Figure 1.
QDs’ external association with yeast. Green signals are expressed Vps10-GFP and red signals are CdSe/ZnS QDs. The size bar is equivalent to 5 μm. (A) QDs attach to the surface of yeast cells. (B) Percent of cells (% Frequency) that had QDs attached to mother site, daughter/budding site, or at the neck/both mother and daughter site.
Figure 1.
QDs’ external association with yeast. Green signals are expressed Vps10-GFP and red signals are CdSe/ZnS QDs. The size bar is equivalent to 5 μm. (A) QDs attach to the surface of yeast cells. (B) Percent of cells (% Frequency) that had QDs attached to mother site, daughter/budding site, or at the neck/both mother and daughter site.
Figure 2.
QDs’ intracellular trafficking. Cells expressing GFP-2PH or Abp1-GFP were cultured in SD-HIS for 0 h, 3 h, 6 h, or 24 h with or without QDs (4 μg/mL). The images show different channels (in order: GFP, RFP, and merge) for control on the left and QD-treated on the right. The size bar is equivalent to 5 μm. (A) Confocal microscopy images of the plasma membrane reference marker GFP-2PH (top) and confocal image of the early endocytosis reference marker (bottom). Images B and C show the graphs of Pearson’s correlation coefficient of QDs with different reference markers. (B) Graph for QDs and plasma membrane colocalization and (C) graph for QDs and the early endocytosis vesicle colocalization. * p < 0.05, ** p < 0.01.
Figure 2.
QDs’ intracellular trafficking. Cells expressing GFP-2PH or Abp1-GFP were cultured in SD-HIS for 0 h, 3 h, 6 h, or 24 h with or without QDs (4 μg/mL). The images show different channels (in order: GFP, RFP, and merge) for control on the left and QD-treated on the right. The size bar is equivalent to 5 μm. (A) Confocal microscopy images of the plasma membrane reference marker GFP-2PH (top) and confocal image of the early endocytosis reference marker (bottom). Images B and C show the graphs of Pearson’s correlation coefficient of QDs with different reference markers. (B) Graph for QDs and plasma membrane colocalization and (C) graph for QDs and the early endocytosis vesicle colocalization. * p < 0.05, ** p < 0.01.
Figure 3.
QDs’ intracellular trafficking. Cells expressing FAPPI-GFP or Vps10-GFP were cultured in SD-HIS for 0 h, 3 h, 6 h, and 24 h with or without QDs (4 μg/mL). The microscopic images were obtained from different channels (in order: GFP, RFP, and merge) for control on the left and QD-treated on the right. The size bar is equivalent to 5 μm. (A) Confocal microscopy images of the late Golgi/trans-Golgi network (top) and confocal image of the late endosome (bottom). Images (B) and (C) showed the graphs of Pearson’s correlation coefficient of QDs with different reference markers. (B) Graph for QDs and FAPPI-GFP colocalization and (C) graph for QDs and the Vps10-GFP colocalization. * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001.
Figure 3.
QDs’ intracellular trafficking. Cells expressing FAPPI-GFP or Vps10-GFP were cultured in SD-HIS for 0 h, 3 h, 6 h, and 24 h with or without QDs (4 μg/mL). The microscopic images were obtained from different channels (in order: GFP, RFP, and merge) for control on the left and QD-treated on the right. The size bar is equivalent to 5 μm. (A) Confocal microscopy images of the late Golgi/trans-Golgi network (top) and confocal image of the late endosome (bottom). Images (B) and (C) showed the graphs of Pearson’s correlation coefficient of QDs with different reference markers. (B) Graph for QDs and FAPPI-GFP colocalization and (C) graph for QDs and the Vps10-GFP colocalization. * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001.
Figure 4.
A pulse and chase experiment to determine the degree of colocalization between red quantum dots (RQDs) and green pinocytosis tracking dye Fm1-43. Fm1-43-labeled cells at 0 °C were incubated at room temperature for varying durations up to 60 min. The size bar is equivalent to 5 μm. (A) Confocal microscopy of the pinocytosis pathway within 60 min (left—untreated, right—treated) and (B) graph of Fm1-43 and QDs colocalization assessed by Pearson’s coefficient.
Figure 4.
