Exploring the Multifaceted Biologically Relevant Roles of circRNAs: From Regulation, Translation to Biomarkers
Abstract
:1. Introduction
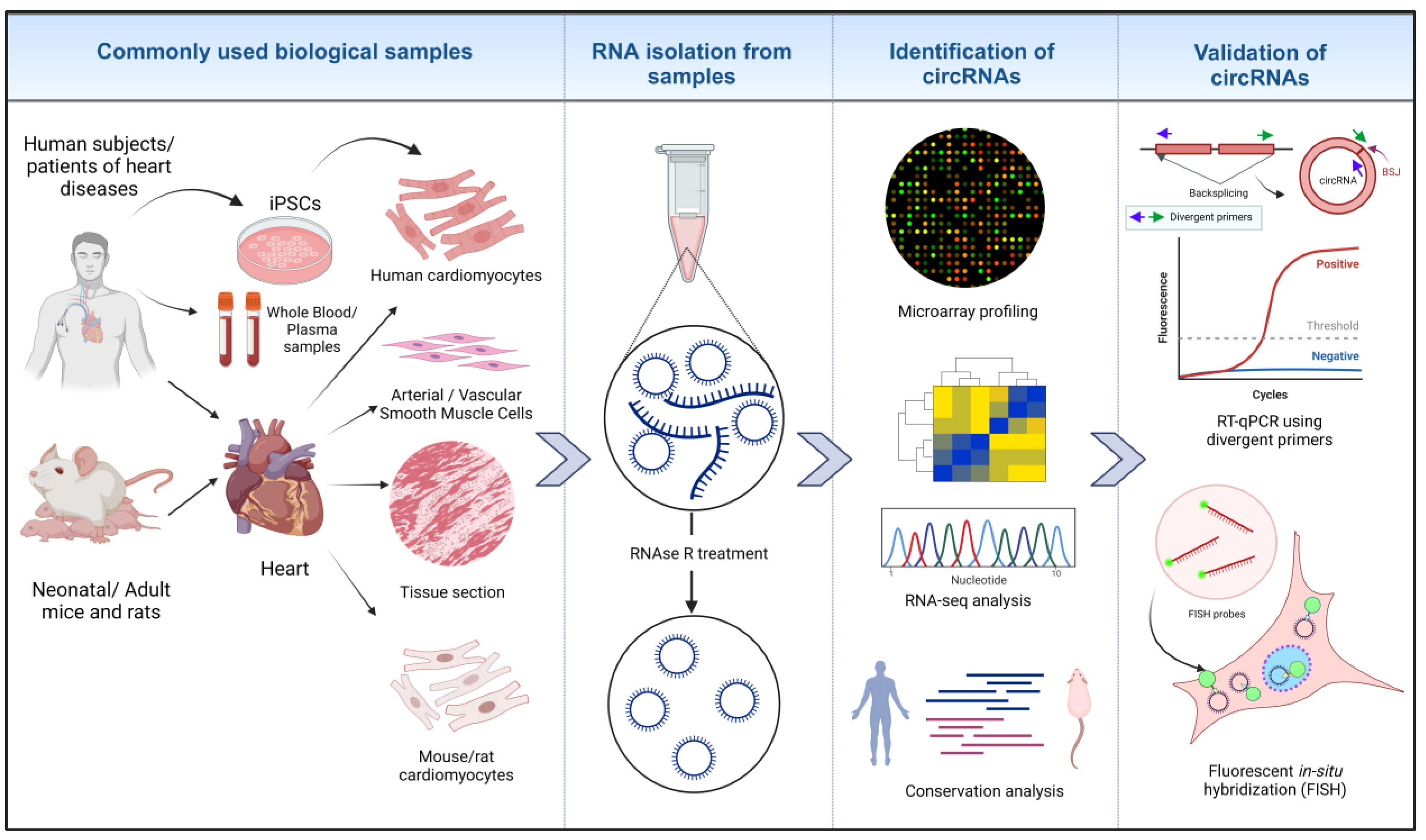
1.1. Methodologies for the Identification of circRNAs
1.2. circRNAs as miRNA Sponges
1.3. circRNA-Protein Interactions
1.4. Translatable circRNAs in Heart
1.5. circRNAs as Biomarkers for Cardiovascular Diseases
2. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Alexander, R.P.; Fang, G.; Rozowsky, J.; Snyder, M.; Gerstein, M.B. Annotating non-coding regions of the genome. Nat. Rev. Genet. 2010, 11, 559–571. [Google Scholar] [CrossRef]
- Mercer, T.R.; Dinger, M.E.; Mattick, J.S. Long non-coding RNAs: Insights into functions. Nat. Rev. Genet. 2009, 10, 155–159. [Google Scholar] [CrossRef]
- Esteller, M. Non-coding RNAs in human disease. Nat. Rev. Genet. 2011, 12, 861–874. [Google Scholar] [CrossRef]
- Zhang, P.; Wu, W.; Chen, Q.; Chen, M. Non-Coding RNAs and their Integrated Networks. J. Integr. Bioinform. 2019, 16, 20190027. [Google Scholar] [CrossRef]
- Wojciechowska, A.; Osiak, A.; Kozar-Kamińska, K. MicroRNA in cardiovascular biology and disease. Adv. Clin. Exp. Med. 2017, 26, 868–874. [Google Scholar] [CrossRef] [PubMed]
- Uchida, S.; Dimmeler, S. Long Noncoding RNAs in Cardiovascular Diseases. Circ. Res. 2015, 116, 737–750. [Google Scholar] [CrossRef] [PubMed]
- Gurha, P. Noncoding RNAs in cardiovascular diseases. Curr. Opin. Cardiol. 2019, 34, 241–245. [Google Scholar] [CrossRef] [PubMed]
- Jusic, A.; Thomas, P.B.; Wettinger, S.B.; Dogan, S.; Farrugia, R.; Gaetano, C.; Tuna, B.G.; Pinet, F.; Robinson, E.L.; Tual-Chalot, S.; et al. Noncoding RNAs in age-related cardiovascular diseases. Ageing Res. Rev. 2022, 77, 101610. [Google Scholar] [CrossRef] [PubMed]
- Fasolo, F.; Di Gregoli, K.; Maegdefessel, L.; Johnson, J.L. Non-coding RNAs in cardiovascular cell biology and atherosclerosis. Cardiovasc. Res. 2019, 115, 1732–1756. [Google Scholar] [CrossRef]
- Chen, L.L.; Yang, L. Regulation of circRNA biogenesis. RNA Biol. 2015, 12, 381–388. [Google Scholar] [CrossRef]
- Wilusz, J.E. A 360° view of circular RNAs: From biogenesis to functions. Wiley Interdiscip. Rev. RNA 2018, 9, e1478. [Google Scholar] [CrossRef] [PubMed]
- Kristensen, L.S.; Andersen, M.S.; Stagsted, L.V.W.; Ebbesen, K.K.; Hansen, T.B.; Kjems, J. The biogenesis, biology and characterization of circular RNAs. Nat. Rev. Genet. 2019, 20, 675–691. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Yang, L.; Chen, L.L. The Biogenesis, Functions, and Challenges of Circular RNAs. Mol. Cell 2018, 71, 428–442. [Google Scholar] [CrossRef] [PubMed]
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak, A.; Maier, L.; Mackowiak, S.D.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013, 495, 333–338. [Google Scholar] [CrossRef] [PubMed]
- Jeck, W.R.; Sorrentino, J.A.; Wang, K.; Slevin, M.K.; Burd, C.E.; Liu, J.; Marzluff, W.F.; Sharpless, N.E. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 2013, 19, 141–157. [Google Scholar] [CrossRef]
- Zhang, X.-O.; Dong, R.; Zhang, Y.; Zhang, J.-L.; Luo, Z.; Zhang, J.; Chen, L.-L.; Yang, L. Diverse alternative back-splicing and alternative splicing landscape of circular RNAs. Genome Res. 2016, 26, 1277–1287. [Google Scholar] [CrossRef] [PubMed]
- Kelly, S.; Greenman, C.; Cook, P.R.; Papantonis, A. Exon Skipping Is Correlated with Exon Circularization. J. Mol. Biol. 2015, 427, 2414–2417. [Google Scholar] [CrossRef]
- Zhang, X.-O.; Wang, H.-B.; Zhang, Y.; Lu, X.; Chen, L.-L.; Yang, L. Complementary Sequence-Mediated Exon Circularization. Cell 2014, 159, 134–147. [Google Scholar] [CrossRef]
- Ivanov, A.; Memczak, S.; Wyler, E.; Torti, F.; Porath, H.T.; Orejuela, M.R.; Piechotta, M.; Levanon, E.Y.; Landthaler, M.; Dieterich, C.; et al. Analysis of intron sequences reveals hallmarks of circular RNA biogenesis in animals. Cell Rep. 2015, 10, 170–177. [Google Scholar] [CrossRef]
- Stagsted, L.V.W.; O’Leary, E.T.; Ebbesen, K.K.; Hansen, T.B. The RNA-binding protein SFPQ preserves long-intron splicing and regulates circRNA biogenesis in mammals. eLife 2021, 10, e63088. [Google Scholar] [CrossRef]
- Zhang, Y.; Xue, W.; Li, X.; Zhang, J.; Chen, S.; Zhang, J.-L.; Yang, L.; Chen, L.-L. The Biogenesis of Nascent Circular RNAs. Cell Rep. 2016, 15, 611–624. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.U.; Agarwal, V.; Guo, H.; Bartel, D.P. Expanded identification and characterization of mammalian circular RNAs. Genome Biol. 2014, 15, 409. [Google Scholar] [CrossRef] [PubMed]
- Salzman, J.; Chen, R.E.; Olsen, M.N.; Wang, P.L.; Brown, P.O. Cell-Type Specific Features of Circular RNA Expression. PLoS Genet. 2013, 9, e1003777. [Google Scholar] [CrossRef]
- Salzman, J.; Gawad, C.; Wang, P.L.; Lacayo, N.; Brown, P.O. Circular RNAs Are the Predominant Transcript Isoform from Hundreds of Human Genes in Diverse Cell Types. PLoS ONE 2012, 7, e30733. [Google Scholar] [CrossRef]
- Meng, S.; Zhou, H.; Feng, Z.; Xu, Z.; Tang, Y.; Li, P.; Wu, M. CircRNA: Functions and properties of a novel potential biomarker for cancer. Mol. Cancer 2017, 16, 94. [Google Scholar] [CrossRef]
- Jeck, W.R.; Sharpless, N.E. Detecting and characterizing circular RNAs. Nat. Biotechnol. 2014, 32, 453–461. [Google Scholar] [CrossRef] [PubMed]
- Xiao, M.-S.; Wilusz, J.E. An improved method for circular RNA purification using RNase R that efficiently removes linear RNAs containing G-quadruplexes or structured 3′ ends. Nucleic Acids Res. 2019, 47, 8755–8769. [Google Scholar] [CrossRef]
- Suzuki, H. Characterization of RNase R-digested cellular RNA source that consists of lariat and circular RNAs from pre-mRNA splicing. Nucleic Acids Res. 2006, 34, e63. [Google Scholar] [CrossRef]
- Panda, A.C. Circular RNAs Act as miRNA Sponges. In Circular RNAs: Biogenesis and Functions; Xiao, J., Ed.; Springer: Singapore, 2018; pp. 67–79. [Google Scholar]
- Hansen, T.B.; Jensen, T.I.; Clausen, B.H.; Bramsen, J.B.; Finsen, B.; Damgaard, C.K.; Kjems, J. Natural RNA circles function as efficient microRNA sponges. Nature 2013, 495, 384–388. [Google Scholar] [CrossRef]
- Zhou, W.Y.; Cai, Z.R.; Liu, J.; Wang, D.S.; Ju, H.Q.; Xu, R.H. Circular RNA: Metabolism, functions and interactions with proteins. Mol. Cancer 2020, 19, 172. [Google Scholar] [CrossRef]
- Ulshöfer, C.J.; Pfafenrot, C.; Bindereif, A.; Schneider, T. Methods to study circRNA-protein interactions. Methods 2021, 196, 36–46. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, Z. Efficient backsplicing produces translatable circular mRNAs. RNA 2015, 21, 172–179. [Google Scholar] [CrossRef]
- Pamudurti, N.R.; Bartok, O.; Jens, M.; Ashwal-Fluss, R.; Stottmeister, C.; Ruhe, L.; Hanan, M.; Wyler, E.; Perez-Hernandez, D.; Ramberger, E.; et al. Translation of CircRNAs. Mol. Cell 2017, 66, 9–21.e27. [Google Scholar] [CrossRef]
