremic Retention Solutes on Immune Cells. Toxins 2020, 12, 300. [Google Scholar] [CrossRef] [PubMed]
- Hénaut, L.; Candellier, A.; Boudot, C.; Grissi, M.; Mentaverri, R.; Choukroun, G.; Brazier, M.; Kamel, S.; Massy, Z.A. New Insights into the Roles of Monocytes/Macrophages in Cardiovascular Calcification Associated with Chronic Kidney Disease. Toxins 2019, 11, 529. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, H.Y.; Yoo, T.H.; Cho, J.Y.; Kim, H.C.; Lee, W.W. Indoxyl sulfate-induced TNF-alpha is regulated by crosstalk between the aryl hydrocarbon receptor, NF-kappaB, and SOCS2 in human macrophages. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2019, 33, 10844–10858. [Google Scholar] [CrossRef] [Green Version]
- Wakamatsu, T.; Yamamoto, S.; Ito, T.; Sato, Y.; Matsuo, K.; Takahashi, Y.; Kaneko, Y.; Goto, S.; Kazama, J.J.; Gejyo, F.; et al. Indoxyl Sulfate Promotes Macrophage IL-1beta Production by Activating Aryl Hydrocarbon Receptor/NF-kappa/MAPK Cascades, but the NLRP3 inflammasome Was Not Activated. Toxins 2018, 10, 124. [Google Scholar] [CrossRef] [Green Version]
- Groh, L.; Keating, S.T.; Joosten, L.A.B.; Netea, M.G.; Riksen, N.P. Monocyte and macrophage immunometabolism in atherosclerosis. Semin. Immunopathol. 2018, 40, 203–214. [Google Scholar] [CrossRef]
- Yoon, B.R.; Oh, Y.J.; Kang, S.W.; Lee, E.B.; Lee, W.W. Role of SLC7A5 in Metabolic Reprogramming of Human Monocyte/Macrophage Immune Responses. Front. Immunol. 2018, 9, 53. [Google Scholar] [CrossRef] [Green Version]
- Yu, W.; Wang, Z.; Zhang, K.; Chi, Z.; Xu, T.; Jiang, D.; Chen, S.; Li, W.; Yang, X.; Zhang, X.; et al. One-Carbon Metabolism Supports S-Adenosylmethionine and Histone Methylation to Drive Inflammatory Macrophages. Mol. Cell 2019, 75, 1147–1160.e1145. [Google Scholar] [CrossRef]
- Palsson-McDermott, E.M.; O’Neill, L.A.J. Targeting immunometabolism as an anti-inflammatory strategy. Cell Res. 2020, 30, 300–314. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, C.Y.; Mauro, C. Similarities in the Metabolic Reprogramming of Immune System and Endothelium. Front. Immunol. 2017, 8, 837. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kelly, B.; O’Neill, L.A. Metabolic reprogramming in macrophages and dendritic cells in innate immunity. Cell Res. 2015, 25, 771–784. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bonacina, F.; Baragetti, A.; Catapano, A.L.; Norata, G.D. The Interconnection Between Immuno-Metabolism, Diabetes, and CKD. Curr. Diabetes Rep. 2019, 19, 21. [Google Scholar] [CrossRef]
- Sato, E.; Mori, T.; Mishima, E.; Suzuki, A.; Sugawara, S.; Kurasawa, N.; Saigusa, D.; Miura, D.; Morikawa-Ichinose, T.; Saito, R.; et al. Metabolic alterations by indoxyl sulfate in skeletal muscle induce uremic sarcopenia in chronic kidney disease. Sci. Rep. 2016, 6, 36618. [Google Scholar] [CrossRef]
- Stenvinkel, P.; Ketteler, M.; Johnson, R.J.; Lindholm, B.; Pecoits-Filho, R.; Riella, M.; Heimburger, O.; Cederholm, T.; Girndt, M. IL-10, IL-6, and TNF-alpha: Central factors in the altered cytokine network of uremia—The good, the bad, and the ugly. Kidney Int. 2005, 67, 1216–1233. [Google Scholar] [CrossRef] [Green Version]
