Using In Silico Approach for Metabolomic and Toxicity Prediction of Alternariol
Abstract
:1. Introduction
2. Results
2.1. Metabolomics Profile of Alternariol
2.2. Physicochemical Properties, Pharmacokinetic Predictions and Drug Likeness
2.3. Prediction of Toxicity
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Prediction of Alternariol Metabolites
5.2. Prediction of Physicochemical Properties, Pharmacokinetic Predictions and Drug Likeness
5.3. Prediction of Toxicity
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- López, P.; Venema, D.; Mol, H.; Spanjer, M.; de Stoppelaar, J.; Pfeiffer, E.; de Nijs, M. Alternaria toxins and conjugates in selected foods in the netherlands. Food Control 2016, 69, 153–159. [Google Scholar] [CrossRef]
- Fraeyman, S.; Croubels, S.; Devreese, M.; Antonissen, G. Emerging fusarium and alternaria mycotoxins: Occurrence, toxicity and toxicokinetics. Toxins 2017, 9, 228. [Google Scholar] [CrossRef] [PubMed]
- Awuchi, C.G.; Ondari, E.N.; Ogbonna, C.U.; Upadhyay, A.K.; Baran, K.; Okpala, C.O.R.; Korzeniowska, M.; Guiné, R.P.F. Mycotoxins affecting animals, foods, humans, and plants: Types, occurrence, toxicities, action mechanisms, prevention, and detoxification strategies—A revisit. Foods 2021, 10, 1279. [Google Scholar] [CrossRef]
- Meena, M.; Samal, S. Alternaria host-specific (hsts) toxins: An overview of chemical characterization, target sites, regulation and their toxic effects. Toxicol. Rep. 2019, 6, 745–758. [Google Scholar] [CrossRef]
- Crudo, F.; Varga, E.; Aichinger, G.; Galaverna, G.; Marko, D.; Dall’Asta, C.; Dellafiora, L. Co-occurrence and combinatory effects of alternaria mycotoxins and other xenobiotics of food origin: Current scenario and future perspectives. Toxins 2019, 11, 640. [Google Scholar] [CrossRef] [PubMed]
- Hickert, S.; Bergmann, M.; Ersen, S.; Cramer, B.; Humpf, H.U. Survey of alternaria toxin contamination in food from the german market, using a rapid hplc-ms/ms approach. Mycotoxin Res. 2016, 32, 7–18. [Google Scholar] [CrossRef]
- Walravens, J.; Mikula, H.; Rychlik, M.; Asam, S.; Devos, T.; Njumbe Ediage, E.; Diana Di Mavungu, J.; Jacxsens, L.; Van Landschoot, A.; Vanhaecke, L.; et al. Validated uplc-ms/ms methods to quantitate free and conjugated alternaria toxins in commercially available tomato products and fruit and vegetable juices in belgium. J. Agric. Food Chem. 2016, 64, 5101–5109. [Google Scholar] [CrossRef]
- EFSA; Arcella, D.; Eskola, M.; Gómez Ruiz, J.A. Dietary exposure assessment to alternaria toxins in the european population. EFSA J. 2016, 14, e04654. [Google Scholar]