- Liu, H.; Hao, W.; Yang, J.; Zhang, Y.; Wang, X.; Zhang, C. Emerging roles and potential clinical applications of translatable circular RNAs in cancer and other human diseases. Genes Dis. 2023, 10, 1994–2012. [Google Scholar] [CrossRef]
- Lin, H.-H.; Chang, C.-Y.; Huang, Y.-R.; Shen, C.-H.; Wu, Y.-C.; Chang, K.-L.; Lee, Y.-C.; Lin, Y.-C.; Ting, W.-C.; Chien, H.-J.; et al. Exon junction complex mediates the cap-independent translation of circular RNA. Mol. Cancer Res. 2023, 21, 1220–1233. [Google Scholar] [CrossRef]
- Kristensen, L.S.; Jakobsen, T.; Hager, H.; Kjems, J. The emerging roles of circRNAs in cancer and oncology. Nat. Rev. Clin. Oncol. 2022, 19, 188–206. [Google Scholar] [CrossRef] [PubMed]
- Ghafouri-Fard, S.; Hussen, B.M.; Taheri, M.; Ayatollahi, S.A. Emerging role of circular RNAs in breast cancer. Pathol.-Res. Pract. 2021, 223, 153496. [Google Scholar] [CrossRef] [PubMed]
- Long, F.; Li, L.; Xie, C.; Ma, M.; Wu, Z.; Lu, Z.; Liu, B.; Yang, M.; Zhang, F.; Ning, Z.; et al. Intergenic CircRNA Circ_0007379 Inhibits Colorectal Cancer Progression by Modulating miR-320a Biogenesis in a KSRP-Dependent Manner. Int. J. Biol. Sci. 2023, 19, 3781–3803. [Google Scholar] [CrossRef]
- Chen, S.; Chen, C.; Hu, Y.; Song, G.; Shen, X. The diverse roles of circular RNAs in pancreatic cancer. Pharmacol. Ther. 2021, 226, 107869. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Zhu, Q. Circular RNAs: Emerging roles and new insights in human cancers. Biomed. Pharmacother. 2023, 165, 115217. [Google Scholar] [CrossRef]
- Najafi, S.; Aghaei Zarch, S.M.; Majidpoor, J.; Pordel, S.; Aghamiri, S.; Fatih Rasul, M.; Asemani, Y.; Vakili, O.; Mohammadi, V.; Movahedpour, A.; et al. Recent insights into the roles of circular RNAs in human brain development and neurologic diseases. Int. J. Biol. Macromol. 2023, 225, 1038–1048. [Google Scholar] [CrossRef]
- Xu, D.; Ma, X.; Sun, C.; Han, J.; Zhou, C.; Chan, M.T.V.; Wu, W.K.K. Emerging roles of circular RNAs in neuropathic pain. Cell Prolif. 2021, 54, e13139. [Google Scholar] [CrossRef]
- Mehta, S.L.; Dempsey, R.J.; Vemuganti, R. Role of circular RNAs in brain development and CNS diseases. Prog. Neurobiol. 2020, 186, 101746. [Google Scholar] [CrossRef]
- Liu, X.; Xiao, H.; Peng, X.; Chai, Y.; Wang, S.; Wen, G. Identification and comprehensive analysis of circRNA–miRNA–mRNA regulatory networks in osteoarthritis. Front. Immunol. 2023, 13, 1050743. [Google Scholar] [CrossRef]
- Mohanapriya, R.; Akshaya, R.L.; Selvamurugan, N. A regulatory role of circRNA-miRNA-mRNA network in osteoblast differentiation. Biochimie 2022, 193, 137–147. [Google Scholar] [CrossRef]
- Yang, C.; Ni, B.; Li, C.; Sun, W.; Wang, Z.; Wang, H.; Hou, X.; Yan, S.; Wang, X.; Xu, D. circRNA_17725 Promotes Macrophage Polarization towards M2 by Targeting FAM46C to Alleviate Arthritis. Mediat. Inflamm. 2023, 2023, 6818524. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Liu, Y. circRNA_0016624 could sponge miR-98 to regulate BMP2 expression in postmenopausal osteoporosis. Biochem. Biophys. Res. Commun. 2019, 516, 546–550. [Google Scholar] [CrossRef] [PubMed]
- Zheng, S.; Zhang, X.; Odame, E.; Xu, X.; Chen, Y.; Ye, J.; Zhou, H.; Dai, D.; Kyei, B.; Zhan, S.; et al. CircRNA-Protein Interactions in Muscle Development and Diseases. Int. J. Mol. Sci. 2021, 22, 3262. [Google Scholar] [CrossRef] [PubMed]
- Das, A.; Das, A.; Das, D.; Abdelmohsen, K.; Panda, A.C. Circular RNAs in myogenesis. Biochim. Biophys. Acta (BBA)-Gene Regul. Mech. 2020, 1863, 194372. [Google Scholar] [CrossRef]
- Mahmoudi, E.; Cairns, M.J. CircRNA and Ageing. Subcell. Biochem. 2023, 102, 249–270. [Google Scholar] [CrossRef]
- Wei, L.; Liu, L.; Bai, M.; Ning, X.; Sun, S. CircRNAs: Versatile players and new targets in organ fibrosis. Cell Commun. Signal. 2023, 21, 90. [Google Scholar] [CrossRef] [PubMed]
- Hosaka, T.; Tsuji, H.; Kwak, S. Roles of Aging, Circular RNAs, and RNA Editing in the Pathogenesis of Amyotrophic Lateral Sclerosis: Potential Biomarkers and Therapeutic Targets. Cells 2023, 12, 1443. [Google Scholar] [CrossRef]
- Liu, J.; Deng, Z.; Yu, Z.; Zhou, W.; Yuan, Q. The circRNA circ-Nbea participates in regulating diabetic encephalopathy. Brain Res. 2022, 1774, 147702. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Jiang, Z.; Yu, M.; Yang, G. Roles of circular RNAs in regulating the self-renewal and differentiation of adult stem cells. Differentiation 2020, 113, 10–18. [Google Scholar] [CrossRef] [PubMed]
- Arthurs, A.L.; Jankovic-Karasoulos, T.; Smith, M.D.; Roberts, C.T. Circular RNAs in Pregnancy and the Placenta. Int. J. Mol. Sci. 2022, 23, 4551. [Google Scholar] [CrossRef] [PubMed]
- Ye, Y.; Zhang, L.; Hu, T.; Yin, J.; Xu, L.; Pang, Z.; Chen, W. CircRNA_103765 acts as a proinflammatory factor via sponging miR-30 family in Crohn’s disease. Sci. Rep. 2021, 11, 565. [Google Scholar] [CrossRef] [PubMed]
- Xia, X.; Tang, X.; Wang, S. Roles of CircRNAs in Autoimmune Diseases. Front. Immunol. 2019, 10, 639. [Google Scholar] [CrossRef] [PubMed]
- Altesha, M.A.; Ni, T.; Khan, A.; Liu, K.; Zheng, X. Circular RNA in cardiovascular disease. J. Cell. Physiol. 2019, 234, 5588–5600. [Google Scholar] [CrossRef] [PubMed]
- Ju, J.; Song, Y.-N.; Chen, X.-Z.; Wang, T.; Liu, C.-Y.; Wang, K. circRNA is a potential target for cardiovascular diseases treatment. Mol. Cell. Biochem. 2022, 477, 417–430. [Google Scholar] [CrossRef]
- Tan, W.L.; Lim, B.T.; Anene-Nzelu, C.G.; Ackers-Johnson, M.; Dashi, A.; See, K.; Tiang, Z.; Lee, D.P.; Chua, W.W.; Luu, T.D.; et al. A landscape of circular RNA expression in the human heart. Cardiovasc. Res. 2017, 113, 298–309. [Google Scholar] [CrossRef]
- Tong, X.; Zhao, X.; Dang, X.; Kou, Y.; Kou, J. circRNA, a novel diagnostic biomarker for coronary heart disease. Front. Cardiovasc. Med. 2023, 10, 1070616. [Google Scholar] [CrossRef] [PubMed]
- Werfel, S.; Nothjunge, S.; Schwarzmayr, T.; Strom, T.-M.; Meitinger, T.; Engelhardt, S. Characterization of circular RNAs in human, mouse and rat hearts. J. Mol. Cell. Cardiol. 2016, 98, 103–107. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Liu, B. Circular RNA in Diseased Heart. Cells 2020, 9, 1240. [Google Scholar] [CrossRef] [PubMed]
- American Heart Association. What Is Cardiovascular Disease? Available online: https://www.heart.org/en/health-topics/consumer-healthcare/what-is-cardiovascular-disease (accessed on 5 May 2023).
- World Health Organization. Cardiovascular Diseases (CVDs). Available online: https://www.who.int/news-room/fact-sheets/detail/cardiovascular-diseases-(cvds) (accessed on 22 August 2023).
- National Health Service (NHS). Cardiovascular Disease. Available online: https://www.nhs.uk/conditions/cardiovascular-disease/ (accessed on 22 August 2023).