A pulse and chase experiment to determine the degree of colocalization between red quantum dots (RQDs) and green pinocytosis tracking dye Fm1-43. Fm1-43-labeled cells at 0 °C were incubated at room temperature for varying durations up to 60 min. The size bar is equivalent to 5 μm. (A) Confocal microscopy of the pinocytosis pathway within 60 min (left—untreated, right—treated) and (B) graph of Fm1-43 and QDs colocalization assessed by Pearson’s coefficient.
Figure 5.
Growth assay to determine the growth rate of yeast in the presence of QDs in SD-HIS. (A) Growth assay for yeast cells at 6 h after QD treatment. (B) Growth assay for yeast cells at 24 h after QD treatment. (C) A 6 h kinetic growth curve of yeast in the presence of quantum dots in various concentrations (0 μL/mL, 4 μg/mL, 12 μg/mL, 25 μg/mL, and 50 μg/mL). (D) A 24 h kinetic growth curve of yeast in the presence of quantum dots in various concentrations (0 μL/mL, 4 μg/mL, 12 μg/mL, 25 μg/mL, and 50 μg/mL). *** p < 0.001, **** p < 0.0001.
Figure 5.
Growth assay to determine the growth rate of yeast in the presence of QDs in SD-HIS. (A) Growth assay for yeast cells at 6 h after QD treatment. (B) Growth assay for yeast cells at 24 h after QD treatment. (C) A 6 h kinetic growth curve of yeast in the presence of quantum dots in various concentrations (0 μL/mL, 4 μg/mL, 12 μg/mL, 25 μg/mL, and 50 μg/mL). (D) A 24 h kinetic growth curve of yeast in the presence of quantum dots in various concentrations (0 μL/mL, 4 μg/mL, 12 μg/mL, 25 μg/mL, and 50 μg/mL). *** p < 0.001, **** p < 0.0001.
Figure 6.
The distribution of GFP-Snc1 on the plasma membrane. For each, 30 cells were counted. The size bar is equivalent to 5 μm. (A) Images of cells with different polarization at 6 h post-QD treatment. (B) The percentage of cells showed the polarized distribution of GFP-Snc1 on the plasma membrane for 1 h after treatment. (C) The percentage of cells showed the polarized distribution of GFP-Snc1 on the plasma membrane 3 h after treatment. (D) The percentage of cells showed the polarized distribution of GFP-Snc1 on the plasma membrane at 6 h after treatment. (E) The percentage of cells showed the polarized distribution of GFP-Snc1 on the plasma membrane at 24 h after treatment. * p < 0.05.
Figure 6.
The distribution of GFP-Snc1 on the plasma membrane. For each, 30 cells were counted. The size bar is equivalent to 5 μm. (A) Images of cells with different polarization at 6 h post-QD treatment. (B) The percentage of cells showed the polarized distribution of GFP-Snc1 on the plasma membrane for 1 h after treatment. (C) The percentage of cells showed the polarized distribution of GFP-Snc1 on the plasma membrane 3 h after treatment. (D) The percentage of cells showed the polarized distribution of GFP-Snc1 on the plasma membrane at 6 h after treatment. (E) The percentage of cells showed the polarized distribution of GFP-Snc1 on the plasma membrane at 24 h after treatment. * p < 0.05.
Figure 7.
RNAseq analysis. (A) A volcano plot showing altered genes in RNA-Seq analysis with red dots indicating genes that have a fold change larger than 1.25 and less than −1.25 with p−value < 0.05. (B–E) Bar graphs represent classes of genes that were differentially expressed due to the treatment of QDs. Each bar is the number of differentially expressed genes in a protein class or associated with a biological process. (B) Downregulated genes sorted by protein class. (C) Upregulated genes sorted by protein class. (D) Downregulated genes are sorted by associated biological processes. (E) Upregulated genes sorted by associated biological processes.
Figure 7.
RNAseq analysis. (A) A volcano plot showing altered genes in RNA-Seq analysis with red dots indicating genes that have a fold change larger than 1.25 and less than −1.25 with p−value < 0.05. (B–E) Bar graphs represent classes of genes that were differentially expressed due to the treatment of QDs. Each bar is the number of differentially expressed genes in a protein class or associated with a biological process. (B) Downregulated genes sorted by protein class. (C) Upregulated genes sorted by protein class. (D) Downregulated genes are sorted by associated biological processes. (E) Upregulated genes sorted by associated biological processes.