- Tousoulis, D.; Oikonomou, E.; Economou, E.K.; Crea, F.; Kaski, J.C. Inflammatory cytokines in atherosclerosis: Current therapeutic approaches. Eur. Heart J. 2016, 37, 1723–1732. [Google Scholar] [CrossRef] [Green Version]
- Pioli, P.A.; Jensen, A.L.; Weaver, L.K.; Amiel, E.; Shen, Z.; Shen, L.; Wira, C.R.; Guyre, P.M. Estradiol attenuates lipopolysaccharide-induced CXC chemokine ligand 8 production by human peripheral blood monocytes. J. Immunol. 2007, 179, 6284–6290. [Google Scholar] [CrossRef] [Green Version]
- Kaufmann, A.; Salentin, R.; Gemsa, D.; Sprenger, H. Increase of CCR1 and CCR5 expression and enhanced functional response to MIP-1 alpha during differentiation of human monocytes to macrophages. J. Leukoc. Biol. 2001, 69, 248–252. [Google Scholar]
- Lubbers, R.; van Essen, M.F.; van Kooten, C.; Trouw, L.A. Production of complement components by cells of the immune system. Clin. Exp. Immunol. 2017, 188, 183–194. [Google Scholar] [CrossRef] [Green Version]
- Gistera, A.; Hansson, G.K. The immunology of atherosclerosis. Nat. Rev. Nephrol. 2017, 13, 368–380. [Google Scholar] [CrossRef] [PubMed]
- Kon, V.; Linton, M.F.; Fazio, S. Atherosclerosis in chronic kidney disease: The role of macrophages. Nat. Rev. Nephrol. 2011, 7, 45–54. [Google Scholar] [CrossRef] [Green Version]
- Patra, K.C.; Hay, N. The pentose phosphate pathway and cancer. Trends Biochem. Sci. 2014, 39, 347–354. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Baardman, J.; Verberk, S.G.S.; Prange, K.H.M.; van Weeghel, M.; van der Velden, S.; Ryan, D.G.; Wust, R.C.I.; Neele, A.E.; Speijer, D.; Denis, S.W.; et al. A Defective Pentose Phosphate Pathway Reduces Inflammatory Macrophage Responses during Hypercholesterolemia. Cell Rep. 2018, 25, 2044–2052.e2045. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Viaene, L.; Annaert, P.; de Loor, H.; Poesen, R.; Evenepoel, P.; Meijers, B. Albumin is the main plasma binding protein for indoxyl sulfate and p-cresyl sulfate. Biopharm. Drug Dispos. 2013, 34, 165–175. [Google Scholar] [CrossRef]
- Pei, J.; Juni, R.; Harakalova, M.; Duncker, D.J.; Asselbergs, F.W.; Koolwijk, P.; Hinsbergh, V.V.; Verhaar, M.C.; Mokry, M.; Cheng, C. Indoxyl Sulfate Stimulates Angiogenesis by Regulating Reactive Oxygen Species Production via CYP1B1. Toxins 2019, 11, 454. [Google Scholar] [CrossRef] [Green Version]
- Patel, A.A.; Zhang, Y.; Fullerton, J.N.; Boelen, L.; Rongvaux, A.; Maini, A.A.; Bigley, V.; Flavell, R.A.; Gilroy, D.W.; Asquith, B.; et al. The fate and lifespan of human monocyte subsets in steady state and systemic inflammation. J. Exp. Med. 2017, 214, 1913–1923. [Google Scholar] [CrossRef]
- Carvalho, B.S.; Irizarry, R.A. A framework for oligonucleotide microarray preprocessing. Bioinformatics 2010, 26, 2363–2367. [Google Scholar] [CrossRef]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic. Acids. Res. 2015, 43, e47. [Google Scholar] [CrossRef]
- Team, R.C. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2008. [Google Scholar]
- Kolde, R. Pheatmap: Pretty Heatmaps (Version 1.0.8). 2013. Available online: https://rdrr.io/cran/pheatmap (accessed on 11 December 2015).