- Lindahl, B. Acute coronary syndrome—The present and future role of biomarkers. Clin. Chem. Lab. Med. 2013, 51, 1699–1706. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Liang, D. Letter to the Editor: There may be a misunderstanding about the function of circular RNA as miRNA sponges. Hepatology 2023, 77, E102. [Google Scholar] [CrossRef] [PubMed]
- Li, H.M.; Ma, X.L.; Li, H.G. Intriguing circles: Conflicts and controversies in circular RNA research. Wiley Interdiscip. Rev. RNA 2019, 10, e1538. [Google Scholar] [CrossRef]
- Nielsen, A.F.; Bindereif, A.; Bozzoni, I.; Hanan, M.; Hansen, T.B.; Irimia, M.; Kadener, S.; Kristensen, L.S.; Legnini, I.; Morlando, M.; et al. Best practice standards for circular RNA research. Nat. Methods 2022, 19, 1208–1220. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Zhou, Y.; Jia, E.; Liu, Z.; Pan, M.; Bai, Y.; Zhao, X.; Ge, Q. Comparative analysis of circular RNA enrichment methods. RNA Biol. 2022, 19, 55–67. [Google Scholar] [CrossRef]
- Glažar, P.; Papavasileiou, P.; Rajewsky, N. circBase: A database for circular RNAs. RNA 2014, 20, 1666–1670. [Google Scholar] [CrossRef]
- Chen, X.; Han, P.; Zhou, T.; Guo, X.; Song, X.; Li, Y. circRNADb: A comprehensive database for human circular RNAs with protein-coding annotations. Sci. Rep. 2016, 6, 34985. [Google Scholar] [CrossRef]
- Liu, Y.C.; Li, J.R.; Sun, C.H.; Andrews, E.; Chao, R.F.; Lin, F.M.; Weng, S.L.; Hsu, S.D.; Huang, C.C.; Cheng, C.; et al. CircNet: A database of circular RNAs derived from transcriptome sequencing data. Nucleic Acids Res. 2016, 44, D209–D215. [Google Scholar] [CrossRef]
- Xia, S.; Feng, J.; Lei, L.; Hu, J.; Xia, L.; Wang, J.; Xiang, Y.; Liu, L.; Zhong, S.; Han, L.; et al. Comprehensive characterization of tissue-specific circular RNAs in the human and mouse genomes. Brief. Bioinform. 2017, 18, 984–992. [Google Scholar] [CrossRef] [PubMed]
- Panda, A.C.; Gorospe, M. Detection and Analysis of Circular RNAs by RT-PCR. Bio-Protocol 2018, 8, e2775. [Google Scholar] [CrossRef]
- Koppula, A.; Abdelgawad, A.; Guarnerio, J.; Batish, M.; Parashar, V. CircFISH: A Novel Method for the Simultaneous Imaging of Linear and Circular RNAs. Cancers 2022, 14, 428. [Google Scholar] [CrossRef] [PubMed]
- Seitz, H. Redefining MicroRNA Targets. Curr. Biol. 2009, 19, 870–873. [Google Scholar] [CrossRef] [PubMed]
- Salmena, L.; Poliseno, L.; Tay, Y.; Kats, L.; Pandolfi, P.P. A ceRNA Hypothesis: The Rosetta Stone of a Hidden RNA Language? Cell 2011, 146, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Zhong, Y.; Wang, X.; Shen, J.; An, W. Advances in Circular RNA and Its Applications. Int. J. Med. Sci. 2022, 19, 975–985. [Google Scholar] [CrossRef]
- Panda, A.C.; Abdelmohsen, K.; Gorospe, M. RT-qPCR Detection of Senescence-Associated Circular RNAs. In Oncogene-Induced Senescence: Methods and Protocols; Nikiforov, M.A., Ed.; Humana Press: New York, NY, USA, 2017; pp. 79–87. [Google Scholar]
- Zhang, X.; Wang, S.; Wang, H.; Cao, J.; Huang, X.; Chen, Z.; Xu, P.; Sun, G.; Xu, J.; Lv, J.; et al. Circular RNA circNRIP1 acts as a microRNA-149-5p sponge to promote gastric cancer progression via the AKT1/mTOR pathway. Mol. Cancer 2019, 18, 20. [Google Scholar] [CrossRef]
- Pang, Q.; Lin, X.; Sun, J.; Hu, J.; Dai, S.; Shen, Y.; Xu, M.; Xu, J. Comprehensive Analysis of Circular RNA Expression in ceRNA Networks and Identification of the Effects of hsa_circ_0006867 in Keloid Dermal Fibroblasts. Front. Mol. Biosci. 2022, 9, 800122. [Google Scholar] [CrossRef]
- Chen, J.; Chen, T.; Zhu, Y.; Li, Y.; Zhang, Y.; Wang, Y.; Li, X.; Xie, X.; Wang, J.; Huang, M.; et al. circPTN sponges miR-145-5p/miR-330-5p to promote proliferation and stemness in glioma. J. Exp. Clin. Cancer Res. 2019, 38, 398. [Google Scholar] [CrossRef] [PubMed]
- Yang, R.; Wang, R.C. Research Techniques Made Simple: Studying Circular RNA in Skin Diseases. J. Investig. Dermatol. 2021, 141, 2313–2319.e2311. [Google Scholar] [CrossRef]
- Caño-Carrillo, S.; Lozano-Velasco, E.; Castillo-Casas, J.M.; Sánchez-Fernández, C.; Franco, D. The Role of ncRNAs in Cardiac Infarction and Regeneration. J. Cardiovasc. Dev. Dis. 2023, 10, 123. [Google Scholar] [CrossRef]
- Mao, Y.Y.; Wang, J.Q.; Guo, X.X.; Bi, Y.; Wang, C.X. Circ-SATB2 upregulates STIM1 expression and regulates vascular smooth muscle cell proliferation and differentiation through miR-939. Biochem. Biophys. Res. Commun. 2018, 505, 119–125. [Google Scholar] [CrossRef]
- Zeng, Z.; Xia, L.; Fan, S.; Zheng, J.; Qin, J.; Fan, X.; Liu, Y.; Tao, J.; Li, K.; Ling, Z.; et al. Circular RNA CircMAP3K5 Acts as a MicroRNA-22-3p Sponge to Promote Resolution of Intimal Hyperplasia Via TET2-Mediated Smooth Muscle Cell Differentiation. Circulation 2021, 143, 354–371. [Google Scholar] [CrossRef] [PubMed]
- Luo, C.; Ling, G.X.; Lei, B.F.; Feng, X.; Xie, X.Y.; Fang, C.; Li, Y.G.; Cai, X.W.; Zheng, B.S. Circular RNA PVT1 silencing prevents ischemia-reperfusion injury in rat by targeting microRNA-125b and microRNA-200a. J. Mol. Cell. Cardiol. 2021, 159, 80–90. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.F.; Xu, L.L.; Xie, B.; Ding, H.G.; Fang, F.; Fang, Q. Hsa-circ-0068566 inhibited the development of myocardial ischemia reperfusion injury by regulating hsa-miR-6322/PARP2 signal pathway. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 6980–6993. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Zhao, P.; Sun, L.; Lu, Y.; Zhu, W.; Zhang, J.; Xiang, C.; Mao, Y.; Chen, Q.; Zhang, F. Overexpression of circRNA SNRK targets miR-103-3p to reduce apoptosis and promote cardiac repair through GSK3β/β-catenin pathway in rats with myocardial infarction. Cell Death Discov. 2021, 7, 84. [Google Scholar] [CrossRef]
- Huang, S.; Li, X.; Zheng, H.; Si, X.; Li, B.; Wei, G.; Li, C.; Chen, Y.; Liao, W.; Liao, Y.; et al. Loss of Super-Enhancer-Regulated circRNA Nfix Induces Cardiac Regeneration After Myocardial Infarction in Adult Mice. Circulation 2019, 139, 2857–2876. [Google Scholar] [CrossRef] [PubMed]
- Hall, I.F.; Climent, M.; Quintavalle, M.; Farina, F.M.; Schorn, T.; Zani, S.; Carullo, P.; Kunderfranco, P.; Civilini, E.; Condorelli, G.; et al. Circ_Lrp6, a Circular RNA Enriched in Vascular Smooth Muscle Cells, Acts as a Sponge Regulating miRNA-145 Function. Circ. Res. 2019, 124, 498–510. [Google Scholar] [CrossRef] [PubMed]
- Geng, H.-H.; Li, R.; Su, Y.-M.; Xiao, J.; Pan, M.; Cai, X.-X.; Ji, X.-P. The Circular RNA Cdr1as Promotes Myocardial Infarction by Mediating the Regulation of miR-7a on Its Target Genes Expression. PLoS ONE 2016, 11, e0151753. [Google Scholar] [CrossRef] [PubMed]
- Guo, Z.; Cao, Q.; Zhao, Z.; Song, C. Biogenesis, Features, Functions, and Disease Relationships of a Specific Circular RNA: CDR1as. Aging Dis. 2020, 11, 1009. [Google Scholar] [CrossRef] [PubMed]
- Hansen, T.B.; Wiklund, E.D.; Bramsen, J.B.; Villadsen, S.B.; Statham, A.L.; Clark, S.J.; Kjems, J. miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA. EMBO J. 2011, 30, 4414–4422. [Google Scholar] [CrossRef] [PubMed]
- Mester-Tonczar, J.; Winkler, J.; Einzinger, P.; Hasimbegovic, E.; Kastner, N.; Lukovic, D.; Zlabinger, K.; Spannbauer, A.; Traxler, D.; Batkai, S.; et al. Association between Circular RNA CDR1as and Post-Infarction Cardiac Function in Pig Ischemic Heart Failure: Influence of the Anti-Fibrotic Natural Compounds Bufalin and Lycorine. Biomolecules 2020, 10, 1180. [Google Scholar] [CrossRef]
- Chen, C.; Shen, H.; Huang, Q.; Li, Q. The Circular RNA CDR1as Regulates the Proliferation and Apoptosis of Human Cardiomyocytes Through the miR-135a/HMOX1 and miR-135b/HMOX1 Axes. Genet. Test. Mol. Biomark. 2020, 24, 537–548. [Google Scholar] [CrossRef]
- Ma, C.; Gu, R.; Wang, X.; He, S.; Bai, J.; Zhang, L.; Zhang, J.; Li, Q.; Qu, L.; Xin, W.; et al. circRNA CDR1as Promotes Pulmonary Artery Smooth Muscle Cell Calcification by Upregulating CAMK2D and CNN3 via Sponging miR-7-5p. Mol. Ther. Nucleic Acids 2020, 22, 530–541. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Q.; Bao, C.; Guo, W.; Li, S.; Chen, J.; Chen, B.; Luo, Y.; Lyu, D.; Li, Y.; Shi, G.; et al. Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat. Commun. 2016, 7, 11215. [Google Scholar] [CrossRef]
- Ni, H.; Li, W.; Zhuge, Y.; Xu, S.; Wang, Y.; Chen, Y.; Shen, G.; Wang, F. Inhibition of circHIPK3 prevents angiotensin II-induced cardiac fibrosis by sponging miR-29b-3p. Int. J. Cardiol. 2019, 292, 188–196. [Google Scholar] [CrossRef]
- Wang, W.; Zhang, S.; Xu, L.; Feng, Y.; Wu, X.; Zhang, M.; Yu, Z.; Zhou, X. Involvement of circHIPK3 in the pathogenesis of diabetic cardiomyopathy in mice. Diabetologia 2021, 64, 681–692. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhao, R.; Shen, C.; Liu, W.; Yuan, J.; Li, C.; Deng, W.; Wang, Z.; Zhang, W.; Ge, J.; et al. Exosomal CircHIPK3 Released from Hypoxia-Induced Cardiomyocytes Regulates Cardiac Angiogenesis after Myocardial Infarction. Oxidative Med. Cell. Longev. 2020, 2020, 8418407. [Google Scholar] [CrossRef]
- Wang, Y.; Zhao, R.; Liu, W.; Wang, Z.; Rong, J.; Long, X.; Liu, Z.; Ge, J.; Shi, B. Exosomal circHIPK3 Released from Hypoxia-Pretreated Cardiomyocytes Regulates Oxidative Damage in Cardiac Microvascular Endothelial Cells via the miR-29a/IGF-1 Pathway. Oxidative Med. Cell. Longev. 2019, 2019, 7954657. [Google Scholar] [CrossRef]
- Bai, M.; Pan, C.L.; Jiang, G.X.; Zhang, Y.M.; Zhang, Z. CircHIPK3 aggravates myocardial ischemia-reperfusion injury by binding to miRNA-124-3p. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 10107–10114. [Google Scholar]
- Zhang, W.-B.; Qi, Y.-F.; Xiao, Z.-X.; Chen, H.; Liu, S.-H.; Li, Z.-Z.; Zeng, Z.-F.; Wu, H.-F. CircHIPK3 Regulates Vascular Smooth Muscle Cell Calcification Via the miR-106a-5p/MFN2 Axis. J. Cardiovasc. Transl. Res. 2022, 15, 1315–1326. [Google Scholar] [CrossRef] [PubMed]
- Pan, J.; Xu, Z.; Guo, G.; Xu, C.; Song, Z.; Li, K.; Zhong, K.; Wang, D. Circ_nuclear factor I X (circNfix) attenuates pressure overload-induced cardiac hypertrophy via regulating miR-145-5p/ATF3 axis. Bioengineered 2021, 12, 5373–5385. [Google Scholar] [CrossRef] [PubMed]