Figure 8.
Model of important upregulated genes (blue) and downregulated genes (red) upon CdSe/ZnS QDs treatment, as shown by RNA-seq data. Upregulated genes include genes associated with the mitochondria structure and function (PET111, PET112, and SLS1), genes involved in the vesicular trafficking (DID2, COS10, and TRS85), a gene involved in cell wall integrity (ZRG8), transcription regulation (MGA2), and vacuole structure (VAC7). Important downregulated genes include genes involved in translation (RPL30, RPL29, RPL25, and RPS13), mitochondria and metabolism-related genes (ATP20, TIM12, and TIM10), transcription factor genes (GAT2, CIN5, STP4, and SKO1), cell wall integrity genes (DAN1 and DAN4), endocytosis regulation gene (APS2), and vacuole-associated genes (VMA9, VMA11, VMA21, and LCL2). Created with BioRender.com.
Figure 8.
Model of important upregulated genes (blue) and downregulated genes (red) upon CdSe/ZnS QDs treatment, as shown by RNA-seq data. Upregulated genes include genes associated with the mitochondria structure and function (PET111, PET112, and SLS1), genes involved in the vesicular trafficking (DID2, COS10, and TRS85), a gene involved in cell wall integrity (ZRG8), transcription regulation (MGA2), and vacuole structure (VAC7). Important downregulated genes include genes involved in translation (RPL30, RPL29, RPL25, and RPS13), mitochondria and metabolism-related genes (ATP20, TIM12, and TIM10), transcription factor genes (GAT2, CIN5, STP4, and SKO1), cell wall integrity genes (DAN1 and DAN4), endocytosis regulation gene (APS2), and vacuole-associated genes (VMA9, VMA11, VMA21, and LCL2). Created with BioRender.com.
Figure 9.
Inefficient endocytosis and prolonged path duration of the endocytosis vesicle resulted from the slowed detachment of Abp1-GFP. (A) Kymograph of endocytic vesicle formation showing inefficient endocytosis for groups treated with 25 μg/mL of CdSe/ZnS at 0 h, 3 h, 6 h, and 24 h post-treatment. (B) Prolonged lifetime of Abp1-GFP on the plasma membrane. (C) Lifetime of Abp1-GFP post-membrane detachment. ** p < 0.01, **** p < 0.0001.
Figure 9.
Inefficient endocytosis and prolonged path duration of the endocytosis vesicle resulted from the slowed detachment of Abp1-GFP. (A) Kymograph of endocytic vesicle formation showing inefficient endocytosis for groups treated with 25 μg/mL of CdSe/ZnS at 0 h, 3 h, 6 h, and 24 h post-treatment. (B) Prolonged lifetime of Abp1-GFP on the plasma membrane. (C) Lifetime of Abp1-GFP post-membrane detachment. ** p < 0.01, **** p < 0.0001.
Figure 10.
Actin filament integrity assay. (A) Cells were treated with an increasing concentration of QDs for 6 h and stained with Actistain 488. Cells were then visualized using the confocal microscope. The left column is a flat, 2D image taken at the mid-plane of cells when cells are in focus. The right columns are the projected pictures from 3D images that allow the visualization of all cables inside yeast cells. Each row signifies images from a treatment group, in the top-down order: control, 4 μg/mL, 12 μg/mL, 25 μg/mL, and 50 μg/mL. The size bar is equivalent to 5 μm. (B) The percentage of cells with abnormal actin cable was quantified and graphed. A total of 30 cells were used for each sample. * p < 0.05, ** p < 0.01.
Figure 10.
Actin filament integrity assay. (A) Cells were treated with an increasing concentration of QDs for 6 h and stained with Actistain 488. Cells were then visualized using the confocal microscope. The left column is a flat, 2D image taken at the mid-plane of cells when cells are in focus. The right columns are the projected pictures from 3D images that allow the visualization of all cables inside yeast cells. Each row signifies images from a treatment group, in the top-down order: control, 4 μg/mL, 12 μg/mL, 25 μg/mL, and 50 μg/mL. The size bar is equivalent to 5 μm. (B) The percentage of cells with abnormal actin cable was quantified and graphed. A total of 30 cells were used for each sample. * p < 0.05, ** p < 0.01.