- Alexander, J.; Benford, D.; Boobis, A.; Ceccatelli, S.; Cottrill, B.; Cravedi, J.-P.; Doerge, D.; Dogliotti, E.; Edler, L.; Farmer, P.; et al. Scientific opinion on the risks for animal and public health related to the presence of alternaria toxins in feed and food—efsa panel on contaminants in the food chain (contam). EFSA J. 2011, 9, 1–97. [Google Scholar]
- Johnson, C.H.; Patterson, A.D.; Idle, J.R.; Gonzalez, F.J. Xenobiotic metabolomics: Major impact on the metabolome. Annu. Rev. Pharmacol. Toxicol. 2012, 52, 37–56. [Google Scholar] [CrossRef]
- Zhang, Z.; Nie, D.; Fan, K.; Yang, J.; Guo, W.; Meng, J.; Zhao, Z.; Han, Z. A systematic review of plant-conjugated masked mycotoxins: Occurrence, toxicology, and metabolism. Crit. Rev. Food Sci. Nutr. 2020, 60, 1523–1537. [Google Scholar] [CrossRef]
- Juan-García, A.; Juan, C.; Tolosa, J.; Ruiz, M.-J. Effects of deoxynivalenol, 3-acetyl-deoxynivalenol and 15-acetyl-deoxynivalenol on parameters associated with oxidative stress in hepg2 cells. Mycotoxin Res. 2019, 35, 197–205. [Google Scholar] [CrossRef]
- Balázs, A.; Faisal, Z.; Csepregi, R.; Kőszegi, T.; Kriszt, B.; Szabó, I.; Poór, M. In vitro evaluation of the individual and combined cytotoxic and estrogenic effects of zearalenone, its reduced metabolites, alternariol, and genistein. Int. J. Mol. Sci. 2021, 22, 6281. [Google Scholar] [CrossRef]
- Puntscher, H.; Cobankovic, I.; Marko, D.; Warth, B. Quantitation of free and modified alternaria mycotoxins in european food products by lc-ms/ms. Food Control 2019, 102, 157–165. [Google Scholar] [CrossRef]
- Johnson, W.W.; Yamazaki, H.; Shimada, T.; Ueng, Y.-F.; Guengerich, F.P. Aflatoxin b1 8,9-epoxide hydrolysis in the presence of rat and human epoxide hydrolase. Chem. Res. Toxicol. 1997, 10, 672–676. [Google Scholar] [CrossRef] [PubMed]
- Tiessen, C.; Ellmer, D.; Mikula, H.; Pahlke, G.; Warth, B.; Gehrke, H.; Zimmermann, K.; Heiss, E.; Fröhlich, J.; Marko, D. Impact of phase i metabolism on uptake, oxidative stress and genotoxicity of the emerging mycotoxin alternariol and its monomethyl ether in esophageal cells. Arch. Toxicol. 2017, 91, 1213–1226. [Google Scholar] [CrossRef] [PubMed]
- Pfeiffer, P.; Goedecke, W.; Obe, G. Mechanisms of DNA double-strand break repair and their potential to induce chromosomal aberrations. Mutagenesis 2000, 15, 289–302. [Google Scholar] [CrossRef]
- Dellafiora, L.; Warth, B.; Schmidt, V.; Del Favero, G.; Mikula, H.; Fröhlich, J.; Marko, D. An integrated in silico/in vitro approach to assess the xenoestrogenic potential of alternaria mycotoxins and metabolites. Food Chem. 2018, 248, 253–261. [Google Scholar] [CrossRef]