- Tian, M.; Xue, J.; Dai, C.; Jiang, E.; Zhu, B.; Pang, H. CircSLC8A1 and circNFIX can be used as auxiliary diagnostic markers for sudden cardiac death caused by acute ischemic heart disease. Sci. Rep. 2021, 11, 4695. [Google Scholar] [CrossRef] [PubMed]
- Jing, X.; Wu, S.; Liu, Y.; Wang, H.; Huang, Q. Circular RNA Sirtuin1 represses pulmonary artery smooth muscle cell proliferation, migration and autophagy to ameliorate pulmonary hypertension via targeting microRNA-145-5p/protein kinase-B3 axis. Bioengineered 2022, 13, 8759–8771. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Liu, H.; Yu, H.; Zhou, Y.; Zhang, J.; Xin, W.; Li, Y.; He, S.; Ma, C.; Zheng, X.; et al. Circular RNA Calm4 Regulates Hypoxia-Induced Pulmonary Arterial Smooth Muscle Cells Pyroptosis via the Circ-Calm4/miR-124-3p/PDCD6 Axis. Arterioscler. Thromb. Vasc. Biol. 2021, 41, 1675–1693. [Google Scholar] [CrossRef]
- Zhang, J.; Li, Y.; Qi, J.; Yu, X.; Ren, H.; Zhao, X.; Xin, W.; He, S.; Zheng, X.; Ma, C.; et al. Circ-calm4 Serves as an miR-337-3p Sponge to Regulate Myo10 (Myosin 10) and Promote Pulmonary Artery Smooth Muscle Proliferation. Hypertension 2020, 75, 668–679. [Google Scholar] [CrossRef]
- Wang, Z.-Y.; Liu, X.-X.; Deng, Y.-F. Negative feedback of SNRK to circ-SNRK regulates cardiac function post-myocardial infarction. Cell Death Differ. 2022, 29, 709–721. [Google Scholar] [CrossRef] [PubMed]
- Lim, T.B.; Aliwarga, E.; Luu, T.D.A.; Li, Y.P.; Ng, S.L.; Annadoray, L.; Sian, S.; Ackers-Johnson, M.A.; Foo, R.S.-Y. Targeting the highly abundant circular RNA circSlc8a1 in cardiomyocytes attenuates pressure overload induced hypertrophy. Cardiovasc. Res. 2019, 115, 1998–2007. [Google Scholar] [CrossRef]
- Li, Y.; Ren, S.; Xia, J.; Wei, Y.; Xi, Y. EIF4A3-Induced circ-BNIP3 Aggravated Hypoxia-Induced Injury of H9c2 Cells by Targeting miR-27a-3p/BNIP3. Mol. Ther. Nucleic Acids 2020, 19, 533–545. [Google Scholar] [CrossRef]
- Xu, W.; Qian, L.; Yuan, X.; Lu, Y. Regulation of a Novel CircTRRAP/miR-761/MAP3K2 CeRNA Cascade in Inflammation, Apoptosis, and Oxidative Stress in Human AC16 Cardiomyocytes under Hypoxia Conditions. Int. Heart J. 2023, 64, 442–452. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Li, Z.; Wang, J.; Chen, H.; He, R.; Wu, H. CircTRRAP Knockdown Has Cardioprotective Function in Cardiomyocytes via the Signal Regulation of miR-370-3p/PAWR Axis. Cardiovasc. Ther. 2022, 2022, 7125602. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Guo, K. CircRbms1 knockdown alleviates hypoxia-induced cardiomyocyte injury via regulating the miR-742-3p/FOXO1 axis. Cell. Mol. Biol. Lett. 2022, 27, 31. [Google Scholar] [CrossRef] [PubMed]
- Jin, L.; Zhang, Y.; Jiang, Y.; Tan, M.; Liu, C. Circular RNA Rbms1 inhibited the development of myocardial ischemia reperfusion injury by regulating miR-92a/BCL2L11 signaling pathway. Bioengineered 2022, 13, 3082–3092. [Google Scholar] [CrossRef]
- Liang, Y.; Jie, H.; Liu, Q.; Li, C.; Xiao, R.; Xing, X.; Sun, J.; Yu, S.; Hu, Y.; Xu, G.-h. Knockout of circRNA single stranded interacting protein 1 (circRBMS1) played a protective role in myocardial ischemia-reperfusion injury though inhibition of miR-2355-3p/Mammalian Sterile20-like kinase 1 (MST1) axis. Bioengineered 2022, 13, 12726–12737. [Google Scholar] [CrossRef]
- Li, Y.; Wang, B. Circular RNA circCHFR downregulation protects against oxidized low-density lipoprotein-induced endothelial injury via regulation of microRNA-15b-5p/growth arrest and DNA damage inducible gamma. Bioengineered 2022, 13, 4481–4492. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Zhao, L.; Zhao, Y.; Fei, J.; Zhang, W. Circular RNA circ_0029589 regulates proliferation, migration, invasion, and apoptosis in ox-LDL-stimulated VSMCs by regulating miR-424-5p/IGF2 axis. Vasc. Pharmacol. 2020, 135, 106782. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, J.B.; Li, T.; Hu, X.M.; Ning, M.; Gao, W.Q.; Lang, Y.H.; Zheng, W.F.; Wei, J. Circ_CHFR expedites cell growth, migration and inflammation in ox-LDL-treated human vascular smooth muscle cells via the miR-214-3p/Wnt3/β-catenin pathway. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 3282–3292. [Google Scholar]
- Yang, L.; Yang, F.; Zhao, H.; Wang, M.; Zhang, Y. Circular RNA circCHFR Facilitates the Proliferation and Migration of Vascular Smooth Muscle via miR-370/FOXO1/Cyclin D1 Pathway. Mol. Ther.-Nucleic Acids 2019, 16, 434–441. [Google Scholar] [CrossRef]
- Wang, X.; Cheng, Z.; Xu, J.; Feng, M.; Zhang, H.; Zhang, L.; Qian, L. Circular RNA Arhgap12 modulates doxorubicin-induced cardiotoxicity by sponging miR-135a-5p. Life Sci. 2021, 265, 118788. [Google Scholar] [CrossRef]
- Miao, R.; Qi, C.; Fu, Y.; Wang, Y.; Lang, Y.; Liu, W.; Zhang, Y.; Zhang, Z.; Liu, A.; Chai, H.; et al. Silencing of circARHGAP12 inhibits the progression of atherosclerosis via miR-630/EZH2/TIMP2 signal axis. J. Cell. Physiol. 2022, 237, 1057–1069. [Google Scholar] [CrossRef]
- Lu, G.-F.; Geng, F.; Deng, L.-P.; Lin, D.-C.; Huang, Y.-Z.; Lai, S.-M.; Lin, Y.-C.; Gui, L.-X.; Sham, J.S.K.; Lin, M.-J. Reduced CircSMOC1 Level Promotes Metabolic Reprogramming via PTBP1 (Polypyrimidine Tract-Binding Protein) and miR-329-3p in Pulmonary Arterial Hypertension Rats. Hypertension 2022, 79, 2465–2479. [Google Scholar] [CrossRef]
- Kong, P.; Yu, Y.; Wang, L.; Dou, Y.-Q.; Zhang, X.-H.; Cui, Y.; Wang, H.-Y.; Yong, Y.-T.; Liu, Y.-B.; Hu, H.-J.; et al. circ-Sirt1 controls NF-κB activation via sequence-specific interaction and enhancement of SIRT1 expression by binding to miR-132/212 in vascular smooth muscle cells. Nucleic Acids Res. 2019, 47, 3580–3593. [Google Scholar] [CrossRef] [PubMed]
- Ding, P.; Ding, Y.; Tian, Y.; Lei, X. Circular RNA circ_0010283 regulates the viability and migration of oxidized low-density lipoprotein-induced vascular smooth muscle cells via an miR-370-3p/HMGB1 axis in atherosclerosis. Int. J. Mol. Med. 2020, 46, 1399–1408. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Liu, Y.; Zhang, W.; Huang, J.a.; Jiang, J.; Wang, R.; Zeng, D. circ-BPTF serves as a miR-486-5p sponge to regulate CEMIP and promotes hypoxic pulmonary arterial smooth muscle cell proliferation in COPD. Acta Biochim. Biophys. Sin. 2023, 55, 438–448. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Sun, L.; Hao, M.; Liu, X. Circ-SWT1 Ameliorates H2O2—Induced Apoptosis, Oxidative Stress and Endoplasmic Reticulum Stress in Cardiomyocytes via miR-192-5p/SOD2 Axis. Cardiovasc. Toxicol. 2022, 22, 378–389. [Google Scholar] [CrossRef]
- Yang, X.; Li, X.; Zhong, C.; Peng, J.; Pang, J.; Peng, T.; Wan, W.; Li, X. Circular RNA circPHKA2 Relieves OGD-Induced Human Brain Microvascular Endothelial Cell Injuries through Competitively Binding miR-574-5p to Modulate SOD2. Oxidative Med. Cell. Longev. 2021, 2021, 3823122. [Google Scholar] [CrossRef] [PubMed]
- Gan, J.; Yuan, J.; Liu, Y.; Lu, Z.; Xue, Y.; Shi, L.; Zeng, H. Circular RNA_101237 mediates anoxia/reoxygenation injury by targeting let-7a-5p/IGF2BP3 in cardiomyocytes. Int. J. Mol. Med. 2019, 45, 451–460. [Google Scholar] [CrossRef]
- Sun, Y.; Zhang, S.; Yue, M.; Li, Y.; Bi, J.; Liu, H. Angiotensin II inhibits apoptosis of mouse aortic smooth muscle cells through regulating the circNRG-1/miR-193b-5p/NRG-1 axis. Cell Death Dis. 2019, 10, 362. [Google Scholar] [CrossRef]
- Jiang, J.; Gao, G.; Pan, Q.; Liu, J.; Tian, Y.; Zhang, X. Circular RNA circHIPK3 is downregulated in diabetic cardiomyopathy and overexpression of circHIPK3 suppresses PTEN to protect cardiomyocytes from high glucose-induced cell apoptosis. Bioengineered 2022, 13, 6272–6279. [Google Scholar] [CrossRef]
- Das, A.; Sinha, T.; Shyamal, S.; Panda, A.C. Emerging role of circular rna–protein interactions. Non-Coding RNA 2021, 7, 48. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Ge, Y.; Wang, X.; Yin, W.; Zhu, X.; Wang, J.; Qiao, S. Circ-USP9X interacts with EIF4A3 to promote endothelial cell pyroptosis by regulating GSDMD stability in atherosclerosis. Clin. Exp. Hypertens. 2023, 45, 2186319. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Wang, J.; Feng, J.; Zhou, G.; Jiang, Y.; Luo, C.; Cheng, Z.; Li, J. Circ-JA760602 promotes the apoptosis of hypoxia-induced cardiomyocytes by transcriptionally suppressing BCL2. Int. J. Dev. Biol. 2023, 67, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Zhu, C.; Wang, B.; Zheng, H.; McAlister, V.; Lacefield, J.C.; Quan, D.; Mele, T.; Greasley, A.; Liu, K.; et al. Circular RNA Foxo3 in cardiac ischemia-reperfusion injury in heart transplantation: A new regulator and target. Am. J. Transplant. 2021, 21, 2992–3004. [Google Scholar] [CrossRef] [PubMed]
- Shao, Y.; Li, M.; Yu, Q.; Gong, M.; Wang, Y.; Yang, X.; Liu, L.; Liu, D.; Tan, Z.; Zhang, Y.; et al. CircRNA CDR1as promotes cardiomyocyte apoptosis through activating hippo signaling pathway in diabetic cardiomyopathy. Eur. J. Pharmacol. 2022, 922, 174915. [Google Scholar] [CrossRef] [PubMed]
- Wu, N.; Xu, J.; Du, W.W.; Li, X.; Awan, F.M.; Li, F.; Misir, S.; Eshaghi, E.; Lyu, J.; Zhou, L.; et al. YAP Circular RNA, circYap, Attenuates Cardiac Fibrosis via Binding with Tropomyosin-4 and Gamma-Actin Decreasing Actin Polymerization. Mol. Ther. 2021, 29, 1138–1150. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Huang, S.; Wei, G.; Sun, Y.; Li, C.; Si, X.; Chen, Y.; Tang, Z.; Li, X.; Chen, Y.; et al. CircRNA Samd4 induces cardiac repair after myocardial infarction by blocking mitochondria-derived ROS output. Mol. Ther. 2022, 30, 3477–3498. [Google Scholar] [CrossRef]
- Pang, P.; Si, W.; Wu, H.; Wang, C.; Liu, K.; Jia, Y.; Zhang, Z.; Zhang, F.; Kong, X.; Yang, Y.; et al. The circular RNA circHelz enhances cardiac fibrosis by facilitating the nuclear translocation of YAP1. Transl. Res. 2023, 257, 30–42. [Google Scholar] [CrossRef]
- Ding, F.; Lu, L.; Wu, C.; Pan, X.; Liu, B.; Zhang, Y.; Wang, Y.; Wu, W.; Yan, B.; Zhang, Y.; et al. circHIPK3 prevents cardiac senescence by acting as a scaffold to recruit ubiquitin ligase to degrade HuR. Theranostics 2022, 12, 7550–7566. [Google Scholar] [CrossRef]
- Du, W.W.; Yang, W.; Chen, Y.; Wu, Z.-K.; Foster, F.S.; Yang, Z.; Li, X.; Yang, B.B. Foxo3 circular RNA promotes cardiac senescence by modulating multiple factors associated with stress and senescence responses. Eur. Heart J. 2017, 38, 1402–1412. [Google Scholar] [CrossRef]