- Mootha, V.K.; Lindgren, C.M.; Eriksson, K.F.; Subramanian, A.; Sihag, S.; Lehar, J.; Puigserver, P.; Carlsson, E.; Ridderstråle, M.; Laurila, E.; et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 2003, 34, 267–273. [Google Scholar] [CrossRef]
- Liberzon, A.; Birger, C.; Thorvaldsdóttir, H.; Ghandi, M.; Mesirov, J.P.; Tamayo, P. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 2015, 1, 417–425. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bindea, G.; Mlecnik, B.; Hackl, H.; Charoentong, P.; Tosolini, M.; Kirilovsky, A.; Fridman, W.H.; Pagès, F.; Trajanoski, Z.; Galon, J. ClueGO: A Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 2009, 25, 1091–1093. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bindea, G.; Galon, J.; Mlecnik, B. CluePedia Cytoscape plugin: Pathway insights using integrated experimental and in silico data. Bioinformatics 2013, 29, 661–663. [Google Scholar] [CrossRef] [PubMed]




| Probe ID | Symbol | Gene Name | Fold Change (log2) | ||
|---|---|---|---|---|---|
| ESRD | IS | ||||
| Top 50 Up-Regulated Genes | 16967771 | CXCL8 | C-X-C motif chemokine ligand 8 | 6.5945 | 0.3171 |
| 17000793 | CD14 | CD14 molecule | 6.1733 | 0.3099 | |
| 17058719 | NCF1C | neutrophil cytosolic factor 1C pseudogene | 4.9855 | 0.7591 | |
| 16716590 | MYOF | myoferlin | 4.7301 | 0.3701 | |
| 16857736 | MCEMP1 | mast cell expressed membrane protein 1 | 4.5776 | 0.6207 | |
| 16854486 | DSC2 | desmocollin 2 | 4.4608 | 0.3290 | |
| 16952874 | CCR1 | C-C motif chemokine receptor 1 | 4.3715 | 0.3022 | |
| 16869666 | ADGRE3 | adhesion G protein-coupled receptor E3 | 4.2049 | 0.7200 | |
| 16891082 | CYP27A1 | cytochrome P450 family 27 subfamily A member 1 | 4.0764 | 1.2708 | |
| 16709333 | TCF7L2 | transcription factor 7 like 2 | 3.9883 | 0.9611 | |
| 16684056 | FGR | FGR proto-oncogene, Src family tyrosine kinase | 3.8943 | 0.4057 | |
| 16768297 | DUSP6 | dual specificity phosphatase 6 | 3.8745 | 0.3807 | |
| 17086496 | DAPK1 | death associated protein kinase 1 | 3.8445 | 0.4779 | |
| 17059955 | PDK4 | pyruvate dehydrogenase kinase 4 | 3.5273 | 0.4756 | |
| 17096030 | FBP1 | fructose-bisphosphatase 1 | 3.4529 | 0.4545 | |
| 17046911 | NCF1B | neutrophil cytosolic factor 1B pseudogene | 3.3574 | 1.0349 | |
| 16896561 | CYP1B1 | cytochrome P450 family 1 subfamily B member 1 | 3.2051 | 1.5738 | |
| 16684144 | PTAFR | platelet activating factor receptor | 3.1942 | 0.3886 | |
| 17000724 | HBEGF | heparin binding EGF like growth factor | 2.8582 | 0.8480 | |
| 17044442 | SNX10 | sorting nexin 10 | 2.8334 | 0.3116 | |
| 16732985 | SLC37A2 | solute carrier family 37 member 2 | 2.7402 | 0.3424 | |
| 16658864 | PGD | phospho-gluconate dehydrogenase | 2.6875 | 0.7136 | |
| 17111711 | VSIG4 | V-set and immunoglobulin domain containing 4 | 2.6333 | 0.6154 | |
| 16805474 | ARRDC4 | arrestin domain containing 4 | 2.5744 | 0.3588 | |
| 16777685 | FLT3 | fms related tyrosine kinase 3 | 2.5176 | 0.4135 | |
| 17053697 | LOC644090 | uncharacterized LOC644090 | 2.4605 | 0.6179 | |
| 16991125 | NDST1 | N-deacetylase and N-sulfotransferase 1 | 2.3802 | 0.3088 | |
| 16669196 | CD101 | CD101 molecule | 2.2198 | 0.5474 | |
| 17063722 | CLEC5A | C-type lectin domain containing 5A | 2.2079 | 0.6715 | |