- Hemmerich, J.; Ecker, G.F. In silico toxicology: From structure-activity relationships towards deep learning and adverse outcome pathways. Comput. Mol. Sci. 2020, 10, e1475. [Google Scholar] [CrossRef]
- Moroy, G.; Martiny, V.Y.; Vayer, P.; Villoutreix, B.O.; Miteva, M.A. Toward in silico structure-based admet prediction in drug discovery. Drug Discov. Today 2012, 17, 44–55. [Google Scholar] [CrossRef]
- Van de Waterbeemd, H.; Gifford, E. Admet in silico modelling: Towards prediction paradise? Nat. Rev. Drug Discov. 2003, 2, 192–204. [Google Scholar] [CrossRef] [PubMed]
- Dellafiora, L.; Oswald, I.P.; Dorne, J.-L.; Galaverna, G.; Battilani, P.; Dall’Asta, C. An in silico structural approach to characterize human and rainbow trout estrogenicity of mycotoxins: Proof of concept study using zearalenone and alternariol. Food Chem. 2020, 312, 126088. [Google Scholar] [CrossRef] [PubMed]
- Daina, A.; Michielin, O.; Zoete, V. Swissadme: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef] [PubMed]
- Pires, D.E.V.; Blundell, T.L.; Ascher, D.B. Pkcsm: Predicting small-molecule pharmacokinetic and toxicity properties using graph-based signatures. J. Med. Chem. 2015, 58, 4066–4072. [Google Scholar] [CrossRef] [PubMed]
- Agahi, F.; Juan, C.; Font, G.; Juan-García, A. In silico methods for metabolomic and toxicity prediction of zearalenone, α-zearalenone and β-zearalenone. Food Chem. Toxicol. 2020, 146, 111818. [Google Scholar] [CrossRef] [PubMed]
- Domínguez-Villa, F.X.; Durán-Iturbide, N.A.; Ávila-Zárraga, J.G. Synthesis, molecular docking, and in silico adme/tox profiling studies of new 1-aryl-5-(3-azidopropyl)indol-4-ones: Potential inhibitors of SARS-CoV-2 main protease. Bioorganic Chem. 2021, 106, 104497. [Google Scholar] [CrossRef] [PubMed]
- Mvondo, J.; Matondo, A.; Mawete, D.; Bambi, S.-M.; Mbala, B.; Lohohola, P. In silico ADME/t properties of quinine derivatives using swissadme and pkcsm webservers. Int. J. Trop. Dis. Health 2021, 42, 1–12. [Google Scholar] [CrossRef]
- Lipinski, C.A.; Lombardo, F.; Dominy, B.W.; Feeney, P.J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 1997, 23, 3–25. [Google Scholar] [CrossRef]
- Lin, J.H.; Yamazaki, M. Role of p-glycoprotein in pharmacokinetics: Clinical implications. Clin. Pharmacokinet. 2003, 42, 59–98. [Google Scholar] [CrossRef]