- Du, W.W.; Yang, W.; Liu, E.; Yang, Z.; Dhaliwal, P.; Yang, B.B. Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res. 2016, 44, 2846–2858. [Google Scholar] [CrossRef] [PubMed]
- Sun, G.; Shen, J.-F.; Wei, X.-F.; Qi, G.-X. Circular RNA Foxo3 Relieves Myocardial Ischemia/Reperfusion Injury by Suppressing Autophagy via Inhibiting HMGB1 by Repressing KAT7 in Myocardial Infarction. J. Inflamm. Res. 2021, 14, 6397–6407. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.; Wang, X.; Zhang, L.; Zhu, X.; Bai, J.; He, S.; Mei, J.; Jiang, J.; Guan, X.; Zheng, X.; et al. Super Enhancer-Associated Circular RNA-CircKrt4 Regulates Hypoxic Pulmonary Artery Endothelial Cell Dysfunction in Mice. Arterioscler. Thromb. Vasc. Biol. 2023, 43, 1179–1198. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Yu, P.; Wang, J.; Xu, G.; Wang, T.; Feng, J.; Bei, Y.; Xu, J.; Wang, H.; Das, S.; et al. Downregulation of circ-ZNF609 Promotes Heart Repair by Modulating RNA N6-Methyladenosine-Modified Yap Expression. Research 2022, 2022, 9825916. [Google Scholar] [CrossRef]
- Zhou, L.-Y.; Zhai, M.; Huang, Y.; Xu, S.; An, T.; Wang, Y.-H.; Zhang, R.-C.; Liu, C.-Y.; Dong, Y.-H.; Wang, M.; et al. The circular RNA ACR attenuates myocardial ischemia/reperfusion injury by suppressing autophagy via modulation of the Pink1/FAM65B pathway. Cell Death Differ. 2019, 26, 1299–1315. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Wang, J.; Feng, J.; Zhou, J.; Hou, L.; Gao, Y.; Cheng, Z. Circ-TLR4 promotes cardiac hypertrophy through recruiting FUS to stabilize TLR4 mRNA. J. Interv. Card. Electrophysiol. 2022, 65, 153–163. [Google Scholar] [CrossRef] [PubMed]
- Garikipati, V.N.S.; Verma, S.K.; Cheng, Z.; Liang, D.; Truongcao, M.M.; Cimini, M.; Yue, Y.; Huang, G.; Wang, C.; Benedict, C.; et al. Circular RNA CircFndc3b modulates cardiac repair after myocardial infarction via FUS/VEGF-A axis. Nat. Commun. 2019, 10, 4317. [Google Scholar] [CrossRef]
- Cao, S.; Li, C.; Li, L.; Zhou, G.; Jiang, Y.; Feng, J. Circular RNA hsa_circ_0000848 Regulates Cardiomyocyte Proliferation and Apoptosis Under Hypoxia via Recruiting ELAVL1 and Stabilizing SMAD7 mRNA. Anatol. J. Cardiol. 2022, 26, 189–197. [Google Scholar] [CrossRef]
- Guo, M.; Yan, R.; Ji, Q.; Yao, H.; Sun, M.; Duan, L.; Xue, Z.; Jia, Y. IFN regulatory Factor-1 induced macrophage pyroptosis by modulating m6A modification of circ_0029589 in patients with acute coronary syndrome. Int. Immunopharmacol. 2020, 86, 106800. [Google Scholar] [CrossRef]
- Gao, X.-Q.; Liu, C.-Y.; Zhang, Y.-H.; Wang, Y.-H.; Zhou, L.-Y.; Li, X.-M.; Wang, K.; Chen, X.-Z.; Wang, T.; Ju, J.; et al. The circRNA CNEACR regulates necroptosis of cardiomyocytes through Foxa2 suppression. Cell Death Differ. 2022, 29, 527–539. [Google Scholar] [CrossRef]
- Gong, X.; Tian, M.; Cao, N.; Yang, P.; Xu, Z.; Zheng, S.; Liao, Q.; Chen, C.; Zeng, C.; Jose, P.A.; et al. Circular RNA circEsyt2 regulates vascular smooth muscle cell remodeling via splicing regulation. J. Clin. Investig. 2021, 131, e147031. [Google Scholar] [CrossRef] [PubMed]
- Huang, M.; Dong, Y.; Sun, G.; Yu, Y. Circ-Sirt1 inhibits vascular smooth muscle cells proliferation via the c-Myc/cyclin B1 axis. Cell Biol. Int. 2022, 46, 628–636. [Google Scholar] [CrossRef] [PubMed]
- Ju, J.; Li, X.-M.; Zhao, X.-M.; Li, F.-H.; Wang, S.-C.; Wang, K.; Li, R.-F.; Zhou, L.-Y.; Liang, L.; Wang, Y.; et al. Circular RNA FEACR inhibits ferroptosis and alleviates myocardial ischemia/reperfusion injury by interacting with NAMPT. J. Biomed. Sci. 2023, 30, 45. [Google Scholar] [CrossRef]
- Heumüller, A.W.; Jones, A.N.; Mourão, A.; Klangwart, M.; Shi, C.; Wittig, I.; Fischer, A.; Muhly-Reinholz, M.; Buchmann, G.K.; Dieterich, C.; et al. Locus-Conserved Circular RNA cZNF292 Controls Endothelial Cell Flow Responses. Circ. Res. 2022, 130, 67–79. [Google Scholar] [CrossRef]
- Liu, Y.; Gao, J.; Xu, M.; Zhou, Q.; Zhang, Z.; Ye, J.; Li, R. Circular RNA circ-RCCD promotes cardiomyocyte differentiation in mouse embryo development via recruiting YY1 to the promoter of MyD88. J. Cell. Mol. Med. 2022, 26, 3616–3627. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.-Y.; Kuo, H.-C. The emerging roles and functions of circular RNAs and their generation. J. Biomed. Sci. 2019, 26, 29. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Li, F.; He, A.T.; Yang, B.B. Circular RNAs: Expression, localization, and therapeutic potentials. Mol. Ther. 2021, 29, 1683–1702. [Google Scholar] [CrossRef]
- Chen, C.-K.; Cheng, R.; Demeter, J.; Chen, J.; Weingarten-Gabbay, S.; Jiang, L.; Snyder, M.P.; Weissman, J.S.; Segal, E.; Jackson, P.K.; et al. Structured elements drive extensive circular RNA translation. Mol. Cell 2021, 81, 4300–4318.e4313. [Google Scholar] [CrossRef]
- Du, W.W.; Xu, J.; Yang, W.; Wu, N.; Li, F.; Zhou, L.; Wang, S.; Li, X.; He, A.T.; Du, K.Y.; et al. A Neuroligin Isoform Translated by circNlgn Contributes to Cardiac Remodeling. Circ. Res. 2021, 129, 568–582. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Chen, L.-W.; Huang, Z.-Q.; Guo, J.-S.; Li, H.; Shan, Y.; Chen, Z.-R.; Yan, Y.-M.; Zhu, J.-N.; Guo, H.-M.; et al. Suppression of the Inhibitory Effect of circ_0036176-Translated Myo9a-208 on Cardiac Fibroblast Proliferation by miR-218-5p. J. Cardiovasc. Transl. Res. 2022, 15, 548–559. [Google Scholar] [CrossRef]
- Koch, A.; Aguilera, L.; Morisaki, T.; Munsky, B.; Stasevich, T.J. Quantifying the dynamics of IRES and cap translation with single-molecule resolution in live cells. Nat. Struct. Mol. Biol. 2020, 27, 1095–1104. [Google Scholar] [CrossRef] [PubMed]
- Macejak, D.G.; Sarnow, P. Internal initiation of translation mediated by the 5′ leader of a cellular mRNA. Nature 1991, 353, 90–94. [Google Scholar] [CrossRef] [PubMed]
- van Heesch, S.; Witte, F.; Schneider-Lunitz, V.; Schulz, J.F.; Adami, E.; Faber, A.B.; Kirchner, M.; Maatz, H.; Blachut, S.; Sandmann, C.-L.; et al. The Translational Landscape of the Human Heart. Cell 2019, 178, 242–260.e229. [Google Scholar] [CrossRef] [PubMed]
- Ingolia, N.T.; Ghaemmaghami, S.; Newman, J.R.S.; Weissman, J.S. Genome-Wide Analysis In Vivo of Translation with Nucleotide Resolution Using Ribosome Profiling. Science 2009, 324, 218–223. [Google Scholar] [CrossRef] [PubMed]
- Siede, D.; Rapti, K.; Gorska, A.A.; Katus, H.A.; Altmüller, J.; Boeckel, J.N.; Meder, B.; Maack, C.; Völkers, M.; Müller, O.J.; et al. Identification of circular RNAs with host gene-independent expression in human model systems for cardiac differentiation and disease. J. Mol. Cell. Cardiol. 2017, 109, 48–56. [Google Scholar] [CrossRef] [PubMed]
- Jahn, C.; Bär, C.; Thum, T. CircSlc8a1, breaking a vicious circle in cardiac hypertrophy. Cardiovasc. Res. 2019, 115, 1946–1947. [Google Scholar] [CrossRef]
- Pan, Z.; Cai, J.; Lin, J.; Zhou, H.; Peng, J.; Liang, J.; Xia, L.; Yin, Q.; Zou, B.; Zheng, J.; et al. A novel protein encoded by circFNDC3B inhibits tumor progression and EMT through regulating Snail in colon cancer. Mol. Cancer 2020, 19, 71. [Google Scholar] [CrossRef]
- Xu, J.; Du, W.W.; Wu, N.; Li, F.; Li, X.; Xie, Y.; Wang, S.; Yang, B.B. The circular RNA circNlgnmediates doxorubicin-inducedcardiac remodeling and fibrosis. Mol. Ther.-Nucleic Acids 2022, 28, 175–189. [Google Scholar] [CrossRef]
- Li, S.; Mason, C.E. The Pivotal Regulatory Landscape of RNA Modifications. Annu. Rev. Genom. Hum. Genet. 2014, 15, 127–150. [Google Scholar] [CrossRef]
- Yang, Y.; Fan, X.; Mao, M.; Song, X.; Wu, P.; Zhang, Y.; Jin, Y.; Yang, Y.; Chen, L.-L.; Wang, Y.; et al. Extensive translation of circular RNAs driven by N6-methyladenosine. Cell Res. 2017, 27, 626–641. [Google Scholar] [CrossRef]
- Zhang, L.; Hou, C.; Chen, C.; Guo, Y.; Yuan, W.; Yin, D.; Liu, J.; Sun, Z. The role of N6-methyladenosine (m6A) modification in the regulation of circRNAs. Mol. Cancer 2020, 19, 105. [Google Scholar] [CrossRef]
- Zhou, C.; Molinie, B.; Daneshvar, K.; Pondick, J.V.; Wang, J.; Van Wittenberghe, N.; Xing, Y.; Giallourakis, C.C.; Mullen, A.C. Genome-Wide Maps of m6A circRNAs Identify Widespread and Cell-Type-Specific Methylation Patterns that Are Distinct from mRNAs. Cell Rep. 2017, 20, 2262–2276. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.G.; Chen, R.; Ahmad, S.; Verma, R.; Kasturi, S.P.; Amaya, L.; Broughton, J.P.; Kim, J.; Cadena, C.; Pulendran, B.; et al. N6-Methyladenosine Modification Controls Circular RNA Immunity. Mol. Cell 2019, 76, 96–109.e109. [Google Scholar] [CrossRef] [PubMed]
- Di Timoteo, G.; Dattilo, D.; Centrón-Broco, A.; Colantoni, A.; Guarnacci, M.; Rossi, F.; Incarnato, D.; Oliviero, S.; Fatica, A.; Morlando, M.; et al. Modulation of circRNA Metabolism by m6A Modification. Cell Rep. 2020, 31, 107641. [Google Scholar] [CrossRef] [PubMed]
- Kumari, R.; Ranjan, P.; Suleiman, Z.G.; Goswami, S.K.; Li, J.; Prasad, R.; Verma, S.K. mRNA modifications in cardiovascular biology and disease: With a focus on m6A modification. Cardiovasc. Res. 2022, 118, 1680–1692. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Wang, Y.; An, Y.; Wang, J.; Gao, Y. The Particular Expression Profiles of Circular RNA in Peripheral Blood of Myocardial Infarction Patients by RNA Sequencing. Front. Cardiovasc. Med. 2022, 9, 810257. [Google Scholar] [CrossRef]
- Legnini, I.; Di Timoteo, G.; Rossi, F.; Morlando, M.; Briganti, F.; Sthandier, O.; Fatica, A.; Santini, T.; Andronache, A.; Wade, M.; et al. Circ-ZNF609 Is a Circular RNA that Can Be Translated and Functions in Myogenesis. Mol. Cell 2017, 66, 22–37.e29. [Google Scholar] [CrossRef]
- Yu, P.; Wang, J.; Xu, G.-e.; Zhao, X.; Cui, X.; Feng, J.; Sun, J.; Wang, T.; Spanos, M.; Lehmann, H.I.; et al. RNA m6A-Regulated circ-ZNF609 Suppression Ameliorates Doxorubicin-Induced Cardiotoxicity by Upregulating FTO. JACC Basic Transl. Sci. 2023, 8, 677–698. [Google Scholar] [CrossRef]
- Ouyang, X.; He, Z.; Fang, H.; Zhang, H.; Yin, Q.; Hu, L.; Gao, F.; Yin, H.; Hao, T.; Hou, Y.; et al. A protein encoded by circular ZNF609 RNA induces acute kidney injury by activating the AKT/mTOR-autophagy pathway. Mol. Ther. 2023, 31, 1722–1738. [Google Scholar] [CrossRef]