Figure 11.
Actin cable integrity assay. (A) Abp140-GFP-expressing cells were treated with QDs and visualized over time using the confocal microscope. The size bar is equivalent to 5 μm. (B) Actin cables of 30 cells were assessed for their structure. *** p < 0.001.
Figure 11.
Actin cable integrity assay. (A) Abp140-GFP-expressing cells were treated with QDs and visualized over time using the confocal microscope. The size bar is equivalent to 5 μm. (B) Actin cables of 30 cells were assessed for their structure. *** p < 0.001.
Figure 12.
Graphs show the level of GFP-tagged actin-associated proteins. (A) F-actin stained with Actistain488 dye, (B) Pfy1-GFP, (C) GFP-Cof1, and (D) Crn1-GFP. ** p < 0.01, *** p < 0.001, **** p < 0.0001.
Figure 12.
Graphs show the level of GFP-tagged actin-associated proteins. (A) F-actin stained with Actistain488 dye, (B) Pfy1-GFP, (C) GFP-Cof1, and (D) Crn1-GFP. ** p < 0.01, *** p < 0.001, **** p < 0.0001.
Figure 13.
A recovery assay was performed where cells exposed to QDs were moved to a QD-free media for 3 h and the morphology of the actin cable was assessed. (A) Images of the actin cable when stained with Actistain 488 and visualized with the confocal microscope. The left column is a flat, 2D image taken at the mid-plane of cells when cells are in focus. The right column is the projected pictures from 3D images that allow the visualization of all cables inside yeast cells. Each row signifies images from a treatment group, in the top-down order: control, 4 μg/mL, 12 μg/mL, 25 μg/mL, and 50 μg/mL. The size bar is equivalent to 5 μm. (B) The percentage of cells with abnormal actin cable was quantified and graphed. A total of 30 cells were used for each sample. * p < 0.05.
Figure 13.