- Neha, M.; Kishore, K.P.; Benjamin, D.M.; Amanda, M.; Juliet, M.M.; Sabrinia, C.; Joseph, W.P.; Andrew, D.W. Predicting volume of distribution in humans: Performance of in silico methods for a large set of structurally diverse clinical compounds. Drug Metab. Dispos. 2021, 49, 169. [Google Scholar]
- Shargel, L.; Wu-Pong, S.; Yu, A.B.C. Chapter 10. Physiologic drug distribution and protein binding. In Applied Biopharmaceutics & Pharmacokinetics, 6th ed.; The McGraw-Hill Companies: New York, NY, USA, 2012. [Google Scholar]
- Filimonov, D.A.; Lagunin, A.; Gloriozova, T.; Rudik, A.V.; Druzhilovskii, D.S.; Pogodin, P.; Poroikov, V. Prediction of the biological activity spectra of organic compounds using the pass online web resource. Chem. Heterocycl. Compd. 2014, 50, 444–457. [Google Scholar] [CrossRef]
- Rudik, A.V.; Bezhentsev, V.M.; Dmitriev, A.V.; Druzhilovskiy, D.S.; Lagunin, A.A.; Filimonov, D.A.; Poroikov, V.V. Metatox: Web application for predicting structure and toxicity of xenobiotics’ metabolites. J. Chem. Inf. Model. 2017, 57, 638–642. [Google Scholar] [CrossRef]
- Scheibenzuber, S.; Hoffmann, T.; Effenberger, I.; Schwab, W.; Asam, S.; Rychlik, M. Enzymatic synthesis of modified alternaria mycotoxins using a whole-cell biotransformation system. Toxins 2020, 12, 264. [Google Scholar] [CrossRef] [PubMed]
- Soukup, S.T.; Kohn, B.N.; Pfeiffer, E.; Geisen, R.; Metzler, M.; Bunzel, M.; Kulling, S.E. Sulfoglucosides as novel modified forms of the mycotoxins alternariol and alternariol monomethyl ether. J. Agric. Food Chem. 2016, 64, 8892–8901. [Google Scholar] [CrossRef] [PubMed]
- Pfeiffer, E.; Schmit, C.; Burkhardt, B.; Altemöller, M.; Podlech, J.; Metzler, M. Glucuronidation of the mycotoxins alternariol and alternariol-9-methyl ether in vitro: Chemical structures of glucuronides and activities of human udp-glucuronosyltransferase isoforms. Mycotoxin Res. 2009, 25, 3–10. [Google Scholar] [CrossRef]
- Burkhardt, B.; Pfeiffer, E.; Metzler, M. Absorption and metabolism of the mycotoxins alternariol and alternariol-9-methyl ether in caco-2 cells in vitro. Mycotoxin Res. 2009, 25, 149–157. [Google Scholar] [CrossRef]
- Appel, B.; Gottmann, J.; Schäfer, J.; Bunzel, M. Absorption and metabolism of modified mycotoxins of alternariol, alternariol monomethyl ether, and zearalenone in caco-2 cells. Cereal Chem. 2021, 98, 109–122. [Google Scholar] [CrossRef]
- Fehr, M.; Pahlke, G.; Fritz, J.; Christensen, M.O.; Boege, F.; Altemöller, M.; Podlech, J.; Marko, D. Alternariol acts as a topoisomerase poison, preferentially affecting the iialpha isoform. Mol. Nutr. Food Res. 2009, 53, 441–451. [Google Scholar] [CrossRef]