- Abe, N.; Hiroshima, M.; Maruyama, H.; Nakashima, Y.; Nakano, Y.; Matsuda, A.; Sako, Y.; Ito, Y.; Abe, H. Rolling Circle Amplification in a Prokaryotic Translation System Using Small Circular RNA. Angew. Chem. Int. Ed. 2013, 52, 7004–7008. [Google Scholar] [CrossRef] [PubMed]
- Abe, N.; Matsumoto, K.; Nishihara, M.; Nakano, Y.; Shibata, A.; Maruyama, H.; Shuto, S.; Matsuda, A.; Yoshida, M.; Ito, Y.; et al. Rolling Circle Translation of Circular RNA in Living Human Cells. Sci. Rep. 2015, 5, 16435. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Ali, M.M.; Brook, M.A.; Li, Y. Rolling Circle Amplification: Applications in Nanotechnology and Biodetection with Functional Nucleic Acids. Angew. Chem. Int. Ed. 2008, 47, 6330–6337. [Google Scholar] [CrossRef] [PubMed]
- Xiong, F.; Mao, R.; Zhang, L.; Zhao, R.; Tan, K.; Liu, C.; Xu, J.; Du, G.; Zhang, T. CircNPHP4 in monocyte-derived small extracellular vesicles controls heterogeneous adhesion in coronary heart atherosclerotic disease. Cell Death Dis. 2021, 12, 948. [Google Scholar] [CrossRef] [PubMed]
- Memczak, S.; Papavasileiou, P.; Peters, O.; Rajewsky, N. Identification and Characterization of Circular RNAs As a New Class of Putative Biomarkers in Human Blood. PLoS ONE 2015, 10, e0141214. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.; Wu, J.; Han, P.; Zhao, Z.; Song, X. Circular RNA expression profiles and features in human tissues: A study using RNA-seq data. BMC Genom. 2017, 18, 680. [Google Scholar] [CrossRef] [PubMed]
- He, A.T.; Liu, J.; Li, F.; Yang, B.B. Targeting circular RNAs as a therapeutic approach: Current strategies and challenges. Signal Transduct. Target. Ther. 2021, 6, 185. [Google Scholar] [CrossRef] [PubMed]
- Xin, R.; Gao, Y.; Wang, R.; Kadash-Edmondson, K.E.; Liu, B.; Wang, Y.; Lin, L.; Xing, Y. isoCirc catalogs full-length circular RNA isoforms in human transcriptomes. Nat. Commun. 2021, 12, 266. [Google Scholar] [CrossRef]
- Wawrzyniak, O.; Zarębska, Ż.; Kuczyński, K.; Gotz-Więckowska, A.; Rolle, K. Protein-Related Circular RNAs in Human Pathologies. Cells 2020, 9, 1841. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Zhang, X.O.; Jiang, T.; Cai, L.; Huang, X.; Liu, Q.; Li, D.; Lu, A.; Liu, Y.; Xue, W.; et al. Comprehensive identification of alternative back-splicing in human tissue transcriptomes. Nucleic Acids Res. 2020, 48, 1779–1789. [Google Scholar] [CrossRef]
- Shen, Z.; Shao, Y.L.; Liu, W.; Zhang, Q.; Yuan, L. Prediction of Back-splicing sites for CircRNA formation based on convolutional neural networks. BMC Genom. 2022, 23, 581. [Google Scholar] [CrossRef]
- Li, M.; Ding, W.; Sun, T.; Tariq, M.A.; Xu, T.; Li, P.; Wang, J. Biogenesis of circular RNAs and their roles in cardiovascular development and pathology. FEBS J. 2018, 285, 220–232. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.-K. Circular RNAs as a promising biomarker for heart disease. Biomed. Pharmacother. 2022, 156, 113935. [Google Scholar] [CrossRef] [PubMed]
- Vausort, M.; Salgado-Somoza, A.; Zhang, L.; Leszek, P.; Scholz, M.; Teren, A.; Burkhardt, R.; Thiery, J.; Wagner, D.R.; Devaux, Y. Myocardial Infarction-Associated Circular RNA Predicting Left Ventricular Dysfunction. J. Am. Coll. Cardiol. 2016, 68, 1247–1248. [Google Scholar] [CrossRef]
- Zhu, X.; Wang, Y.; Mo, R.; Chong, H.; Cao, C.; Fan, F.; Zhou, Q.; Wang, D. Left Atrial Appendage Circular RNAs Are New Predictors of Atrial Fibrillation Recurrence After Surgical Ablation in Valvular Atrial Fibrillation Patients. Heart Surg. Forum 2021, 24, E968–E976. [Google Scholar] [CrossRef]
- Chen, Y.; Ouyang, T.; Yin, Y.; Fang, C.; Tang, C.-E.; Luo, J.; Luo, F. Analysis of infiltrated immune cells in left atriums from patients with atrial fibrillation and identification of circRNA biomarkers for postoperative atrial fibrillation. Front. Genet. 2022, 13, 1003366. [Google Scholar] [CrossRef] [PubMed]
- Stefanizzi, F.M.; Zhang, L.; Salgado-Somoza, A.; Dankiewicz, J.; Stammet, P.; Hassager, C.; Wise, M.P.; Friberg, H.; Cronberg, T.; Hundt, A.; et al. Circular RNAs to predict clinical outcome after cardiac arrest. Intensive Care Med. Exp. 2022, 10, 41. [Google Scholar] [CrossRef] [PubMed]
- Costa, M.C.; Calderon-Dominguez, M.; Mangas, A.; Campuzano, O.; Sarquella-Brugada, G.; Ramos, M.; Quezada-Feijoo, M.; Pinilla, J.M.G.; Robles-Mezcua, A.; del Aguila Pacheco-Cruz, G.; et al. Circulating circRNA as biomarkers for dilated cardiomyopathy etiology. J. Mol. Med. 2021, 99, 1711–1725. [Google Scholar] [CrossRef] [PubMed]
- Heil, B.; Wilson Tang, W.H. Biomarkers: Their potential in the diagnosis and treatment of heart failure. Clevel. Clin. J. Med. 2015, 82, S28–S35. [Google Scholar] [CrossRef]
- Sun, C.; Ni, M.; Song, B.; Cao, L. Circulating Circular RNAs: Novel Biomarkers for Heart Failure. Front. Pharmacol. 2020, 11, 560537. [Google Scholar] [CrossRef]
- Sygitowicz, G.; Sitkiewicz, D. Involvement of circRNAs in the Development of Heart Failure. Int. J. Mol. Sci. 2022, 23, 14129. [Google Scholar] [CrossRef]
- Tian, M.; Cao, Z.; Pang, H. Circular RNAs in Sudden Cardiac Death Related Diseases: Novel Biomarker for Clinical and Forensic Diagnosis. Molecules 2021, 26, 1155. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Wang, Z.; Sun, L.; Zheng, D.; Hu, B.; Li, N.; Shao, G. Hsa_circ_0000437 upregulates and promotes disease progression in rheumatic valvular heart disease. J. Clin. Lab. Anal. 2022, 36, e24197. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q.; Boeckel, J.N.; Yao, J.; Zhao, J.; Bai, Y.; Lv, Y.; Hu, M.; Meng, D.; Xie, Y.; Yu, P.; et al. Diagnosis of acute myocardial infarction using a combination of circulating circular RNA cZNF292 and clinical information based on machine learning. MedComm 2023, 4, e299. [Google Scholar] [CrossRef]
- Dinh, P.; Peng, J.; Tran, T.; Wu, D.; Tran, C.; Dinh, T.; Pan, S. Identification of hsa_circ_0001445 of a novel circRNA-miRNA-mRNA regulatory network as potential biomarker for coronary heart disease. Front. Cardiovasc. Med. 2023, 10, 1104223. [Google Scholar] [CrossRef] [PubMed]
- Liang, G.; Chen, S.; Xin, S.; Dong, L. Overexpression of hsa_circ_0001445 reverses oxLDL-induced inhibition of HUVEC proliferation via SRSF1. Mol. Med. Rep. 2021, 24, 507. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.; Xu, H.; Zhang, W.; Chen, Z.; Li, W.; Ke, W. Circular RNA circCCDC9 alleviates ischaemic stroke ischaemia/reperfusion injury via the Notch pathway. J. Cell. Mol. Med. 2020, 24, 14152–14159. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Liu, C.; Sun, Y.-N.; Zhu, C.-Y.; Xu, S.-S.; Shan, K.; Zhang, S.-J.; Yan, B.; Lu, Y. Targeting choroidal vascular dysfunction via inhibition of circRNA-FoxO1 for prevention and management of myopic pathology. Mol. Ther. 2021, 29, 2268–2280. [Google Scholar] [CrossRef]
- Qiu, Y.; Xie, X.; Lin, L. circFOXO3 protects cardiomyocytes against radiation-induced cardiotoxicity. Mol. Med. Rep. 2020, 23, 177. [Google Scholar] [CrossRef]
- Das, A.; Shyamal, S.; Sinha, T.; Mishra, S.S.; Panda, A.C. Identification of Potential circRNA-microRNA-mRNA Regulatory Network in Skeletal Muscle. Front. Mol. Biosci. 2021, 8, 762185. [Google Scholar] [CrossRef]
- Piwecka, M.; Glažar, P.; Hernandez-Miranda, L.R.; Memczak, S.; Wolf, S.A.; Rybak-Wolf, A.; Filipchyk, A.; Klironomos, F.; Cerda Jara, C.A.; Fenske, P.; et al. Loss of a mammalian circular RNA locus causes miRNA deregulation and affects brain function. Science 2017, 357, eaam8526. [Google Scholar] [CrossRef]
- Singh, S.; Sinha, T.; Panda, A.C. Regulation of microRNA by circular RNA. Wiley Interdiscip. Rev. RNA 2023, e1820. [Google Scholar] [CrossRef]
- Goodman, W.G.; Salusky, I.B. Non-invasive assessments of cardiovascular disease in patients with renal failure. Curr. Opin. Nephrol. Hypertens. 2001, 10, 365–369. [Google Scholar] [CrossRef] [PubMed]
- Huang, A.; Zheng, H.; Wu, Z.; Chen, M.; Huang, Y. Circular RNA-protein interactions: Functions, mechanisms, and identification. Theranostics 2020, 10, 3503–3517. [Google Scholar] [CrossRef]
- Liu, C.X.; Chen, L.L. Circular RNAs: Characterization, cellular roles, and applications. Cell 2022, 185, 2016–2034. [Google Scholar] [CrossRef]
- Denzler, R.; Agarwal, V.; Stefano, J.; Bartel, D.P.; Stoffel, M. Assessing the ceRNA Hypothesis with Quantitative Measurements of miRNA and Target Abundance. Mol. Cell 2014, 54, 766–776. [Google Scholar] [CrossRef]
- Lyngbakken, M.N.; Myhre, P.L.; Røsjø, H.; Omland, T. Novel biomarkers of cardiovascular disease: Applications in clinical practice. Crit. Rev. Clin. Lab. Sci. 2019, 56, 33–60. [Google Scholar] [CrossRef]
- Tousoulis, D.; Papageorgiou, N. Cardiovascular Disease: From Diagnosis to Therapy. Curr. Med. Chem. 2020, 27, 4438–4439. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Zhang, H.; Huang, S.; Yin, L.; Wang, F.; Luo, P.; Huang, H. Epigenetic regulation in cardiovascular disease: Mechanisms and advances in clinical trials. Signal Transduct. Target. Ther. 2022, 7, 200. [Google Scholar] [CrossRef]
- Lu, Q.; Li, Y.; Lou, J.; Li, P.; Gu, Y.; Wang, X. Circ-CHFR modulates the proliferation, migration, and invasion of ox-LDL-induced human aorta vascular smooth muscle cells through the miR-214-3p/PAPPA axis. Clin. Hemorheol. Microcirc. 2022, 80, 399–412. [Google Scholar] [CrossRef] [PubMed]
- Fuentes-Arderiu, X. What is a biomarker? It’s time for a renewed definition. Clin. Chem. Lab. Med. 2013, 51, 1689–1690. [Google Scholar] [CrossRef] [PubMed]
- Morrow, D.A.; de Lemos, J.A. Benchmarks for the Assessment of Novel Cardiovascular Biomarkers. Circulation 2007, 115, 949–952. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Liu, H. Factors influencing degradation kinetics of mRNAs and half-lives of microRNAs, circRNAs, lncRNAs in blood in vitro using quantitative PCR. Sci. Rep. 2022, 12, 7259. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Yang, T.; Xiao, J. Circular RNAs: Promising Biomarkers for Human Diseases. EBioMedicine 2018, 34, 267–274. [Google Scholar] [CrossRef] [PubMed]
- Arnaiz, E.; Sole, C.; Manterola, L.; Iparraguirre, L.; Otaegui, D.; Lawrie, C.H. CircRNAs and cancer: Biomarkers and master regulators. Semin. Cancer Biol. 2019, 58, 90–99. [Google Scholar] [CrossRef] [PubMed]