A recovery assay was performed where cells exposed to QDs were moved to a QD-free media for 3 h and the morphology of the actin cable was assessed. (A) Images of the actin cable when stained with Actistain 488 and visualized with the confocal microscope. The left column is a flat, 2D image taken at the mid-plane of cells when cells are in focus. The right column is the projected pictures from 3D images that allow the visualization of all cables inside yeast cells. Each row signifies images from a treatment group, in the top-down order: control, 4 μg/mL, 12 μg/mL, 25 μg/mL, and 50 μg/mL. The size bar is equivalent to 5 μm. (B) The percentage of cells with abnormal actin cable was quantified and graphed. A total of 30 cells were used for each sample. * p < 0.05.
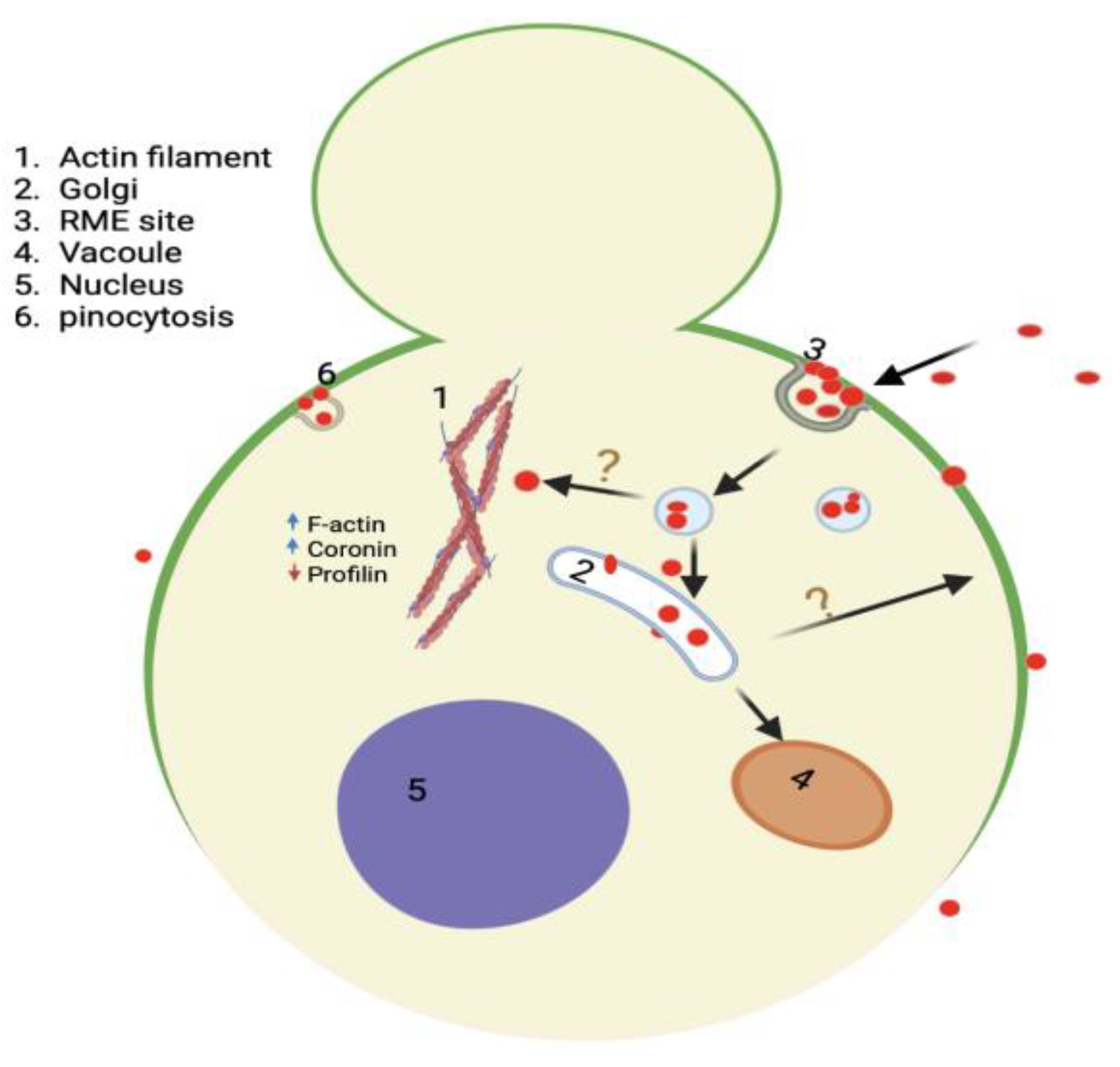
Figure 14.
Model summarizing the interaction and impact of QDs in yeast, including their mode of entry mainly by endocytosis, and their trafficking to the Golgi. The model also shows multiple impacts QDs have on yeast cells, such as loss of polarized GFP-Snc1 membrane distribution and alteration of actin dynamic regulatory protein to support actin polymerization and bundling. Created with BioRender.com.
Figure 14.
Model summarizing the interaction and impact of QDs in yeast, including their mode of entry mainly by endocytosis, and their trafficking to the Golgi. The model also shows multiple impacts QDs have on yeast cells, such as loss of polarized GFP-Snc1 membrane distribution and alteration of actin dynamic regulatory protein to support actin polymerization and bundling. Created with BioRender.com.
Table 1.
Yeast strains used in this study.
Table 1.
Yeast strains used in this study.
| Strain | Status | Media | Genotype |
|---|
| S288C | Existing | YPD | MATα SUC2 mal met gal2 CUP1 flo1 flo8-1 hap1 |
| Abp1-GFP | Existing | SD-HIS | MATα ABP1-GFP-HIS3 his3∆1 leu2∆ met15∆ ura3∆ |
| GFP-Snc1 | Existing | SD-URA | MATα his3∆1 leu2∆ met15∆ ura3∆ SNC1-GFP-URA |
| GFP-2PH | Existing | SD-URA | MATα his3∆ura∆leu∆trp∆lys∆ GFP-2PH (PLC)-URA3 |
| Vps10-GFP | Existing | SD-HIS | MATα his3∆1 leu2∆ met15∆ ura3∆ pRS426 GFP-PH FAPPI |
| Abp140-GFP | This study | SD-HIS | MATα his3∆1 leu2∆0 met15∆0 ura3∆0 Abp140-GFP-HIS |
| GFP-Cof1 | This study | SD-URA | MATα his3∆1 leu2∆0 met15∆0 ura3∆0 GFP-Cof1-URA |
Table 2.