- EFSA. Scientific opinion on the risks for human and animal health related to the presence of modified forms of certain mycotoxins in food and feed. EFSA J. 2014, 12, 3916. [Google Scholar]
- McDonnell, A.M.; Dang, C.H. Basic review of the cytochrome p450 system. J. Adv. Pract. Oncol. 2013, 4, 263–268. [Google Scholar]
- Lootens, O.; Vermeulen, A.; Croubels, S.; De Saeger, S.; Van Bocxlaer, J.; De Boevre, M. Possible mechanisms of the interplay between drugs and mycotoxins—is there a possible impact? Toxins 2022, 14, 873. [Google Scholar] [PubMed]
- Guengerich, F.P. A history of the roles of cytochrome p450 enzymes in the toxicity of drugs. Toxicol. Res. 2021, 37, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Pahlke, G.; Tiessen, C.; Domnanich, K.; Kahle, N.; Groh, I.A.M.; Schreck, I.; Weiss, C.; Marko, D. Impact of alternaria toxins on cyp1a1 expression in different human tumor cells and relevance for genotoxicity. Toxicol. Lett. 2016, 240, 93–104. [Google Scholar] [CrossRef]
- Schreck, I.; Deigendesch, U.; Burkhardt, B.; Marko, D.; Weiss, C. The alternaria mycotoxins alternariol and alternariol methyl ether induce cytochrome p450 1a1 and apoptosis in murine hepatoma cells dependent on the aryl hydrocarbon receptor. Arch. Toxicol. 2012, 86, 625–632. [Google Scholar] [CrossRef] [PubMed]
- Fliszár-Nyúl, E.; Ungvári, O.; Dombi, Á.; Ozvegy-Laczka, C.; Poór, M. Interactions of mycotoxin alternariol with cytochrome p450 enzymes and oatp transporters. Metabolites 2023, 13, 45. [Google Scholar] [CrossRef]
- Catania, V.A.; Sánchez Pozzi, E.J.; Luquita, M.G.; Ruiz, M.L.; Villanueva, S.S.M.; Jones, B.; Mottino, A.D. Co-regulation of expression of phase ii metabolizing enzymes and multidrug resistance-associated protein 2. Ann. Hepatol. 2004, 3, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Pan, L.L.; Yang, Y.; Hui, M.; Wang, S.; Li, C.Y.; Zhang, H.; Ding, Y.H.; Fu, L.; Diao, X.X.; Zhong, D.F. Sulfation predominates the pharmacokinetics, metabolism, and excretion of forsythin in humans: Major enzymes and transporters identified. Acta Pharmacol. Sin. 2021, 42, 311–322. [Google Scholar] [CrossRef]
- Pavicich, M.A.; De Boevre, M.; Vidal, A.; Iturmendi, F.; Mikula, H.; Warth, B.; Marko, D.; De Saeger, S.; Patriarca, A. Fate of free and modified alternaria mycotoxins during the production of apple concentrates. Food Control 2020, 118, 107388. [Google Scholar] [CrossRef]
- Zwickel, T.; Kahl, S.M.; Klaffke, H.; Rychlik, M.; Müller, M.E.H. Spotlight on the underdogs—An analysis of underrepresented alternaria mycotoxins formed depending on varying substrate, time and temperature conditions. Toxins 2016, 8, 344. [Google Scholar] [CrossRef]