- Vea, A.; Llorente-Cortes, V.; de Gonzalo-Calvo, D. Circular RNAs in Blood. Adv. Exp. Med. Biol. 2018, 1087, 119–130. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Yang, Q.; He, A.T.; Yang, B.B. Circular RNAs in cancer: Limitations in functional studies and diagnostic potential. Semin. Cancer Biol. 2021, 75, 49–61. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Luo, J.; Yang, W.; Ye, W.C. CircRNAs in colorectal cancer: Potential biomarkers and therapeutic targets. Cell Death Dis. 2023, 14, 353. [Google Scholar] [CrossRef]
- Das, A.; Das, D.; Panda, A.C. Quantification of Circular RNAs Using Digital Droplet PCR. J. Vis. Exp. 2022, 187, e64464. [Google Scholar] [CrossRef]
- Li, T.; Shao, Y.; Fu, L.; Xie, Y.; Zhu, L.; Sun, W.; Yu, R.; Xiao, B.; Guo, J. Plasma circular RNA profiling of patients with gastric cancer and their droplet digital RT-PCR detection. J. Mol. Med. 2018, 96, 85–96. [Google Scholar] [CrossRef]
- Peng, J.; Li, F.; Xu, X.; Hu, S. Single-Cell Analysis of circRNA Using ddPCR. Methods Mol. Biol. 2023, 2689, 169–177. [Google Scholar] [CrossRef] [PubMed]
- Gezer, U.; Bronkhorst, A.J.; Holdenrieder, S. The Clinical Utility of Droplet Digital PCR for Profiling Circulating Tumor DNA in Breast Cancer Patients. Diagnostics 2022, 12, 3042. [Google Scholar] [CrossRef] [PubMed]





| circRNA | Original Gene | miRNA(s) Sponged/mRNA Target | Number of miRNA Binding Sites | Disease/Biological Context | Biological Sample | circRNA Expression | miRNA Expression | Refs. |
|---|---|---|---|---|---|---|---|---|
| CDR1as | Cerebellar-degeneration-related protein 1 (CDR1) exons | miR-7/PARP and SP1 | 73 | MI | Mouse tissue and cardiomyocytes | Upregulated | Upregulated | [95] |
| miR-7-5p/CAMK2D and CNN3 | 8 | Pulmonary hypertension (PH) | HPASMCs | Upregulated | Decreased | [100] | ||
| miR-135a and b/HMOX1 | 1 | CHF-associated proliferation and apoptosis | CHF patient plasma, HCMs, and AC16 cell lines | Upregulated | Decreased | [99] | ||
| CircHIPK3 | Homeodomain-interacting protein kinases (HIPK3) exon 2 | miR-106a-5p/MFN2 | 1 | Atherosclerosis | AS patient tissue, blood samples, and VSMC | Low expression | High expression | [107] |
| miR-29a/IGF-1 and VEGF | 1 | MI | Mouse cardiac microvascular endothelial cells (CMVECs) | Upregulated | Upregulated | [104,105] | ||
| miRNA-124-3p | 1 | Myocardial ischemia/reperfusion (IR) injury | HCM cells | Highly expressed | Negatively correlated with circRNA expression | [106] | ||
| circNFIX | Nuclear Factor IX exon 2 | miR-214/Gsk3β | 3 | MI and cardiac regeneration | Mouse cardiomyocytes | Overexpressed in human, rats and mice | Negatively correlated with circRNA expression | [93] |
| miR-145-5p/ATF3 | 1 | Cardiac hypertrophy | Patient plasma, neonatal mouse cardiomyocytes, mouse model | Downregulated | Negatively correlated with circRNA expression | [108] | ||
| circSlc8a1 | Sodium–calcium exchanger gene Slc8a1 | miR-133a | 17 | Cardiac hypertrophy | Neonatal mouse cardiomyocytes | Unchanged during myocardial stress response, but overexpression leads to heart failure | Unaffected by circRNA expression and lower expression is associated with disease development | [114] |
| circ_Lrp6 | Lipoprotein receptor 6 (Lrp6) | miR-145/ITGβ8, FASCIN, and KLF4 | At least 7 | VSMC growth, differentiation, and homeostasis | Mouse and human VSCMs | Enriched in VSMCs | Not correlated with circRNA expression | [94] |
| circ-SNRK | Sucrose nonfermenting 1-related kinase (SNRK) exon 1-2 | miR-33/SNRK | 7 | Heart failure (HF) and associated hypoxia | Primary neonatal rat cardiomyocytes and HF rat model | Decreased | Unchanged | [113] |
| Circ_BNIP3 (hsa_circ_0005972) | BCL2 interaction protein 3 (BNIP3) gene chr10:133784141-133787447 | miR-27a-3p/BNIP3, and 2 more unexplored miRNAs, miR-27b-3p and miR-128-3p | 1 | Hypoxia induced cardiac myocyte injury in ischemia and acute myocardial infarction (AMI) | Rat H9c2 cells | Upregulated | Downregulated while the other 2 were unchanged | [115] |
| circTRRAP (hsa_circ_0081241) | Transformation/transcription domain-associated protein (TRRAP) gene chr7:98495363-98506585 | miR-761/MAP3K2 | 1 | Hypoxia induced cardiomyocyte injury in AMI | AC16 human cardiomyocytes | Overexpressed | Downregulated | [116] |
| miR-370-3p/PAWR | 1 | Hypoxia induced cardiomyocyte injury in AMI | AC16 human cardiomyocytes | Upregulated | Negatively correlated | [117] | ||
| CircRbms1 (mmu_circ_0001022, hsa_circ_0002136) | RNA binding motif single-stranded interacting protein 1 (Rbms1) gene | miR-742-3p/FOXO1 | 1 | Hypoxia induced cardiomyocyte injury in MI | MI mouse model tissues and mouse cardiomyocytes (H9c2) | Upregulated | Negatively correlated with circRNA expression | [118] |
| miR-92a/BCL2L11 | 1 | AMI | MI mouse model and mouse cardiomyocytes (H9c2) | Overexpressed | Decreased | [119] | ||
| miR-2355-3p/MST1 | 1 | I/R injury | H/R induced HCMs | Increased | Decreased | [120] | ||
| circCHFR (circ_0029589) | Checkpoint with forkhead and ring finger domains (CHFR) gene chr12:133428203-133430159 | miR-15b-5p/GADD45G | 1 | Atherosclerosis | ox-LDL) induced HUVECs | Upregulated | Downregulated | [121] |
| miR-424-5p/IGF2 | 1 | Atherosclerosis | Ox-LDL-treated human VSMCs | Upregulated | Downregulated | [122] | ||
| miR-214-3p/Wnt3 | 1 | Atherosclerosis | Ox-LDL-treated human VSMCs | Upregulated | Negatively correlated | [123] | ||
| miR-370/FOXO1 | 1 | Atherosclerosis | Ox-LDL-treated human VSMCs | Upregulated | Negatively correlated | [124] | ||
| circArhgap12 | Rho GTPase activating protein 12 (ARHGAP12) gene exon 3 and exon 2 | miR-135a-5p/ADCY1, and 7 more miRNAs | 1 | Doxorubicin-induced cardiotoxicity | Mouse cardiomyocytes | Upregulated | Upregulated while 7 others were unchanged | [125] |
| miR-630/EZH2 | 1 | Atherosclerosis | Mouse aortic smooth muscle cells (MASMCs) | Upregulated | miR-630 was negatively correlated and miR-610 was unchanged, but not used for sponging validation | [126] | ||
| circSMOC1 | Modular calcium-binding protein 1 (SMOC1) gene exon 4 to exon 7 | miR-329-3p/PDHB | 1 | Pulmonary hypertension | Rat pulmonary artery smooth muscle cells | Downregulated | Unchanged | [127] |
| circSirtuin1 or circSirt1 | Sirtuin 1 (SIRT1) exon 2 to exon-7 | miR-132/212/Sirt1 | 3 | Atherosclerosis | Human and rat arterial tissues and VSMCs | Low expression | Negatively correlated with circRNA expression | [128] |
| miR-145-5p/Akt3 | 1 | Pulmonary hypertesion | Rat model for PH and Human PASMCs | Increased | Decreased | [110] | ||
| circ-calm4 | Calmodulin 4 gene single exon | miR-337-3p/Myosin 10 | 17 | Pulmonary hypertension | Mouse pulmonary artery smooth muscle cells (PASMCs) | Upregulated | Downregulated | [112] |
| miR-124-3p/PDCD6 | 2 | Pulmonary hypertension and vascular smooth muscle cell pyroptosis | PH mouse model and cultured pulmonary artery smooth muscle cells (PASMCs) | Upregulated | Downregulated | [111] | ||
| circ_0010283 | Ubiquitin protein ligase E3 component n-recognin 4 (UBR4) gene chr1:19449326-19480433 | miR-370-3p/HMGB1 | 1 | Atherosclerosis | ox-LDL-induced HVSMCs | Upregulated | Downregulated | [129] |
| circ-BPTF | Bromodomain PHD finger transcription factor (BPTF) gene exons 21 to 27 | miR-486-5p/CEMIP | 2 | Chronic obstructive pulmonary disease (COPD) | Human PASMCs | Upregulated | Downregulated | [130] |
| circ-SWT1 (hsa_circ_0015677) | SWT1 RNA endoribonuclease homolog gene chr1: 185,153,374–185,200,840 | miR-192-5p/SOD2 | 2 | H2O2 induced apoptosis in AMI | Human AC16 cardiomyocytes | Downregulated | Upregulated | [131] |
| circPHKA2 (hsa_circ_0090002) | phosphorylase kinase regulatory subunit alpha 2 (PHKA2) exons 2-29 | miR-574-5p/SOD2 | 2 | Acute ischemic stroke (AIS) | Patient blood, immortalized HBMECs | Downregulated | Upregulated | [132] |
| circRNA_101237 | Cyclin-dependent kinase 8 (CDK8) gene exon 10 to exon 12 | let-7a-5p/IGF2BP3 | 2 | Anoxia/reoxygenation (A/R) induced cardiomyocyte death | A/R-treated mouse cardiomyocytes | Upregulated | Unaffected by circRNA expression | [133] |
| circNRG-1 | Neuregulin-1 (NRG-1) gene | miR-193b-5p/NRG-1 | 3 | VSMC proliferation in vascular remodeling | Ang-II-treated mouse aortic smooth muscle cells (MASMCs) | Downregulated | Upregulated | [134] |
| circRNA-Protein Interaction | circRNA | Interacting Protein | Nature of Interaction | Consequence of Interaction | Cell/Tissue | Physiology/Disease | Refs. |
|---|---|---|---|---|---|---|---|
| Regulating mRNA stability/expression | Circ-USP9x | ElF4A3 | Direct binding in the cytoplasm | Increased stability of GSDMD mRNA, leading to AS-associated pyropstosis | Ox-LDL treated HUVECs | Atherosclerosis | [137] |
| CircZNF609 | YTHDF3 | Direct binding in the cytoplasm | Competitive binding with YAP mRNA to YTHDF3 and modulation of YAP expression to promote heart repair | AC16 human cardiomyocyte | Cardiac repair | [149] | |
| Autophagy-related circular RNA (ACR mmu_circRNA_006636) | Dnmt3B | Direct binding | Inhibition of DNA methylation of Pink1 promoter, thus blocking the binding of Dnmt3B to suppress cardiac autophagy and I/R injury | Mouse cardiomyocytes | I/R injury of cardiomyocytes in MI | [150] | |
| CircSMOC1 | PTBP1 | Direct binding in the nucleus | Competitive inhibition of pyruvate kinase M 1 (PKM1) pre-mRNA to promote the expression of PKM2 and regulate glycolysis | Rat pulmonary artery smooth muscle cells | Pulmonary vascular remodeling and arterial hypertension | [127] | |
| Circ-TLR4 | FUS | Direct binding in the cytoplasm | Promotion of TLR4 mRNA stability in the disease | Human cardiomyocytes | Cardiac hypertrophy | [151] | |
| CircFndc3b | FUS | Unclear interaction | Interaction with FUS is suggested to regulate FUS mRNA stability and regulate VEGF expression | Rat cardiomyocytes and mouse cardiac endothelial cells | Cardiac repair | [152] | |
| Circ_SMAD7 (hsa_circ_0000848) | ELAVL1 | Unclear interaction | Increased stability of the MI suppressor SMAD7 mRNA, reducing disease effects | HPC-CMs and H9c2 cardiomyocytes | Apoptosis of hypoxia induced cardiomyocytes | [153] | |
| CircFoxo3 | KAT7 | Unclear interaction | Suppressing the expression of MI associated factor, HMGB1 by inhibiting KAT7 and reducing the enrichment of H3 lysine acetylation and RNA pol II at the HMGB1 promoter | MI rat model and H9c2 rat cardiomyocytes | MI induced Myocardial injury | [147] | |