| 16875723 | TMEM150B | transmembrane protein 150B | 2.1721 | 0.3627 | |
| 16748788 | MGST1 | microsomal glutathione S-transferase 1 | 2.1668 | 0.8541 | |
| 16867784 | C3 | complement C3 | 2.1475 | 0.5078 | |
| 17121820 | USP32 | ubiquitin specific peptidase 32 | 2.1419 | 0.3092 | |
| 16929920 | H1F0 | H1 histone family member 0 | 2.0673 | 0.3007 | |
| 16848070 | SLC16A6 | solute carrier family 16 member 6 | 2.0182 | 0.4231 | |
| 16839524 | MIR22HG | MIR22 host gene | 2.0173 | 0.4588 | |
| 16774427 | LACC1 | laccase domain containing 1 | 1.9373 | 0.7574 | |
| 16729298 | ACER3 | alkaline ceramidase 3 | 1.9074 | 0.4178 | |
| 16894335 | ADAM17 | ADAM metallopeptidase domain 17 | 1.9073 | 0.3308 | |
| 17026267 | TNF | tumor necrosis factor | 1.8353 | 0.6968 | |
| 17013520 | SASH1 | SAM and SH3 domain containing 1 | 1.8301 | 0.6865 | |
| 17115636 | G6PD | glucose-6-phosphate dehydrogenase | 1.8165 | 0.4642 | |
| 16683445 | FUCA1 | alpha-L-fucosidase 1 | 1.7958 | 1.5007 | |
| 16876849 | ASAP2 | ArfGAP with SH3 domain, ankyrin repeat and PH domain 2 | 1.7922 | 0.3036 | |
| 16852871 | SERPINB2 | serpin family B member 2 | 1.7359 | 2.2714 | |
| 17058978 | GSAP | gamma-secretase activating protein | 1.7231 | 0.3256 | |
| 17071162 | CPQ | carboxypeptidase Q | 1.7211 | 0.3579 | |
| 16968447 | GPAT3 | glycerol-3-phosphate acyltransferase 3 | 1.7197 | 0.3522 | |
| 16720318 | TALDO1 | trans-aldolase 1 | 1.6930 | 0.3785 | |
| 16688929 | GNG5 | G protein subunit gamma 5 | 1.6846 | 0.3409 | |
| Top 50 Down-Regulated Genes | 16781893 | YME1L1 | YME1 like 1 ATPase | −5.1212 | −0.3594 |
| 16798132 | SNORD116-1 | small nucleolar RNA, C/D box 116-1 | −4.8499 | −0.8151 | |
| 17074313 | DEFA1B | defensin alpha 1B | −4.7542 | −0.3063 | |
| 16761201 | CD69 | CD69 molecule | −4.1940 | −0.3732 | |
| 16781830 | TRAJ17 | T cell receptor alpha joining 17 | −3.9577 | −0.3544 | |
| 16877297 | TRIB2 | tribbles pseudo-kinase 2 | −3.6828 | −0.3169 | |
| 16906571 | STAT4 | signal transducer and activator of transcription 4 | −3.6785 | −0.3412 | |
| 16798206 | SNORD116-20 | small nucleolar RNA, C/D box 116-20 | −3.3709 | −0.3708 | |
| 16672669 | LY9 | lymphocyte antigen 9 | −3.2701 | −0.3066 | |
| 16748205 | CLEC2D | C-type lectin domain family 2 member D | −3.2426 | −0.3819 | |
| 16761631 | DUSP16 | dual specificity phosphatase 16 | −3.1818 | −0.4038 | |
| 17110670 | PIM2 | Pim-2 proto-oncogene, serine/threonine kinase | −2.9802 | −0.6422 | |
| 16912130 | CST7 | cystatin F | −2.5494 | −0.3388 | |
| 17092767 | MLLT3 | MLLT3, super elongation complex subunit | −2.4775 | −0.7344 | |
| 16720085 | IFITM1 | interferon induced transmembrane protein 1 | −2.4475 | −0.3137 | |
| 17056849 | TRGV3 | T cell receptor gamma variable 3 | −2.4407 | −0.8184 | |
| 17005858 | HIST1H2AI | histone cluster 1 H2A family member i | −2.2799 | −0.4418 | |
| 16995717 | SNORD72 | small nucleolar RNA, C/D box 72 | −2.2604 | −0.7078 | |
| 17016503 | HIST1H3I | histone cluster 1 H3 family member i | −2.2455 | −0.4036 | |
| 16764564 | LIMA1 | LIM domain and actin binding 1 | −2.2305 | −0.3310 | |
| 16934045 | PIK3IP1 | phosphoinositide-3-kinase interacting protein 1 | −2.2022 | −0.4481 | |