- Puntscher, H.; Aichinger, G.; Grabher, S.; Attakpah, E.; Krüger, F.; Tillmann, K.; Motschnig, T.; Hohenbichler, J.; Braun, D.; Plasenzotti, R.; et al. Bioavailability, metabolism, and excretion of a complex alternaria culture extract versus altertoxin ii: A comparative study in rats. Arch. Toxicol. 2019, 93, 3153–3167. [Google Scholar] [CrossRef]
- Hayes, J.D.; McLellan, L.I. Glutathione and glutathione-dependent enzymes represent a co-ordinately regulated defence against oxidative stress. Free Radic. Res. 1999, 31, 273–300. [Google Scholar] [CrossRef]
- Fernández-Blanco, C.; Font, G.; Ruiz, M.J. Oxidative stress of alternariol in caco-2 cells. Toxicol. Lett. 2014, 229, 458–464. [Google Scholar] [CrossRef] [PubMed]
- Li, H.S.; Zhou, Y.N.; Li, L.; Li, S.F.; Long, D.; Chen, X.L.; Zhang, J.B.; Feng, L.; Li, Y.P. Hif-1α protects against oxidative stress by directly targeting mitochondria. Redox Biol. 2019, 25, 101109. [Google Scholar] [CrossRef] [PubMed]
- Bitew, M.; Desalegn, T.; Demissie, T.B.; Belayneh, A.; Endale, M.; Eswaramoorthy, R. Pharmacokinetics and drug-likeness of antidiabetic flavonoids: Molecular docking and dft study. PLoS ONE 2021, 16, e0260853. [Google Scholar] [CrossRef] [PubMed]
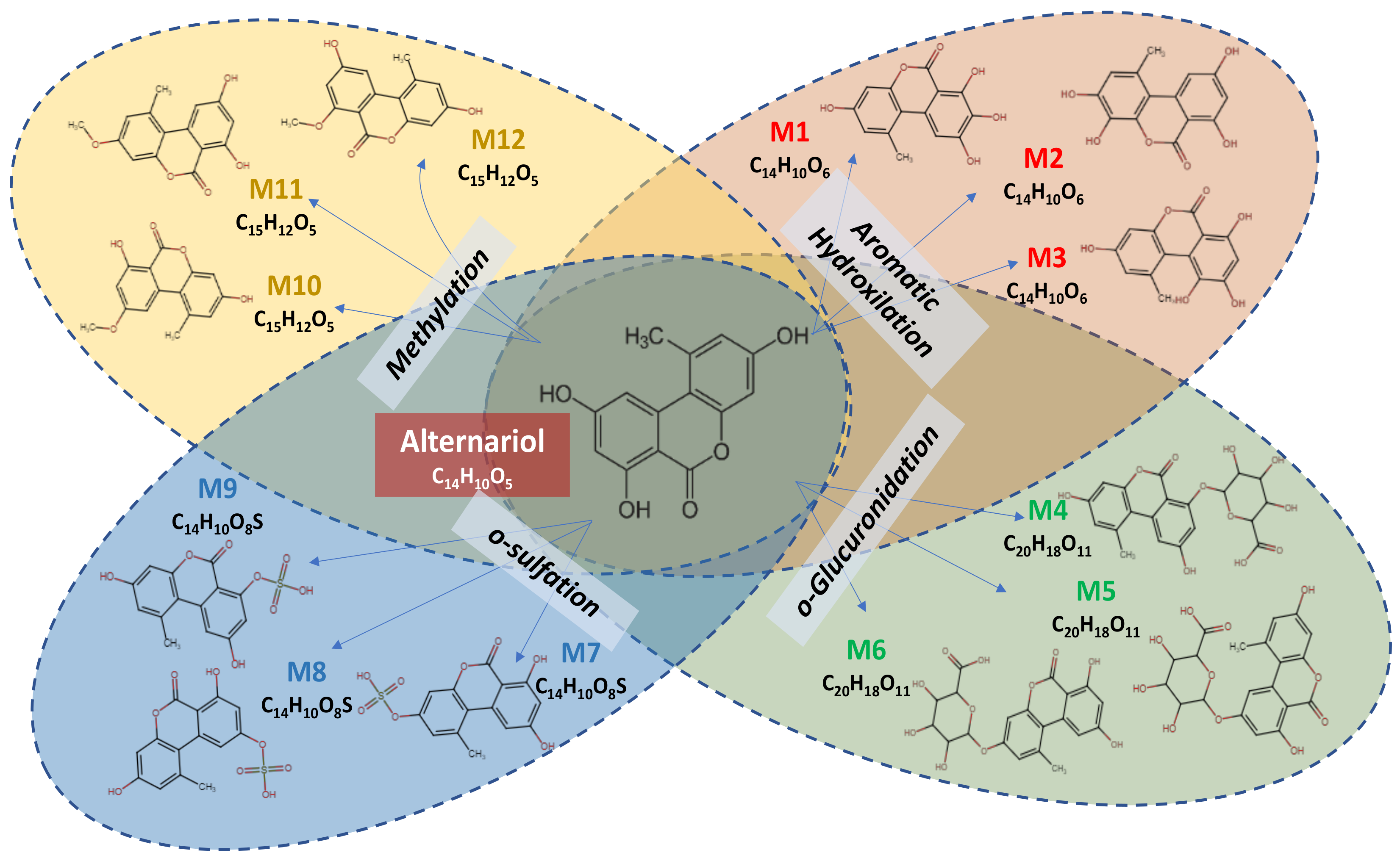





| Parent Compound/Metabolite | Canonical SMILES |
|---|---|
| Alternariol (AOH) | CC1=CC(O)=CC2=C1C1=CC(O)=CC(O)=C1C(=O)O2 |
| Aromatic Hydroxylation | |
| Metabolite (M1) | CC1=CC(O)=CC2=C1C1=CC(O)=C(O)C(O)=C1C(=O)O2 |
| Metabolite (M2) | CC1=CC(O)=C(O)C2=C1C1=CC(O)=CC(O)=C1C(=O)O2 |
| Metabolite (M3) | CC1=CC(O)=CC2=C1C1=C(O)C(O)=CC(O)=C1C(=O)O2 |
| O-Glucuronidation | |
| Metabolite (M4) | CC1=CC(O)=CC2=C1C1=CC(O)=CC(OC3OC(C(O)C(O)C3O)C(O)=O)=C1C(=O)O2 |
| Metabolite (M5) | CC1=CC(O)=CC2=C1C1=CC(OC3OC(C(O)C(O)C3O)C(O)=O)=CC(O)=C1C(=O)O2 |
| Metabolite (M6) | CC1=CC(OC2OC(C(O)C(O)C2O)C(O)=O)=CC2=C1C1=CC(O)=CC(O)=C1C(=O)O2 |
| O-Sulfation | |
| Metabolite (M7) | CC1=CC(OS(O)(=O)=O)=CC2=C1C1=CC(O)=CC(O)=C1C(=O)O2 |
| Metabolite (M8) | CC1=CC(O)=CC2=C1C1=CC(OS(O)(=O)=O)=CC(O)=C1C(=O)O2 |
| Metabolite (M9) | CC1=CC(O)=CC2=C1C1=CC(O)=CC(OS(O)(=O)=O)=C1C(=O)O2 |
| Methylation | |
| Metabolite (M10) | COC1=CC(O)=C2C(=O)OC3=CC(O)=CC(C)=C3C2=C1 |
| Metabolite (M11) | COC1=CC(C)=C2C(OC(=O)C3=C(O)C=C(O)C=C23)=C1 |
| Metabolite (M12) | COC1=CC(O)=CC2=C1C(=O)OC1=CC(O)=CC(C)=C21 |
| Parent Compound/Metabolite | MW (g/mol) (≤500) | HBA (≤10) | HBD (≤5) | cLogP (<5) | MR (40–130) | n-ROTB | TPSA (Å2) |
|---|---|---|---|---|---|---|---|