Mean size and zeta potential of QDs in water, nutrient-lacking histidine media (−His +Glu), and yeast peptone dextrose media (YPD) using DLS. The polydispersity index (PDI) value is also provided. PDI that is less than 0.1 represents a highly monodisperse sample, usually a clean standard. In contrast, a PDI value greater than 0.7 indicates that the sample has a very broad size distribution and may have agglomerations.
Table 2.
Mean size and zeta potential of QDs in water, nutrient-lacking histidine media (−His +Glu), and yeast peptone dextrose media (YPD) using DLS. The polydispersity index (PDI) value is also provided. PDI that is less than 0.1 represents a highly monodisperse sample, usually a clean standard. In contrast, a PDI value greater than 0.7 indicates that the sample has a very broad size distribution and may have agglomerations.
| | Sample | QD + Water | QD + HIS | QD + YPD |
|---|
| Mean Intensity | PDI | Mean Intensity | PDI | Mean Intensity | PDI |
|---|
| Size (nm) | Run-1 | 17.2 | 0.168 | 1372.5 | 0.994 | 178.0 | 0.309 |
| Run-2 | 15.6 | 0.106 | 906.9 | 0.882 | 169.7 | 0.122 |
| Run-3 | 17.6 | 0.061 | 2091.2 | 0.856 | 184.1 | 0.240 |
| Avg | 16.8 | 0.112 | 1456.9 | 0.911 | 177.3 | 0.224 |
| ZP (mV) | Run-1 | −49.2 | 3.4 | 0.7 |
| Run-2 | −49.5 | 3.2 | 0.6 |
| Run-3 | −49 | 3.3 | 0.9 |
| Avg | −49.23 | 3.30 | 0.73 |
Table 3.
Agglomeration assay. QDs incubated in water with a pH ranging from 1 to 7 and incubated for 24 h. The samples were spun to isolate largely agglomerated QDs at the bottom; the supernatant was then tested using ICP-OES.
Table 3.
Agglomeration assay. QDs incubated in water with a pH ranging from 1 to 7 and incubated for 24 h. The samples were spun to isolate largely agglomerated QDs at the bottom; the supernatant was then tested using ICP-OES.
| | Average Cd Content (ppm) |
|---|
| Control + QDs | 2.038 |
| pH 1 + QDs | 0.977 |
| pH 2 + QDs | 1.926 |
| pH 3 + QDs | 2.032 |
| pH 4 + QDs | 1.920 |
| pH 5 + QDs | 2.111 |
| pH 6 + QDs | 2.112 |
| pH 7 + QDs | 2.093 |
Table 4.
Agglomeration assay. QDs in SD-HIS and YPD with natural pH (4.86 and 5.57) and neutral pH (pH 7). The samples were spun to isolate largely agglomerated QDs at the bottom; the supernatant was then tested using ICP-OES.
Table 4.
Agglomeration assay. QDs in SD-HIS and YPD with natural pH (4.86 and 5.57) and neutral pH (pH 7). The samples were spun to isolate largely agglomerated QDs at the bottom; the supernatant was then tested using ICP-OES.
| | Average Cd Content (ppm) |
|---|
| YPD pH 5.57 | 1.049 |
| YPD pH 7 | 1.107 |
| −His + Glu pH 4.86 | 0.852 |
| −His + Glu pH 7 | 1.069 |
Table 5.
Upregulated genes that showed the impact of red CdSe/ZnS QDs on membrane trafficking, metabolism, and cellular processes. Genes are sorted by protein class.
Table 5.