| CircHIPK3 | PTEN | Unclear interaction | Decrease in the levels of PTEN mRNA and protein-suppressing cardiomyocyte apoptosis | AC16 human cardiomyocytes | High-glucose-induced cell apoptosis in diabetic cardiomyopathy | [135] | |
| Expression regulation of circRNA by protein | Circ_0029589 (circCHFR) | IFN regulatory factor 1 (IRF1) | m6A modification | Promotion of m6A modifications to the circRNA by IRF1 via m6A methyltransferase METTL3 to downregulate the circRNA in the disease | Human-PBMC-derived macrophages from CAD patients | Acute coronary syndrome and Atherosclerosis | [154] |
| Circ_BNIP3 (hsa_circ_0005972) | ElF4A3 | Binding to the parent mRNA | ElF4A3 bound to the upstream site of BNIP3 mRNA to induce circBNIP3 expression | Rat H9c2 cells | Hypoxia induced cardiomyocyte injury | [115] | |
| circSNRK | NOVA alternative splicing regulator 1 | Direct binding to flanking introns of pre-circ-SNRK | Inducing circSNRK expression by promoting alternative splicing mediated by the competitive binding of a 55 kDa SNRK peptide | Primary neonatal rat cardiomyocytes and HF rat model | HF and associated hypoxia | [113] | |
| Nuclear translocation or Cytoplasmic sequestration of protein | circHelz | YAP1 | Direct binding in the cytoplasm | Promotion of nuclear localization of YAP1 to promote growth and proliferation | Mouse CFs | Cardiac Fibrosis | [143] |
| Circ-JA760602 | EGR1 and E2F1 | Direct binding in the cytoplasm | Inhibited nuclear translocation of EGR1 and E2F1 to suppress transcriptional activation of BCL2 | AC16 human cardiomyocytes | Hypoxia induced AMI and associated apoptosis | [138] | |
| Necroptosis-associated circRNA (CNEACR) | Histone deacetylases 7 (HDAC7) | Direct binding in the cytoplasm | Restriction of the nuclear import of HDAC7 to attenuate FoxA2 transcription and prevent myocardial damage | Mouse cardiomyocytes | Necroptotic death and I/R injury of cardiomyocytes in MI | [155] | |
| CircEsyt2 | PolyC-binding protein 1 (PCBP1) | Direct binding in the cytoplasm | Inhibition of nuclear translocation of PCBP1 to regulate p53 pre-mRNA splicing | Human aortic smooth muscle cells (HASMCs) and mouse VSMCs | Arterial remodeling | [156] | |
| Circ-Sirt1 | Cardiac myosin binding protein-C (c-Myc) | Direct binding in the cytoplasm | Promotion of cytoplasmic sequestration of VSCM-proliferation-associated c-Myc to prevent PDGF-BB-induced binding of c-Myc to the cyclin B1 promoter | Rat VSMCs | Restenosis and neointimal formation after injury | [157] | |
| Protein stability | Ferroptosis-associated circRNA (circFEACR) | NAMPT | Direct binding in the cytoplasm | Increased half-life and stability of NAMPT following cycloheximide treatment without affecting mRNA levels | Mouse cardiomyocytes | Ferroptosis inhibition in myocardial I/R injury | [158] |
| Protein degradation | CDR1as | MST1 | Unclear interaction | Increase in ubiquitination and subsequent degradation of MST1, thus activating Hippo-signaling pathway and promoting apoptosis of cardiomyocytes | Mouse cardiomyocytes | Diabetic cardiomyopathy | [140] |
| circNFIX | Y-box binding protein 1 (Ybx1) | Direct binding in the cytoplasm | Promotion of Ybx1 degradation through ubiquitinoylation by competing with E3 ubiquitin ligase Nedd41, influencing the interaction between Ybx1 and Nedd4l | Mouse cardiomyocytes | Cardiac regeneration | [93] | |
| Protein phosphorylation | CircFoxo3 | Foxo3 | Direct binding | Inhibition of Foxo3 protein phosphorylation at Ser253 via AKT and regulating I/R injury | HL-1 mouse atrial cardiomyocytes | Transplantation induced I/R injury | [139] |
| Protein complex formation/Protein scaffolds | circYap (hsa_circ_0002320) | Tropomyosin-4 and Gamma-Actin | Direct binding | Enhancing the binding between TPM4 and ACTG to form complexes to inhibit actin polymerization | AC16 human cardiomyocytes and pressure-overload mouse model | Cardiac fibrosis | [141] |
| CircHIPK3 | HuR and β-TrCP | Direct binding in the cytoplasm | Post-transcriptional regulation and localization of HuR, increasing its association with E3 ubiquitin ligase β-TrCP, promoting its degradation and cardiac senescence | CircHIPK3 KO mouse and mouse cardiomyocytes | Cardiac senescence and aging | [144] | |
| cZNF292/cZfp292 | Syndesmos (SDOS) | Direct binding | Enhancing the interaction between SDC4 and SDOS, thus influencing endothelial cell flow response | Mouse-model-derived aortic endothelium and retinal blood vessel and HUVECs | Endothelial cell morphology | [159] | |
| Mitochondrial recruitment of protein | circSamd4 | Vcp protein | Direct binding near the mitochondria | Promotion of mitochondrial localization of Vcp to repress Vdac1 and reactive oxygen species | Mouse fetal and neonatal cardiomyocytes | Cardiac repair and regeneration | [142] |
| Transcriptional recruitment of protein | circ-RCCD | YY1 transcription factor | Direct binding to YY1 and MyD88 in the cytoplasm | Promoting nuclear localization of YY1 to MyD88 promoter to inhibit its expression and facilitate cell differentiation | Mouse cardiac tissues and cardiomyocyte | Heart development and cardiomyocyte differentiation | [160] |
| Multifaceted functionality | CircKrt4 | Transcriptional activator protein Pur-alpha (Pura) | Direct binding in the nucleus and cytoplasm | Regulation of endothelial-to-mesenchymal transition by modulating the interaction between Pura and N-cadherin | Mouse pulmonary artery endothelial cell (PAEC) | Pulmonary hypertension | [148] |
| Glycerol kinase (Glpk) | Direct binding in the nucleus and cytoplasm | Regulating the mitochondrial translocation of Glpk | |||||
| RNA-binding-motif protein 25 (RBM25) | Binding to Krt4 pre-mRNA | Promoting alternative splicing and cyclization of Krt4 gene, thereby inducing circKrt4 expression |
| circRNA | Translated Product | Product Size | Translation Mechanism | Cardiovascular Disease/Physiology | Cell/Tissue/Location | Functional Importance | Refs. |
|---|---|---|---|---|---|---|---|
| circNlgn | Nlgn173 | 173aa | Unclear | Cardiac remodeling and cardiac fibrosis | AC16 human cardiomyocytes, Mouse primary cardiomyocytes, CFs and transgenic mouse model | CF growth and cardiomyocyte survival | [164,173] |
| circ_0036176 | Myo9a-208 | 208aa | IRES | Cardiac remodeling | Human Ac16 cardiomyocytes | Inhibition of CF proliferation through the suppression of cyclin/Rb pathway | [165] |
| circFNDC3B | circFNDC3B-218aa | 218aa | IRES | Unknown | Human CC cell lines | Colon cancer cell proliferation, migration, invasion and epithelial to mesenchymal transition | [152,172] |
| CircZNF609 | ZNF609-250aa | 250aa | m6A modification/IRES | Unknown | Mouse and human myoblasts, HeLa, HEK293T | Regulation of myoblast proliferation and ischemic AKI | [179,182,184] |
| circNPHP4 | - | - | Possible IRES | Coronary heart atherosclerotic disease | HCAECs | Unknown | [188] |
| circ_Lrp6 | - | - | ORF detected | Vascular smooth muscle cell activity. | Human VSMCs | Regulation of VSMC activity | [94] |
| CDR1as | - | - | Suggested | Unknown | Human heart | Unknown | [168] |
| circRNA | Disease/Physiology | Cell/Tissue/Location | Expression | Function | Refs. |
|---|---|---|---|---|---|
| circRNA MICRA | LV dysfunction in MI | Human peripheral blood | Low expression | Uncharacterized | [198] |
| circ 81906-RYR2 | AF | Left atrial appendage | Upregulated | Uncharacterized | [199] |
| hsa_circ_0006314 and hsa_circ_0055387 | POAF | Whole blood | Upregulated | Uncharacterized | [200] |
| circNFAT5 | Cardiac arrest | Whole blood | Upregulated | Uncharacterized | [201] |
| hsa_circ_0000437 | Rheumatic valvular heart disease (RVHD) | RVHD plasma samples. | Higher expression | Promotion of cell proliferation and migration, inhibition of apoptosis | [207] |
| CircSLC8A1 | Sudden cardiac death (SCD) caused by acute ischemic heart disease (IHD) | IHD rat and H9c2 cell models | Upregulated | Regulation of I/R injury and cardiomyocyte apoptosis | [109] |
| circNFIX | Elevated at an early stage of ischemia and later downregulated | Cardiomyocyte apoptosis | |||
| cZNF292 | Acute MI | Whole blood, HUVECs, and hESC-CM | Upregulated | Regulation of HUVEC AND hESC-CM apoptosis | [208] |
| circSMARCA5 (hsa_circ_0001445) | Coronary heart disease (CHD) | Peripheral blood leukocytes | Downregulated | Implicated interaction with several miRNAs | [209] |
| Atherosclerosis | Ox-LDL-treated HUVECs | Downregulated | SRSF1/β–catenin pathway influencing cell proliferation | [210] | |
| circCCDC9 | Ischemic stroke and I/R injury | Transient middle cerebral artery occlusion (tMCAO) mice model | Decreased | Notch signaling pathway | [211] |
| circNPHP4 | Coronary heart atherosclerotic disease | CAD-related monocytes | Upregulated | Heterogeneous cell adhesion of coronary artery endothelial cells | [188] |
| circFoxO1 | Myopic choroidal vascular dysfunction | RF/6A cells | Upregulated | Inhibition of endothelial effects of angiogenesis | [212] |
| circFoxo3 | Radiation-induced cardiotoxicity and heart damage | AC16 human cardiomyocytes | Upregulated | Protection against cardiotoxicity induced by radiation | [213] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hoque, P.; Romero, B.; Akins, R.E.; Batish, M. Exploring the Multifaceted Biologically Relevant Roles of circRNAs: From Regulation, Translation to Biomarkers. Cells 2023, 12, 2813. https://doi.org/10.3390/cells12242813
Hoque P, Romero B, Akins RE, Batish M. Exploring the Multifaceted Biologically Relevant Roles of circRNAs: From Regulation, Translation to Biomarkers. Cells. 2023; 12(24):2813. https://doi.org/10.3390/cells12242813
Chicago/Turabian StyleHoque, Parsa, Brigette Romero, Robert E Akins, and Mona Batish. 2023. "Exploring the Multifaceted Biologically Relevant Roles of circRNAs: From Regulation, Translation to Biomarkers" Cells 12, no. 24: 2813. https://doi.org/10.3390/cells12242813
APA StyleHoque, P., Romero, B., Akins, R. E., & Batish, M. (2023). Exploring the Multifaceted Biologically Relevant Roles of circRNAs: From Regulation, Translation to Biomarkers. Cells, 12(24), 2813. https://doi.org/10.3390/cells12242813