| Alternariol (AOH) | 258.23 | 5 | 3 | 71.03 | 0 | 90.9 | |
| Aromatic Hydroxylation | |||||||
| Metabolite (M1) | 274.23 | 6 | 4 | 1.71 | 73.05 | 0 | 111.1 |
| Metabolite (M2) | 1.72 | ||||||
| Metabolite (M3) | 1.46 | ||||||
| O-Glucuronidation | |||||||
| Metabolite (M4) | 434.35 | 11 * | 6 * | 0.06 | 103.79 | 3 | 187.1 |
| Metabolite (M5) | 0.31 | ||||||
| Metabolite (M6) | 0.31 | ||||||
| O-Sulfation | |||||||
| Metabolite (M7) | 338.29 | 8 | 3 | 1.58 | 81.22 | 3 | 142.6 |
| Metabolite (M8) | 1.57 | ||||||
| Metabolite (M9) | 1.38 | ||||||
| Methylation | |||||||
| Metabolite (M10) | 272.25 | 5 | 2 | 2.55 | 75.49 | 1 | 79.9 |
| Metabolite (M11) | 2.55 | ||||||
| Metabolite (M12) | 2.55 |
| Model Name/Parameters | AOH | Aromatic Hydroxilation | O-Glucuronidation | O-Sulfation | Methylation | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| M1 | M2 | M3 | M4 | M5 | M6 | M7 | M8 | M9 | M10 | M11 | M12 | ||
| Absorption | |||||||||||||
| Water solubility (log mol/L) | −2.982 | −3.059 | −3.151 | −3.072 | −2.892 | −2.894 | −2.894 | −3.158 | −3.118 | −2.938 | −3.5 | −3.293 | −3.388 |
| Caco2 permeability (log Paap in 10−6 cm/s) | 1.025 | 0.838 | 0.815 | 0.818 | −0.745 | −0.699 | −0.885 | 0.388 | 0.635 | 0.717 | 1.057 | 0.952 | 0.9 |
| Intestinal absorption (human) % | 95.473 | 73.662 | 81.84 | 76.718 | 18.049 | 18.043 | 14.941 | 45.397 | 48.67 | 47.87 | 95.627 | 95.804 | 97.087 |
| Skin Permeability (log Kp) | −2.745 | −2.735 | −2.735 | −2.735 | −2.735 | −2.735 | −2.735 | −2.735 | −2.735 | −2.735 | −2.739 | −2.737 | −2.747 |
| P-glycoprotein substrate | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes |
| P-glycoprotein I inhibitor | No | No | No | No | No | No | No | No | No | No | No | No | No |
| P-glycoprotein II inhibitor | No | No | No | No | No | No | No | No | No | No | No | No | No |
| Distribution | |||||||||||||
| VDss (human) (log L/kg) | −0.032 | 0.064 | 0.144 | 0.215 | −0.891 | −0.928 | −1.032 | −0.557 | −0.306 | −0.402 | −0.009 | −0.061 | 0.208 |
| Fraction unbound (human) (Fu) | 0.14 | 0.083 | 0.11 | 0.112 | 0.22 | 0.202 | 0.193 | 0.163 | 0.176 | 0.223 | 0.125 | 0.164 | 0.164 |
| BBB permeability (log BB) | −0.965 | −1.32 | −1.218 | −1.325 | −1.736 | −1.647 | −1.834 | −1.556 | −1.397 | −1.48 | −0.107 | −0.225 | −0.16 |
| CNS permeability (log PS) | −2.247 | −2.557 | −2.499 | −2.498 | −4.452 | −4.438 | −4.551 | −3.76 | −3.565 | −3.5 | −2.236 | −2.24 | −2.218 |
| Metabolism | |||||||||||||
| CYP2D6 substrate | No | No | No | No | No | No | No | No | No | No | No | No | No |