Upregulated genes that showed the impact of red CdSe/ZnS QDs on membrane trafficking, metabolism, and cellular processes. Genes are sorted by protein class.
| Upregulated Genes |
|---|
| Vesicular Trafficking (6) | DID2, COS10, VAC7, ECM21, TRS85, SND1 |
| Zinc Ion Homeostasis (3) | IZH4, IZH1, IZH2 |
| Transcription Factors (5) | GAT2, CIN5, HAP4, SKO1, STP4 |
| Enzymes and Protein Kinases (10) | TOS3, MPS1, BIO2, ENA2, ARO10, NFL1, CIP1, DPB2, SAP1, AFT1 |
| Mitochondria (3) | PET111, PET112, SLS1 |
| Cell Wall Organization (1) | ZRG8 |
| Eisosome (2) | SEG2, SEG1 |
| Fatty Acid Metabolism (10) | OLE1, FAA4, MGA2, PBL2, ICT1, NEM1, TDA4, TDA4, HMG2, OSH2, SCT1 |
Table 6.
A list of downregulated genes showed the impact of red CdSe/ZnS QDs on membrane trafficking, cell wall organization, cytoskeleton, metabolism, and protein translation. Genes are sorted by protein class.
Table 6.
A list of downregulated genes showed the impact of red CdSe/ZnS QDs on membrane trafficking, cell wall organization, cytoskeleton, metabolism, and protein translation. Genes are sorted by protein class.
| Downregulated Genes |
|---|
| Ribosomal large subunits (28) | RPL46A, 37A, 26B, 23B, 31A, 37B, 22B, 38, 43B, 33B, 22B, 43A, 33A, 39, 30, 29, 25, 42B, 35A, 40A, 14A, 24A, 35B, 28, 42A, 34B, 7B, 14B |
| Ribosomal small subunits (18) | RPS28A, 29A, 29B, 23B, 28B, 30B, 27B, 10B, 17A, 13, 25A, 27A, 25B, 31, 21A, 19A, 21B, 22A |
| Ribosome associated (7) | RPS30A, TMA7, TMA10, TMA16, YET2, MRT4, FAP7 |
| Mitochondria function (32) | ATP20, ATP18, ATP19, ATP15, TIM8, TIM12, TIM10, TIM11, COA2, FMP45, TMH18, RSM19, MIX14, MRPL50, MRPL37, MRPS16, MTC3, EMI1, CMC1, BOL1, COA4, MIC12, MRP10, MRP2, COX7, FIS1, MRPS12, MRPL51, TRX1, HSP10, OAC1, QCR8 |
| RNA processing (13) | LSM3, NOP10, RDS3, IST3, HUB1, YSF3, LSM7, DIB1, POP5, POP8, LSM6, APQ12, SMX3, |
| Stress response (21) | ROQ1, GRX1, HOR7, ATX1, MHF1, MHF2, SRX1, TMA10, GRE1, DDR2, POP8, HMF1, HSP12, SPR28, MEI5, SPO19, PAU20, PAU19, AUS1, FMP16, SDD2 |
| Cell wall integrity (4) | DAN4, DAN1, FMP45, TIR4 |
| Sporulation and meiosis (7) | BNS1, SHU2, EMI1, SPR28, MEI5, SPO19, FMP45 |
| Protein folding/chaperone/Degradation (7) | PFD1, GIM3, ROQ1, RUB1, HSP10, POC4, EMC6 |
| Transporter (10) | PHO89, AUS1, PMP3, SIT1, OAC1, DRE7, HXT6, VMA11, VMA9, VMA21 |
| Vacuole (4) | VMA9, VMA11, VMA21, LCL2 |
| Vesicular trafficking and Secretion (21) | YOS1, TRX1, SNX3, APS2, EFM4, MLC2, DYN2, APR10, TCA17, LDB18, SFT1/NCE101, YSY6, SBH2, SRT1, GOT1, SSA3, KSH1, SPC2, SSS1, OST1 |
| Transcription (13) | MBF1, TFB5, FYV5, HMRA1, SPT4, HHT1, AHC2, RPB11, SRB6. RPC11, RPO26, RPC10, YFR1-2 |
| Nuclear import and export (3) | DIF1, ECM1, SYO1 |
| Membrane lipid synthesis and homeostasis (6) | CHO1, OST1, SRT1, TSC3, OPI11, OPI3 |
| Enzymes (20) | FCY1, CTF8, TSC3, RNH203, PBI2, OPI1, VMA9, VMA21, WIP1, BNA1, BNA2, SSA3, REE1, HMRA1, URA10, RPC10, POC4, CFD1, HIS6, DFR1 |
| Replication (1) | CTF8 |
| Apoptosis (1) | SDD2 |