| 16900737 | ANKRD36B | ankyrin repeat domain 36B | −2.0806 | −0.3064 | |
| 17016386 | HIST1H3D | histone cluster 1 H3 family member d | −2.0589 | −0.3624 | |
| 17122454 | ANKRD36 | ankyrin repeat domain 36 | −2.0333 | −0.5181 | |
| 17121094 | SRP54-AS1 | SRP54 antisense RNA 1 (head to head) | −1.9509 | −0.4420 | |
| 16756286 | C12orf75 | chromosome 12 open reading frame 75 | −1.8645 | −0.3758 | |
| 16798248 | SNHG14 | small nucleolar RNA host gene 14 | −1.8228 | −0.3124 | |
| 17122912 | ANKRD20A5P | ankyrin repeat domain 20 family member A5, pseudogene | −1.8010 | −0.3463 | |
| 16686060 | SLC2A1 | solute carrier family 2 member 1 | −1.7904 | −0.3997 | |
| 16968314 | PRDM8 | PR/SET domain 8 | −1.7855 | −0.5214 | |
| 17016383 | HIST1H4D | histone cluster 1 H4 family member d | −1.7560 | −0.3952 | |
| 16837296 | ARHGAP27P2 | Rho GTPase activating protein 27 pseudogene 2 | −1.7467 | −0.4159 | |
| 17005560 | HIST1H4C | histone cluster 1 H4 family member c | −1.7216 | −0.3526 | |
| 16849349 | TNRC6C-AS1 | TNRC6C antisense RNA 1 | −1.6941 | −0.6821 | |
| 17016360 | HIST1H4B | histone cluster 1 H4 family member b | −1.6893 | −0.5007 | |
| 17011708 | SLC16A10 | solute carrier family 16 member 10 | −1.6648 | −0.4770 | |
| 16780585 | GPR18 | G protein-coupled receptor 18 | −1.6552 | −0.4941 | |
| 17116745 | P2RY8 | P2Y receptor family member 8 | −1.6285 | −0.4951 | |
| 16987361 | ARSK | arylsulfatase family member K | −1.6196 | −0.3051 | |
| 16869324 | PRDX2 | peroxiredoxin 2 | −1.6134 | −0.4093 | |
| 16980528 | LRBA | LPS responsive beige-like anchor protein | −1.6071 | −0.3149 | |
| 16838300 | TMC8 | transmembrane channel like 8 | −1.5657 | −0.3583 | |
| 17100888 | LOC102723630 | uncharacterized LOC102723630 | −1.5554 | −0.4080 | |
| 16956983 | CBLB | Cbl proto-oncogene B | −1.5203 | −0.5545 | |
| 16930173 | APOBEC3D | apolipoprotein B mRNA editing enzyme catalytic subunit 3D | −1.5054 | −0.4054 | |
| 16985518 | PIK3R1 | phosphoinositide-3-kinase regulatory subunit 1 | −1.5016 | −0.5210 | |
| 17113606 | septin 6 | septin 6 | −1.5012 | −0.4799 | |
| 16993173 | LOC728554 | THO complex 3 pseudo-gene | −1.4969 | −0.3166 | |
| 16889160 | PLCL1 | phospholipase C like 1 (inactive) | −1.4891 | −0.3684 | |
| 17013197 | PEX3 | peroxisomal biogenesis factor 3 | −1.4474 | −0.3493 | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, H.Y.; Lee, S.J.; Hwang, Y.; Lee, G.H.; Yoon, C.E.; Kim, H.C.; Yoo, T.-H.; Lee, W.-W. Indoxyl Sulfate-Mediated Metabolic Alteration of Transcriptome Signatures in Monocytes of Patients with End-Stage Renal Disease (ESRD). Toxins 2020, 12, 621. https://doi.org/10.3390/toxins12100621
Kim HY, Lee SJ, Hwang Y, Lee GH, Yoon CE, Kim HC, Yoo T-H, Lee W-W. Indoxyl Sulfate-Mediated Metabolic Alteration of Transcriptome Signatures in Monocytes of Patients with End-Stage Renal Disease (ESRD). Toxins. 2020; 12(10):621. https://doi.org/10.3390/toxins12100621
Chicago/Turabian StyleKim, Hee Young, Su Jeong Lee, Yuri Hwang, Ga Hye Lee, Chae Eun Yoon, Hyeon Chang Kim, Tae-Hyun Yoo, and Won-Woo Lee. 2020. "Indoxyl Sulfate-Mediated Metabolic Alteration of Transcriptome Signatures in Monocytes of Patients with End-Stage Renal Disease (ESRD)" Toxins 12, no. 10: 621. https://doi.org/10.3390/toxins12100621