| CYP3A4 substrate | No | No | No | No | No | No | No | No | No | No | No | No | No |
| CYP1A2 inhibitor | Yes | Yes | Yes | Yes | No | No | No | Yes | Yes | No | Yes | Yes | Yes |
| CYP2C19 inhibitor | No | No | No | No | No | No | No | No | No | No | Yes | Yes | Yes |
| CYP2C9 inhibitor | Yes | Yes | No | No | No | No | No | No | No | No | No | Yes | Yes |
| CYP2D6 inhibitor | No | No | No | No | No | No | No | No | No | No | No | No | No |
| CYP3A4 inhibitor | No | No | No | No | No | No | No | No | No | No | No | No | No |
| Excretion | |||||||||||||
| Total Clearance (log mL/min/kg) | 0.723 | 0.658 | 0.676 | 0.717 | 0.77 | 0.785 | 0.815 | 0.841 | 0.848 | 0.831 | 0.841 | 0.82 | 0.79 |
| Renal OCT2 substrate | No | No | No | No | No | No | No | No | No | No | No | No | No |
| Toxicity | AOH | Reaction/Metabolites | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Aromatic Hydroxilation | O-Glucuronidation | O-Sulfation | Methylation | ||||||||||
| M1 | M2 | M3 | M4 | M5 | M6 | M7 | M8 | M9 | M10 | M11 | M12 | ||
| AMES toxicity | Yes | Yes | No | Yes | No | No | No | No | No | No | No | Yes | No |
| Max. tolerated dose (human) | 0.697 | 0.94 | 0.832 | 0.888 | 0.681 | 0.693 | 0.654 | 0.976 | 0.945 | 0.918 | 0.489 | 0.765 | 0.472 |
| Oral Rat Acute Toxicity (LD50) | 2.82 | 2.611 | 2.515 | 2.586 | 2.438 | 2.441 | 2.466 | 2.582 | 2.63 | 2.509 | 2.918 | 3.001 | 3.036 |
| Oral Rat Chronic Toxicity (LOAEL) | 1.51 | 2.747 | 2.675 | 2.625 | 4.409 | 4.101 | 3.945 | 1.917 | 2.169 | 2.158 | 1.285 | 0.98 | 0.944 |
| T. pyriformis toxicity | 0.335 | 0.301 | 0.305 | 0.297 | 0.285 | 0.285 | 0.285 | 0.285 | 0.285 | 0.285 | 0.339 | 0.349 | 0.348 |
| Minnow toxicity | 1.512 | 1.235 | 1.54 | 2.128 | 3.918 | 3.476 | 4.205 | 0.414 | 1.586 | 1.927 | 0.332 | −0.352 | 0.319 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Marin, D.E.; Taranu, I. Using In Silico Approach for Metabolomic and Toxicity Prediction of Alternariol. Toxins 2023, 15, 421. https://doi.org/10.3390/toxins15070421
Marin DE, Taranu I. Using In Silico Approach for Metabolomic and Toxicity Prediction of Alternariol. Toxins. 2023; 15(7):421. https://doi.org/10.3390/toxins15070421
Chicago/Turabian StyleMarin, Daniela Eliza, and Ionelia Taranu. 2023. "Using In Silico Approach for Metabolomic and Toxicity Prediction of Alternariol" Toxins 15, no. 7: 421. https://doi.org/10.3390/toxins15070421
APA StyleMarin, D. E., & Taranu, I. (2023). Using In Silico Approach for Metabolomic and Toxicity Prediction of Alternariol. Toxins, 15(7), 421. https://doi.org/10.3390/toxins15070421






